Abstract
Background/Rationale:
Experimental studies support the role of neurotransmitter genes in dementia risk, but human studies utilizing single variants in candidate genes have had limited success.
Methods:
We used the gene-based testing program Versatile Gene-based Association Study to assess whether aggregate variation across 6 neurotransmitter pathways influences risk of cognitive decline in 8159 cognitively normal elderly (≥65 years old) adults from 3 community-based cohorts.
Results:
Common genetic variation in
Conclusion:
Genetic variation analyses and gene expression data suggest that
Introduction
The prevalence of dementia will rise as the world’s population ages; however, the risk factors and predictors of cognitive decline in cognitively normal older individuals are not fully understood. Dementia represents the clinical manifestation of a spectrum of neurodegenerative disorders that are likely caused by multiple factors. Despite variable underlying neuropathological etiologies, many forms of dementia are characterized by progressive cognitive decline, and early changes in cognitive function are often required for diagnosis under neurodegenerative conditions such as Alzheimer’s disease (AD). 1 and frontotemporal dementia. 2 Twin and sibling studies suggest that cognitive phenotypes such as working memory, episodic memory, and risk of dementia are heritable. 3 –7 Although recent research has broadened our understanding of the genetic underpinnings of cognitive decline and dementia, much remains to be understood. 8,9
Evidence from studies in humans and rodents suggests that neurotransmitter function declines in aging and that these changes could be associated with risk of dementia and other disorders.
10
–14
Current knowledge of genetic variation within neurotransmitter genes as risk factors for dementia is heavily derived from single-variant candidate gene studies and has often rendered mixed results. For example, common variants in
Factors that impact global cognition may provide unique insight into the early pathological changes that underlie declines in cognitive function and mechanisms that may be helpful for combating it. Given their key roles in relaying synaptic signals throughout different networks of the brain critical for cognition, neurotransmitters are clear candidates for influencing cognitive aging. In addition to suggestive human genetic findings, research in animal models supports the role of components within neurotransmitter pathways as contributing to cognitive function in aging. For example, given the role depression may play as a risk factor for dementia, 20 many studies have examined candidate genes related to neurotransmitter or neuroendocrine pathways for roles in cognition and risk of neurodegenerative disorders leading to dementia such as AD. Further, the serotonin transporter length polymorphism (5-HTTLPR) has been associated with poorer memory function in older adults 21 and with risk of dementia. 22 Norepinephrine (adrenergic system) signaling is associated with better performance on memory tasks and is suggested to play a protective role against AD pathology, 23 –25 and extensive loss of norepinephrine projecting cells is often seen in AD. 23 Anticholinergic medications have been linked to cognitive decline. 26 One review suggests the excitatory glutamatergic system as another strong candidate system for risk of late-life depression and dementia that may relate to changes in signal transduction in older age. 27
Taken together, these findings suggest that multiple neurotransmitter pathways may contribute to risk for dementia, and that by investigating genes rather than single polymorphisms the authors may better understand the role these functional systems play in cognition during aging. Finally, there are many approved drugs that modulate neurotransmitter levels and are tolerated in dementia,
28
which makes treatment based on a gene-based analysis more tractable. For example, the drug memantine is already used to treat AD and is an
Methods and Materials
Cohort Descriptions and Cognitive Measures
Three cohorts comprising a total of 8159 participants with cognitive and genetic data were used in our study. In all 3 cohorts, we required that individuals had genome-wide SNP genotypes, ≥2 cognitive test scores available, and were of European ancestry and unrelated based on SNP data. All participants in all studies provided written informed consent, and the institutional review boards at recruitment sites and the study-coordinating center (University of California, San Francisco) approved all aspects of this study.
Discovery Cohort—Health, Aging, and Body Composition
Our discovery cohort was the Health, Aging, and Body Composition (Health ABC) study, a prospective cohort study of 3075 community dwelling black and white men and women living in Memphis, Tennessee or Pittsburgh, Pennsylvania, and aged 70 to 79 years at recruitment in 1997. Eligible participants reported no difficulty with daily living activities, walking a quarter of a mile, or climbing 10 steps without resting at baseline. All participants were free from life-threatening cancer diagnoses and planned to live in the study area for at least 3 years. The cohort has been described in more detail in previous publications. 30
In the Health ABC cohort, we used 2 different measures of cognitive function: the Modified Mini-Mental State Examination (3MS), a measure of global cognition, 31 and the Digit Symbol Substitution Test (DSST), a measure of processing speed. 32 In Health ABC, the 3MS was administered in years 1, 3, 5, and 8 and the DSST was administered in years 1, 5, and 8. For phenotype generation, 3MS and DSST repeated measures from across the research period were used to calculate participant-specific slopes and intercepts for each test. As in prior work, we generated cognitive test score slopes as best linear unbiased predictions using a linear mixed model with random slopes and intercepts (Stata10, StataCorp, College Station, Texas). 33 This type of modeling enables estimation of population-level fixed effects (changes in cognitive function across the entire cohort) as well as individual-level random effects (each participant’s deviation from the overall group). This approach allows each participant to have a unique intercept and trajectory (slope describing a change in cognitive function). Estimated slopes were used as a quantitative trait in association analysis with genetic variants as described below. Covariate adjustments were introduced at the genetic analysis stage, as detailed below.
