Abstract
The inhibitor of kappa B kinase epsilon is overexpressed in glioma and plays antiapoptotic role via activating nuclear factor-kappa B. microRNA-98 can suppress glioma, modulate the activities of nuclear factor-kappa B, and bind to the 3′-untranslated region of inhibitor of kappa B kinase epsilon messenger RNA. This study was aimed to investigate the modulation of inhibitor of kappa B kinase epsilon/nuclear factor-kappa B by microRNA-98 in glioma. The results indicated that microRNA-98 was downregulated in glioma cell lines and human glioma tissues. Overexpression of microRNA-98 in U87MG and T98G glioma cells significantly increased the apoptosis induced by ultraviolet irradiation and suppressed nuclear factor-kappa B luciferase activity, nuclear factor-kappa B p50 subunit expression, and B-cell lymphoma-2 (Bcl-2) expression in glioma cells. Silencing inhibitor of kappa B kinase epsilon decreased the expression of nuclear factor-kappa B p50 subunit and the luciferase activity of nuclear factor-kappa B, while the nuclear factor-kappa B activity could be significantly retrieved when inhibitor of kappa B kinase epsilon was expressed in microRNA-98-transfected cells. These findings indicated that microRNA-98 could promote apoptosis of glioma cells via inhibiting inhibitor of kappa B kinase epsilon/nuclear factor-kappa B signaling and presented a novel regulatory pathway of microRNA-98 by direct suppression of inhibitor of kappa B kinase epsilon/nuclear factor-kappa B expression in glioma cells.
Introduction
Aberrant constitutive activation of nuclear factor-kappa B (NF-κB), a transcriptional factor known as a pivotal mediator in inflammation and cancer, has been identified in glioma, 1 which is the one of most common and intractable malignant tumors in brain. 2 Classical NF-κB functions as a heterodimer composed of the DNA-binding subunit NF-κB (p50) and the transactivation subunit RelA (p65). 3 Previous studies have confirmed the association of NF-κB overstimulation with multiple aspects of glioma genesis, 4 –7 while NF-κB inhibition showed antitumor effects in glioma as well as a variety of other cancers. 8,9 The inhibitor of kappa B kinase epsilon (IKBKE, also known as IKKε/IKKi) has been identified as an inducible IκB kinase protein leading to NF-κB activation via phosphorylating IκB. 10,11 The IKBKE also functions as an oncogenic kinase overexpressed in many kinds of cancers, including breast cancer, ovarian cancer, and prostate cancer. 12 –14 Previous study has indicated that the expression of IKBKE is elevated in glioma cell lines and human glioma tissues and plays an important role in the prevention of cell apoptosis in glioma, 13 acting as an activator of NF-κB pathway in breast cancer and glioma. 13,14 These studies suggest that IKBKE/NF-kB pathway plays an important role in glioma pathology; however, how IKBKE expression is modulated in cancer has not been elucidated.
MicroRNAs (miRNAs), a family of small noncoding RNAs, can bind to the 3′-untranslated region (3′-UTR) of messenger RNAs (mRNAs), leading to mRNA degradation and/or translational repression. 15 Recent studies have characterized miRNAs as mediators of NF-κB/IKBKE pathway, which involves in the immune responses, cell proliferation and death, and inflammation as well as the development and progression of human cancers. 16,17 miR-98, a member of let-7 miRNA family, can downregulate NF-κB activation in immune response through targeting the 3′-UTR of cytokine-inducible Src homology 2-containing protein (CIS) in cholangiocytes. 18 Overexpression of miR-98 has been found to be involved in suppression of glioma and may modulate the activities of NF-κB. 19,20 In addition, miR-98 is predicted to bind to the 3′-UTR of IKBKE mRNA by 3 databases (TargetScan, MiRanda, and PicTar). All these data urge us to delineate whether miR-98 could actually target IKBKE and proceed to promote apoptosis of glioma cells via modulating the activities of NF-κB in glioma.
