Abstract
The previously published data on the association between the cytochrome P450 1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms and lung cancer risk have remained controversial. Hence, we performed a meta-analysis to investigate the association between cytochrome P450 1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms and lung cancer risk under different inheritance models. A total of 22 studies were identified, including 2881 cases and 3653 controls for Leu432Val polymorphism (from 13 studies), 3009 cases and 3887 controls for Asn453Ser polymorphism (from 5 studies), and 1301 cases and 2045 controls for Ala119Ser polymorphism (from 4 studies). Overall, significant association was observed between cytochrome P450 1B1 Leu432Val polymorphism and lung cancer risk (dominant model: odds ratio = 1.29, 95% confidence interval = 1.08-1.53; recessive model: odds ratio = 1.21, 95% confidence interval = 1.05-1.39; additive model: odds ratio = 1.43, 95% confidence interval = 1.21-1.69) when all the eligible studies were pooled into the meta-analysis. In the further stratified and sensitivity analyses, significantly increased lung cancer risk was also observed in caucasians and smokers. No significant association was observed between cytochrome P450 1B1Asn453Ser and Ala119Ser polymorphisms and lung cancer risk in overall analysis. In summary, this meta-analysis suggests that cytochrome P450 1B1Leu432Val polymorphism is associated with increased lung cancer risk in caucasians and smokers.
Introduction
Lung cancer is a major cause of cancer-related death worldwide, and the overall survival rate is still extremely poor. 1 The exact mechanism of lung cancer is still under investigation. Epidemiological studies have demonstrated tobacco smoking as well as environmental tobacco smoke in nontobacco users as the major risk factor for the development of lung cancer. 2,3 However, only 10% to 15% of lifelong smokers develop lung cancer indicating that genetic factors may play an important role in determining the susceptibility to lung cancer. 4 -6 Polymorphisms in metabolic enzymes, including phase II detoxifying enzymes, have been studied to investigate the association with the risk of lung cancer.
Cytochrome P450 1B1 (CYP1B1) gene is located on chr2p22-p21, which is involved in the metabolic activation of polycyclic aromatic hydrocarbons (PAHs) including benzo(a)pyrene and dimethylbenz(a)anthracene but with a product distribution that is distinct from CYP1A1. 7,8 Several lines of evidence suggest that CYP1B1 plays a role in carcinogenesis. CYP1B1 is commonly overexpressed in human malignancies 9,10 and activates a variety of carcinogens. For example, CYP1B1 catalyzes both the formation of dihydrodiols of specific PAHs and their subsequent oxidation to carcinogenic dihydrodiol epoxides. 11 In humans, CYP1B1 is genetically polymorphic and more than 50 single-nucleotide polymorphisms (SNPs) have been reported so far, of which certain deleterious mutations are associated with primary congenital glaucoma. 12 Of the most common SNPs of CYP1B1 gene, 4 have been reported to result in amino acid substitutions, including Arg by Gly at codon 48(rs10012), Ala by Ser at codon 119 (rs1056827), Leu by Val at codon 432 (rs1056836), and Asn by Ser at codon 453 (rs1800440).
To date, a lot of studies have reported the association between CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms and lung cancer risk, but the results remained controversial. 13 -25 Some original studies suggested that CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms were associated with lung cancer risk, but others had different opinions. In order to explore the association between CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms and lung cancer risk, a meta-analysis was conducted to summarize the data.
Materials and Methods
Identification and Eligibility of Relevant Studies
A comprehensive literature search was performed using the PubMed and EMBASE database for relevant articles published (the last search update was February 16, 2015) with the following key words “CYP1B1”, “cytochrome P-450 1B1”, or “cytochrome P450 1B1”, “polymorphism”, “variant”, or “mutation”, and “lung.” In addition, studies were identified by a manual search of the reference lists of reviews and retrieved studies. All eligible studies were retrieved, and their bibliographies were checked for other relevant publications.
Inclusion Criteria
The included studies have to meet the following criteria: (1) only the case–control studies or cohort studies were considered; (2) evaluated the CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms and the risk of lung cancer; and (3) the genotype distribution of CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms in cases and controls were described in details, and the results were expressed as odds ratio (OR) and corresponding 95% confidence interval (95% CI). Major reasons for exclusion of studies were as follows: (1) not for cancer research, (2) only case population, and (3) duplicate of previous publication.
