Abstract
Background:
Leucine zipper protein (LUZP) plays key roles in development. Overexpression of LUZP was documented in several types of solid tumors. In this study, expression of LUZP messenger RNA (LUZP mRNA) in human hepatocellular carcinoma (HCC) was examined, and the correlations of LUZP mRNA level with patients’ characteristics and prognosis were also investigated.
Methods:
Total RNA was extracted from HCC and paired noncancerous liver tissues of 77 patients. Expression of LUZP mRNA in the tissues was determined by real-time quantitative reverse transcriptase polymerase chain reaction. Using average LUZP mRNA level in noncancerous liver tissues as the cutoff, patients with HCC were categorized into high-expression group and low-expression group. Correlations of LUZP mRNA with clinical parameters were analyzed. Overall survival of the patients in the 2 groups was analyzed by Kaplan-Meier method.
Results:
The LUZP mRNA level was significantly higher in HCC samples than in the noncancerous liver tissues (1.87 ± 0.11 vs 0.58 ± 0.05, P < .01). Significant differences were found between the 2 groups in terms of portal vein invasion, Tumor Lymph Node Metastasis (TNM) stage, and recurrence of HCC. The current study failed to find significant differences between the 2 groups in clinical characteristics such as age, gender, lymph node metastasis, hepatitis B virus infection, family HCC history, and alcohol intake. Overall survival in high-expression group was 12 months while that in the low-expression group was 34 months (P = .03).
Conclusion:
The LUZP mRNA is a prognostic indicator in HCC, and overexpression is associated with poor prognosis in patients with HCC.
Introduction
Hepatocellular carcinoma (HCC) is the fifth most common cancer and the third leading cause of cancer-related death worldwide. 1 Surgical resection of the tumor offers a chance for cure but does not achieve optimal long-term survival, mainly due to recurrence and metastasis. 2 In many cases, particularly in China, HCC develops on a background of an underlying liver disease such as chronic hepatitis, alcoholism, and exposure to hepatotoxins. 3,4 Risk factors such as hepatitis B virus (HBV) and hepatitis C virus (HCV) infection may cause genetic or epigenetic disorder, thus causing development of HCC. 5,6
Leucine zipper protein (LUZP), also known as LUZP 1, is a protein containing leucine zipper motif. The leucine zipper motif is a super-secondary structure that functions as a dimerization domain. 7,8 A single leucine zipper consists of multiple leucine residues at approximately 7-residue intervals, which forms an amphipathic α-helix with a hydrophobic region running along one side. This hydrophobic region provides an area for dimerization, allowing the motifs to “zip” together. Furthermore, the hydrophobic leucine region is absolutely required for DNA binding. Some leucine zipper motif containing proteins such as leucine zipper downregulated in cancer 1 have an important role in the progression of some cancers, but the exact function of the LUZP-encoded protein is not known. It has been documented that LUZP is predominantly expressed in the brain. 9 In mice, LUZP affects neural tube closure. The LUZP-deficient embryos display cranial neural tube closure defect during brain development. 10 Overproduction or mutation of LUZP may cause cancer. Abnormal expression of LUZP protein is found in many types of tumor such as breast cancer, kidney cancer, cervical cancer, and pancreatic cancer. 11 –14
In this study, we examined expression of LUZP messenger RNA (LUZP mRNA) in tumor versus adjacent noncancerous liver tissues in 77 patients with HCC who received surgical resection. Potential correlations of expression of LUZP mRNA with clinicopathological characteristics of HCC (eg, TNM staging and survival) were also analyzed.
Materials and Methods
Ethics Statement
The study protocol was approved by the Ethics Committee of Provincial Hospital Affiliated to Shandong University. All the participants provided their written informed consent for inclusion in the data analysis and publication of the article.
