Abstract
As pathogenic (micro)organisms constitute a biological hazard, they have been categorized into classes of risk (CR). The CR have been defined for human pathogens by the World Health Organization and for animal pathogens by the World Organization for Animal Health. A few countries have also established their own definitions and lists of pathogenic (micro)organisms with their class of risk. The aim of this paper is to propose an improved approach to strengthen the final result obtained for the classification of ill-defined or uncategorized newly (re)emerging human or animal pathogen. This approach has been tested and validated with highly pathogenic avian influenza (HPAI) A H5N1 viruses. In this one health approach, first, a questionnaire containing questions related to the criteria for biological risk assessment of a microorganism was sent to human and animal health professionals and stakeholders. After that, aggregated summary results were communicated to participating experts, after which a meeting was organized to confront the different opinions and address any remaining uncertainties. We show that a semi-quantitative approach based on expert knowledge elicitation allows circumventing limitations from purely qualitative models by confronting and critically assessing opinions of experts from different fields of expertise.
Keywords
Introduction
The intrinsic properties of a pathogenic (micro)organisms contribute to the adverse effects it could have on human or animal health. The World Health Organization (WHO) and many countries have defined criteria for the classification of (micro)organisms into 4 classes of risk (CR), taking into account the severity of the disease that the pathogens may cause in immuno-competent humans, their ability to spread among the population, and the availability of prophylaxis or efficient treatment. 1 Similarly animal pathogens are classified according to the definitions of the World Organization for Animal Health (OIE), which also considers economic aspects of diseases. 2
Many national regulations aiming at protecting human health and the environment against harmful effects of pathogenic (micro)organisms refer to lists in which these (micro)organisms are classified into CR. This is also the case in Belgium, where the legislation provides classification lists of human and animal pathogens (3 decrees, 1 per region).3 -5 Classification is often directly referring to containment measures or even containment levels. The Belgian classification system only considers the inherent characteristics of the (micro)organisms and not their intended use, making the risk classification independent of containment measures. This means that a clear distinction is made between the biological CR of the pathogen and the CR of the activity. The class of risk of the activity depends both on the biological CR of the pathogen and the type of manipulations carried out within the laboratory or animal facility (eg, diagnosis, research, animal experiments). In a risk assessment, both need to be considered to define containment level and specific safety measures that should be adopted to protect human health and the environment. Hence, the CR of the activity may be equivalent to the CR of the micro-organism, or it may be higher or even lower.
Classification lists should ideally be dynamic and regularly updated in light of new scientific knowledge and emergence of new pathogens. In the current classification lists published in the Belgian regional decrees and revised in 2008, influenza A, B, and C viruses were classified into CR2 for human and CR3 for animals. Given the diversity of influenza viruses and the emergence of new strains, a revision of the classification was deemed necessary.
Influenza viruses from the Orthomyxoviridae family consist of 7 genera; only viruses of the A genus are known to infect birds as well as mammals and have been responsible for several pandemics since the 16th century. 6 The 3 major pandemics were the Spanish flu (1918-1919), Asian flu (1957), and Hong Kong flu (1968-1969), which resulted in a large number of deaths. 7 The 1918 (H1N1) pandemic has been recorded as the worst pandemic in history. It infected 500 million people globally and killed 50 to 100 million people worldwide. 