Data Extraction
Information was carefully extracted from all eligible studies independently by 2 investigators according to the inclusion criteria listed earlier. The following data were collected from each study: first author’s name, year of publication, country, ethnicity, source of controls (population-based controls and hospital-based controls), sample size, and numbers of cases and controls in the CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser genotypes whenever possible. Ethnicity was categorized as “Caucasian”, “Asian”, and “African”. When one study did not state which ethnic groups was included or if it was impossible to separate participants according to phenotype, the sample was termed as “mixed population”. Articles that reported different ethnic groups and different countries or locations were considered different study samples for each category cited earlier.
Statistical Analysis
Crude ORs together with their corresponding 95% confidence intervals (95% CIs) were used to assess the strength of association between the CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms and lung cancer risk. The pooled ORs were calculated for dominant model (Leu432Val: Leu/Val + Val/Val vs Leu/Leu, Asn453Ser: Asn/Ser + Ser/Ser vs Asn/Asn, and Ala119Ser: Ala/Ser + Ser/Ser vs Ala/Ala), recessive model (Leu432Val: Leu/Val + Leu/Leu vs Val/Val, Asn453Ser: Asn/Ser + Asn/Asn vs Ser/Ser, and Ala119Ser: Ala/Ser + Ala/Ala vs Ser/Ser), and additive model (Leu432Val: Leu/Leu vs Val/Val, Asn453Ser: Asn/Asn vs Ser/Ser, and Ala119Ser: Ala/Ala vs Ser/Ser), respectively. Between-study heterogeneity was assessed by calculating Q-statistic (heterogeneity was considered statistically significant if P < 0.10) 26 and quantified using the I 2 value, Venice criteria 27 for the I 2 test included: “I 2 < 25% represents no heterogeneity, I 2 = 25% to 50% represents moderate heterogeneity, I 2 = 50% to 75% represents large heterogeneity, and I 2 > 75% represents extreme heterogeneity”. If results were not heterogeneous, the pooled ORs were calculated by the fixed effect model (we used the Q-statistic, which represents the magnitude of heterogeneity between studies). 28 Otherwise, a random effect model was used when the heterogeneity between studies was significant. 29 We also conducted subgroup analysis by ethnicity (Caucasian and Asian) and source of controls. Moreover, sensitivity analysis was performed by excluding a single study each time. In addition, we also ranked studies according to sample size, and then repeated this meta-analysis. Sample size was classified according to a minimum of 200 participants and those with fewer than 200 participants. 30 Hardy-Weinberg equilibrium (HWE) was calculated using the goodness-of-fit test, and deviation was considered when P < .01. Begg funnel plots 31 and Egger linear regression test 32 were used to assess publication bias. A meta-regression analysis was carried out to identify the major sources of between-studies variation in the results, using the log of the ORs from each study as dependent variables, and ethnicity, source of controls, and sample size as the possible sources of heterogeneity. All the calculations were performed using STATA version 10.0 (STATA Corporation, College Station, Texas).
Results
Literature Search and Meta-Analysis Databases
Figure 1 provides a flow chart for this meta-analysis. A total of 72 publications were identified, among which 33 irrelevant articles were excluded. Thus, 39 publications were eligible. Another 26 articles among these publications were excluded because they were review articles, case reports, and other polymorphisms of CYP1B1. In addition, the study of Cote et al 13 was excluded because patients were included in the study of Cote et al. 17 As summarized in Table 1, 12 publications with 22 case–control studies were included in the meta-analysis, including 2881 cases and 3653 controls for CYP1B1 Leu432Val polymorphism (from 13 studies), 3009 cases and 3887 controls for CYP1B1 Asn453Ser polymorphism (from 5 studies), and 1301 cases and 2045 controls for CYP1B1 Ala119Ser polymorphism (from 4 studies). Seven studies included enough information in a subgroup analysis by smoking status. 14,15,18 -22 The distribution of genotypes in the controls was consistent with Hardy-Weinberg equilibrium in all studies. All the cases were pathologically confirmed.