In this study, we demonstrated that miR-98 expression was decreased in glioma cell lines and human glioma tissues, while exogenous overexpression of miR-98 in glioma cells enhanced cell apoptosis through decreasing the activation of NF-κB and its downstream effector Bcl-2 and caspase-3. We also verified that miR-98 negatively regulated IKBKE/NF-κB signaling pathway in glioma cells, which could be via directly targeting 3′-UTR of IKBKE mRNA.
Materials and Methods
Clinical Specimens
The biopsy specimens with adjacent noncancerous brain tissues were collected from 20 glioma patients during surgery in 2013 at the First Affiliated Hospital of Sun Yat-sen University. The patients included 3 cases in grade 1, 4 cases in grade 2, 7 cases in grade 3, and 6 cases in grade 4 by pathological diagnosis and grading of glioma, according to the World Health Organization classification. 21 This study was approved by the Clinical Research and Experimental Animal Ethics Committee of the hospital. Prior written informed consents and approval were obtained from all the patients involved in this study.
Cell Lines and Culture
Normal human astrocytes (NHAs) were purchased from the ScienCell Research Laboratories (Carlsbad, California) as control. Glioma cell lines A172, LN-18, U-251MG, LN-308, LN-382, LN-428, LN-444, LN-464, U-87MG, and T98G were kindly provided by Dr Shi-Yuan Cheng at the Department of Pathology, University of Pittsburgh (Pittsburgh, Pennsylvania). All cell lines were passaged for no more than 6 generations after receipt or resuscitation, and so authentication from the sources was accepted. Cells were incubated in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 5% fetal bovine serum at 37°C and 5% CO2.
RNA Extraction and miRNA Real-Time Reverse Transcription Polymerase Chain Reaction
Total RNA was extracted from cultured cells and specimens of patients using TRIzol (Invitrogen, Carlsbad, California) and used to synthesize complementary DNA using SuperScript II Reverse Transcriptase (Invitrogen), according to the manufacturer’s instructions. The Bulge-Loop miR-98 primers (RiboBio Co Ltd, Guangzhou, China) were forward 5′-UGAGGUAGUAAGUUGUAUUGUU-3′ and reverse 5′-AACAAUACAACUUCUACCUCA-3′, resulting in production of 200 bp length sequence. Real-time reverse transcription polymerase chain reaction (RT-PCR) was performed with the CFX384 system (Bio-Rad Laboratories, Berkeley, California) using SYBR Green (Roche Applied Science, Mannheim, Germany) for 10 minutes of initial denaturation at 95°C, followed by 39 cycles of 15-second denaturation at 95°C, 1-minutes annealing at 60°C, and 1-minutes extension at 60°C. The threshold cycles (Ct) were calculated automatically using the CFX manager software (Bio-Rad Laboratories). The expression of miR-98 was defined based on the Ct, and relative expression levels were calculated as 2−[(Ct of miR-98) − (Ct of U6)] after normalization to the expression level of U6 small nuclear RNA.
Western Blot Analysis
Western blot was performed and total protein extracts were prepared from tissues and cell lines according to a standard method as described previously. 22 Cellular nuclear protein subfractions were extracted using a commercial extraction kit (Active Motif, Tokyo, Japan), according to the protocol provided by the manufacturer. The following primary antibodies were used: anti-IKBKE (1:500; Sigma-Aldrich, St Louis, Missouri), anti-α-tubulin (1:1000; Sigma-Aldrich), anti-Elongation Factor 1 Alpha (EF-1a) antibody (1:1000; Upstate, Temecula, California), anti-NF-κB p50 (1:500; Santa Cruz Biotechnology, Santa Cruz, California), and anti-Bcl-2 and anti-caspase-3 (1:1000; Cell Signaling Technology, Danvers, Massachusetts). The secondary antibodies included goat anti-rabbit IgG-HRP and rabbit anti-mouse IgG-horse radish peroxidase (HRP).
Transfection
Both MiR-98 mimic and miR-98 inhibitors were designed as described previously. 14 Both U87MG and T98G cells were seeded in 6-well plates and transfected with miR-98 mimic, miR-98 inhibitor (RiboBio Co Ltd), IKBKE–small interfering RNA (siRNA), or miR-98 plus IKBKE at a final concentration of 50 nmol/L using the Lipofectamine 2000 reagent (Invitrogen), according to the manufacturer instructions. After 72 hours of transfection, cells were split and RNA was preserved in TRIzol reagent (Invitrogen).
