Abstract
Background:
Long noncoding RNAs have been shown to play crucial roles in cancer biology, while the long noncoding RNA landscapes of pancreatic ductal adenocarcinoma have not been completely characterized. We aimed to determine whether long noncoding RNA could serve as early diagnostic biomarkers for pancreatic ductal adenocarcinoma.
Method:
We conducted a genome-wide microarray analysis on pancreatic ductal adenocarcinoma and their adjacent noncancerous tissues from 8 Chinese patients.
Results:
A total of 3352 significantly differentially expressed long noncoding RNAs were detected. Of total, 1249 long noncoding RNAs were upregulated and 2103 were downregulated (fold change ≥2,
Conclusion:
Our study provided a genome-wide survey of dysregulated long noncoding RNAs and long noncoding RNA/messenger RNA co-regulation networks in pancreatic ductal adenocarcinoma tissue. These dysregulated long noncoding RNA/messenger RNA networks could be used as biomarkers to provide early diagnosis of pancreatic ductal adenocarcinoma or its subtype, predict prognosis, and evaluate treatment efficacy.
Keywords
Introduction
Pancreatic ductal adenocarcinoma (PDAC), an aggressive and fatal malignancy, is one of the most common malignancies and ranks #4 leading causes of carcinoma-related death. Because it is very difficult to diagnose, especially at their early stage, the morbidity and mortality of PDAC remain high. Most patients have already presented with serious local invasion and/or distant metastasis when PDAC is first diagnosed and thereby missing the optimal timing for curative surgical resection. 1,2 The overall survival rate was less than 5%. 3,4
Long noncoding RNAs (lncRNAs), a subset of noncoding RNA transcripts that are longer than 200 nucleotides in length, 5 are extensively distributed in the genome. 6 They have been suggested to participate in various biological processes, such as epigenetic regulation, chromosome imprinting, cell-cycle control, transcription, translation, splicing, and cell differentiation, 7 -10 and therefore show clinical significance. 11 Dysregulation of lncRNAs contribute to a variety of diseases by altering the expression of lncRNA target genes. 12 -18 For instance, HOTAIR, an oncogenic lncRNA, was associated with the pathogenesis and progression of breast cancer, 19 colorectal cancer, 20 and pancreatic cancer. 21 With the development of transcriptome-sequencing technologies, a considerable amount of novel human lncRNAs have been identified and annotated. However, the molecular mechanisms and biological functions of the vast majority of these annotated lncRNAs remain unknown.
With the development of next-generation sequencing, genome-wide transcriptome profiling has become an effective approach to detect novel lncRNAs in various diseases status. Dysregulation of lncRNAs has been observed in several human cancers, such as osteosarcoma,
22,23
hepatocellular carcinoma,
24
gastric cancer,
25
breast cancer,
26
and endometrial cancer.
27
Recent studies have further revealed that lncRNAs could potentially serve as diagnostic and prognostic biomarkers. For example, a set of 6 lncRNAs has been shown to be independent prognostic factors for glioblastoma after adjust for other clinical variables.
28
Seventeen differentially expressed (DE) lncRNAs, named “SubSigLnc-17”can not only discriminate 2 subtypes of diffused large B-cell lymphoma (DLBCL)—germinal center B-cell-like subtype and activated B-cell-like subtype—but also predict the prognosis of DLBCL.
29
Importantly, lncRNAs have been suggested to participate in pancreatic cancer development and progression by promoting cell growth, migration, invasion, and epithelial–mesenchymal transition.
21,30
In a prospective study, Zhou
Materials and Methods
Patient Recruitment
Eight PDAC patients who did not receive any chemotherapy or other forms of therapy were recruited from Huashan Hospital, Fudan University. All participants provided written informed consent prior to enrollment. All human patient-related protocols were approved by medical ethics committee of Huashan Hospital affiliated to Fudan University. The PDAC tissue and their adjacent noncancerous tissue were obtained surgically. Totally, 16 samples (2 samples/patient) were immediately frozen down in liquid nitrogen and stored in −80°C freezer. Surgically removed tissues were pathologically confirmed with more than 80% viable tumor cells, and clinical data were obtained retrospectively from electronic clinical records.
RNA Extraction
Total RNAs were extracted from 16 samples above using RNeasy Mini Kit (Qiagen, Hilden, Germany) following the manufacturer’s manual. The quantities and integrity were tested by using NanoDrop ND-1000 spectrophotometer (Thermo Fisher Scientific, Waltham, Massachusetts) and standard denaturing agarose gel electrophoresis.
