Abstract
The worldwide high mortality rate of lung cancer could be reduced significantly by its noninvasive early detection. The quantitative analysis of cell-free circulating DNA in plasma presents a potential noninvasive approach for liquid biopsy of tumor. In this study, real-time polymerase chain reaction–based approach was used to quantify free circulating DNA in plasma. The concentration of free circulating DNA was checked using human telomerase reverse transcriptase gene as marker, and amplification status of oncogene RAC-β serine/threonine protein kinase along with the DNA methylation status of tumor suppressor gene (deleted in colorectal cancer) was assessed. The concentration of free circulating DNA in patients with lung cancer (22.8 ng/mL) was found approximately 6 times above than the value detected in controls (2.8 ng/mL). Considerable variation in the
Introduction
Lung cancer is the deadliest of all cancers having highest mortality and incidence rates worldwide. About 25% of the entire cancer-associated deaths are caused by lung cancer, and approximately 1.2 million new lung cancer cases are reported annually, worldwide. 1,2 In Pakistan, the situation is not very different from the world; lung-cancer is a major cancer type in men. Its prevalence rate is 20.3 per million in men and 2.9 per million in women. 3
Unfortunately, greater than 60% of lung cancer-cases are diagnosed at advanced stages after tumors have metastasized, when cure is unlikely. 4,5 Current screening tests available for early diagnosis of lung cancer include sputum cytology test, chest X-ray, and computed tomography (CT) scans, which are not much efficient for earlier diagnosis and increased survival rate. 6,7 These techniques have pitfalls such as high cost, hazardous effects of radiation exposure, reduced sensitivity especially in the case of former smokers, and poor specificity. 8,9
Lung cancer diagnosis has entered into a new era with the finding that tumors also secrete their nucleic acid content, that is, DNA and RNA into circulation. 10 The actual mechanism of free circulating DNA (fcDNA) release is not completely clear, but mostly it is associated with apoptosis of circulating tumor cells leading to the release of nucleic acids into the circulation. 11,12 These circulating nucleic acids can be retrieved from plasma and serum. 10 Therefore, quantifying fcDNA and molecular characterization of early genetic and epigenetic changes would be a valuable tool in screening and managing lung cancer.
For determination of elevated plasma fcDNA levels in patients with lung cancer, different approaches have been applied, and the real-time polymerase chain reaction (PCR)–based assay with a reference gene has been proved to be the most sensitive one. 13 –16 Typical tumor genetic changes, such as mutations, inversions, microsatellite alterations and deletions, 17 and epigenetic changes such as aberrant hypermethylation within the regulatory regions of gene, could be conveniently identified in circulating-DNA using PCR-based methods. 18,19