8 Epidemics and the occasional pandemics result from the segmented nature of the genome and high-frequency mutations during replication in multiple hosts. Each segment codes for one of the viral proteins, which include the major surface glycoproteins hemagglutinin and neuraminidase. Influenza A viruses are further divided into subtypes based on any combination based on the antigenic relationships in the surface glycoproteins, hemagglutinin and neuraminidase. The high mutation rate of type A viruses, a higher degree of variability in its antigenicity, and virulence constitutes a threat each year for humans as it can cause zoonotic infections and adapts easily to humans, leading to a sustained human-to-human transmission, which favors the emergence of novel strains. Human-to-human transmission usually occurs through respiratory droplets. Most of the combinations of influenza A subtypes were isolated from avian species where they widely circulate without causing symptoms and so-called low pathogenic avian influenza (LPAI). In birds, the virus mainly spreads via feces and invasion of virus particles by the intestinal tract. To date, only viruses of H5 and H7 subtype have the potential to cause highly pathogenic avian influenza (HPAI) following mutation of LPAI that are introduced into susceptible species, but not all H5 and H7 viruses are virulent. Not all factors that trigger mutation have been understood; it can sometimes happen rapidly after introduction and other times more slowly. Many factors, such as the strain of virus, species of bird, and environmental conditions, make the understating of bird-to-bird transmission complex.9,10
Outbreaks of HPAI viruses have immediate drastic consequence on the agricultural sector due to their faculty of rapid spread and high mortality rates. But HPAI viruses can also infect humans and cause flu-like symptoms, occasionally resulting in potentially fatal severe respiratory disease. Between 2006 and 2016, HPAI has been detected in at least 30 countries and caused 860 human cases with 454 deaths in at least 16 of these countries.11,12
Within this context and taking into account the continuous and evolving threat of a pandemic, we choose to take HPAI A H5N1 viruses as a case study for the development of a risk classification model aimed at assigning CR to microorganisms for humans and animals in a transparent and traceable manner. This model is based on an expert knowledge elicitation approach. We considered it was a suitable approach to address cases where relevant information for a robust evaluation of the CR of a pathogen is scarce or unavailable and circumvent limitations from purely qualitative models relying only on literature.
Methods
This methodology was based on the Delphi method, which starts with an initial question and uses feedback from an expert panel to reach consensus. Delphi method typically uses a questionnaire. It allows experts to provide their individual contribution, confront their view with the panel of experts, and finally, have the opportunity to revise views. It also allows some degree of anonymity for the individual responses. 13
An online questionnaire was developed and made available through a secure extranet platform (Limesurvey). The link to the questionnaire was sent with an individual password to the participants. Each participant completed the questionnaire individually. The questionnaire contained open-ended questions (with a request for justification and/or scientific references) and multiple-choice questions related to the criteria for biological risk assessment of a microorganism described in the Belgian regional decrees3 -5 and Van Vaerenbergh et al. 14 The questions were grouped into modules, each containing a set of questions related to the severity of the disease, its likelihood to effectively spread within the community, and the availability of prophylactic or therapeutic measures, respectively, for humans and animals (Table 1).
An Overview of the Modules and Questions Included in the Questionnaire.

Some questions were common for humans and animals; others were specific. For animals, the CR of a given pathogen should take into account the most sensitive animal species to this pathogen. The determination of the CR of human pathogens does not take into account the increased susceptibility due to, for example, preexisting diseases, medications, compromised immunity, pregnancy, or breastfeeding. Only adults (18-65 years) are considered, excluding YOPIs (young, old, pregnant, and immune-deficient persons). Multiple-choice questions had 3 possible answers that fit an ordinal scale (type: low < medium < high) except for 5 questions: (1) host range (human and animals – only animals – only humans), (2) genetic stability of the virus (type: low < high), (3) horizontal mode of transmission (type: by vector or needle prick – by ingestion – by contact – airborne), (4) prophylaxis, and (5) treatment (type: not available or not used – available but not 100% efficient – available and highly efficient). At the end of the questionnaire, participants were asked to propose a CR for the HPAI A viruses (type: 1 [non-pathogenic] – 2 – 3 – 4). To minimize ambiguity with regards to the questions and their potential answers, definitions were provided for the following items: host range, infectious dose, potential of survival outside the host, mode of transmission, invasiveness, virulence, and classes of risk (see Supplemental Material in the online version of the journal). Participants were allowed to leave questions unanswered with the 2 additional answers of no opinion or unknown.
The resulting data were first interpreted based on the descriptive results and scientific justification and/or scientific references (qualitative risk assessment).