Microarray and Data Analyses
We utilized Human 46180K lncRNA arrays manufactured by Agilent Technologies (Santa Clara, California) and Sureprint G3 Human lncRNA Chip (ie, BT1000312) for lncRNA and mRNA microarray analysis. These 2 chips have been reported to represent more than 46 506 lncRNAs and 30 656 mRNAs from NCBI RefSeq, UCSC, RNAdb, and newly annotated lncRNAs in the human genome. Each transcript was represented by up to 5 probes to improve statistical confidence. Differentially expressed genes were defined as fold change ≥2,
Total RNA (200 ng) from each sample was reversely transcribed into complementary DNA (cDNA) using an RNA Spike In Kit with one color (Agilent Technologies) in the presence of 0.8 mL of random primer mix and 2 mL of Spike mix. These cDNA samples were then cleaned and labeled in accordance with the one color Agilent Gene Expression Analysis protocol using Low Input Quick-Amp Labeling Kit (Agilent Technologies). These labeled cDNA samples were used as probes to hybridize to microarrays for 17 hours at 65°C using an Agilent Gene Expression Hybridization Kit in hybridization chamber gasket slides (Agilent Technologies).
Gene Function Analysis
We used Database for Annotation, Visualization, and Integrated Discovery (http://david.abcc.ncifcrf.gov/) to perform Gene Ontology (GO) analysis.
15
We further applied the Kyoto Encyclopedia of Genes and Genomes (KEGG) database (http://www.genome.ad.jp/kegg/) and BioCarta (http://www.biocarta.com) to analyze the potential functions of these target genes in the pathways.16,17. The lower the
Long noncoding RNA/mRNA Co-Expression Network
To construct lncRNA/mRNA co-expression network, we calculated the Pearson correlation coefficient and
Real-Time PCR
Real-time polymerase chain reaction (PCR) was performed using the ABI Prism 7300 sequence detection system (Applied Biosystems, Foster City, California). The reaction mixture (20 µL) contained 10 ng cDNA template, sense and antisense primers (200 nM each), and 10 µL 2 × SYBR-Green PCR Mix (TaKaRa, TaKaRa Biomedical Technology, Beijing, China). Real-time PCR was performed in triplicates for each sample, and the specificity of PCR product was tested by dissociation curve. Housekeeping gene β-actin was used as the internal control. 2−△△Ct method was used to quantify the relative expressions of each lncRNA and mRNAs. Data were presented as fold changes of transcripts in PDAC tissue compared to noncancerous tissue. A 2-tailed
Results
DE lncRNAs and mRNA in PDAC Tissues Compared With Adjacent Noncancerous Tissues
A genome-wide investigation of lncRNA and mRNA expressions in 8 pathologically confirmed PDAC tissues and their adjacent noncancerous tissues was performed using Agilent Human 46180K lncRNA arrays and Sureprint G3 Human lncRNA Chip. Clinical characteristics of these 8 patients are shown in Table 1, and none of these patients had metastasis to other part of the body (M0). A total of 63 431 lncRNAs were detected by microarray and were presented in volcano plot for visualization (Figure 1A) and whiskers plots for quantification (Supplemental Figure 1A). We found that the median number of lncRNA detected were similar between PDAC and noncancerous tissue among individuals (Supplemental Figure 1A). A total of 3352 of these lncRNAs were differentially expressed in PDAC tissues (fold change ≥2,
Clinical Characteristics of Enrolled Patients (n = 8).a

a Clinical characteristics of patients enrolled in the present study. American Joint Committee on Cancer (AJCC) TNM system: T: whether the primary tumor has grown outside the pancreas and into nearby organs size; N: tumor spreads to regional lymph nodes. M: cancer has metastasized to other organs of the body. n = 8 patients.