Study flow chart explaining the selection of the 12 articles included in the meta-analysis.
Main Characteristics of All Studies Included in the Meta-Analysis.

Abbreviations: PB, population-based study; HB, hospital-based study; Y, yes; SC, source of controls; HWE, Hardy-Weinberg equilibrium; SNP, single-nucleotide polymorphism.
Meta-Analysis Results
Table 2 lists the main results of the meta-analysis of CYP1B1 Leu432Val polymorphism and lung cancer risk. Overall, significant association was observed in all genetic models (dominant model: OR = 1.29, 95% CI = 1.08-1.53, P h = 0.029, I 2 = 47.6%, Figure 2; recessive model: OR = 1.21, 95% CI = 1.05-1.39, P h = 0.694, I 2 = 0.0%, Figure 3; additive model: OR = 1.43, 95% CI = 1.21-1.69, P h = 0.144, I 2 = 30.9%, Figure 4) when all the eligible studies were pooled into the meta-analysis. And in the subgroup analysis, significantly increased lung cancer risk was observed in Caucasians (dominant model: OR = 1.38, 95% CI = 1.06-1.79, P h = 0.021, I 2 = 65.3%; recessive model: OR = 1.19, 95% CI = 1.02-1.39, P h = 0.288, I 2 = 19.9%; additive model: OR = 1.47, 95% CI = 1.07-2.02, P h = 0.031, I 2 = 62.3%), smokers (dominant model: OR = 1.26, 95% CI = 1.04-1.52, P h = 0.743, I 2 = 0.0%), and nonsmokers (recessive model: OR = 1.37, 95% CI = 1.02-1.85, P h = 0.176, I 2 = 39.2%).

Forest plot of cytochrome P450 1B1 (CYP1B1) Leu432Val polymorphism and lung cancer risk in overall analysis (dominant model).
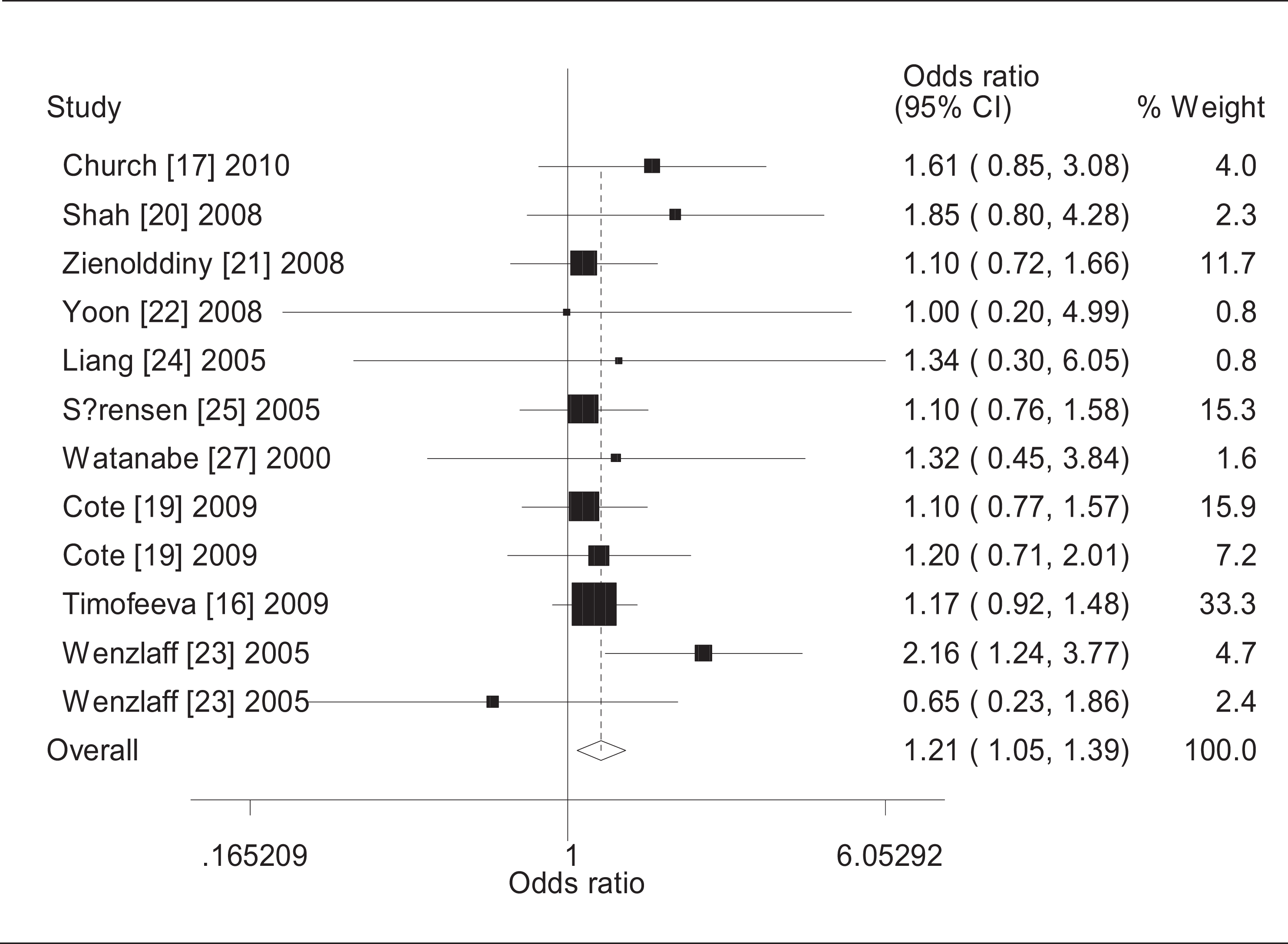
Forest plot of cytochrome P450 1B1 (CYP1B1) Leu432Val polymorphism and lung cancer risk in overall analysis (recessive model).

Forest plot of cytochrome P450 1B1 (CYP1B1) Leu432Val polymorphism and lung cancer risk in overall analysis (additive model).
Stratified Analysis of CYP1B1 Leu432Val Polymorphism on Risk of Lung Cancer.
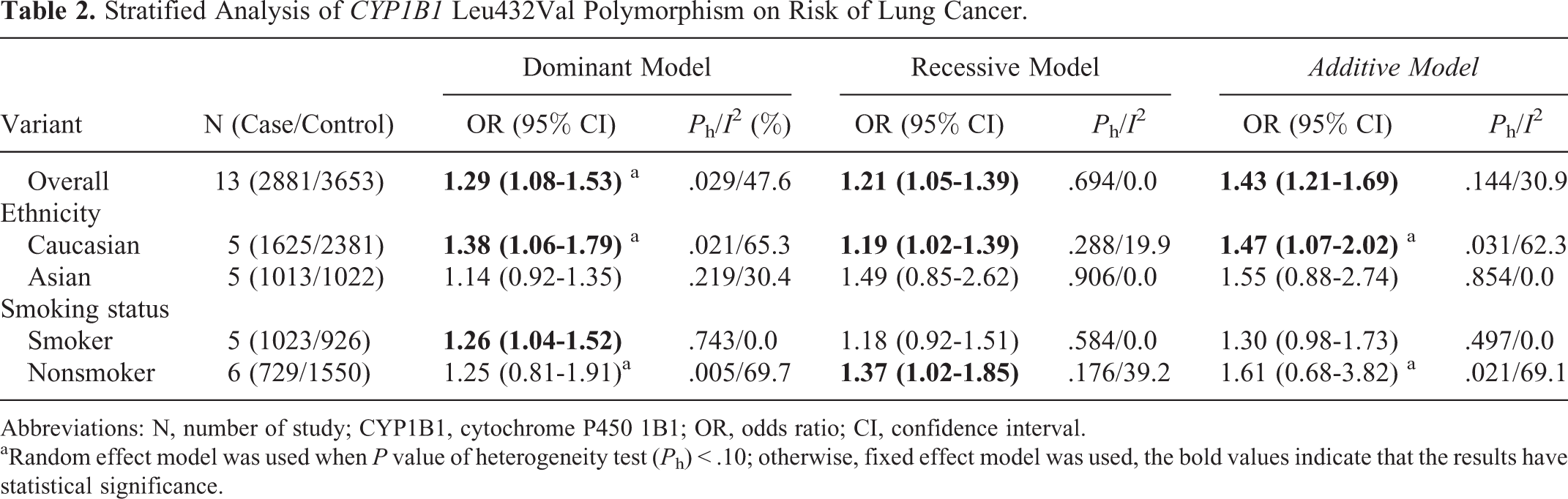
Abbreviations: N, number of study; CYP1B1, cytochrome P450 1B1; OR, odds ratio; CI, confidence interval.
aRandom effect model was used when P value of heterogeneity test (P h) < .10; otherwise, fixed effect model was used, the bold values indicate that the results have statistical significance.