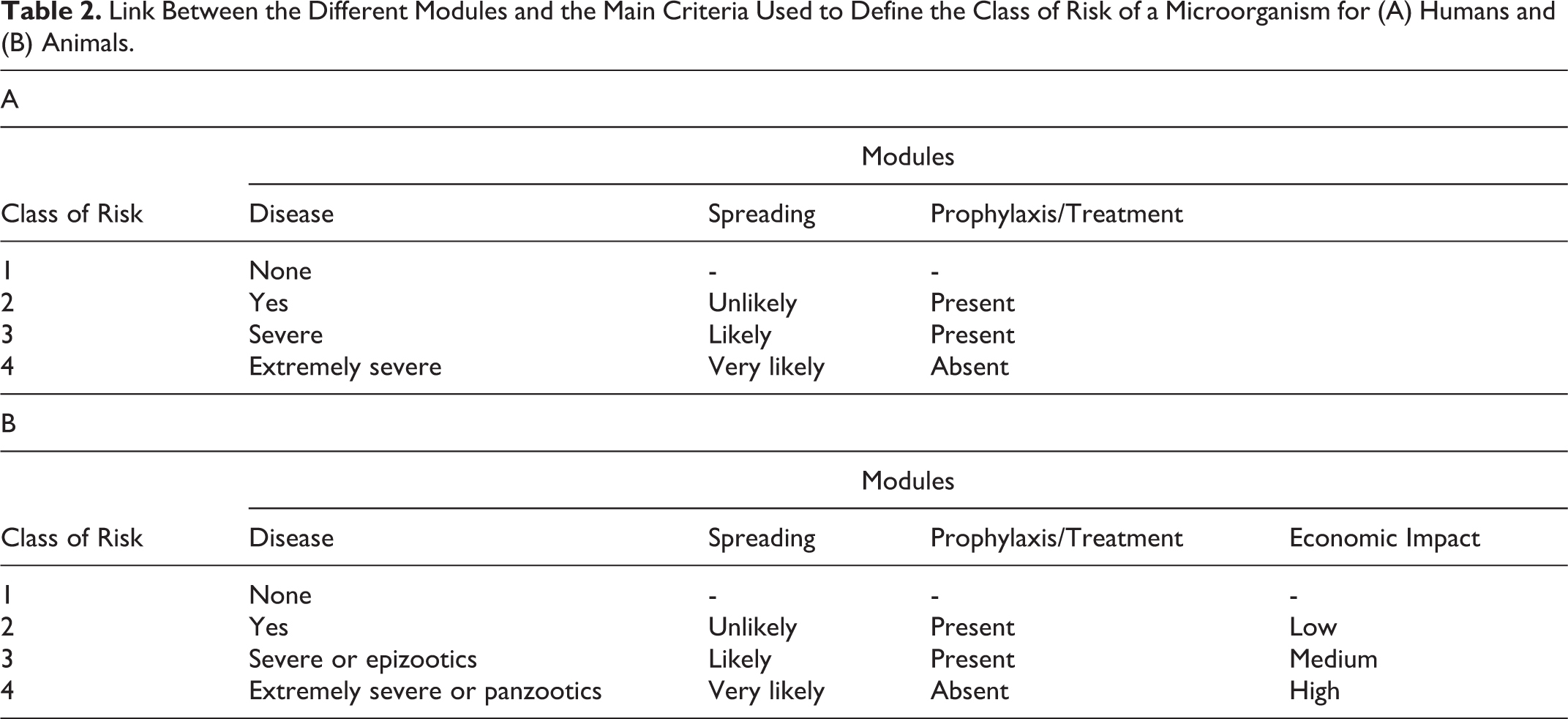
In a second step, an aggregated and anonymous summary of the results was sent to the participants, after which a meeting was organized to address any remaining uncertainties, facilitate a consensus per module based on the different answers provided to each question, and come to an agreement on the final classification of HPAI A H5N1 viruses for humans and animals. To assign a class of risk to HPAI A H5N1 viruses based on the answers to the questions, a matrix was developed to make the link between the different modules and main criteria used to define the class of risk for humans or animals (Table 2).
Link Between the Different Modules and the Main Criteria Used to Define the Class of Risk of a Microorganism for (A) Humans and (B) Animals.
Results
Forty-two Belgian human and animal health professionals with experience in influenza viruses were invited by email to participate in this project. Fifteen effectively answered the questionnaire.