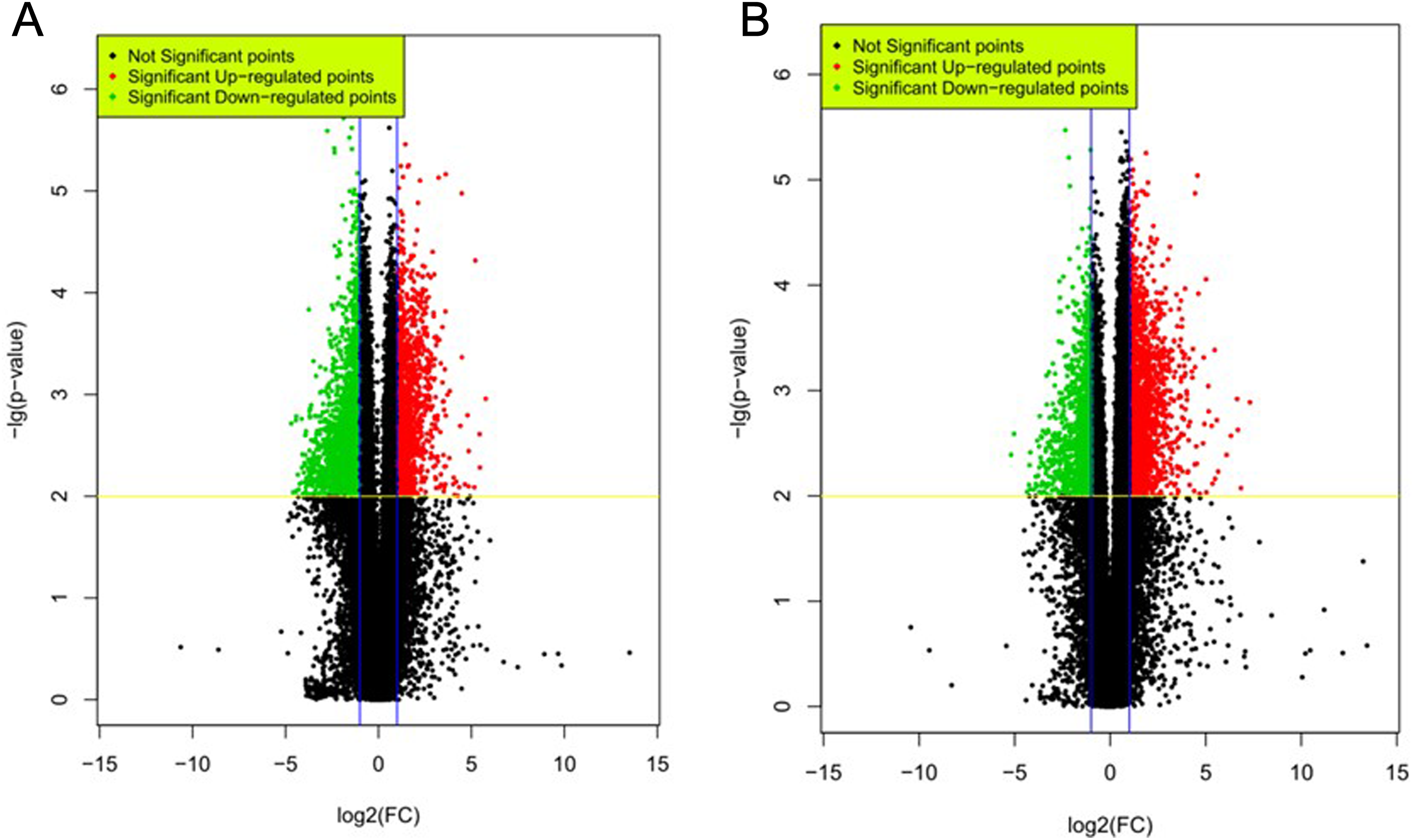
DE lncRNAs and mRNAs in PDAC tissues versus adjacent noncancerous tissue. Shown are the volcano plots of DE lncRNA (A) and mRNA (B). Upregulated and downregulated DE transcripts in PDAC with fold change ≥2,
In parallel, we analyzed the expressions of coding genes in PDAC versus normal tissues. A total of 39 887 genes were detected by microarray in PDAC tissues, and the median number of mRNA detected was comparable between samples (Supplemental Figure 1B). A total of 2717 mRNAs were significantly changed (fold change ≥2,
List of Top 5 Upregulated Coding Genes in PDAC.

Abbreviations: CLDN18, claudin-18; PDAC, pancreatic ductal adenocarcinoma.
List of Top 5 Downregulated Coding Genes in PDAC.

Abbreviation: PDAC, pancreatic ductal adenocarcinoma.