Sample Collection
Seventy-seven patients with HCC were enrolled in the study, who received surgical treatment in the Department of Hepatobiliary Surgery and Center of Liver Transplantation, Shandong Provincial Hospital Affiliated to Shandong University, during September 2010 to January 2012. Diagnosis was confirmed by pathological examination of resected liver tissues. Clinical data were recorded including age, gender, alpha fetoprotein (α-FP) level, tumor size, portal vein invasion, lymph node metastasis, alcohol intake, TNM stage (TNM 3 and 4 as advanced stage), family HCC history, recurrence, HBV infection, HCV infection, history of aflatoxin B1 exposure, and overall survival time. Hepatocellular carcinoma and noncancerous liver samples were collected and stored in liquid nitrogen until use. Noncancerous liver tissue was defined as liver tissue 5 cm or more away from the tumor border. Both cancer and noncancerous liver tissues were verified by a pathologist blinded to the tissue origin using hematoxylin–eosin staining.
RNA Extraction and First Strand Complementary DNA Synthesis
Total RNA was extracted from frozen tissues (30 mg) using AllPrep DNA/RNA Mini Kit (Qiagen, Valencia, California) according to the manufacturer’s instruction. Genomic DNA was removed using DNA-free kit (Applied Biosystems, Carlsbad, California) following the protocol. The integrity of RNA was verified using electrophoresis with 5% agarose/formaldehyde/3-(N-Morpholino) propanesulfonic acid gels followed by ethidium bromide staining and visual inspection under ultraviolet light. Samples with the peaks of 18s and 28s ribosomal RNAs at less than twice the other peak were excluded. Concentration of RNA was measured by Nanodrop ND-2000 spectrophotometer (Thermo Fisher Scientific Inc, Waltham, Massachusetts). The purity of RNA was examined by calculating the ratio of absorbance at 260 nm versus 280 nm. A ratio of ∼2.0 was used as the criterion for “pure” RNA. The quality and integrity of RNA were similar tumor and noncancerous tissue samples. First strand complementary DNA (cDNA) was synthesized from 1 μg of total RNA using high-capacity RNA-to-cDNA kit (Applied Biosystems, Carlsbad, California) according to the manufacturer’s protocol and under the following conditions: 37°C for 60 minutes and 95°C for 5 minutes. First strand cDNA was stored at −20°C until further analysis.
Quantitative Real-Time Polymerase Chain Reaction
Relative level of LUZP mRNA in HCC and adjacent noncancerous liver tissues was determined by quantitative reverse transcriptase polymerase chain reaction (qPCR). β-Actin was used as the internal loading control. The qPCR was performed on an ABI StepOne Plus Real-Time PCR System (Applied Biosystems, Foster City, California). The primer sequence was as follows: LUZP: forward, 5′-CGG AAG TAT TGA CGG AAA GC-3′; reverse, 5′-GCT TGT AAA TTC GGC CAT GT-3′. β-Actin: forward, 5′-GGA CTT CGA GCA AGA GAT GG-3′; reverse, 5′-AGC ACT GTG TTG GCG TAC AG-3′. All samples were run in triplicate. The qPCR amplification was conducted in 20 μL of reaction buffer using ABI Power SYBRGreen PCR Master Mix (Applied Biosystems, Foster City, California) under the following conditions: 95°C for 10 minutes 40 cycles at 95°C for 15 seconds, and 60°C for 1 minute. The melting curve was analyzed for each sample. The amplification products were separated using electrophoresis on 2% agarose gels and visualized by ethidium bromide staining. The expected size of LUZP is 237 bp. The threshold cycle (Ct) was measured in the exponential amplification phase. The amplification plots were analyzed by StepOne v2.2 software (Applied Biosystems, Foster City, California). The Ct values of β-actin were similar across HCC and noncancerous liver tissues. The results were normalized against β-actin and expressed as 2−ΔΔCt.
