Abstract
As gene editing technology continues to evolve at a rapid pace, so must the ability to safeguard against related biosafety hazards, particularly when human health is at issue. Genome editing based on the CRISPR (clustered regularly interspaced short palindromic repeats)–Cas system is a novel and powerful tool for modifying somatic and germ line cells in living organisms. As the technology matures, it is being adopted around the world for research and product development purposes. Yet, there remain challenges with specificity and a potential for unwanted genetic modifications. Here, we review several techniques for pursuing more precise genome editing through CRISPR. We also provide biosafety professionals with current information regarding the state of CRISPR technology and examples of publicly available resources for predicting the likelihood of off-target effects.
Researchers have pursued the ability to easily target and modify specific DNA sequences for decades. In just the past few years, a revolutionary new technology known as
In recent years, researchers have refined their understanding of the underlying biological mechanisms by building a host of genome engineering tools based on the CRISPR system.6
-8 In the commonly used CRISPR-Cas9 system, the Cas9 enzyme from
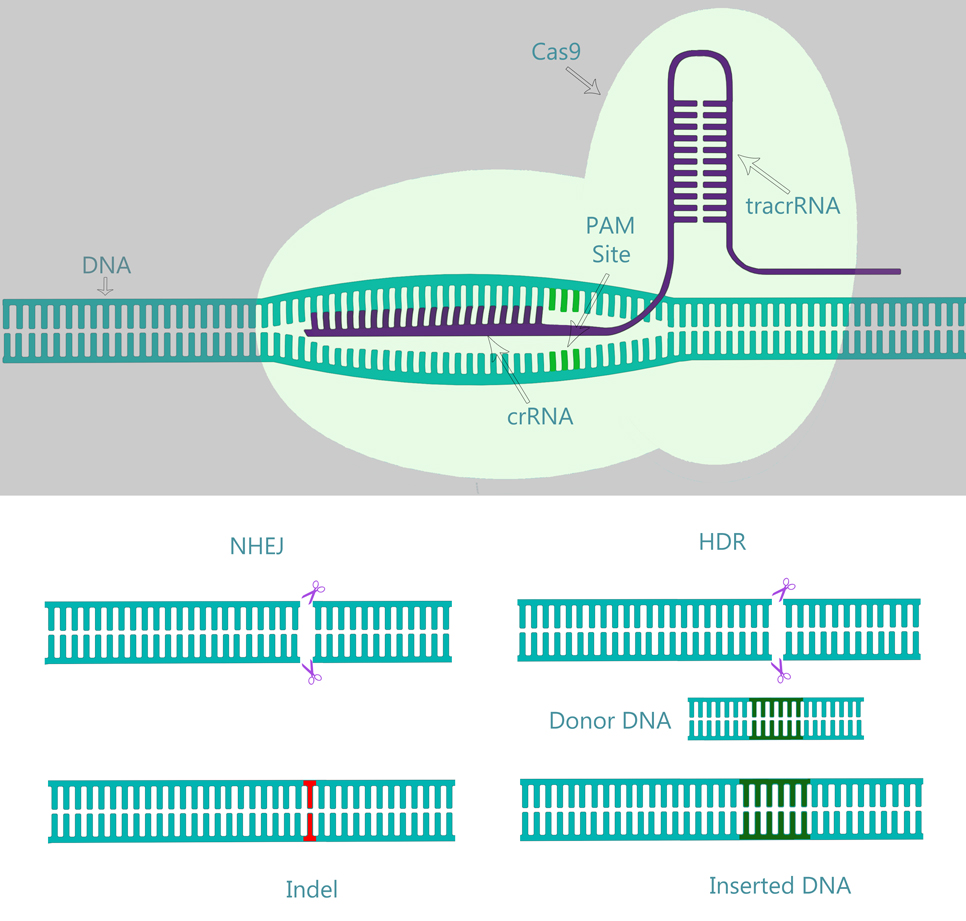
DNA cleavage and repair by the CRISPR (clustered regularly interspaced short palindromic repeats)–Cas9 system. The CRISPR-Cas9 system with a 20-nucleotide gRNA composed of CRISPR RNA and trans-activating CRISPR RNA in complex with Cas9. The crRNA base pairs with the target sequence, and Cas9 binds and cleaves the target sequence upstream of the protospacer adjacent motif (PAM) site. Two common forms of DNA repair mechanisms are shown: nonhomologous end joining (NHEJ), whereby the cell attempts to ligate the cut ends back together, often resulting in insertions or deletions of nucleotides at the repair site (shown in red); homology-directed repair (HDR), whereby repair can be guided by researcher-provided donor DNA, which the cell uses as a template to repair the double-strand break, allowing for gene editing of the target sequence (shown in green). (The color version of this figure is available in the online version at http://journals.sagepub.com/doi/full/10.1177/1535676017694148.)