Replication Cohorts—Study of Osteoporotic Fractures and Osteoporotic Fractures in Men
Our replication cohorts were the Study of Osteoporotic Fractures (SOF) and Osteoporotic Fractures in Men (MrOS) study. The SOF is a multicenter observational study of 9704 women aged 65 or older started in 1986 to study risk factors for osteoporosis, falls, and fractures. It was expanded to include measures of successful aging and has been previously described in detail. 34 The MrOS study is a multicenter observational study of 5994 older men started in 2000 to study risk factors for osteoporosis, fractures, and prostate cancer susceptibility, which included multiple measures of successful aging and cognitive health. It has previously been described in detail. 35
In accordance with study guidelines, cognitive phenotype generation for SOF and MrOS was conducted by the San Francisco Coordinating Center (SFCC), a nonprofit academic organization that manages the clinical data for multisite studies and clinical trials. For the SOF cohort, the Mini-Mental State Examination (MMSE), 36 an assessment similar to the 3MS, was used as a measure of general cognitive function. The MMSE was administered in years 0, 6, 8, and 10. In the MrOS cohort, the 3MS was used as a measure of general cognitive function. The 3MS was administered in years 0, 4, and 7. For SOF and MrOS data, standardized measures of cognitive function (z scores) were separately calculated in each cohort. Participant-specific slopes were calculated by the SFCC using linear mixed effect regression models (PROC MIXED in SAS, SAS Institute, Cary, North Carolina). 37 In the SOF and MrOS cohorts, fixed effects of site, age, and education level were included in the cognitive score models. As in the Health ABC analyses, random effects for slope and intercept were included. Cognitive decline was scored as a dichotomous trait as the phenotype of interest, defining decline as having a slope >1 standard deviation (SD) from the group mean slope over the assessment period included in the present study.
Genotyping and Quality Control
Genotyping for Health ABC was performed on the Illumina Human1M-Duo BeadChip system by the Center for Inherited Disease Research.
33
Genotyping for SOF and MrOS was conducted on the Illumina Omni1-Quad array at the Broad Institute.
37
For individuals of European American genetic background based on principal components analysis (PCA), genotype imputation was performed using MACH/minimac to increase the number of SNPs covered in each cohort using CEU HapMap Phase 2 release 22 as reference.
38
The SNPs with a minor allele frequency ≥0.01, call rate >95%, and Hardy-Weinberg equilibrium
Gene Selection
Neurotransmitter genes were first identified using pathways identified by the
Gene Expression
To assess the expression patterns of our most promising candidate genes, we analyzed gene expression levels in the Braineac database (http://www.braineac.org/). The database consists of 134 neuropathologically confirmed normal donors from the MRC Sudden Death Brain Bank in Edinburgh, United Kingdom, and the Sun Health Research Institute in Sun City West, Arizona. The cohort is composed of 99 males and 35 females with an average age of 58 (range 16-102). The average postmortem interval (PMI) was 41.7 hours. For each individual in the study, up to 10 brain regions were sampled: cerebellar cortex, frontal cortex, hippocampus, medulla, occipital cortex, putamen, substantia nigra, thalamus, temporal cortex, and intralobular white matter. RNA extraction, quality control steps, and analysis of array data are described elsewhere in detail. 40 In brief, RNA expression was assessed using the Affymetrix Exon 1.0 ST Array following manufacturers’ protocols. Differences in transcript expression across the 10 brain regions were assessed using the default settings in Braineac, which utilizes the maximum number of samples per region as possible based on data availability. To minimize the effects of outlier probes within a gene, Braineac utilizes a Winsorized mean across the available probe sets.
To explore and describe the temporal and spatial patterns of gene expression among our most promising candidate genes, we used the Human Brain Transcriptome project (HBT; http://hbatlas.org/pages/hbtd). The HBT provides transcriptome data for the developing and adult human brain. The study used over 1340 tissue samples from 57 clinically unremarkable individuals with no large-scale genetic abnormalities. The cohort is composed of 31 males and 26 females ranging in age from 5.7 weeks postconception (pcw) to 82 years old. The average PMI was 12.1 hours. Tissue was taken from 6 major areas. We examined the hippocampus, amygdala, striatum, cerebellar cortex, mediodorsal nucleus of the thalamus, and neocortex. Cohort characteristics, RNA extraction, quality control steps, and analysis of array data are described in detail in Kang et al. 41 The developers of the HBT project also used the Affymetrix Exon 1.0 ST Array to measure RNA expression.
Finally, cell type–specific expression data were obtained for selected genes using the Brain RNA-Seq tool (http://www.brainrnaseq.org/). All samples came from elective procedures and all postnatal tissue were deemed normal by both electroencephalography and magnetic resonance imaging. We examined all available cell types including fetal astrocytes, mature astrocytes, neurons, oligodendrocytes, microglia/macrophages, and endothelial cells. Detailed descriptions of the samples and processing techniques used have been previously published. 42 –44
Statistical Analysis
We performed a 2-step analysis. First, univariate genetic associations with cognitive slope (Health ABC) or cognitive decline (SOF and MrOS) were determined for SNPs in candidate genes in each cohort separately. Next, we grouped SNPs within gene regions of interest as defined by genes’ membership in specific neurotransmitter pathways in order to evaluate gene-level contributions to cognition. The 3 cohorts used for this study utilized 2 separate cognitive tests and were mixed with respect to sex composition and study initiation date, so they were not combined for analysis. Qualitative comparisons were made between the cohorts when possible.
Single-Variant Associations With Cognitive Measures
We implemented a regression model in R to test for genetic associations between SNPs and the cognitive phenotype of interest in each cohort. In the Health ABC study, we assessed SNP associations with slope of scores on 3MS or DSST separately using linear regression. Both analyses were adjusted for participants’ baseline age, sex, education level,
Gene-Based Testing
The Versatile Gene-based Association Study (VEGAS) algorithm has been previously described.