Chromosome Distribution of DE lncRNAs and mRNA
Most lncRNAs are likely to carry out their function in nearby coding genes. We therefore investigated the chromosome distribution of those DE lncRNAs and mRNA in human genome. We found that these DE lncRNAs and mRNA were not evenly distributed among chromosomes (Figure 2A left pie chart and 2B). The proportion of DE lncRNA among all annotated lncRNAs on each chromosome (DE lncRNA density) varies among each chromosome, ranging from 8% on chromosome Y to 25.4% on chromosome 16 (Figure 2C). Differentially expressed lncRNA density does not correlate with DE mRNA density on each chromosome (Pearson correlation

Chromosome distribution of DE lncRNA and mRNA in PDAC versus noncancerous tissue. A, Distribution of DE lncRNA and mRNA on each chromosome. B, Number of DE lncRNA or mRNA on each chromosome. C, Number of DE transcripts (lncRNA or mRNA) normalized to total number of annotated transcript (lncRNA or mRNA) on each chromosome. DE indicates differentially expressed; lncRNA, long noncoding RNA; mRNA, messenger RNA; PDAC, pancreatic ductal adenocarcinoma.
Hierarchical Clustering of DE lncRNAs and mRNAs in Pancreatic Adenocarcinoma
To visualize the general expression pattern of DE lncRNA and mRNAs from each patient, we performed hierarchical clustering analysis based on their average expression levels. All DE lncRNAs were grouped into 14 clusters (Figure 3A). Within clusters 1 to 10, DE lncRNAs were generally upregulated in PDAC tissues compared to their adjacent noncancerous tissues except for subject 224 (s224T). This was mainly because these lncRNAs were highly expressed in the noncancerous tissues and did not further increase in PDAC tissue in subject224. Besides, there was a large variability in the expressions of certain lncRNAs among individual PDAC samples. For example, the expression of lnc-MMP3-1 (matrix metalloproteinases (MMPs)) was decreased in PDAC tissue from subject 224, but was increased in PDAC from the other patients (Supplemental Figure 1). Lnc-NFYB-1:1 level was only reduced in PDAC from subject 223 but was increased in the PDAC from the other patients. Differentially expressed lncRNAs in clusters11 to 14 were generally downregulated in PDAC tissues with 2 exceptions, subjects 224 and 225. Although these PDAC samples were pathologically confirmed PDAC cases, the variability of DE lncRNAs among individual PDAC case suggested that these DE lncRNAs could be used as a molecular marker for specific subtype of PDAC.
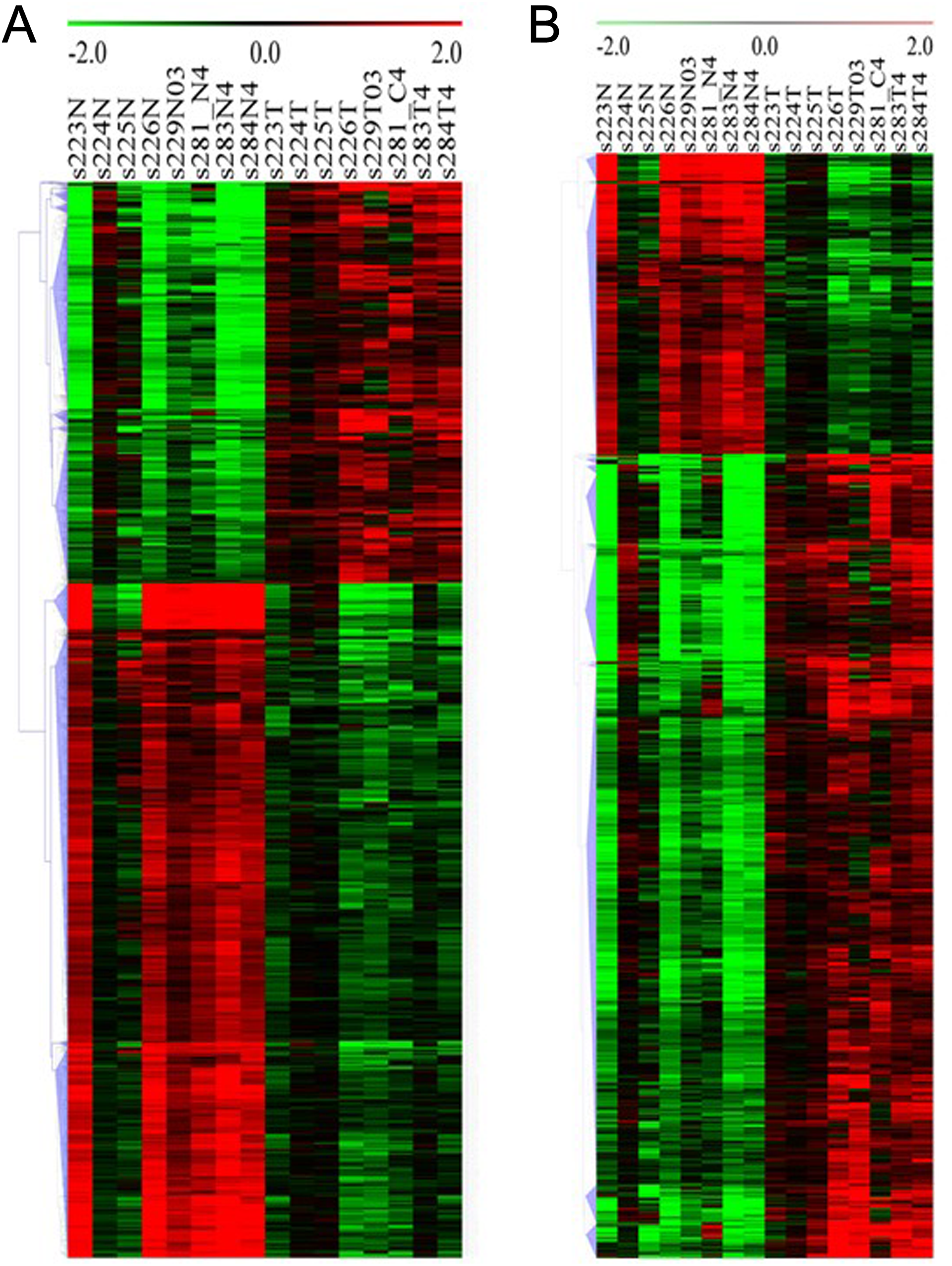
Hierarchical cluster of DE lncRNA (A) and mRNA (B) in human PDAC versus adjacent noncancerous tissue. Clustering was performed based on the average expression levels of transcript from 8 patients. Red: upregulated gene; green: downregulated gene. DE indicates differentially expressed; lncRNA, long noncoding RNA; mRNA, messenger RNA; PDAC, pancreatic ductal adenocarcinoma.