The CRISPR RNA sequence, typically 20 nucleotides in length, is easily modified to match any desired DNA sequence adjacent to a protospacer adjacent motif site. As a result, CRISPR can be used to target and cut DNA sequences of interest. Normal DNA repair mechanisms can be utilized to insert a new DNA sequence, edit the existing sequence, or even knockout the gene altogether. As a result of these capabilities, CRISPR offers a relatively simple, modular, and cost-effective system for performing genome editing, which has been validated in a host of cells and organisms.6 -8 This has led to the development of several impactful applications, many of which are rapidly advancing as CRISPR technology continues to mature.
To address pressing global challenges—such as public health, environmental, and agricultural issues—CRISPR is being used to advance the development of gene drives, which are genetic modifications designed to be self-propagating.9 -11 With this application of CRISPR, a population of mosquitoes, for instance, could be modified to carry a Cas9-based DNA endonuclease cassette, which cuts within the same chromosomal locus, along with a malaria-resistant gene. 9 When these modified mosquitoes reproduce with unmodified mosquitoes, their offspring receive one chromosome containing both the CRISPR cassette (gene drive) and the malaria-resistant gene. The gene drive can then cut the corresponding unmodified chromosome, allowing normal DNA repair processes to repair the cut by using the chromosome containing the gene drive and malaria-resistant gene as a template. This can result in both chromosomes containing the gene drive and malaria-resistant gene. These offspring will then pass on the modified genome to their offspring, and this process is repeated throughout subsequent generations, thereby facilitating widespread distribution of engineered traits. 9
With the improved genome editing capabilities offered by the CRISPR system, researchers may be able to use gene drives to spread almost any desired genomic modification through sexually reproducing populations, theoretically resulting in inheritance of the modification by all members of a population over many generations. Considering the potential for entire wild populations to be modified, unforeseen ecological consequences are of concern. However, the ecological and ethical considerations surrounding the use of gene drives must be balanced against the potential benefits to human health and agriculture, such as controlling the spread of insect-driven diseases (including malaria, dengue, and Zika), countering pesticide and herbicide resistance in plants, and eradicating invasive species causing agricultural and environmental harm.9,12,13
Another application of CRISPR is the development of animal models, which are a valuable resource for medical research.14,15 It is not currently possible to artificially replicate the intricacy and complexity of a living organism, with its many interrelated tissues and organ systems. Therefore, model organisms offer important insights for researchers studying development and diseases, as well as testing the safety and effectiveness of diagnostic procedures and experimental medical treatments.
The use of CRISPR enables faster and more cost-effective creation of genetically tailored preclinical animal models, which can be used to validate new therapies. These animal models also provide a means of studying how genes function
CRISPR is faster and more powerful than traditional methods of generating animal models, allowing researchers to simultaneously modify multiple genes. 14 Mouse strains have been created that express Cas9, bypassing the need to inject Cas9 into animal models when carrying out CRISPR experiments, thereby accelerating researchers’ understanding of disease processes. 16 Furthermore, CRISPR is pushing forward the generation of large animal models to more accurately depict human neurodegenerative diseases, such as Alzheimer disease, Parkinson disease, amyotrophic lateral sclerosis, and Huntington disease. 17 The ability to flexibly produce genetic models will also help researchers understand the cancer genome and pinpoint genes that drive cancer formation and progression.18 -20 Thus, CRISPR provides a means of performing biological investigations and studying gene functions on a large scale.