TUNEL Assay
The cell apoptosis was measured with the DeadEnd Colorimetric TUNEL System (Promega, Madison, Wisconsin) and TdT-mediated dUTP Nick-End labeling (TUNEL) assay, following the manufacturer instructions. In brief, the transfected cells were inoculated in 24-well plates at a density of 5 × 104 cells/well, cultured with DMEM for 24 hours, irradiated with ultraviolet (UV) at 40 J/m2, and examined for apoptosis after 24 hours. The cells were rinsed with 1× phosphate-buffered saline (PBS) once after removal of culture medium, fixed with 4% paraformaldehyde at room temperature for 25 minutes, rinsed with 1× PBS for 3 × 5 minutes, and added with 0.1% Triton X-100 for 2 minutes. After removal of the Triton X-100, the cells were rinsed with PBS once, incubated with 0.3% hydrogen peroxide solution at room temperature for 20 minutes, and rinsed with PBS 3 times. Then, the cells were incubated with 50 μL of biotin labeling solution (composed of TdT 20 μL, biotin-dUTP 480 μL) in 37°C for 1 hour, rinsed with PBS once, added with 0.1 to 0.3 mL of blocking solution for incubation in room temperature for 10 minutes, and rinsed with PBS 3 times. The cells were treated with 50 μL of streptavidin–HRP solution (composed of 10 μL of streptavidin–HRP solution and 490 diluting solution) in room temperature for 30 minutes, rinsed with PBS 3 times, treated with 0.2 to 0.5 mL of DAB solution in room temperature for 15 minutes, and rinsed with PBS 3 times. Finally, the cells were stained with hematoxylin staining solution, rinsed with PBS 3 times, and treated with 95% ethanol for 5 minutes, 100% ethanol for 2 × 3 minutes, and xylene for 2 × 5 minutes. After sealing with coverslip, 10 fields under 400× magnification were selected to account for the apoptotic cells and calculate the apoptosis rate.
Luciferase Reporter Constructs and Luciferase Assays
The full-length sequence of IKBKE-3′-UTR is 744 bp and contains the conserved miR-98 binding sites from 107 to 113 bp. The total region of human IKBKE 3′-UTR was generated by PCR amplification with flanking PstI and SacII at restriction enzyme digestion sites. The annealed oligonucleotides were cloned into the PstI-SacII sites of the pGL3-basic luciferase reporter plasmid (Promega, Madison, Wisconsin). The oligonucleotides harboring IKBKE 3′-UTR with mutations in the seed regions for miR-98 binding were also cloned into pGL3-basic luciferase reporter plasmid. The primers were IKBKE-3′UTR-wt-up, 5′-GGCATCCTGAAGCATTAG-3′, and IKBKE-3′UTR-wt-dn, 5′-GCCACATTTATTCCACTTC-3′. The potential binding site CUACCUCC was substituted by GATGGAG in pGL3-IKBKE-3′UTR-mut. The IKBKE-3′-UTR (wt/mut) and control plasmid were used to examine the quantitative expression of IKBKE-3′UTR. pBabe-puro-Flag-IKBKE was constructed according to previous study. 14 The pNF-κB-luc and control plasmids were purchased from the Clontech Laboratories (Mountain View, California) for quantitatively examining the activity of NF-κB. Cells were cotransfected with reporter plasmid and miR-98 sequence using Lipofectamine 2000 reagent (Invitrogen). Luciferase activity of whole-cell lysates was measured 72 hours after transfection and normalized to the expression level of TK-Renilla using the Dual Luciferase Reporter Assay System (Promega), according to the manufacturer protocols.
Statistical Analysis
The data were expressed as the means ± standard deviation (SD) and analyzed with software SPSS 10.0 (SPSS Inc, Chicago, Illinois). The differences between different groups were compared using Student t tests. P < .05 was set as significant difference level.
