Abstract
Introduction
Over the past few decades, colorectal cancer (CRC) incidence and mortality has steadily declined worldwide; however, CRC is still the most common gastrointestinal malignancy and the second leading cause of cancer-related death. 1 At present, the only curative treatment for CRC is surgical resection, and only modest survival has been reported with chemotherapy. 2 The use of chemotherapy and surgical resection for the treatment of malignant colon cancer is increasing, but the results of these treatments have not considerably improved. Approximately half of CRCs recur, and the patients die within 5 years.2,3 Biomarkers can be used as prognostic indicators, molecular predictive factors, and determiners of targeted therapy.4,5
Colon adenocarcinoma (COAD) has been understood to result from genetic and epigenetic alterations that initiate cancer development and help the progression of cancer.6,7 Oncogenes, which serve as tumor therapeutic targets, promote cancer cell growth and metastasis and have been widely evaluated. 8 Many prognostic markers have been evaluated in COAD; however, most are experimental or have not been prospectively validated.6,9 This study aimed to identify upregulated differentially expressed genes (DEGs) in COAD tumors and nontumor tissues, identify enriched functional pathways, analyze survival outcomes, and search for potential drugs that target the identified DEGs in hope of finding novel and reliable prognostic biomarkers in COAD development.
Materials and Methods
Source of Data
The COAD TGCA (The Cancer Genome Atlas) dataset, updated January 28, 2016, was downloaded (https://cancergenome.nih.gov/). Clinical data and gene expression data including mRNA array and RNA seq data were downloaded using the RTCGAToolbox package in the R software environment. 10 Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) databases GSE41328 and GSE113513 were used for validation of the DEGs identified in the TCGA dataset. Tumor samples and microarray processing data were available for these 2 profiles in the Gene Expression Omnibus database. In total, 10 and 14 pairs of COAD samples were included in GSE41328 and GSE113513, respectively.
Functional Enrichment
The inclusion criteria for identification of upregulated DEGs were set as logFC > 2 and adjusted P < .05. The top 50 upregulated DEGs were selected for heat map and functional enrichment. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway and Gene Ontology (GO) biological process enrichment analysis of upregulated DEGs was conducted using Gene Set Enrichment Analysis (GSEA; http://software.broadinstitute.org/gsea/index.jsp).11,12 To investigate the enrichment of gene sets, upregulated DEGs were uploaded to the Molecular Signatures Database in GSEA. A false discovery rate P value cutoff of <.05 was set as the screening condition. The top 10 KEGG pathways and GO biological processes were presented.
Survival Analysis
Kaplan-Meier survival analysis for upregulated DEGs was conducted in the R software environment using the RTCGAToolbox package 10 and the getSurvival function. All the upregulated DEGs were grouped using median cutoff values into high expression or low expression groups. The R programming script for the survival analysis is attached in the Supplementary Materials.
Analysis of Associated Diseases
After candidate prognostic biomarkers for COAD were detected, the highest scored disease associations for the candidates were identified in the DisGeNET database13,14 (http://www.disgenet.org/web/DisGeNET/menu/home).
Analysis of Targeting Drugs
The DrugBank database (https://www.drugbank.ca/)15,16 was used to identify drugs targeting the candidate biomarkers. The Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP; http://lsp.nwu.edu.cn/tcmsp.php) 17 and the TCM-MESH (http://mesh.tcm.microbioinformatics.org/) 18 databases were used to identify herbs targeting the candidate biomarkers.
Results
Top 50 DEGs in COAD
Using the RTCGAToolbox package 10 in the R software environment, mRNA array gene expression data in tumor and nontumor tissues of patients with COAD in the TCGA dataset were downloaded and analyzed to identify upregulated DEGs. As shown in Figure 1, the top 50 upregulated DEGs between tumor and nontumor samples were identified.

Heat map of the top 50 upregulated differentially expressed genes from the TCGA colorectal carcinoma dataset.
Functional Enrichment of DEGs
Using the GSEA online service, the enrichment of KEGG pathways and GO biological processes in upregulated DEGs was analyzed. As shown in Figure 2, the Wnt signaling pathway, cytokine-cytokine receptor interaction, and pathways in cancer KEGG pathways were enriched in more of the top 50 upregulated DEGs. Tissue development, regulation of cell proliferation, and regulation of multicellular organismal development were the main GO biological processes among the top 50 upregulated DEGs.