Cluster of DE mRNAs revealed that the majority of DE mRNAs were downregulated, which was opposite to DE lncRNA (Figure 3B; Supplemental Figure 2). Moreover, these mRNAs generally demonstrated no, mild, or moderate difference between PDAC and adjacent cancerous tissue in subject 224 and 225 but displayed more profound difference in the other patients. Variabilities between individuals were also observed. For example, the expressions of
GO Enrichment and Pathway Analysis of DE mRNAs in PDAC
To further explore potential targets of these DE lncRNA in PDAC progression, we performed GO enrichment and pathway analysis on DE mRNAs (Figure 4). Two hundred and eighteen GO terms, including biological processes (127), molecular functions (53), and cellular components (38), were detected and targeted by 9649 nearest coding mRNAs. The target genes and detailed information are represented in Supplemental Table 3. Thirty-one GO terms, including 20 biological processes, 4 molecular functions, and 7 cellular components, were determined to be significant with the threshold lg (

GO terms and KEGG pathways significantly enriched in human PDAC. Shown are significantly enriched GO terms (A) and KEGG pathways (B) of DE mRNAs in human PDAC versus adjacent noncancerous tissue. DE indicates differentially expressed; GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; mRNA, messenger RNA; PDAC, pancreatic ductal adenocarcinoma.
Construction of the Co-Expression Network Reveals the Potential Targets (mRNA) of DE lncRNA in PDAC
To further gain insights into the biological functions of lncRNA in the complex biological processes and cellular regulation, DE lncRNA/mRNA co-expression network was constructed to investigate the potential interaction between mRNAs and lncRNAs. As shown in Figure 4A, we detected 393 significant connections (co-expression events) in the network, which was composed of 80 DE lncRNAs and 105 mRNAs (

DE lncRNA and mRNA co-expression networks in human PDAC versus noncancerous tissue. Each arrow represents correlation
As an example, the most remarkably and reliably dysregulated lncRNA in all 8 patients was SH3PXD2A-AS1. This downregulated lncRNA hub was correlated with 11 edges, 8 of which were also significantly and reliably dysregulated in PDAC tissue, including coding UNC-5 netrin receptor A, 13-fold downregulated (
Experimental Validation of DE lncRNAs by qRT-PCR
Finally, we selected7DE lncRNAs that were of highest fold changes and were reliably dysregulated across all 8 patients, including SH3PXD2A-AS1 (highlighted by start), AC006372.4, lnc-BHLHE23-1:1, LYPLAL1-AS1, lnc-RP11-204N11.1.1-3:9, distal-less homeobox 2 antisense RNA-1 (DLX2-AS1), and RP11-680F8.3. We performed experimental validation by reverse transcription quantitative PCR (RT-qPCR; Figure 6). We found that 5 of 7 lncRNAs expression patterns were consistent with microarray data (Figure 6 and yellow stars with red outline in Figure 5). However, changes in LYPLAL1-AS1 and RP11-680F8.3 expressions revealed in qPCR analysis were opposite to microarray analysis (Figure 6 and white stars with red outline in Figure 5).