In the present study, we employed real-time PCR-based approach for quantifying fcDNA in plasma based on amplification of single-copy ubiquitous gene, human telomerase reverse transcriptase (
Materials and Methods
Study Population
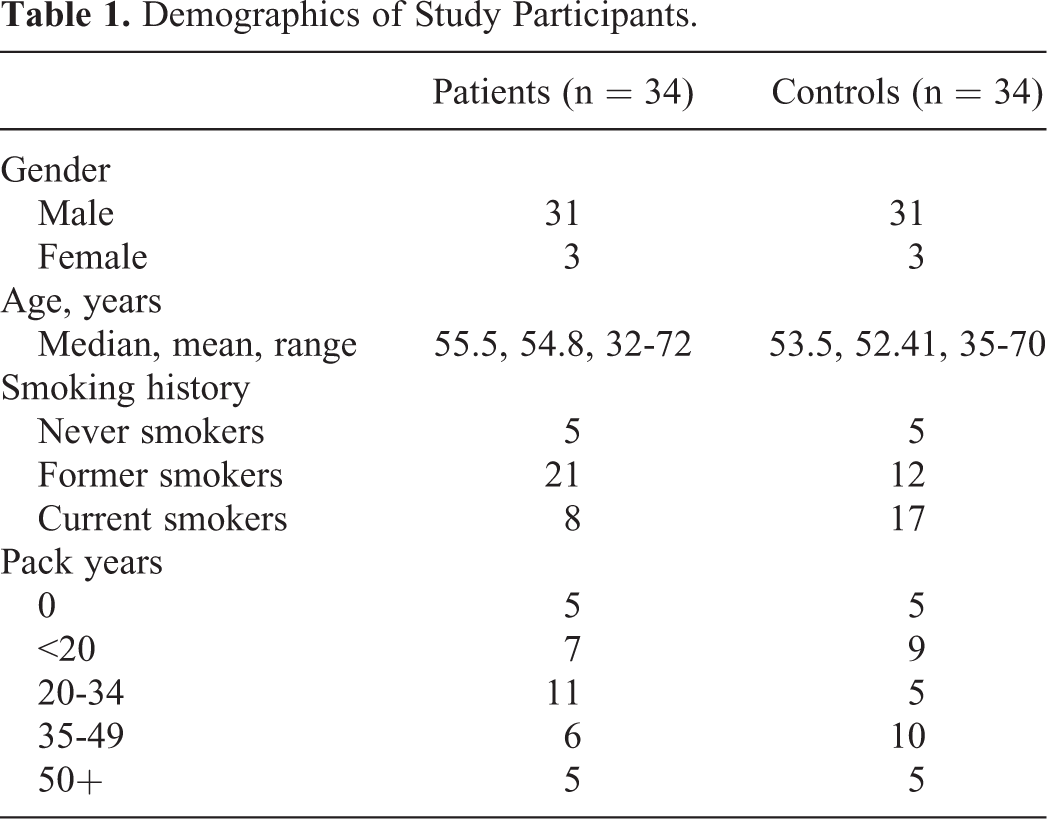
A total of 68 individuals with age ≥30 years were included in this study which comprised of 2 groups, patients with lung cancer and healthy control participants. Complete medical history as well as data regarding age, smoking habits, and histological type of disease were collected. Blood samples of hospitalized patients (n = 34) with non–small cell lung cancer (NSCLC) diagnosed by histopathological technique were collected before the start of anticancer treatment. Cases with primary cancers were only included, while cases with disease relapse or follow-up were excluded from the study population. Healthy controls (n = 34) were selected by sex, age, and smoking habits (like of patients) but without history of any malignant disease. Clinicopathological characteristics of the study-participants are presented in Table 1. Among NSCLC cases, 7 tumors were adenocarcinomas, 19 squamous cell carcinomas (SCCs), and 8 poorly differentiated carcinomas. From each patient and control, 5 mL blood sample was collected in EDTA tubes. Plasma was obtained immediately by centrifugation (2500 rpm, 10 minutes, and 4°C) twice. DNA from plasma was extracted by QIAamp DNA Mini kit (Qiagen, Stockach, Germany), following the manufacturer’s instructions.
Demographics of Study Participants.
Quantification of fcDNA
Quantitative real-time PCR was done for quantifying fcDNA. The assay used the amplification of
Determination of Gene Copy Number
To determine copy number of oncogene
Determination of Aberrant Methylation
First, bisulfite modification of plasma fcDNA was performed with EpiTect Bisulfite kit from Qiagen, following the manufacturer’s-instructions. This bisulfite-modified DNA then served as template for fluorescence-based real-time PCR. The primers used for amplification of the bisulfate-modified promoter region of
Statistical Analysis
For statistical analysis, SPSS Statistics for Windows (version 17.0; SPSS Inc. Chicago, Illinois) was used. The Mann-Whitney
Results
Quantification of fcDNA
The concentration of plasma fcDNA measured using

Box plots of plasma free circulating DNA (fcDNA) concentration (ng/mL) of patients with lung cancer and matched controls, using human telomerase reverse transcriptase
Diagnostic-performance of this fcDNA concentration assay was determined by ROC curve analysis. The ROC was made by plotting the sensitivity against 1-specificity. The AUC defines the capability of assay to distinguish patients from controls. As the area become close to 1, the diagnostic power increases. The AUC was found to be 0.809 (95% CI: 0.708-0.910;
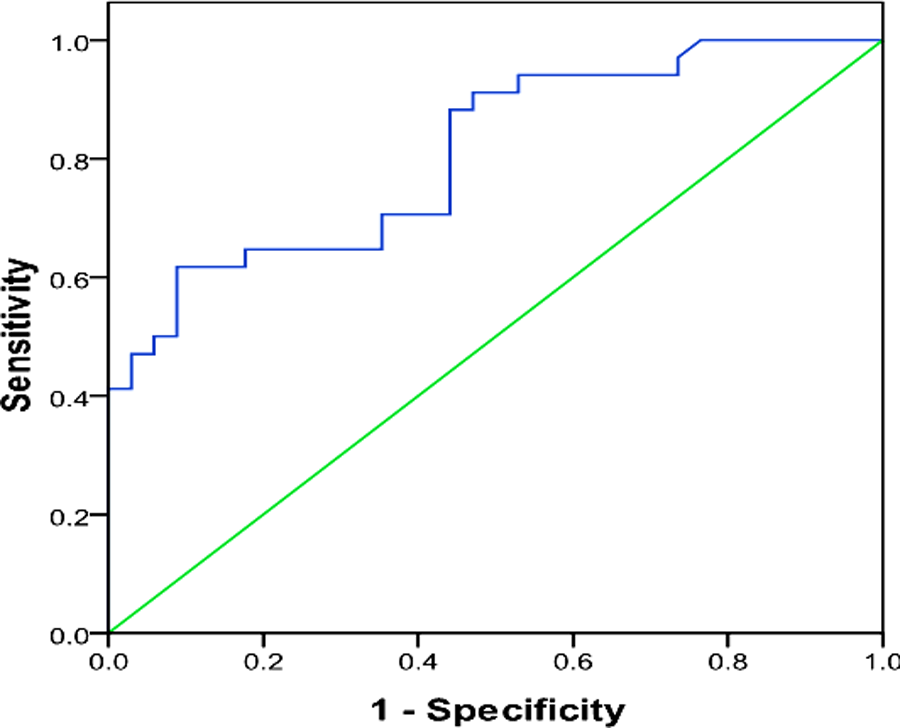
Receiver operating characteristic (ROC) curve for the lung cancer diagnosis by plasma-free circulating DNA (fcDNA) concentration.