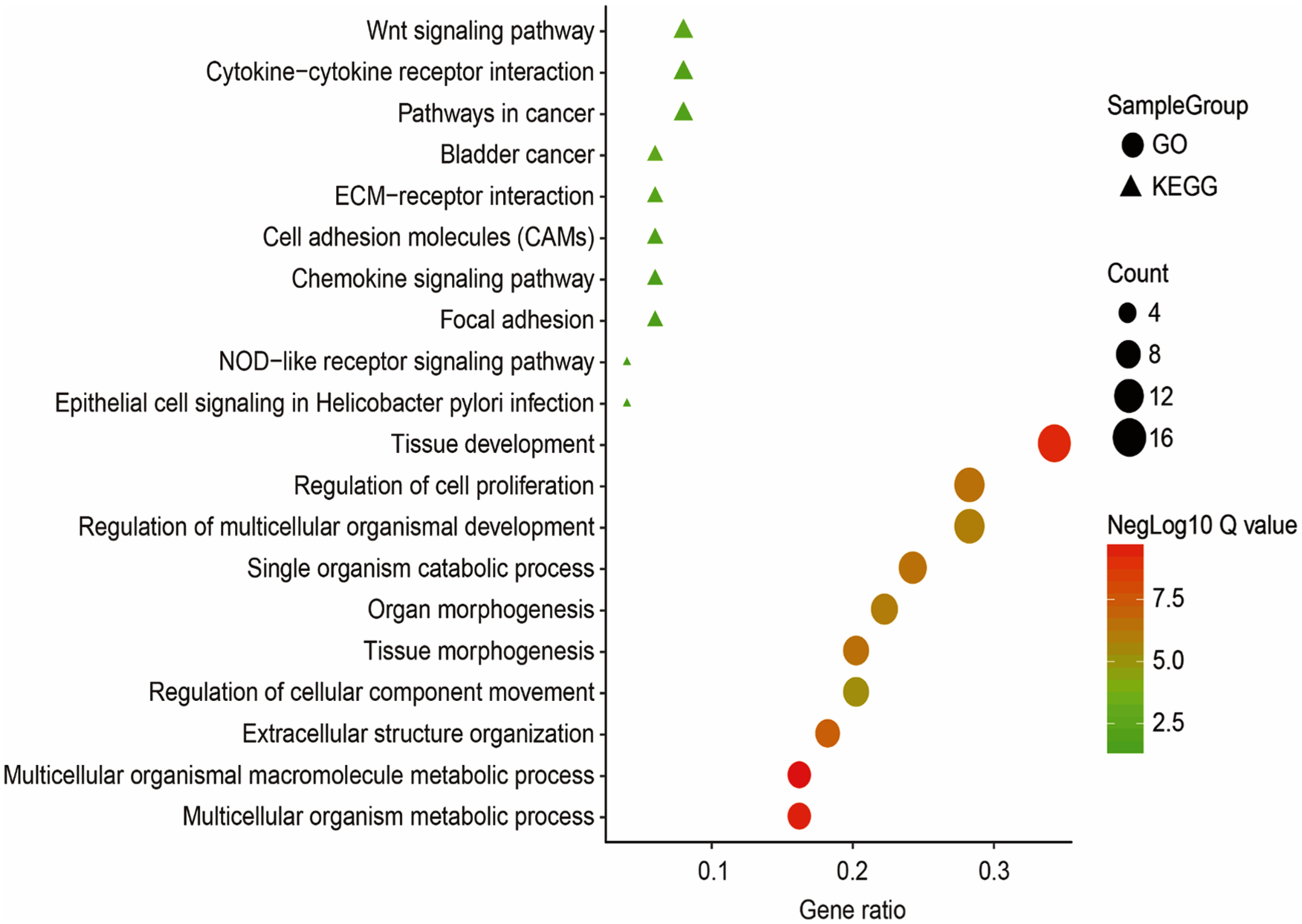
GO biological process and KEGG pathway enrichment of the top 50 upregulated DEGs from the TCGA colorectal carcinoma dataset.