Results
MiR-98 Is Downregulated in Glioma Cell Lines and Glioma Tissues
To investigate the role of miR-98 in glioma, we examined the expression of miR-98 in NHA, human glioma cell lines, and human glioma tissues from patients with glioma using real-time RT-PCR. The results indicated that the expression of miR-98 was significantly decreased at different levels in all tested glioma cell lines (A172, LN-18, U-251MG, LN-308, LN-382, LN-428, LN-444, LN-464, U-87MG, and T98G; n = 20 for each cell line) as compared with that in NHA (n = 6; all P = .001; Figure 1A). Furthermore, we examined the expression of miR-98 in human glioma tissue and adjacent noncancerous tissues from 20 patients. The result revealed that the expression level of miR-98 in glioma tissues was significantly lower than that in noncancerous tissues (P = .017, .001, .001, and .001, respectively; Figure 1B) in gliomas of different grade.

Real-time RT-PCR showing downregulation of miR-98 in glioma cells and clinical glioma specimen. A, Expression of miR-98 in NHA and glioma cell lines, including A172, LN-18, U-251MG, LN-308, LN-382, LN-428, LN-444, LN-464, U-87MG, and T98G. B, Expression of miR-98 in glioma tissues and adjacent noncancerous brain tissues(*P < .05 vs NHA or normal). RT-PCR indicates reverse transcription polymerase chain reaction; NHA, normal human astrocytes.
Upregulation of miR-98 in Glioma Cells Enhances UV-Induced Apoptosis
To determine whether miR-98 could impact the pathological progress of glioma, cell apoptosis induced by UV irradiation was investigated in miR-98-transfected glioma cells. As shown in Figure 2, it was revealed that the percentage of TUNEL-positive cells in miR-98-transfected cells was significantly higher than that in control cells and the percentage of TUNEL-positive cells in miR-98 inhibitor–transfected cells was significantly lower than that in control cells for both U87MG cells (P = .014) and T98G cells (P = .001), suggesting that miR-98 reduces the resistance of glioma cells to UV irradiation and promotes apoptosis in glioma cells (Figure 2).
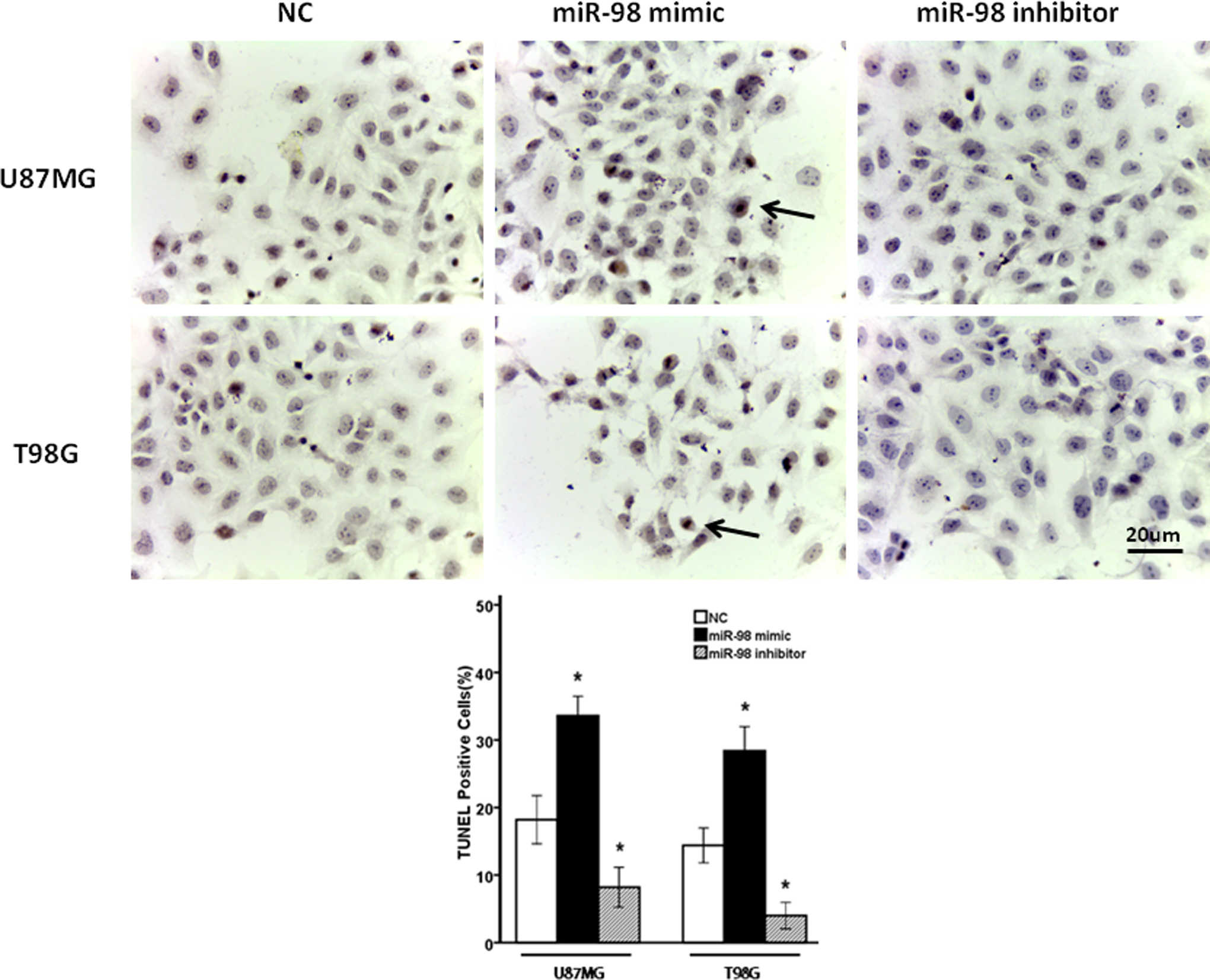
Exogenous expression of miR-98 affected UV-induced apoptosis in glioma cells. The miR-98 mimic and miR-98 inhibitor-transfected glioma cells and their NC were treated by UV irradiation (20 J/m2). After 24 hours, the numbers of TUNEL-positive cells were counted in 5 randomly selected fields. Bar = 20 μm (*P < .05 vs NC). miR-98 indicates microRNA-98; NC, negative controls; UV, ultraviolet.