It was observed that the assay sensitivity tends to increase with lesser cutoff values, although at expense of assay specificity. It can be a preliminary test to discriminate patients from healthy subjects which requires additional verification. Thus, the sensitivity was preferred to specificity to decrease the possibility of false negatives. So a low cutoff-value 1.9 ng/mL was chosen. It provides a sensitivity of 82.4% in identifying patients with lung cancer. Using this cutoff value, test performance is shown in Table 2.
Diagnostic Performance of fcDNA Quantification Using

Abbreviations: DCC, deleted in colorectal cancer; fcDNA, free circulating DNA;
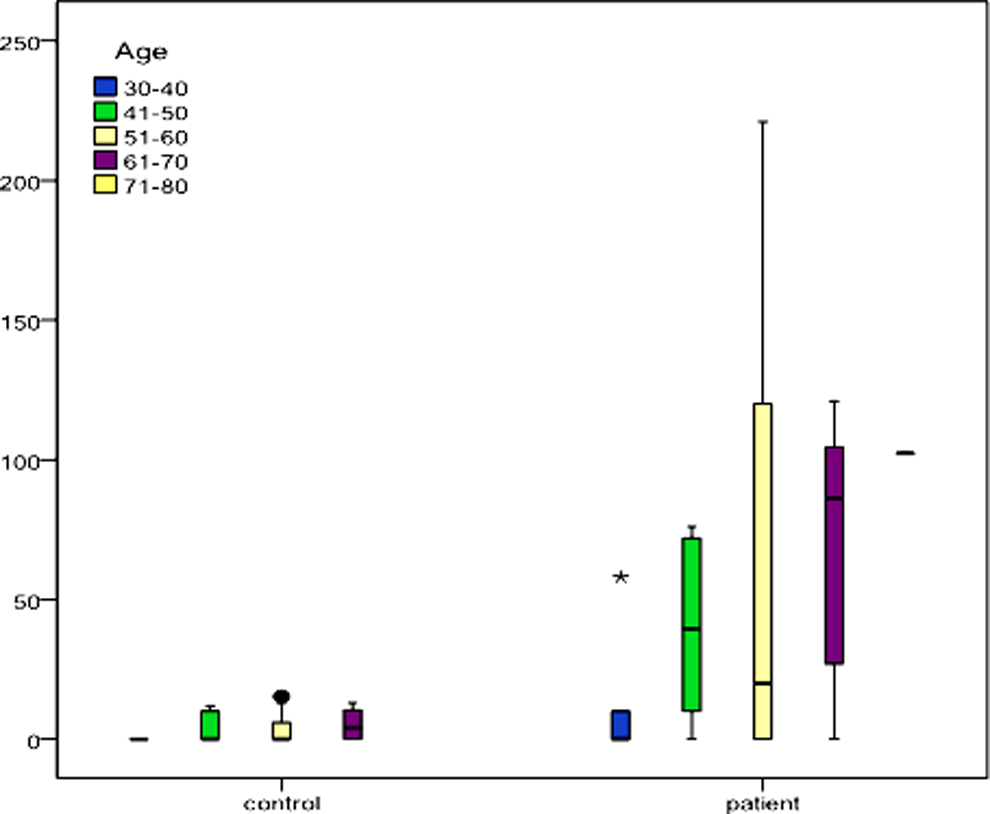
Plasma fcDNA concentration was significantly associated with age (
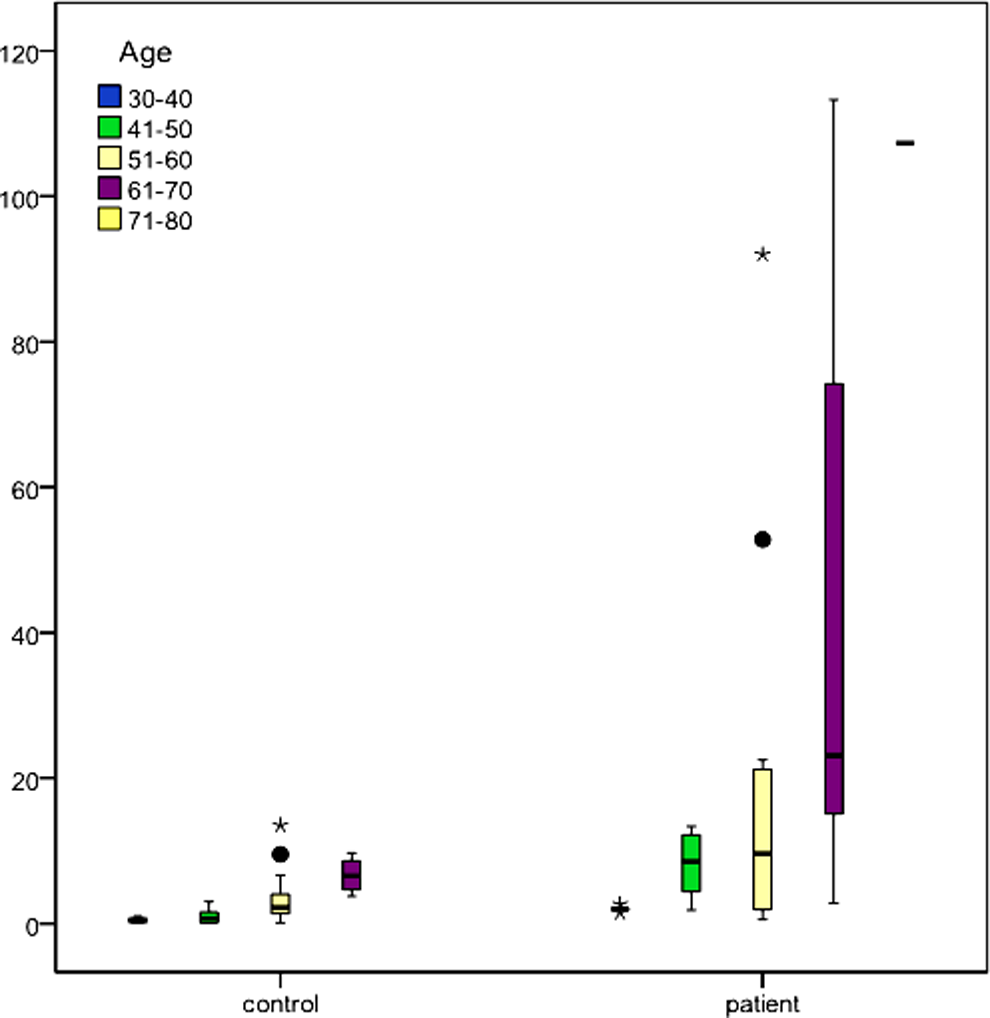
Distribution of free circulating DNA (fcDNA) levels (ng/mL) with respect to age in controls and patients with lung cancer.