Gene therapies are increasingly being pursued as a means of addressing presently incurable diseases, and researchers are already using CRISPR technology to test various experimental treatments
Testing of CRISPR-based gene therapies in humans is now being pursued as well. In June 2016, the first US clinical trial utilizing CRISPR was reviewed by the US National Institutes of Health’s Recombinant DNA Advisory Committee.
27
Scientists conducting the clinical trial, if it receives approval by the US Food and Drug Administration, will obtain T cells from 18 patients with a variety of cancers and genetically modify the cells
CRISPR Biosafety Considerations
Despite its many promising benefits for human health, there are many significant biosafety considerations related to the development and use of CRISPR technology. Protecting laboratory personnel and the community from exposures to infectious agents and biohazardous materials, as well as preventing unintended releases to the environment, remains a key facet of biosafety. As with any emerging technology, this requires understanding the implications of potential exposures, including the potential exposure routes and effects of the materials being used. However, due to the unique challenges posed by genome editing technologies, it is necessary to reevaluate traditional risk assessment procedures to ensure that the right questions are being asked. For instance, how should an investigator evaluate the risk of using a specific gRNA? Similarly, what steps may be taken to prevent unwanted genetic modifications from occurring, as with the testing of gene drives? What are proper bioconfinement methods? Are there special considerations for medical personnel responding to accidents involving CRISPR technology?
Perhaps the primary biosafety issue rapidly gaining real-world relevance relates to the use of CRISPR in gene therapies. A primary challenge for researchers conducting the CRISPR studies in human patients is avoiding unintended genetic modifications, or off-target effects. In the US clinical trial, scientists will perform multiple modifications to the T cells, inserting and removing genes based on their potential to facilitate either protective or harmful effects. 28 Preventing and monitoring for unintended genomic modifications is a primary consideration, and researchers will follow trial participants for 15 years to evaluate potential off-target effects. 28 In fact, the real test of this early stage clinical trial is to simply identify whether CRISPR is safe for human use.
While there are many biosafety considerations relevant to ensuring the safe use of CRISPR technology for genomic-editing research and clinical applications, this article provides insight into just one facet—the potential for genomic off-target effects to lead to unknown health risks. Due to the ease and speed at which genome editing capabilities are being developed, biosafety professionals and scientific investigators need to work collaboratively to design experimental models and practices that are both safe and effective. Although CRISPR technology is still being refined, it is important for biosafety professionals to stay informed about the scientific advancements taking place and the ways in which relevant safety implications are being addressed. Here, we examine several strategies for minimizing unintended off-target genomic effects.
Minimizing Off-Target Effects
To safely employ the many benefits CRISPR offers, researchers need the ability to precisely control the sites at which genomic edits occur. Off-target mutations arise when genomic modifications take place at unintended sequences in the genome. This can occur when the gRNA sequence—designed to direct the genome editing machinery to the DNA sequence intended for modification—is closely matched to another sequence site in the genome, perhaps differing by only a few nucleotides. If the gRNA instead recognizes this closely matched sequence, an off-target genome modification can occur. Thus, the target specificity of CRISPR-Cas9 contends with its propensity for off-target mutagenesis, or off-target genetic modification, and is believed to be affected by a number of factors, the exact parameters of which are still being determined. What follows is a summary of several approaches for improving CRISPR specificity and reducing off-target activity. Many of the materials for carrying out the strategies discussed herein are publicly available through Addgene, a nonprofit plasmid repository.
Strategic gRNA Design
One main strategy for reducing off-target activity and thereby increasing specificity is strategic gRNA design. Typically, several potential gRNAs can accomplish the same gene editing goal, but each gRNA poses its own risk of off-target mutations. Selecting a unique target site that has few genetically similar off-target sites throughout the genome can minimize off-target activity.