The 3′-UTR of IKBKE Is Directly Targeted by miR-98
Our previous study has indicated that the expression of IKBKE is elevated in glioma and plays an important role in preventing apoptosis of glioma cells. 14 Through analysis of the TargetScan (http://www.targetscan.org/) and PicTar (http://pictar.mdc-berlin.de/) algorithms, we examined all computationally predicted target genes of miR-98 and found that IKBKE has conserved 7-mer seed matches with miR-98 in its 3′-UTR. To confirm that IKBKE is the target gene of miR-98 as predicted (Figure 3A), we tested the expression of IKBKE in miR-98-transfected glioma cells using luciferase assays and Western blot analysis. As shown in Figure 3B, the luciferase activity in glioma cells cotransfected with pGL3-IKBKE-3′UTR and miR-98 was significantly decreased in U87MG and T98G cells as compared to that in the control group (P = .001 and .001, respectively), while there was no significant difference in luciferase activity in glioma cells between control group and cotransfection group with pGL3-IKBKE-3′UTR-mut and miR-98 (P = .854 and .937, respectively). It confirmed the miR-98 can recognize and bind to the specific site of IKBKE-3′UTR. Furthermore, we examined the effects of miR-98 transfection on the expression of endogenous IKBKE protein. Western blot analysis showed that the expression levels of endogenous IKBKE protein in miR-98-transfected U87MG and T98G cells were significantly decreased compared with those in control cells (P = .001 and .008, respectively; Figure 3C). Taken together, these results indicated that miR-98 downregulates the expression of IKBKE protein via directly targeting the 3′-UTR of IKBKE.

The miR-98 downregulated IKBKE by targeting IKBEK 3′-UTR which was targeted by miR-98 in glioma cells. A, The IKBKE 3′-UTR contained predicted miR-98-binding sites. B, The effects of miR-98 on the luciferase activity of IKBKE-3′UTR reporter in glioma cells. C, The effect of miR-98 on the expression of IKBKE protein in glioma cells using Western blot (*P < .05 vs pGL3-IKBKE-3′UTR or control). miR-98 indicates microRNA-98; IKBKE, IκB kinase-∊.