46
Briefly, the VEGAS uses the University of California, Santa Cruz genome browser hg18 assembly to assign user supplied SNPs to genes. Gene boundaries were defined as ±50 kb from the 5′ and 3′ end of the untranslated regions, and all SNPs within these boundaries that were analyzed for associations from each data set were included for analysis. The software then takes the full set of markers within a gene and accounts for confounding due to reference population linkage disequilibrium (LD) patterns and gene size. The VEGAS corrects for these confounders using simulations that utilize HapMap LD structures.
46
The VEGAS provides a gene-based test statistic as the sum of all χ2-converted SNP
Results
Cohort Composition
The final Health ABC cohort was comprised of 1598 cognitively normal older individuals who had either the 3MS or DSST available for analysis. There were 1597 individuals with a longitudinal 3MS profile and 1594 individuals with a longitudinal DSST profile. In the SOF cohort, 383 of 3508 women were labeled as showing cognitive decline over the test period. In the MrOS cohort, 384 of 3820 men were labeled as showing cognitive decline over the test period. The average age of participants in the 3 cohorts was in the early to mid-70s (Tables 1 and 2). The Health ABC cohort and MrOS cohorts were highly educated, with a majority of individuals reporting greater than 12 years of education, whereas a majority of the SOF cohort participants reported less than 12 years of education (Tables 1 and 2). Baseline scores on cognitive tests differed between individuals by cognitive trajectory; individuals with a negative slope in the Health ABC cohort or identified as cognitive decliners in the SOF or MrOS cohorts generally did worse compared to those with a positive slope or identified as nondecliners in the SOF or MrOS analyses (Tables 1 and 2).
Primary Health ABC Cohort.
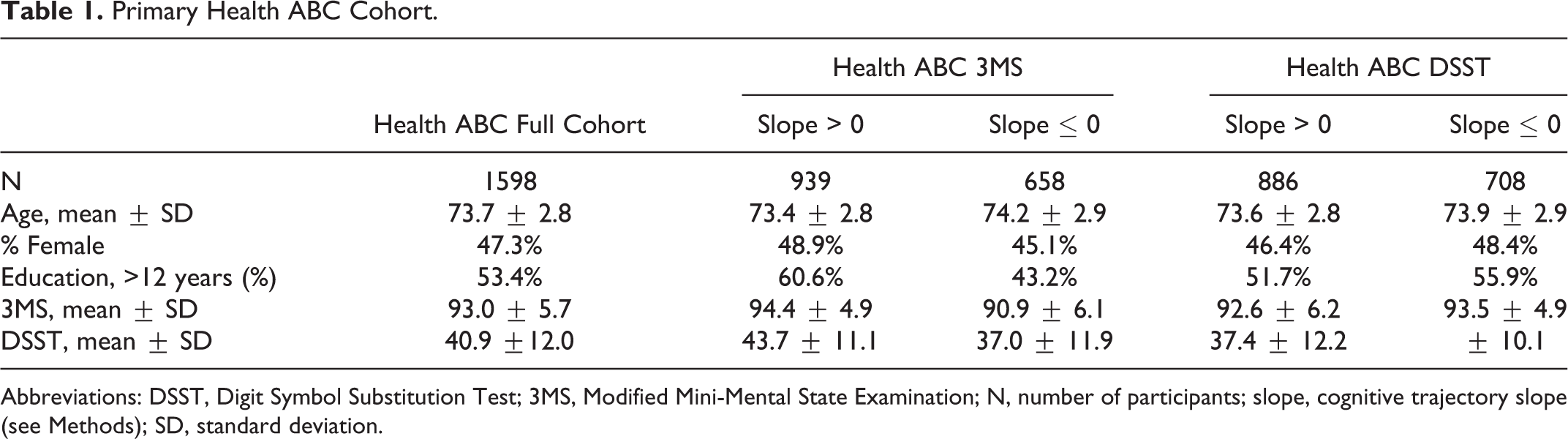
Abbreviations: DSST, Digit Symbol Substitution Test; 3MS, Modified Mini-Mental State Examination; N, number of participants; slope, cognitive trajectory slope (see Methods); SD, standard deviation.
Replication SOF and MrOS Cohorts.
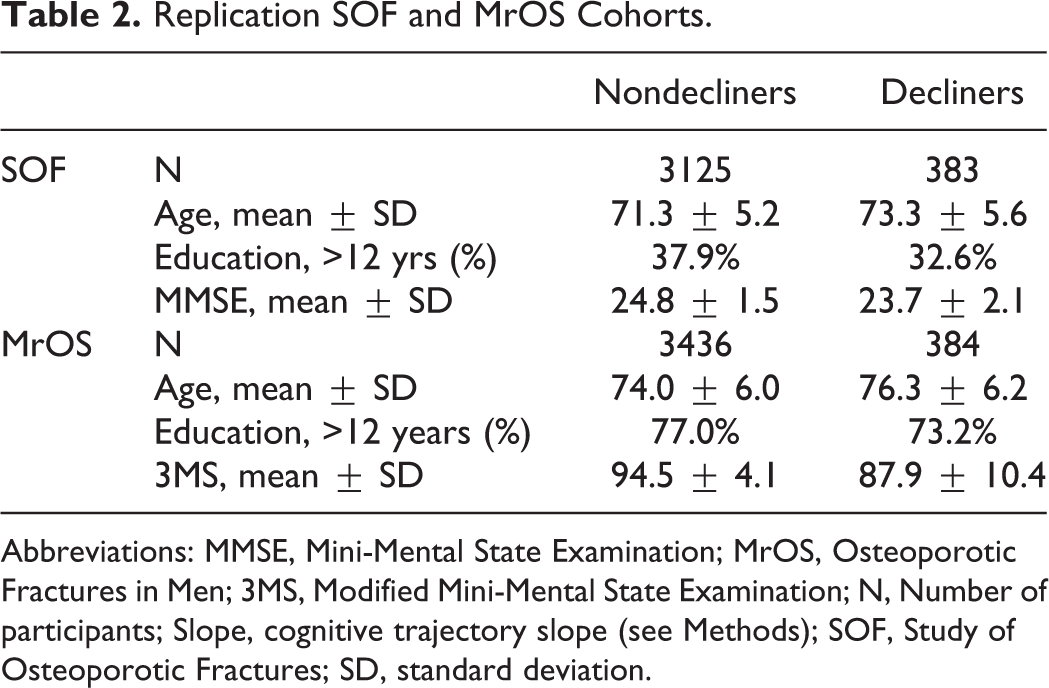
Abbreviations: MMSE, Mini-Mental State Examination; MrOS, Osteoporotic Fractures in Men; 3MS, Modified Mini-Mental State Examination; N, Number of participants; Slope, cognitive trajectory slope (see Methods); SOF, Study of Osteoporotic Fractures; SD, standard deviation.