Quantification of AKT2 Gene Dosage in fcDNA
Quantification of
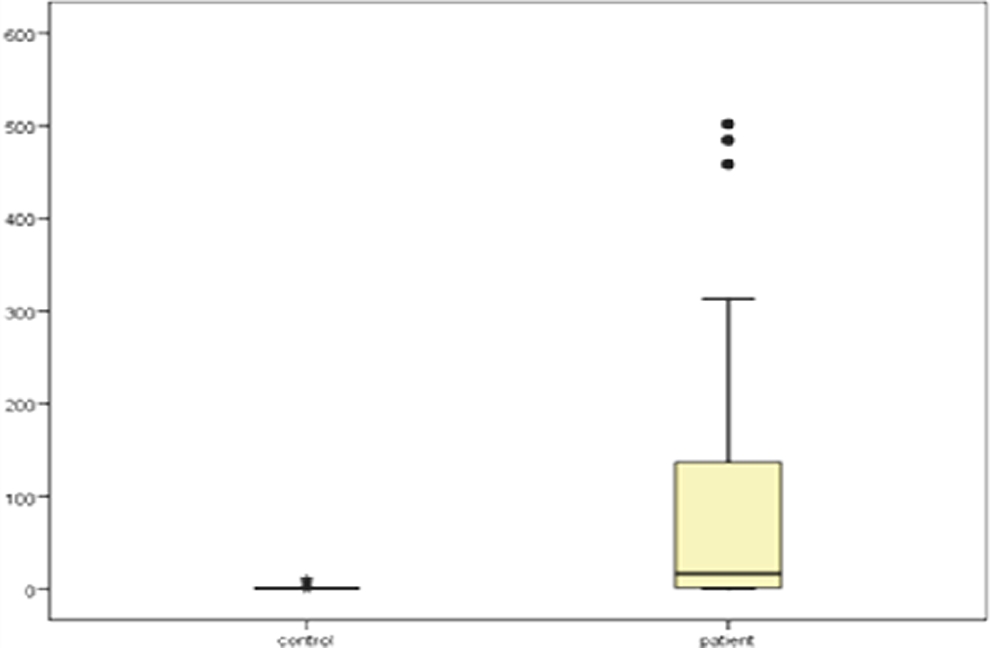
Box plots of plasma free circulating DNA (fcDNA)
The

Receiver operating characteristic (ROC) curve for the lung cancer diagnosis by plasma free circulating DNA (fcDNA)

Box plots of relative methylation values for deleted in colorectal cancer (
Determination of Aberrant Promoter Methylation of DCC
Aberrant promoter methylation of
Aberrant methylation of the

Receiver operating characteristic (ROC) curve for the lung cancer diagnosis by plasma free circulating DNA (fcDNA) deleted in colorectal cancer (
Distribution of
Discussion
Development of noninvasive diagnostic tests for cancer using blood samples is desired since long, as invasive approaches to get tumor tissue are not feasible in all the cases. The evidence for fcDNA in plasma of patients with cancer released from tumor provides this opportunity.
10,24
In lung cancer, quantification of fcDNA and molecular characterization of specific genetic and epigentic changes have been analyzed as diagnostic tool. Several methodologies have been described for quantitative analysis of fcDNA; the use of real-time PCR is most promising because of its high sensitivity and specificity.
13,25
–28
However, circulating DNA analysis has not been standardized to the extent to be used clinically as diagnostic test so far. This is because of variability in sample-processing, fcDNA isolation, and its quantification.
29
Therefore, the development of a reproducible method for clinical-application of this potential diagnostic assay is needed. In this study, the concentration of fcDNA is determined by quantitative PCR approach using
This methodology of plasma fcDNA quantification was found to be very sensitive and specific with effective diagnostic performance as AUC curve approached 0.809 (95% CI: 0.708-0.910;
Previous data about prognostic factors related to fcDNA concentration are contradictory. The correlation of fcDNA with gender, histology, lung cancer stage, and age is unclear yet.
13,27
Our study is also in accordance with other reports, to be deficient in association of plasma fcDNA levels with pathological-factors.
26,27
In our samples, high plasma fcDNA concentrations in any histological subtype of lung cancer were observed without selectivity. A possible reason can be variation in selecting patients, including NSCLC as well as SCLC. A number of-studies report correlation of fcDNA with age,
13,14
excluding others.
27,28
The data we obtained have shown that plasma concentration was significantly correlated with age (
In the present study,
Evidently, quantification of gene copy number changes in plasma by quantitative PCR presents an attractive lung cancer’s diagnostic method. However, small-fcDNA-amount in plasma does not reveal the location of tumor in the lung and also it is not lung cancer specific,
21,35,36
because
DNA methylation, an epigenetic- change capable of inactivating tumor-suppressor-genes presents tumor-biomarker 40 –42 which occurs in distinct CpG-Islands of gene's regulatory-region and can be much conveniently quantified. Also amplification of DNA methylation can be easily performed via PCR even if extracted DNA concentration is low. 43,44 So it can serve as the promising biomarker for cancer detection.
Here, we determined the frequency of tumor suppressor gene
So, the 3 biomarkers panel of fcDNA could be potentially used as a low-cost and less invasive approach for earlier diagnosis of lung cancer or to evaluate suspicious diagnosis cases and their management.
Footnotes
Abbreviations
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