31
Computational, or
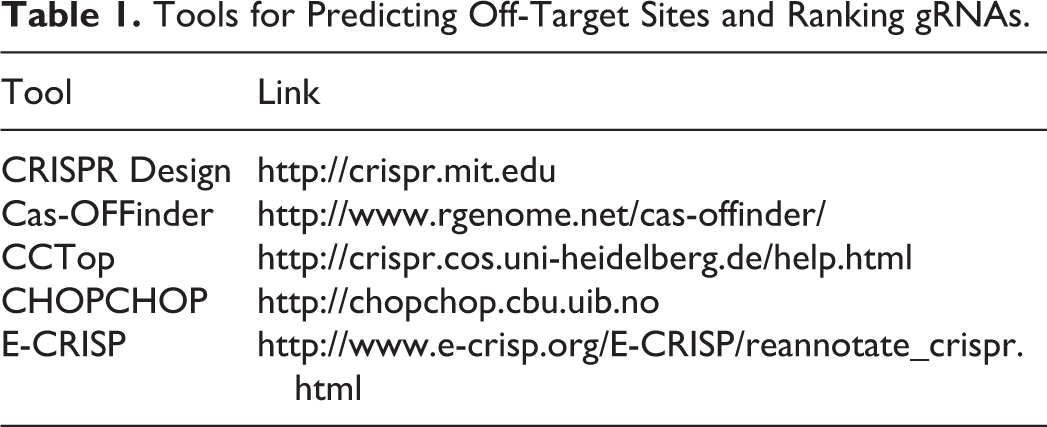
There are many web-based tools available to predict off-target sites and rank gRNAs accordingly, although these applications vary in their sophistication (see Table 1 for a partial list of available resources). Some applications rank gRNAs solely on the basis of sequence similarity between the gRNA and the predicted off-target sites. 33 Other applications generate specificity scores that take into consideration multiple factors expected to influence off-target activity. 33 Such factors may include the number and location of mismatches that the gRNA shares with off-target sites. 33 If a gRNA is very similar to an off-target site, with <3 base mismatches, off-target effects are more likely. 33 Generally, mismatches in the 3′ end of a guide sequence (usually 8-14 bases upstream of the protospacer adjacent motif site) are less tolerable, whereas mismatches toward the 5′ end are more tolerable. 34 Also, off-target effects are sensitive to the amount of gRNA used and the ratio of gRNA to Cas9. 34
Tools for Predicting Off-Target Sites and Ranking gRNAs.

Genome-wide Detection of Off-Target Activity
Computational prediction of off-target activity followed by experimental assessment of the genome at the predicted off-target sites is a routinely used approach for assessing off-target activity. Researchers sequence the genome at predicted off-target sites that have a high probability of mutation. However, this approach leaves open the possibility that other, unpredicted off-target sites will go undetected.
To alleviate these concerns, researchers are analyzing the entire genome for off-target events, not just restricting their analysis to those sites predicted by computational models. The results provided by these deep sequencing-based methods show that sequence homology, by itself, is not an adequate basis for predicting off-target sites.
32
Several assays, such as GUIDE-seq (genome-wide, unbiased identification of DSBs enabled by sequencing)
35
and HTGTS (high-throughput, genome-wide, translocation sequencing),
36
enable researchers to analyze living cells for off-target cleavage sites. Other methods, such as BLESS (direct
Truncated gRNAs
Initially, CRISPR gRNAs were typically designed to have 20 nucleotides of complementarity with a desired target sequence. However, Fu et al hypothesized that some of these 20 nucleotides might be unnecessary and even impede CRISPR’s specificity. 40 After further study, these researchers showed that shorter gRNAs, only 17 or 18 nucleotides in length, functioned comparable to full-length gRNAs but with reduced off-target activity. By reducing the number of nucleotides on the gRNA, they significantly reduced undesired mutagenesis while maintaining on-target editing efficiency. In fact, they showed that truncated gRNAs induced low or undetectable mutagenesis even when the shortened gRNA differed from an off-target site by only one or two mismatches. Thus, it was reasoned that researchers could minimize off-target activity by using truncated gRNAs to target unique sequences in the genome. 40
Cas9 Paired Nickases
Researchers have inactivated one of Cas9’s two cutting domains, creating a Cas9-D10A variant that acts as a nickase.3,41,42 As its name suggests, a nickase only breaks, or “nicks,” one DNA strand, rather than making a DSB (Figure 1). 42 The nickase is directed to the desired target by a coexpressed gRNA. As described by Ran et al, with the paired nickase approach, two gRNAs are used: one targeted to a site on one strand and one targeted to an adjacent site on the opposite strand. 42 Each gRNA individually recruits Cas9-D10A, resulting in nicking of both DNA strands in these adjacent locations. The result is, in effect, a DSB at the intended target site. However, the frequency of off-target mutations is greatly reduced, potentially because any single-strand nicks that occur at off-target sites are likely to be repaired with high fidelity. 42 Ran et al reported that this paired nickase approach improves specificity by up to 1500-fold in cell lines.