Survival Analysis and Expression Comparison
Using the getSurvival function of RTCGAToolbox package in the R software environment, 10 Kaplan-Meier survival analysis was performed. Among the top 50 upregulated DEGs, COAD patients with MYC overexpression in tumor tissues had significantly worse overall survival (OS) than those with low MYC expression (P = .021; Figure 3A). And, high KLK6 was significantly associated with poor OS in COAD patients (P = .047; Figure 3B).
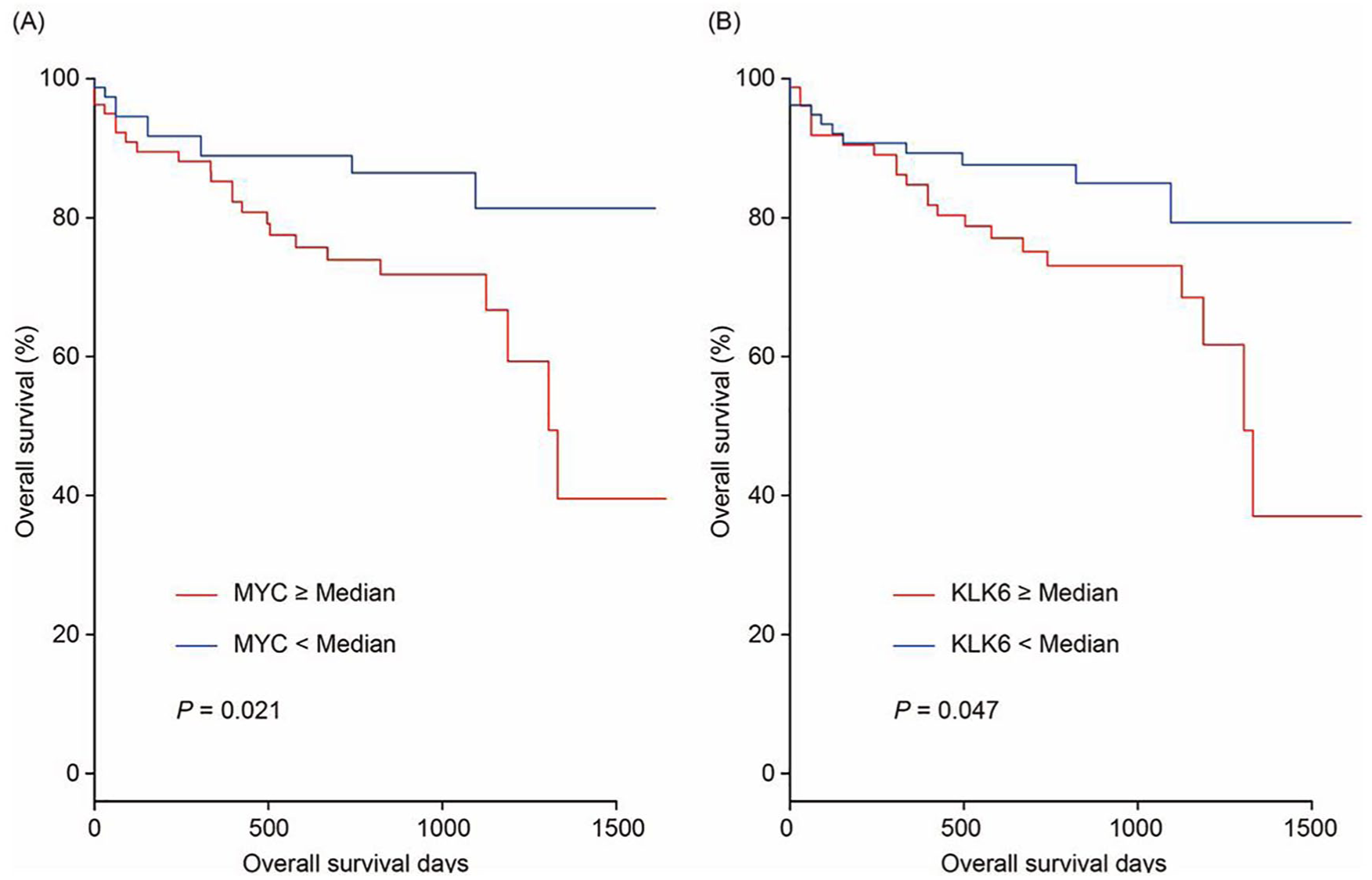
Overall survival of patients with colorectal carcinoma grouped by MYC (A) and KLK6 (B) with median cutoff.