Nuclear Factor-Kappa B Signaling Is Involved in the Pro-Apoptosis of miR-98 in Glioma Cells
To demonstrate the possible pro-apoptosis mechanism of miR-98 in glioma, we further explored the related signaling pathway. As shown in Figure 4A, NF-κB reporter assay indicated that the luciferase activity driven by NF-κB-inducible promoter was significantly decreased in miR-98-transfected U87MG and T98G cells in comparison with that in control (P = .001 and .001, respectively; Figure 4A). Western blot analysis showed that the level of NF-κB p50 subunit was significantly diminished in miR-98-transfected cells (P = .015 and .011, respectively; Figure 4B). Furthermore, the expression level of Bcl-2, the direct transcriptional effector of NF-κB, was decreased in miR-98-transfected U87MG and T98G cells as compared with negative control cells (P = .005 and .006, respectively; Figure 4C)

The NF-κB signaling was downregulated in miR-98-transfected glioma cells. A, The effect of miR-98 on luciferase activity of NF-κB in glioma cells. B, The effect of miR-98 on the expression of NF-κB p50 subunit nuclear translocation assessed by Western blot. C, The effect of miR-98 on the expression of Bcl-2 protein in glioma cells determined by Western blot (*P < .05 vs control). NF-κB indicates nuclear factor-kappa B; miR-98, microRNA-98.
Inhibitor of Kappa B Kinase Epsilon Mediates the miR-98-Induced Repression of NF-κB Activity
To evaluate whether IKBKE is involved in the downregulation of NF-κB activity by miR-98, we conducted IKBKE-silencing experiments using siRNA. As shown in Figure 5A, Western blot analysis showed that the expression of NF-κB p50 protein was decreased in IKBKE-RNAi-treated U87MG and T98G cells (P = .001 and .001, respectively). Luciferase results showed that the luciferase activity driven by NF-κB-inducible promoter was significantly decreased in IKBKE-RNAi-treated U87MG and T98G cells in comparison with those in control cells (P = .001 and .001, respectively; Figure 5B). Importantly, further results showed that the percentages of TUNEL-positive cells cotransfected with miR-98 and IKBKE were significantly lower than those of miR-98-transfected cells (P = .003 and .005, respectively; Figure 6A). Furthermore, NF-κB reporter activation was greatly retrieved when miR-98 and IKBKE were cotransfected into U87MG and T98G cells (P = .011 and .032, respectively; Figure 6B). These results were consistent with the effect of miR-98 and IKBKE cotransfection on the expression of p50 (P = .024 and .038, respectively; Figure 7A), Bcl-2 (P = .021 and .018, respectively), and cleavage and activation of caspase-3 (P = .001 and .008, respectively; Figure 7B), which suggested that miR-98 promotes apoptosis of glioma cells at least in part mediated by IKBKE/NF-κB pathway.

The siRNA decreased the expression of NF-κB p50 subunit and the luciferase activity of NF-κB. A, The expression of NF-κB p50 protein was decreased in IKBKE-RNAi U87MG and T98G cells using Western blot. B, The luciferase activity of NF-κB-dependent reporter gene was decreased in IKBKE-RNAi glioma cells (*P < .05 vs Vector). NF-κB indicates nuclear factor-kappa B; IKBKE, IκB kinase-ε.

The effects of miR-98 and IKBKE cotransfection on UV-induced apoptosis and the luciferase activity of NF-κB-dependent reporter gene in glioma cells. A, miR-98 and IKBKE cotransfected glioma cells and miR-98-transfected glioma cells were treated by UV irradiation (20 J/m2). After 24 hours, the numbers of TUNEL-positive cells were counted in 5 randomly selected fields. The percentages of TUNEL-positive cells cotransfected with miR-98 and IKBKE were significantly lower than those of miR-98-transfected cells. B, The expression of NF-κB-dependent reporter gene was mostly retrieved in miR-98 and IKBKE cotransfected cells, compared with miR-98-transfected cells. Bar = 20 μm (*P < .05 vs miR-98). NF-κB indicates nuclear factor-kappa B; IKBKE, IκB kinase-ε; miR-98, microRNA-98.