Gene Pathway Identification
Using the KEGG and SSDCS databases, we identified a total of 330 genes associated with adrenergic, cholinergic, dopaminergic, GABAergic, glutamatergic, and serotonergic neurotransmitter pathways. A full list of these genes is available in Supplemental Data 1.
Gene-Based Analyses
VEGAS results were available for 315 of the 330 neurotransmitter genes identified through the KEGG and SSDCS databases due to SNP coverage and other factors (sex chromosome genes are not analyzable in version of VEGAS used). A graphical summary of the major findings and replicated genes from the Health ABC analyses as well as the SOF and MrOS cohorts is shown in Figure 1.
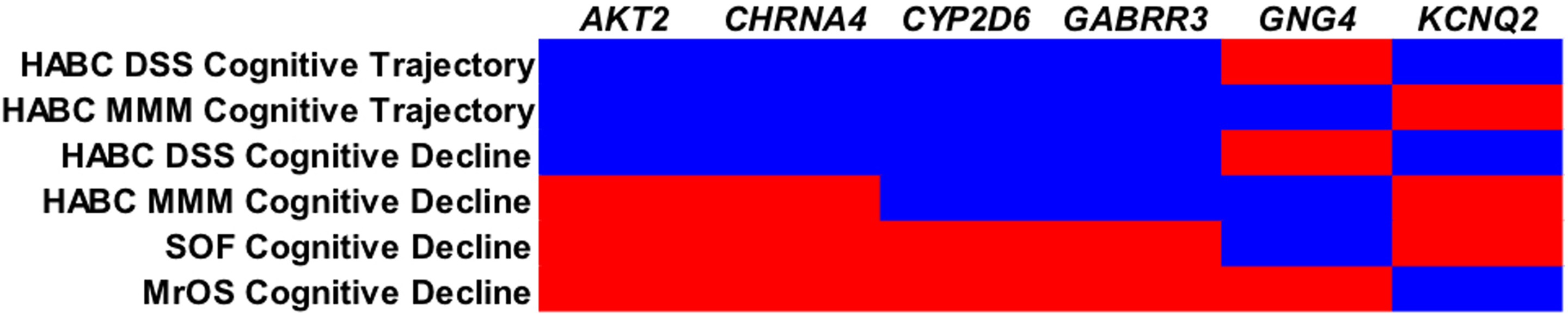
Results from HABC, SOF, and MrOS analyses major findings replicated between the Health ABC, DSST, HABC, 3MS, SOF, and MrOS cohorts are shown. Blue areas were significant and red areas were not significant. DSST, Digit Symbol Substitution Test; HABC, Health ABC; MrOS, Osteoporotic Fractures in Men; 3MS, Modified Mini-Mental State Examination; SOF, Study of Osteoporotic Fractures.