As shown in Figure 4, MYC and KLK6 were overexpressed in tumor tissues compared with nontumor tissues (both P < .0001; Figure 4A). In GSE41328 and GSE113513 validation sets, MYC was upregulated in tumor tissues (both P < .0001; Figure 4B and C). Similarly, KLK6 was also significantly overexpressed in tumor tissues in both GSE41328 and GSE113513 validation sets (P = .0068 and P < .0001, respectively; Figure 4B and C).

MYC and KLK6 levels in nontumor and tumor tissues in the TCGA dataset (A), GSE41328 GEO profile (B), and GSE113513 GEO profile (C).
Associated Diseases and Drugs Targeting MYC and KLK6
Using the DisGeNET online service, MYC and KLK6 were both included in the top-scored disease associations of colonic neoplasms and colorectal neoplasms (Figure 5). In the DrugBank database, when MYC and KLK6 were set as drug targets, nadroparin and benzamidine were identified, respectively. Nadroparin is an approved and investigational MYC inhibitor, and the use of benzamidine for KLK6 is still in experimental stages.
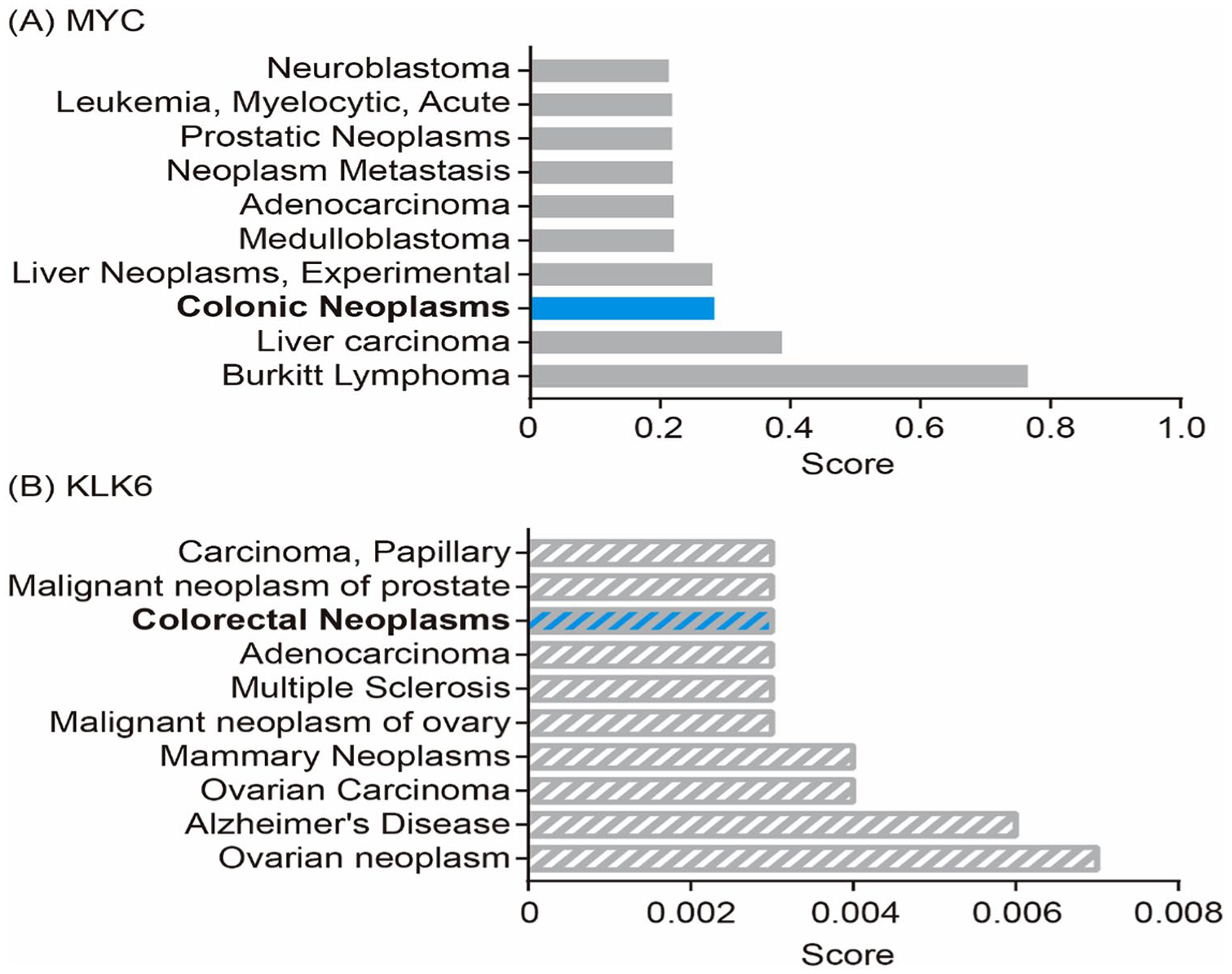
Top-scored disease associations for MYC and KLK6 in the DisGeNET database.
In the TCMSP database, 259 herbs were related to MYC, and no herbs were related to KLK6. In the TCM-MESH database, 61 herbs related to MYC and 15 herbs related to KLK6 were identified. Between the 2 databases, 8 herbs targeting MYC were found in common, including Da Huang (Radix Rhei Et Rhizome), Hu Zhang (Polygoni Cuspidati Rhizoma Et Radix), Huang Lian (Coptidis Rhizoma), Ban Xia (Arum Ternatum Thunb), Tu Fu Ling (Smilacis Glabrae Rhixoma), Lei Gong Teng (Tripterygii Radix), Er Cha (Catechu), and Guang Zao (Choerospondiatis Fructus) (Figure 6).

Identification of herbs targeting MYC and KLK6.