The results obtained for each multiple-choice question are summarized in Table 3. For the questions that received divergent answers, the critical analysis of the results was made difficult due in particular to the diversity in the answers and the fact that several participants (1) did not answer the open-ended questions, (2) failed to give the rationale for their answers, and/or (3) did not appear to have a correct understanding of some questions despite the available definitions.
Aggregated Summary of the Answers to the Questionnaire and Summary of the Points of Discussion and Conclusions at the Meeting with the Experts.
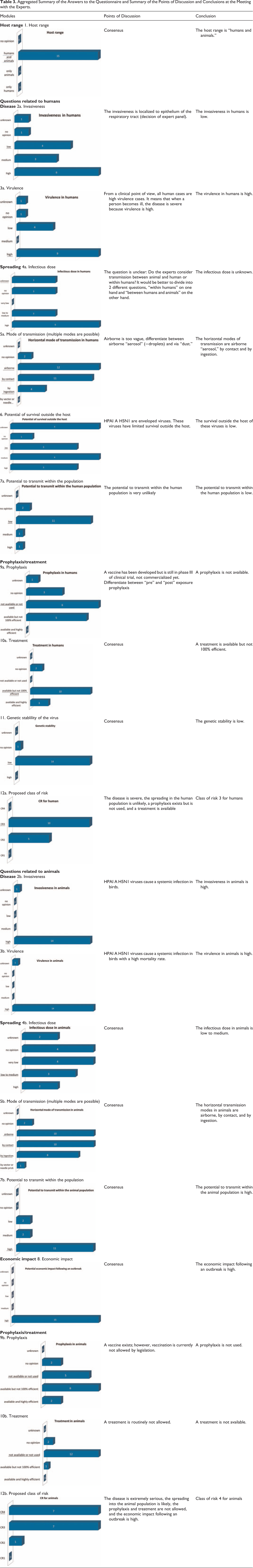
The plenary meeting organized with the participants allowed them to critically discuss the summary of the results and validate the final classification of HPAI A H5N1 viruses for humans and animals. Nine of the 15 participants attended this meeting. The outcome of the exchanges of view on each question during the meeting was sent to all the experts participating in this project. All the experts considered that the host range of HPAI A H5N1 viruses covers humans and animals, and therefore a CR can be assigned for humans and animals. The most sensitive animal species is birds. The subtype of the HPAI A H5N1 viruses determines the characteristics of the viruses. Although the subtype of HPAI A H5N1 viruses was not specified in the survey, all the experts agreed that the classification should be based on the characteristics of the most pathogenic subtype of HPAI A H5N1 viruses, namely, Asian HPAI A H5N1.
With regards to the questions related to human health, the outcome of the meeting can be summarized as follows:
The severity of the disease in humans was hardly discussed. Regarding invasiveness of HPAI A H5N1 viruses, the main targeted tissue is epithelium of the respiratory tract, so the invasiveness was therefore considered as low. All human cases were virulent cases; when a person becomes ill, the disease is severe. Consequently, the disease was considered as a severe disease. Highly pathogenic avian influenza A H5N1 viruses do not appear to be transmitted easily from person to person, and the risk of community-level spread of these viruses remains low. Rare cases of probable human-to-human transmission and no cases of sustained transmission have been reported in humans infected with HPAI A H5N1 viruses. Moreover, these viruses are enveloped viruses and have limited survival outside the host. A prophylaxis is currently not available. Normal flu vaccines that contain circulating strains (influenza A H1N1, H3N2 and influenza B strains) are designed to protect against seasonal flu and not intended to be used against HPAI A H5N1 viruses. In the European Union, the European Commission and the European Medicines Agency (EMA) have put procedures in place to speed up the assessment and authorization of vaccines for use during a flu pandemic. If a pandemic with HPAI A H5N1 viruses occurs, a vaccine against HPAI A H5N1 viruses has already been developed and could be used (Adjupanrix).
15
A treatment based on antivirals (oseltamivir) exists but is not 100% efficient.
16
Based on the results of this process, HPAI A H5N1 viruses were classified in CR3 for humans. Class of risk 3 is defined as microorganisms that can cause a severe disease in humans and present a serious hazard for directly exposed persons; these organisms may present a risk of spreading to the community, but there is usually effective prophylaxis and treatment available.