The Health ABC DSST cognitive trajectory analysis resulted in 20 genes that were significant at a
Health ABC Cognitive Trajectory Analyses.
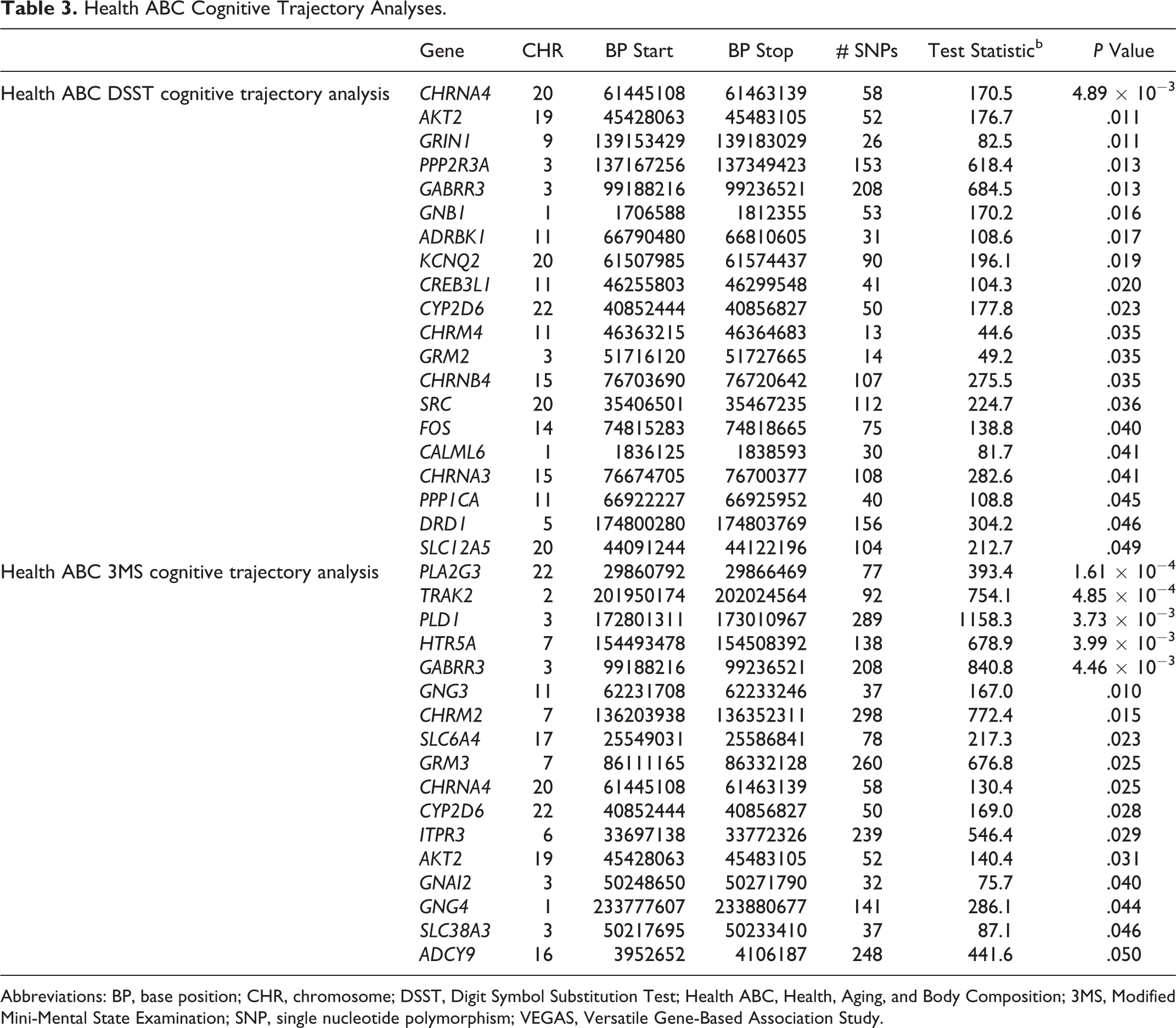
Abbreviations: BP, base position; CHR, chromosome; DSST, Digit Symbol Substitution Test; Health ABC, Health, Aging, and Body Composition; 3MS, Modified Mini-Mental State Examination; SNP, single nucleotide polymorphism; VEGAS, Versatile Gene-Based Association Study.
To test for genes that specifically influence cognitive decline, we restricted our gene-based VEGAS analyses to the Health ABC SNPs with negative β coefficients in the regression model. The Health ABC DSST cognitive decline model resulted in 18 genes that were significant. In the Health ABC 3MS cognitive decline model, 15 of the genes tested were significant at a
Health ABC Cognitive Decline Analyses.
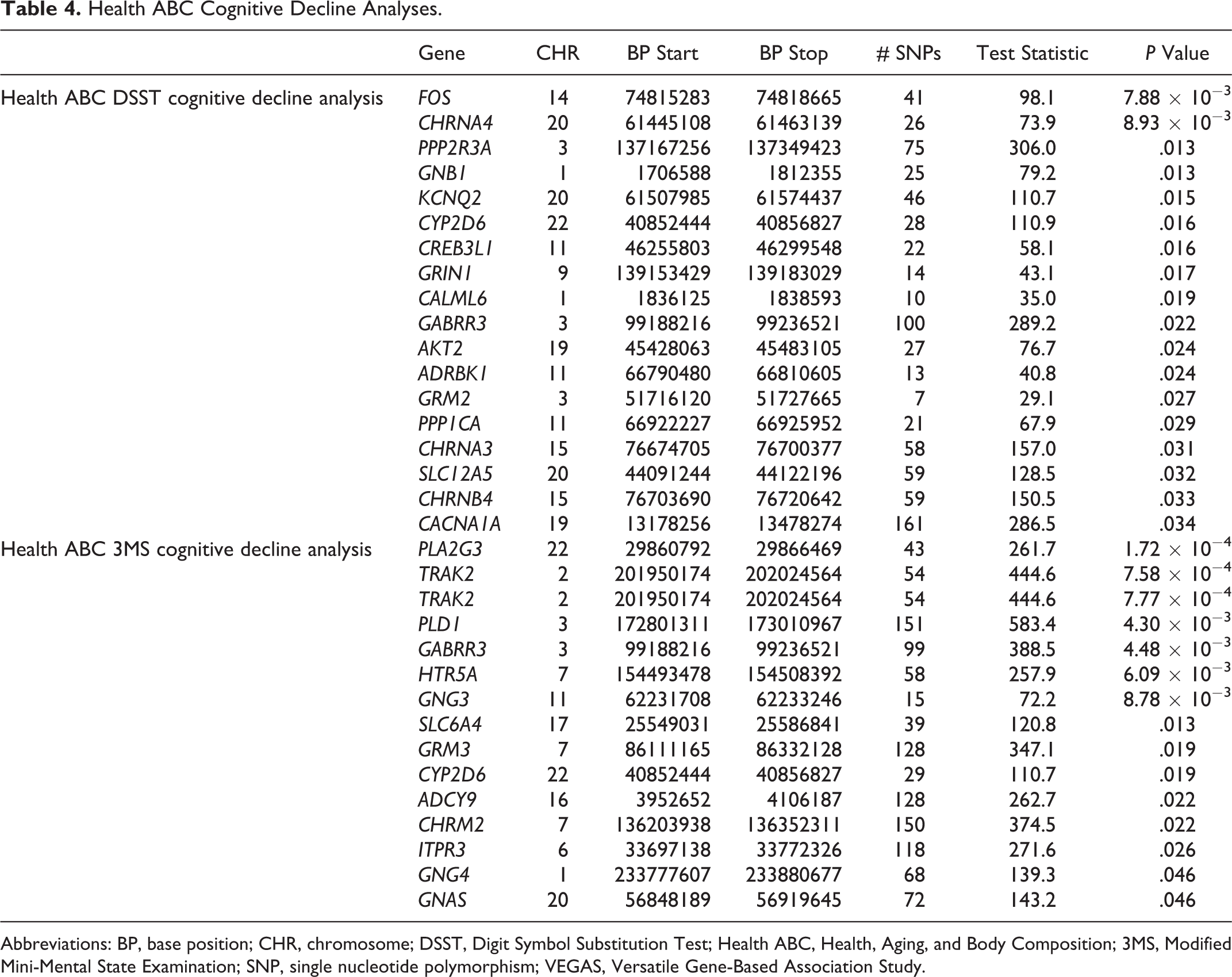
Abbreviations: BP, base position; CHR, chromosome; DSST, Digit Symbol Substitution Test; Health ABC, Health, Aging, and Body Composition; 3MS, Modified Mini-Mental State Examination; SNP, single nucleotide polymorphism; VEGAS, Versatile Gene-Based Association Study.