dCas9 Fused to FokI Nuclease
As described by Bitinaite et al,
eSpCas9
“Enhanced specificity” SpCas9, or eSpCas9, is another tool that can be utilized in genome editing applications to reduce off-target effects throughout the genome, without sacrificing on-target activity. In developing eSpCas9, Slaymaker et al hypothesized that Cas9’s cleavage activity is activated by and dependent on the separation, or unwinding, of the two DNA strands and that strategically weakening Cas9’s strand separation, or helicase, activity would result in reduced off-target effects. 45 In particular, they proposed that off-target cleavage activity takes place when Cas9’s ability to bind to and unwind DNA at an off-target site exceeds the ability of DNA rehybridization forces to bring the two strands back together.
Slaymaker et al devised this hypothesis based on an analysis of SpCas9’s crystal structure in complex with a gRNA and its associated target DNA. The researchers detected a positively charged groove that they believed played a role in stabilizing the nontarget strand as part of the DNA unwinding and cleavage process. By neutralizing positively charged residues within this groove, they were able to weaken Cas9’s ability to interact with the nontarget strand, thereby requiring more exact base pair matching between the gRNA and target DNA. As a result of this more stringent base-pairing requirement, the gRNA is more likely to bind at the correct target site, leading to a substantial improvement in specificity. 45
SpCas9-HF1
Also looking to structural studies of Cas9, Kleinstiver et al developed a high-fidelity variant of Cas9, known as SpCas9-HF1. 46 Their work was based on an “excess energy hypothesis,” which proposed that the SpCas9-gRNA complex contained more energy than was necessary to perform its desired function at its target DNA site. This excess energy was then available for the complex to perform undesired off-target genomic edits. Their strategy was aimed at disrupting certain contacts between SpCas9 and its target DNA site such that the energetics of the complex would diminish its capacity to perform off-target cleavage but still allow for robust on-target activities.
The researchers created multiple variants of SpCas9, each with unique alterations to these contacts, and assessed which variants performed optimally. They found that one of the variants, SpCas9-HF1, was highly functional at on-target sites while exhibiting generally undetectable off-target effects at certain target sites. Other high-fidelity variants exhibited increased specificity and are available in the event that SpCas9-HF1 is not an ideal choice. Thus, the authors provided a highly effective strategy for improving SpCas9 specificity, which can be extended to other Cas9 orthologues and CRISPR-associated nucleases. 46
dCas9 for Base Excision
One final strategy for precise genome editing with reduced off-target activity developed by Komor et al involves fusing dCas9 to a certain enzyme that can correct point mutations, without introducing DSBs. 47 Cellular mechanisms most often respond to DSBs by performing the error-prone nonhomologous end joining DNA repair process, which results in random insertions or deletions (indels) of nucleotides at the repair site, thereby causing a mutation. However, the approach developed by Komor et al directly converts one base to another, without introducing a DSB, thereby avoiding the indel formation that often occurs following a DSB. With further development, this approach has the potential to correct point mutations that are currently causing a host of genetic diseases and reduce undesired genomic modifications. 47
Conclusion
This review provides an overview of some of the applications of CRISPR and the approaches being actively studied to improve its specificity and reduce its potential for off-target effects (Table 2). Most of these studies are focused on using the SpCas9 system because CRISPR systems from other species have been less characterized until now. Understanding the advancements taking place in the CRISPR field is critical to biosafety professionals as it pertains to safety challenges that may arise from accidents, releases, or other exposures to the materials being used. Efforts to maximize CRISPR’s genome-editing efficiency and specificity are also necessary to protect human health and the environment, while allowing humanity to take full advantage of the many benefits this technology promises to one day bestow.