The questions related to animal health triggered fewer issues and led to the following main considerations:
Highly pathogenic avian influenza A H5N1 viruses cause a systemic infection in birds with a high mortality rate. HPAI A H5N1 viruses have killed hundreds of millions of birds.
17
Consequently, the disease is considered as extremely severe. The spreading of these viruses into animal population is likely, and the economic impact following an outbreak is considered as high. A vaccine against HPAI A H5N1 exists; however, vaccination is not used because it is currently not allowed by Belgian legislation because it may mask an outbreak of the disease, thereby delaying detection and increasing the risk of it spreading. A treatment is routinely not allowed. Considering that HPAI A H5N1 viruses have caused extremely serious epizootics in birds, with a very high mortality rate and dramatic economic consequences in the affected farming regions and no medical prophylaxis allowed, these viruses were classified CR4 for animals.
Discussion
The risk classification methodology developed in this project allowed assignment of a CR to a microorganism both for humans and animals through an objective, transparent, traceable, reproducible, and integrated process. This method is based on a one health approach, an approach to design and implement programs, policies, legislation, and research in which multiple sectors communicate and work together to achieve better public health outcomes. 18
The questionnaire developed as a starting point of our methodology was very useful to collect experts’ opinions in a structured and traceable manner and framed according to the criteria used for the risk assessment of a microorganism. However, it was not self-sufficient to draw conclusions, for different reasons:
Although explicit definitions were provided for each question, some of them were considered unclear by some participants and subject to interpretation bias; this shows that the questions and explanatory definitions should be carefully phrased. In some cases, participants did not provide a scientific reasoning and/or scientific references to support their answers. The answers to some questions were influenced by the scientific or professional background (eg, clinicians vs scientists), reflecting differences in the perception of risk. This highlights the importance of involving a multidisciplinary expertise.
The answers to the questionnaire therefore provided an evidence-based framework indicating specific issues requiring further discussion. This was achieved during a meeting gathering together the experts. This meeting allowed gaining further insight in the reason or source of remaining uncertainties around the criteria used for the risk classification of the microorganism. It was an opportunity for the participants to explain their points of view and provide scientific evidence, which, as mentioned before, was sometimes difficult to get via the questionnaire. When uncertainties remained after the discussion, the worst-case scenario was considered. Some participants expressed the views that clearer instructions should be provided in advance to the survey to avoid misinterpretations and stimulate scientific argumentation.
Fifteen of the 42 invited experts effectively participated in this project. The response rate is very difficult to anticipate and depends on several factors, including the availability of the experts. This could be improved in the future by contacting experts on an individual basis and enlarging the panel to international experts. With regards to the face-to-face meeting, experts who cannot attend could be given the opportunity to participate in the discussion via tele- or videoconference.
The final classifications obtained were similar to those proposed previously for wild-type HPAI A H5N1 strains for humans and animals based only on a review of the literature. 19 Although similar conclusions were reached, it is important to note that when relevant information for a robust evaluation of the CR of a pathogen is scarce or unavailable, relying only on literature can introduce bias, stemming from ambiguous results of the papers that were influenced and potentially guided by the research interests of the authors, short-term political agenda, or residuals of historic situations.
We consider that this approach could be used in particular to classify ill-defined or uncategorized newly emerging human or animal pathogen, for example a new re-assortant influenza strain.
Through an expert knowledge elicitation approach, allowing collecting and critically assessing the opinions of experts from different area expertise, the methodology described in this paper circumvents the limitations from purely qualitative models (which are often considered subjective).
Footnotes
Authors’ Note
Aline Baldo and Sarah Welby contributed equally to this work.
Acknowledgments
The authors thank Nicolas Willemarck, Emilie Descamps, Katia Pauwels, and Fanny Coppens (Sciensano) for their useful contribution to this document and all the experts participating in this project. This work received support from the Brussels-Capital Region (IBGE-BIM), the Flemish Region (LNE), and Wallonia (DGARNE).
Ethical Statement
Not applicable.
Statement of Human Rights
Not applicable.
Statement of Informed Consent
All participants gave their informed consent prior to their inclusion in the study.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