Table 3 lists the main results of the meta-analysis of CYP1B1 Asn453Ser polymorphism and lung cancer risk. Overall, no significant association was observed between CYP1B1 Asn453Ser polymorphism and lung cancer risk (dominant model: OR = 0.98, 95% CI = 0.89-1.09, P h = 0.112, I 2 = 46.7%; recessive model: OR = 1.21, 95% CI = 0.96-1.54, P h = 0.480, I 2 = 0.0%; additive model: OR = 1.20, 95% CI = 0.94-1.52, P h = 0.470, I 2 = 0.0%).
Stratified Analysis of CYP1B1 Asn453Ser Polymorphism on Risk of Lung Cancer.
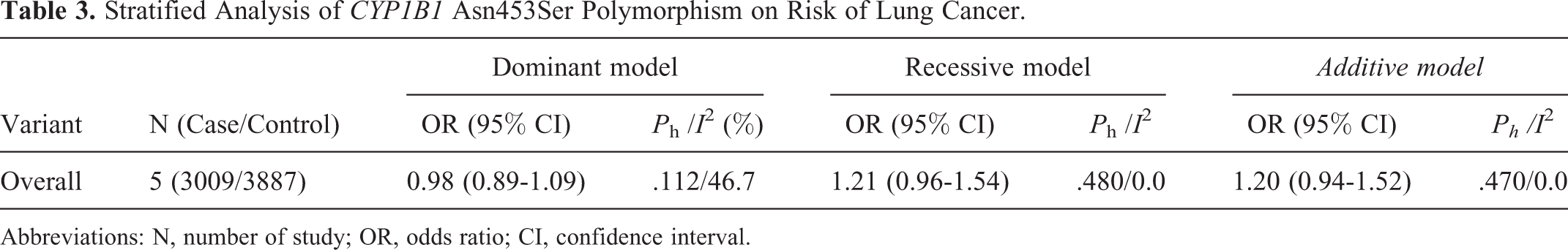
Abbreviations: N, number of study; OR, odds ratio; CI, confidence interval.
Table 4 lists the main results of the meta-analysis of CYP1B1 Ala119Ser polymorphism and lung cancer risk. Overall, no significant association was observed between CYP1B1 Ala119Ser polymorphism and lung cancer risk (dominant model: OR = 1.05, 95% CI = 0.91-1.21, P h = 0.238, I 2 = 29.1%; recessive model: OR = 0.99, 95% CI = 0.53-1.85, P h = 0.013, I 2 = 72.3%; additive model: OR = 1.10, 95% CI = 0.84-1.44, P h = 0.027, I 2 = 67.4%).
Stratified Analysis of CYP1B1 Asn453Ser Polymorphism on Lung Cancer Risk.

Abbreviation: N, number of study; OR, odds ratio; CI, confidence interval.
aRandom effect model was used when P value of heterogeneity test (P h) < .10; otherwise, fixed effect model was used, the bold values indicate that the results have statistical significance.
Sensitivity Analysis
Sensitivity analysis was conducted to determine whether modification of the inclusion criteria of this meta-analysis affected the results. When the study of Wenzlaff et al 21 was excluded, the results were changed between CYP1B1 Leu432Val polymorphism and lung cancer risk in nonsmokers (recessive model: OR = 1.12, 95% CI = 0.78-1.62, P h = 0.536, I 2 = 0.0%). Although the sample size for cases and controls in all eligible studies ranged from 60 to 3897, the corresponding pooled ORs were not qualitatively altered with or without the study of small sample.