Functionally, Da Huang, Hu Zhang, and Huang Lian have heat-clearing and detoxifying effects, Ban Xia has phlegm-resolving and damp-drying effects, Tu Fu Ling and Lei Gong Teng have collateral-unblocking and toxin-relieving effects, and Er Cha and Guang Zao have blood-activating effects (Figure 6). Thus, eliminating pathogenic factors rather than strengthening the body’s resistance might be more effective for cancer treatment targeting MYC.
Discussion
Previous studies have revealed that overexpression of MYC, which is triggered by activation of the Wnt pathway, is considered to be a marker of metastasis and a prognostic factor for good survival.19,20 As a regulator of cell proliferation and apoptosis, 21 elevated expression of MYC mRNA and increased expression of MYC oncoprotein were reported in the majority of CRCs,22,23 induced epithelial-mesenchymal transition, 24 and significantly correlated with advanced tumor stage. 25 Furthermore, primary tumors were significantly more diffusely positive for MYC than metastatic tumors. 23 A previous report showed that MYC on its own failed to have independent prognostic significance but coexpression of nuclear β-catenin and MYC was the strongest marker of impaired prognosis. 26 Our results showed that MYC overexpression in tumors was significantly associated with COAD survival. Hence, the inhibition of MYC is a promising therapeutic strategy for treating COAD. 27 In this analysis, we identified potential drugs targeting MYC, such as nadroparin, and 8 herbs with pathogenic factor–eliminating effects that also target MYC. Nadroparin has both protective and therapeutic effects against colonic inflammation via exerting anti-oxidative and anti-inflammatory effects by modulating nuclear factor E2-related factor-2/heme oxygenase-1 (Nrf2/HO-1) and nuclear factor kappa B (NF-κB) pathways. 28 This might also be the antitumor mechanism of nadroparin in COAD.