We used the SOF and MrOS cohorts to replicate our findings from the Health ABC analyses. In both cohorts, 13 genes were significant.
SOF and MrOS Cognitive Decline Analyses.
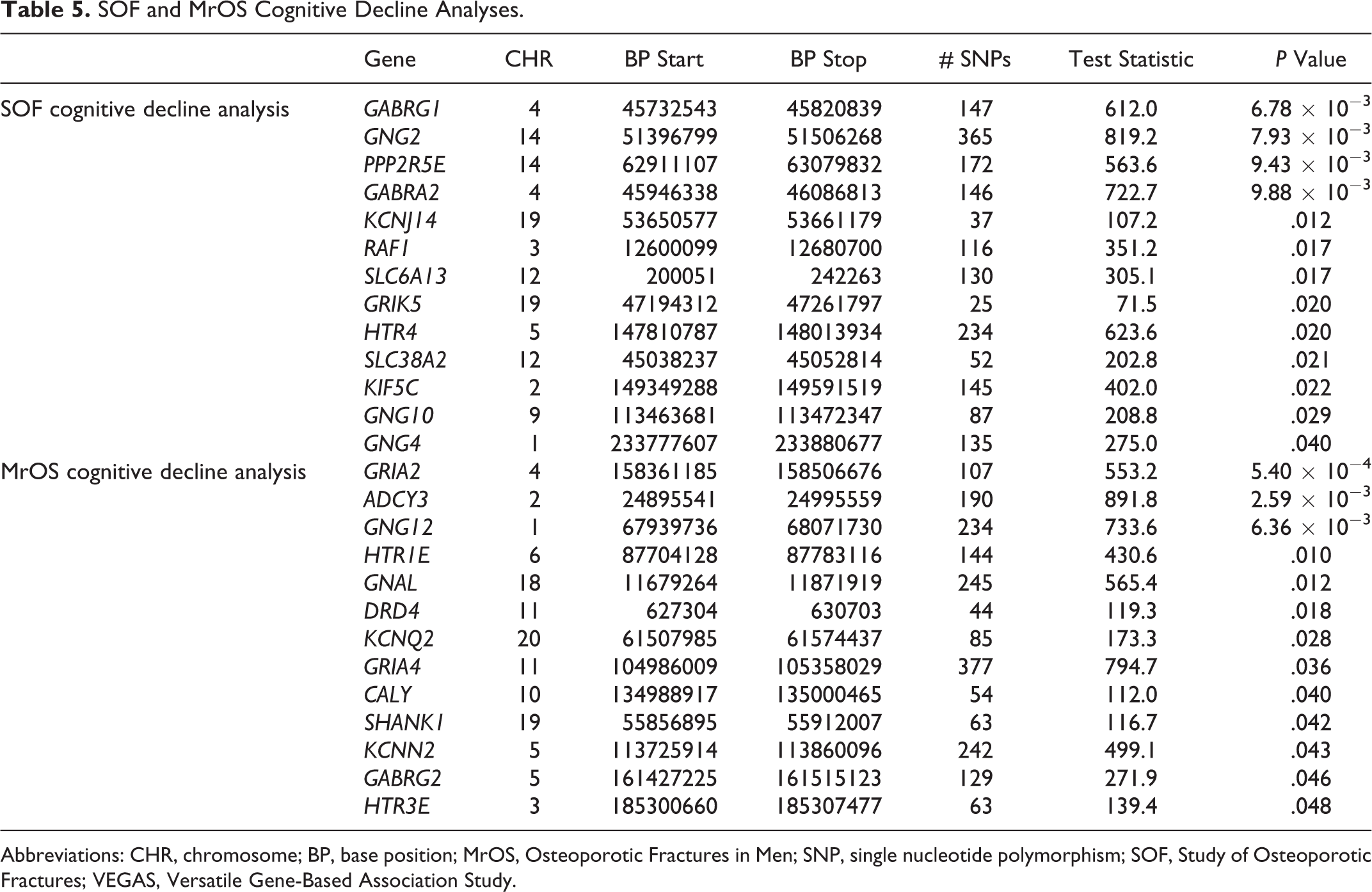
Abbreviations: CHR, chromosome; BP, base position; MrOS, Osteoporotic Fractures in Men; SNP, single nucleotide polymorphism; SOF, Study of Osteoporotic Fractures; VEGAS, Versatile Gene-Based Association Study.