Publication Bias
Both Begg’s funnel plot and Egger’s test were performed to access the publication bias of this meta-analysis. Begg’s funnel plots did not reveal any evidence of obvious asymmetry in the overall meta-analysis (Figures 5 -7). Egger’s test results also suggested no evidence of publication bias in the meta-analysis of CYP1B1 Leu432Val (P = 0.589 for dominant model, P = 0.586 for recessive model, P = 0.722 for additive model), Asn453Ser (P = 0.419 for dominant model, P = 0.329 for recessive model, P = 0.309 for additive model), and Ala119Ser polymorphisms (P = 0.719 for dominant model, P = 0.689 for recessive model, P = 0.759 for additive model), indicating that our results were statistically robust.

Begg funnel plot of the meta-analysis of cytochrome P450 1B1 (CYP1B1) Leu432Val polymorphism and lung cancer risk (recessive model and dominant model).

Begg funnel plot of the meta-analysis of cytochrome P450 1B1 (CYP1B1) Asn453Ser polymorphism and lung cancer risk (recessive model and dominant model).

Begg funnel plot of the meta-analysis of cytochrome P450 1B1 (CYP1B1) Ala119Ser polymorphism and lung cancer risk (recessive model and dominant model).
Discussion
CYP1B1 is commonly overexpressed in human malignancies and activates a variety of carcinogens. For example, CYP1B1 catalyzes both the formation of dihydrodiols of specific PAHs and their subsequent oxidation to carcinogenic dihydrodiol epoxides. The importance of CYP1B1 in chemical carcinogens is well illustrated in animal models in which metabolites of CYP1B1 were shown to induce PCa. 33,34 Furthermore, CYP1B1-null mice, created by targeted gene disruption in embryonic stem cells, were protected from 7, 12-dimethylbenz(a)anthracene-induced malignant lymphomas. 7 Many studies have reported the role of CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser with lung cancer risk, but the results remained controversial. Although 1 study 35 previously performed pooling analysis regarding this issue, there were some mistakes when they collected data of all the eligible studies. 35 In order to derive a more precise estimation of association, we performed the meta-analysis of CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms with lung cancer risk.
Overall, significant association was observed in all genetic models when all the eligible studies were pooled into the meta-analysis of CYP1B1 Leu432Val. In the stratified analysis by ethnicity, we found that CYP1B1 Leu432Val polymorphism was associated with lung cancer risk in Caucasians, but not in Asians. For the same histological types of lung cancer, suggesting a possible role of ethnic difference in genetic background and the environment they lived in. The same gene polymorphisms may play different roles between cancer susceptibility and different ethnic population because cancer is a complicated multigenetic disease and different genetic backgrounds may contribute to the discrepancy. 36 These findings indicate that CYP1B1 Leu432Val polymorphism may be important in specific ethnicity of lung cancer, especially in Caucasians. There is a consensus that tobacco smoking is the major cause of lung cancer and tobacco smoke contains some carcinogens that induce CYP1B1, which may contribute to the development of lung cancer. Several studies have studied the interaction or joint effect of CYP1B1 Leu432Val polymorphism and smoking status on lung cancer. 14,15,18,19,22 In the present meta-analysis, we found that CYP1B1 Leu432Val polymorphism had an increased lung cancer risk in smokers. It is possible that individuals with CYP1B1 Leu432Val polymorphism were more susceptible to smoking-related lung cancer when being exposed to higher level of tobacco smoke. Heterogeneity is a potential problem when interpreting the results of all meta-analyses. After a careful look at our study, we found that the study of Wenzlaff et al 21 was noted to be a major source of heterogeneity for CYP1B1 Leu432Val polymorphism. The reason may be that the study of Wenzlaff et al 21 was only in nonsmokers. When the study of Wenzlaff et al 21 was deleted, the results of CYP1B1 Leu432Val were changed in nonsmokers (recessive model: OR = 1.12, 95% CI = 0.78-1.62). The results indicated nonsmokers may be not associated between CYP1B1 Leu432Val polymorphism and lung cancer risk.