It has been reported that emodin, a phytochemical of Da Huang and Hu Zhang, can inhibit the proliferation of the HT-29 human colon cancer cell line in vitro in a dose-dependent and time-dependent manner. 29 Moreover, emodin inhibits colon cancer cell proliferation, migration and metastasis via activating caspase-3, downregulating of CCR4 and Foxp3, and suppressing STAT3 and the AKT/mTOR signaling pathway.30,31 In addition, the Hu Zhang phytochemical resveratrol can inhibit the proliferation of HCT-15 cells and induce cell apoptosis by arresting the cells in S phase. 32 Berberine (a phytochemical of Huang Lian) can significantly inhibit the proliferation of Caco-2 cells by inducing cell cycle arrest and cell apoptosis. 33 Total alkaloids extracted from Huang Lian can inhibit tumor formation induced by 1-2 dimethylhydrazine (DMH) combined with dextran sodium sulfate solution (DSS). 34 Banxia Xiexin Decotion was found to suppress colon carcinogenesis induced by DMH/DSS by stopping colitis from developing into colonic carcinoma. 35 Celastrol, a phytochemical from Lei Gong Teng, inhibited proliferation of SW620 cells by downregulating p-AKT, NF-κB, and survivin, and activating caspase-7, caspase-3, and PARP. Additionally, celastrol induced the expression of reactive oxygen species, apoptosis, depolarization of the mitochondrial membrane, and cell cycle arrest at G2/M in SW620 cells. 36 Triptolide can induce apoptosis of HCT16 cells by inhibiting Bcl-2, increasing Bax, and promoting the activation of caspase-3 37 ; it can also induce autophagy in CT26 colon cancer cells. 38 No publications on the antitumor effects of Tu Fu Ling, Er Cha, and Guang Zao on colon cancer were found. We suggest more research on these herbs, both in compounds and as ingredients for COAD treatments.
Aberrant expression of KLK6 was reported in several human cancers, including breast, 39 pancreatic, 40 gastric, 41 and colon cancer. 42 High expression of KLK6 mRNA correlated with serosal invasion, liver metastasis, advanced Duke’s stage, and a poor prognosis for patients with CRC. 42 KLK6 mRNA was significantly upregulated in highly invasive tumors and tumors with advanced TNM stage and was shown to predict poor OS and disease-free survival in CRC. 43 Another study by Vakrakou et al also proved that KLK6 expression correlates significantly with increasing tumor stage and histological grade and is a significant factor for OS and disease-free survival in COAD. 44 Interestingly, KLK6 levels in adenomas were significantly higher than those in either the cancerous or non-cancerous tissues examined. Strong KLK6 immunostaining was seen in glandular cells and inflammatory cells of adenomas.44,45 Our results confirmed the predictive value of KLK6 for COAD outcomes in agreement with previous publications. As an inhibitor of KLK6, benzamidine showed antitumor activity in human promyelocytic leukemia cells 46 and B-lymphoid human tumor cells. 47 However, the use of benzamidine to target KLK6 is still in experimental stages, and no herbs and ingredients were identified as related to KLK6 in COAD.
Even though MYC was a promising anticancer target of traditional Chinese medicine including Ban Xia, Da Huang, and Lei Gong Teng in many human malignancies,48-50 no evidence of these drugs and herbs acting on CRC including COAD through targeting of MYC an KLK6 was identified. Our results indicated that MYC and KLK6, which are overexpressed in tumor tissues, may represent potential molecular biomarkers for unfavorable prognosis in COAD. Understanding the biological function and mechanisms of MYC and KLK6 in colorectal tissue may help delineate their roles in colorectal physiology and the pathology of CRCs. Unfortunately, MYC and KLK6 were examined at the transcription level, not at the protein level. Additionally, no experimental mechanisms of these genes were investigated. We suggest that future basic research and clinical studies should focus on the associations between these genes and COAD development and progression.
Supplemental Material
R_script_supplementary_materials – Supplemental material for Identification of Prognostic Biomarkers and Drugs Targeting Them in Colon Adenocarcinoma: A Bioinformatic Analysis
Supplemental material, R_script_supplementary_materials for Identification of Prognostic Biomarkers and Drugs Targeting Them in Colon Adenocarcinoma: A Bioinformatic Analysis by Shu Dong, Zhimin Ding, Hao Zhang and Qiwen Chen in Integrative Cancer Therapies
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Natural Science Foundation of China (81503524/H2708) and Foundation from Shanghai Science and Technology Committee (14401932400).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