The effects of miR-98 and IKBKE cotransfection on p50, caspase-3, and Bcl-2. The effects on the expression of (A) p50 in U87MG and T98G cells and (B) caspase-3 and Bcl-2 in U87MG and T98G cells (*P < .05 vs miR-98). IKBKE indicates IκB kinase-ε; miR-98, microRNA-98.
Discussion
In this study, we demonstrated that miR-98 was downregulated in glioma cell lines and human glioma tissues, while overexpression of miR-98 induced apoptosis of glioma cells and decreased Bcl-2 expression in glioma cells. The 3′-UTR of IKBKE mRNA was found to contain the binding cite of miR-98, while miR-98 reduced the luciferase activities and expression of IKBKE in glioma cells. In addition, IKBKE-RNAi reduced the expression of NF-κB p50 subunit and the luciferase activities of NF-κB but reversed the inhibition of NF-κB by miR-98 in glioma cells. These results indicated that miR-98 mediates the apoptosis of glioma cells through IKBKE/NF-κB and the downstream of Bcl-2 and caspase-3.
As the most common malignant tumor in brain, glioma is still intractable although there are many advancements in the studying of its mechanisms. Accumulating evidence indicates that miRNAs function as oncogenes or tumor suppressors, 16,17,23 while overexpression of miR-98 is found to suppress glioma cells. 19,20 Consistently, in this study, we found that miR-98 was decreased in glioma cell lines and human glioma tissues, while overexpression of miR-98 could enhance the apoptosis of glioma cells and decrease the expression of Bcl-2, which is a protein determining the apoptosis of cells. 24 These results suggest that miR-98 may play an important role in the pathology of glioma through modulation of apoptosis.
The mechanism of miR-98 involvement in glioma is related to the activities of IKBKE. The IKBKE is the computational predicted target of miR-98 (TargetScan, MiRanda, and PicTar), and our previous study demonstrated that IKBKE is overexpressed in glioma and contributes to the resistance of glioma cells to apoptosis. 14 In this study, we showed that miR-98 specifically targeted the highly conserved sequence of 3′-UTR of IKBKE mRNA in glioma cells and the expression level of IKBKE protein was reduced remarkably in miR-98-transfected glioma cells. These results lead to the hypothesis that miR-98 may be involved in the deteriorating process of glioma as the negative regulator of IKBKE.
As well known, NF-κB is constitutively activated in various kinds of cancers including glioma and contributes to multiple aspects of tumorigenesis including proliferation, apoptosis, metastasis, and angiogenesis. 25,26 Previous studies indicated that IKBKE is an activator of NF-κB in breast cancer 13 and suppresses the apoptosis of glioma cells through NF-κB, 14 while miR-98 can downregulate the activation of NF-κB. 18 In this study, we also found that downregulation of IKBKE by siRNA decreased the expression of NF-κB p50 subunit and the luciferase activity of NF-κB in glioma, which was in accordance with our previous results. 14 While further study showed that when IKBKE was overexpressed in miR-98-transfected cells, NF-κB activity was greatly retrieved, apoptosis cells were significantly decreased, the expression of NF-κB p50 and Bcl-2 was increased, and cleavage and activation of caspase-3 was decreased. These results suggest that miR-98 downregulates NF-κB activity and promotes glioma cell apoptosis mostly by suppressing the expression of IKBKE.
In conclusion, this study revealed a mechanism through which miR-98 could promote glioma cell apoptosis via targeting IKBKE/NF-κB pathway, which may be a novel strategy for enhancing the expression of miR-98 in glioma cells to suppress tumor progression and improve chemotherapeutic sensitivity. Therefore, the results have potential value for treating the intractable glioma in clinic.
Footnotes
Abbreviations
Declaration of Conflicting Interests
The author(s) declared no potential conflict of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China (Grant Nos. 81001121 and 81472351).