Gene Expression Analyses
Gene expression analyses were conducted for
We next evaluated the extent to which
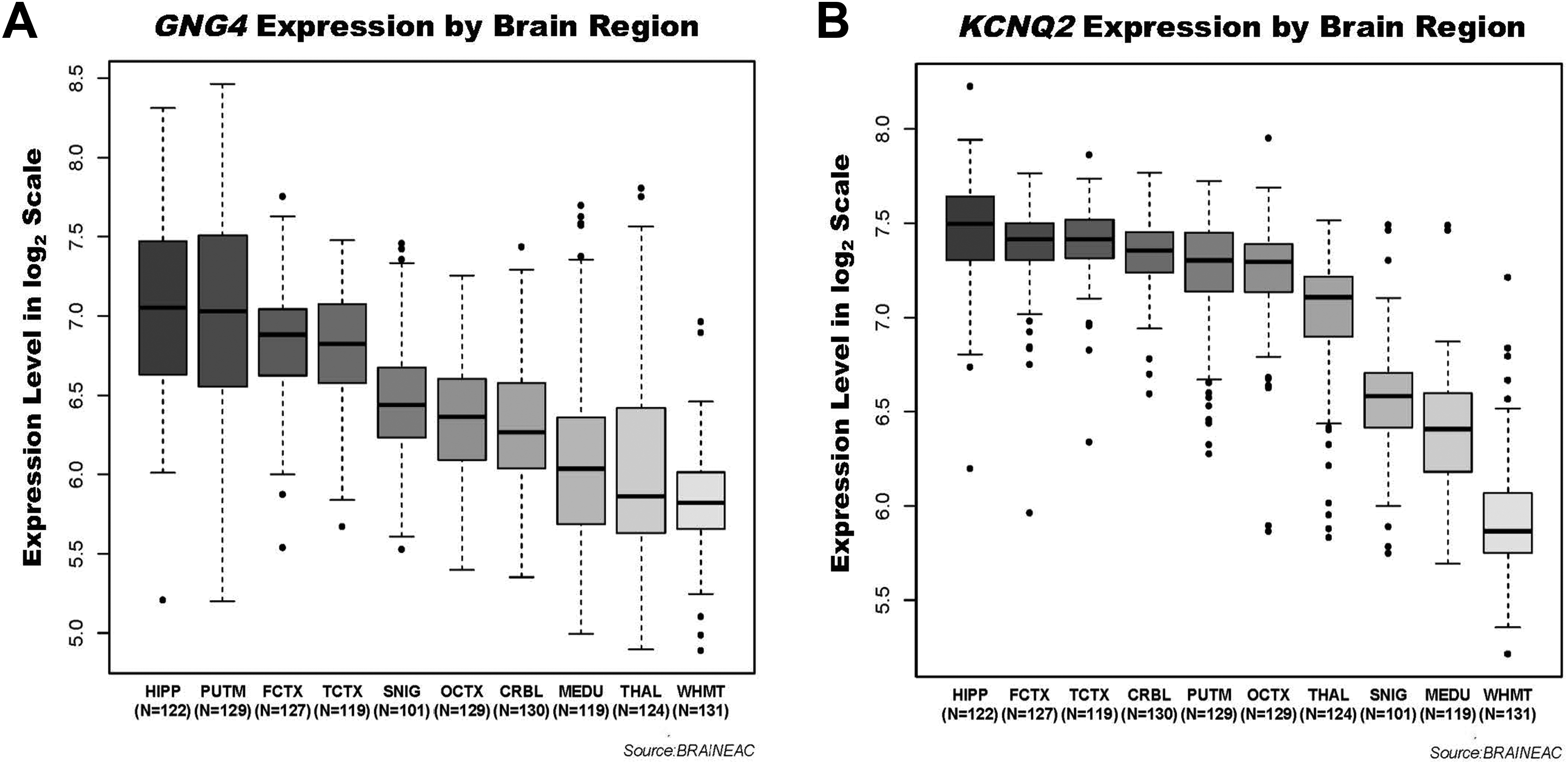
To explore and qualitatively assess the temporal expression patterns of
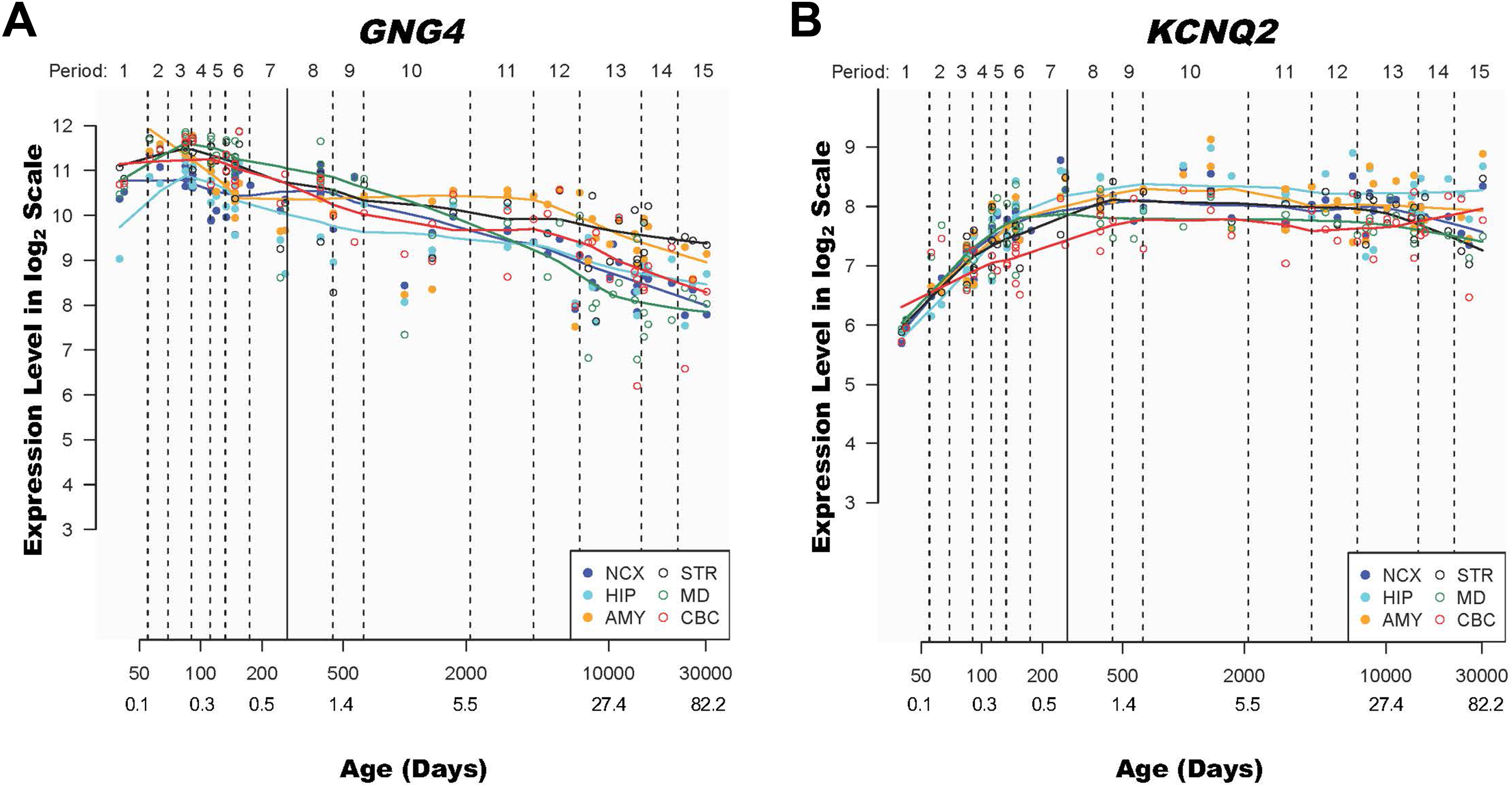
Discussion
Our findings implicate neurotransmitter pathway genes
Of note, both genes showed highest expression in fetal astrocytes. Although speculative, these findings could potentially suggest that genetic variation in
We specifically tested the role of neurotransmitter pathway genes in cognitive decline because their protein products are more accessible as therapeutic targets than many other proteins implicated in cognitive decline. As candidate genes,
This study represents an important step forward in the utilization of gene-based testing to identify neurotransmitter pathway gene candidates that may predispose healthy individuals to cognitive decline, which may ultimately put them at greater risk of (or be an early indicator of) dementia. Our study benefits from its use of 3 well-established and characterized community-based cohorts. As community-based studies, Health ABC, SOF, and MrOS studies are designed to be more representative of the general population. Our findings are supported by gene expression evidence from pathologically confirmed normal individuals. However, our study cannot confirm causality or infer directionality in the relationship between
In this study, we used gene-based testing to identify an association between
Supplemental Material
Supplemental_Figure_1 - Neurotransmitter Pathway Genes in Cognitive Decline During Aging: Evidence for GNG4 and KCNQ2 Genes
Supplemental_Figure_1 for Neurotransmitter Pathway Genes in Cognitive Decline During Aging: Evidence for
Supplemental Material
Supplementary_Data_1 - Neurotransmitter Pathway Genes in Cognitive Decline During Aging: Evidence for GNG4 and KCNQ2 Genes
Supplementary_Data_1 for Neurotransmitter Pathway Genes in Cognitive Decline During Aging: Evidence for