Strategies for Minimizing CRISPR-Cas9 Off-Target Effects.
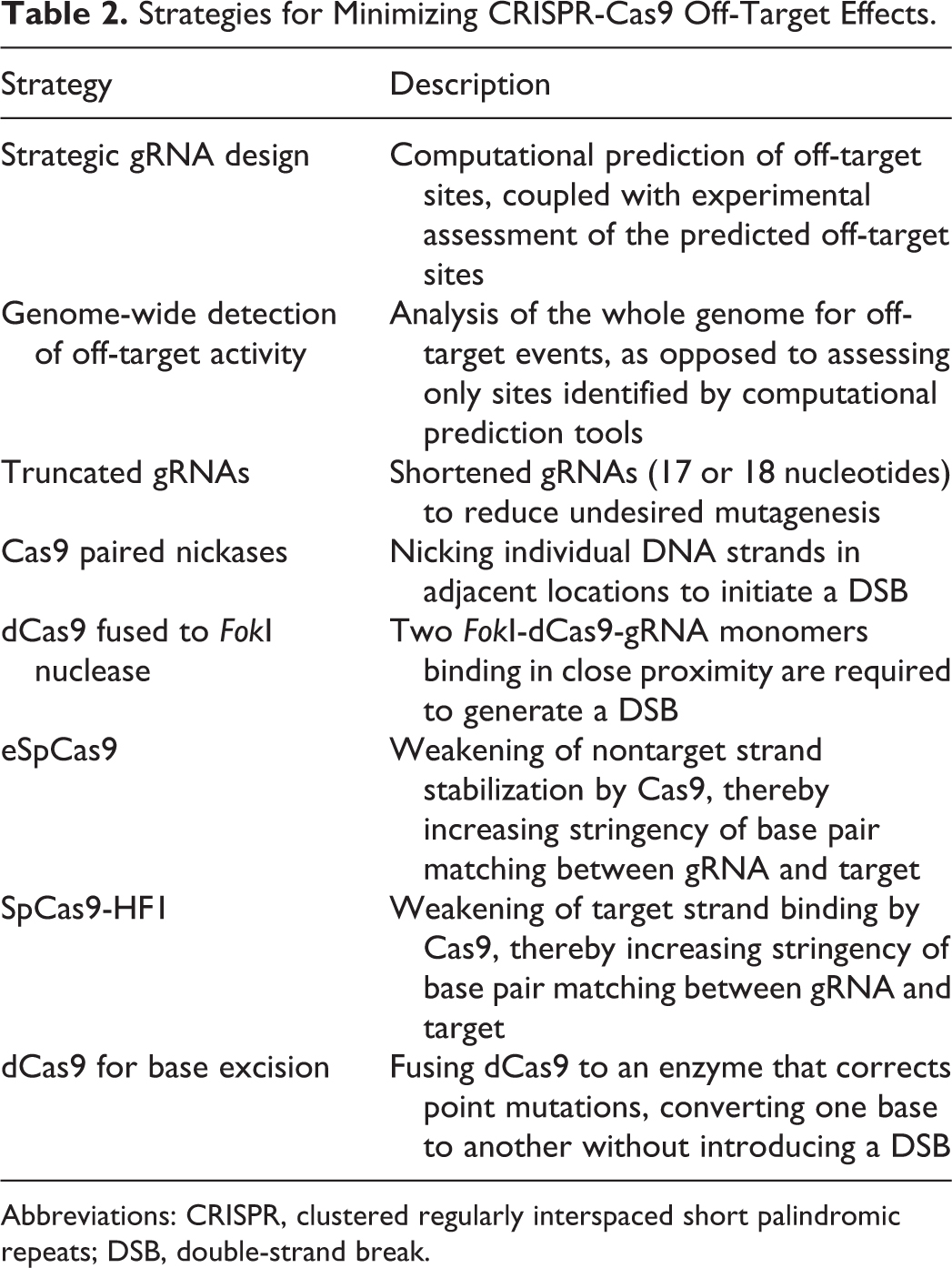

Abbreviations: CRISPR, clustered regularly interspaced short palindromic repeats; DSB, double-strand break.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