Validation of important DE lncRNAs by qRT-PCR. n = 8 patients, *
Discussion
The functional significance of lncRNAs has been widely recognized, 42 and recent studies have demonstrated that lncRNAs play important roles in carcinogenesis. 43,44 Dysregulated lncRNAs could be a major cause of oncogenesis or a molecular markers for cancer diagnosis, risk stratification, and monitor of therapeutics. 45,46,21,30 Genome-wide microarray survey of lncRNA may also facilitate our overall understanding of carcinogenesis mechanisms when combined with mRNAs profiling. However, lncRNA landscapes and lncRNA/mRNA co-regulation networks have not been investigated in PDAC from Chinese patients. In the present study, we detected 3352 significantly dysregulated lncRNAs, ∼60% of which were downregulated. In contrast to lncRNAs, >60% DE mRNAs were upregulated in PDAC tissue. Correlation analysis further revealed that the majority of lncRNA and mRNAs were negatively correlated, suggesting that lncRNAs were likely to serve as negative regulators of their targeting coding genes. Gene Ontology and KEGG pathway analysis identified essential pathways that underlie the mechanisms of PDAC tumorigenesis. Dysregulated mRNAs were enriched in pathways such as extracellular matrix receptor interaction, focal adhesion, phagosome, and regulation of actin cytoskeleton (Figure 4), all of which has been reported to be involved in tumor formation and progression. 42
We further found that these DE lncRNAs and mRNAs were unevenly distributed in human genomes. Morbidity of PDAC was indicated to be different in gender, and the incidence of PDAC was more common in women than in men. 47 We showed that the total number or the density of DE transcripts was significantly greater on chromosome X than chromosome Y, suggesting that dysregulated lncRNAs and mRNAs on sex chromosome X could be responsible for increased disease morbidity in women. Hierarchical clustering analysis revealed high variability of dysregulated lncRNAs in each patient. This indicates that the lncRNA molecular signatures may facilitate disease subtype classification, disease risk stratification, and prognosis prediction in pathologically confirmed PDAC cases.
One of the top and most reliably upregulated DE lncRNA in PDAC was SH3PXD2A antisense RNA-1 (SH3PXD2A-AS1; ∼40 fold upregulated). Its opposite strand encodes a protein coding gene named
LncRNAs/mRNAs co-expression network analysis revealed that SH3PXD2A-AS1, the most significantly upregulated lncRNA, is the most remarkable hub with 11 correlated edges. The correlated coding mRNAs included several genes that were implicated in cancer tumorigenesis. For example,
There are several limitations in the present study: (1) Limited sample size in the present study does not allow us to test the values of DE lncRNAs as the biomarker for early PDAC diagnosis or risk stratification. Further observational or prospective population studies with larger sample size is needed; (2) Future molecular characterization of important DE lncRNAs in PDAC tumorigenesis is warranted.
Conclusion
Our genome-wide survey of dysregulated lncRNAs and lncRNA/mRNA co-expression networks in Chinese PDAC tissue revealed hundreds of dysregulated lncRNA and mRNA. These could be used as biomarkers to guide the diagnosis of subtypes of PDAC, predict prognosis, and evaluate treatment efficacy in patients with PDAC.
Chen Jin and Deliang Fu conceived the study; Sijie Hao, Lie Yao designed the experiments; and Sijie Hao, Lie Yao, Jiaxin Huang, Hang He, Feng Yang, and Yang Di conducted the experiments and analyzed the data.
Footnotes
Abbreviations
Acknowledgment
The authors would like to thank all participating patients.
Declaration of Conflicting Interests
The author(s) declared no potential conflict of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a grant from Shanghai Municipal Commission of Health and Family Planning, China (Grant No. 20134Y081)
Supplemental Material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