Footnotes
Acknowledgments
The authors thank the Heath, Aging, and Body Composition (Health ABC), SOF, and MrOS research participants for their long-standing dedication. The SOF is supported by the National Institutes of Health (NIH) funding. The National Institute on Aging (NIA) provides support under the following grant numbers: R01 AG005407, R01 AR35582, R01 AR35583, R01 AR35584, R01 AG005394, R01 AG027574, and R01 AG027576. The MrOS Study is supported by the NIH funding. The following institutes provided support: the NIA, the National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS), the National Center for Advancing Translational Sciences, and NIH Roadmap for Medical Research under the following grant numbers: U01 AG027810, U01 AG042124, U01 AG042139, U01 AG042140, U01 AG042143, U01 AG042145, U01 AG042168, U01 AR066160, and UL1 TR000128. The NIAMS provides funding for the MrOS ancillary study “GWAS in MrOS and SOF” under the grant number RC2 AR058973. The NIAMS provides funding for the MrOS ancillary study “replication of candidate gene associations and bone strength phenotype in MrOS” under the grant number R01 AR051124. The Health ABC study was supported by NIA contracts N01AG62101, N01AG62103, and N01AG62106. The genome-wide association study was funded by NIA grant 1R01AG032098-01A1 to Wake Forest University Health Sciences and genotyping services were provided by the Center for Inherited Disease Research (CIDR). The CIDR is fully funded through a federal contract from the NIH to The Johns Hopkins University, contract number HHSN268200782096C.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Primary support for data analyses was provided by NIA K24 AG031155 (KY), NIA K01 AG049152 (JSY), the Larry L. Hillblom Foundation 2012-A-015-FEL and 2016-A-005-SUP (JSY), an AFTD Susan Marcus Memorial Fund Clinical Research Grant (JSY), and the Tau Consortium (JSY).
Supplemental Material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