We have read with great interest the article “Current evidence on the relationship between CYP1B1 polymorphisms and lung cancer risk: a meta-analysis” published online in June 2011 issue of “Molecular Biology Reports”. 35 Xu et al 35 included 10 case–control studies for Leu432Val, 4 case–control studies for Asn453Ser, and 3 case–control studies for Ala119Ser. Their meta-analysis suggested that the CYP1B1 Leu432Val and Ala119Ser genotypes were low-penetrance risk factors for developing lung cancer. However, the data reported by Xu et al 35 for the study by Liang et al 22 do not seem in line with the data provided by Liang et al 22 in their original publication. The numbers reported by Xu et al 35 for Leu/Leu, Leu/Val, Val/Val, in cases and controls, respectively, are 158-66-3 and 154-69-4. Interestingly enough, after carefully studying the data presented by Liang et al, 22 the frequencies that we have retrieved on the 227 cases and 227 controls were 154-69-4 and 158-66-3, respectively. Adopting the same search strategy as Xu et al, 35 we identified 3 eligible studies, 15,19,24 which have not been included in the meta-analysis of Xu et al 35 for Leu432Val, one eligible study, 15 which has not been included in the meta-analysis of Xu et al 35 for Asn453Ser, and 2 eligible studies 15,19 that have not been included in the meta-analysis of Xu et al 35 for Ala119Ser. Worthy of note, these 6 studies included 2029 samples. Importantly, of the most common SNPs of CYP1B1 gene, 4 have been reported to result in amino acid substitutions, including Arg by Gly at codon 48 (CYP1B1*2), Ala by Ser at codon 119 (CYP1B1*2), Leu by Val at codon 432 (CYP1B1*3), and Asn by Ser at codon 453 (CYP1B1*4). We found that the study of Shah et al 18 provides only the combined data of CYP1B1*2 ((Arg48Gly and Ala119Ser) but does not separately provide the data of Arg48Gly and Ala119Ser. Hence, the study of Shah et al 18 should be excluded in the meta-analysis of Xu et al 35 for Ala119Ser and Arg48Gly polymorphism. In conclusion, we thought that the ongoing uncertainty still exists and the conclusion by Xu et al 35 was not entirely credible. We have also read with great interest the recent meta-analysis by Chen et al, 37 the study of Chen et al 37 included 9 published studies for Leu432Val. However, the study of Cote et al 13 should be excluded because their populations overlapped with another included study. 17 Hence, the study of Chen et al 37 had only 8 published studies. Although 2 studies 35,37 previously performed pooling analysis regarding this issue, several published studies were not included in these analysis and additional original studies with larger sample sizes have been published since then. Having analyzed an almost 2-fold larger number of studies than the previous meta-analysis, 35 our results seem to confirm and establish the trend in the meta-analysis of the CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms that the data according to the previous meta-analysis 35,37 had indicated. Our meta-analysis indicates that CYP1B1 Leu432Val shows an increased risk of lung cancer in caucasians and smokers, and Asn453Ser and Ala119Ser may not be associated with lung cancer risk.
There are several limitations in this meta-analysis. First, the controls were not uniformly defined. Although most of them were common populations, some controls were population based and other controls were hospital based. Hence, nondifferential misclassification bias is possible. Second, in the subgroup, analysis may have had insufficient statistical power to check an association, Third, we were also unable to examine the interactions among gene and environment, and lack of the original data of the included studies limited our further evaluation of potential interactions, which may be an important component of the association between CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms, environment, and risk of lung cancer. Fourth, it was much difficult to get all the articles published in various languages. Finally, our results were based on unadjusted published estimates. Because of data limitations, such as age and alcohol consumption, and so on, we were unable to adjust them. Our meta-analysis also has several strengths. First, a systematic review of the association of CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms with lung cancer risk is statistically more powerful than any single study. Second, the quality of eligible studies included in the current meta-analysis was satisfactory and met our inclusion criterion. Third, we did not detect any publication bias indicating that the whole pooled results should be unbiased.
In summary, this meta-analysis suggests that CYP1B1 Leu432Val is associated with increased lung cancer susceptibility among Caucasians, and there is an interaction between CYP1B1 Leu432Val polymorphism and smoking, and Asn453Ser and Ala119Ser may be not associated with lung cancer risk. However, a study with the larger sample size is needed to further evaluate gene–environment interaction on CYP1B1 Leu432Val, Asn453Ser, and Ala119Ser polymorphisms and risk of lung cancer.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