Immunohistochemical Analysis
Immunohistochemical analysis was performed as described. 15 Briefly, 5-μm-thick paraffin-embedded tissue sections were deparaffinized with xylene and rehydrated with a graded series of ethanol followed by antigen retrieval with 0.1 mol/L sodium citrate buffer (pH 6.0). Expression of Luzp was detected with a rabbit polyclonal antibody against human luzp (ab97855, 1:1000; Abcam, Cambridge, United Kingdom). Two pathologists, unknown to the type of the sections, independently examined the photographs of immunohistochemical-stained sections under a camera mounted on a Keyence BZ-8000 digital microscope (Keyence, Osaka, Japan). A semiquantitative immunoreactivity score was used to evaluate the expression of luzp. 16
Patient Stratification and FollowUp
Using the average LUZP mRNA level in noncancerous liver tissue samples as the cutoff, patients were stratified into 2 groups, a high-expression group and a low-expression group. The follow-up data were obtained from the medical records and direct communication with the patients or their relatives. The follow-up period was defined as the time from the date of surgery to the date of patient’s death or the last follow-up in October 2013.
Statistical Methods
All statistical analyses were carried out using SPSS version 13.0 for Windows (SPSS Inc., Illinois). Quantitative values were presented as mean ± standard deviation. Correlation analysis between clinical variables and expression of LUZP was carried out using Fisher exact and chi-square tests. The distribution curves of survival and time of high-expression and low-expression groups were analyzed using the Kaplan-Meier method, and a log-rank test was used to compute the differences between the curves. Prognostic value of expression of LUZP was examined with a multivariate analysis, using the Cox proportional hazards regression model. A P value of less than .05 was set as statistically significant.
Results
The median age of the 77 patients was 48.5 years (range: 24-73 years). Sixty-seven patients were men (67 of 77, 87%), and the remaining(10 of 77, 13%) were women. None of the patients had HCV infection and had not been exposed to aflatoxin B1. All patients were followed up till the article was prepared or till their death. All deaths were associated with recurrence of HCC.
The relative level of LUZP mRNA in HCC tissue was 1.87 ± 0.11 (the ratio against β-actin mRNA level) while that in paired noncancerous liver tissue was 0.58 ± 0.05 (P < .01; Figure 1). Using the average level of LUZP mRNA in paired noncancerous liver tissues as cutoff, which was 0.24 ± 0.02, a total of 49 patients were grouped into high-expression group in which the level of LUZP mRNA was above the cutoff. The other 28 patients were grouped in the low-expression group in which the level of LUZP mRNA was below the cutoff (Table 1). Expression of luzp in the cancer and noncancerous liver tissues was verified by immunohistochemical staining. The results showed that expression of luzp was significantly higher in HCC compared to the matched noncancerous tissue that was consistent with expression of LUZP mRNA (P < .01; Figure 2).
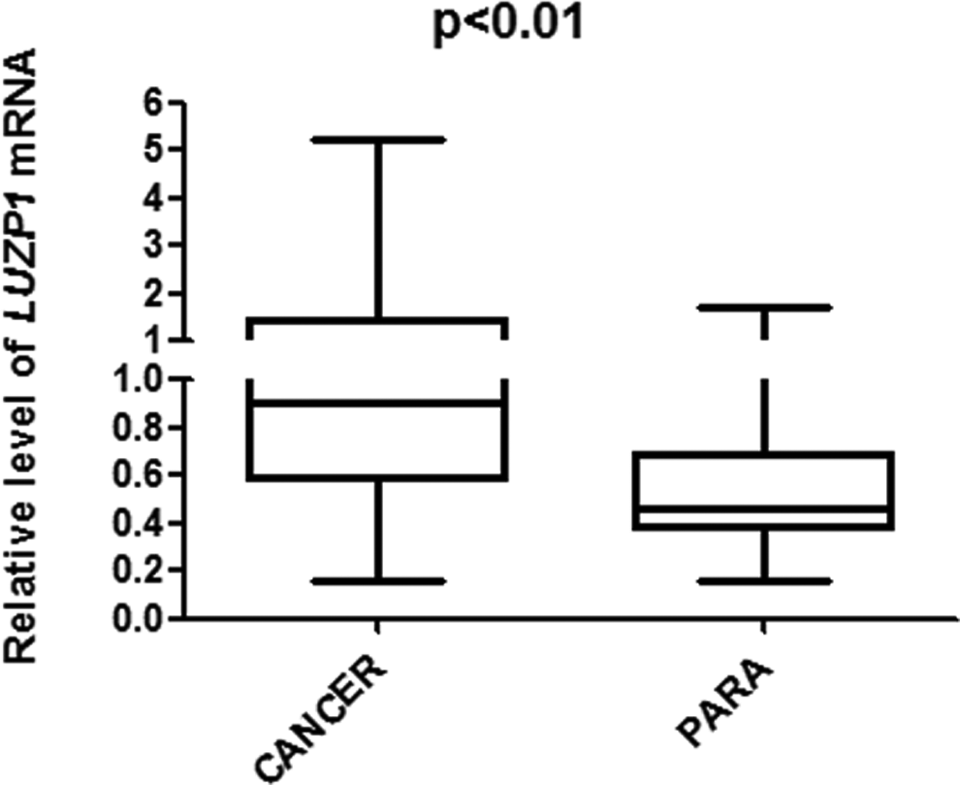
LUZP mRNA in HCC and paired noncancerous liver tissues. LUZP mRNA was detected in HCC and noncancerous liver tissues with qPCR in 77 patients with HCC. Data are presented as the abundance relative to β-actin mRNA. LUZP mRNA in HCC tissues was significantly higher than that in the noncancerous liver tissues (P < .01). LUZP indicates leucine zipper protein; mRNA, messenger RNA; qPCR, quantitative reverse transcriptase polymerase chain reaction; HCC, hepatocellular carcinoma.
Correlations of LUZP mRNA Expression With the Clinicopathological Features of HCC.

Abbreviations: HCC, hepatocellular carcinoma; mRNA, messenger RNA; α-FP, alpha fetoprotein.

Expression of luzp in HCC and paired noncancerous liver tissues. Expression of luzp in HCC and noncancerous liver tissues was verified by immunohistochemistry. Results showed that luzp in HCC tissues was significantly higher than that in the noncancerous liver tissues (P < .01). A, Representative luzp-positive expression in HCC tissues. B, Representative luzp-negative expression in noncancerous tissues. Scale bar, 200 μm and magnified ×200. C, The distribution of the difference of luzp staining (▵IRS = IRS T − IRS N). P values were calculated with the Wilcoxon test. IRS indicates immunoreactivity score; HCC, hepatocellular carcinoma.
Significant differences were found between the 2 groups in terms of portal vein invasion, TNM stage, and recurrence of HCC. Portal vein invasion was found in 23 patients in the high-expression group and 15 patients in the low-expression group (P = .02). With regard to the cancer stage, we found that 30 patients in the high-expression group were in advanced stage (TNM3/4) and the other 19 patients were in TNM1 and TNM2 stages when they considered for surgical treatment. In the low-expression group, 3 patients were in TNM3/4 stage and 25 patients were in stage TNM1/2 (P < .0001). Recurrence of HCC was found in 40 patients in the high-expression group, while only 11 patients had recurrence in the low-expression group during the follow-up (P = .003; Table 1).
No significant differences were found between the 2 groups concerning age, gender, lymphnode metastasis, HBV infection, α-FP level, tumor diameter, family history of HCC, and alcohol intake (Table 1).
Overall survival times were analyzed with Kaplan-Meier method. The survival time was 32 months in the LUZP low-expression group while it was 12 months in the high-expression group (Figure 3). The results showed that the patients in the high-expression group have a short survival time than those in the low-expression group (P = .03, hazard ratio; 1.88, 95% confidence interval; 1.08-3.30).

Kaplan-Meier survival analysis stratified by expression of LUZP. Overall survival was compared between the 2 groups of the patients with HCC using the Kaplan-Meier curves. The overall survival of patients with high LUZP expression was significantly lower than that in patients with low expression (P = .03). LUZP indicates leucine zipper protein; HCC, hepatocellular carcinoma.
Discussion
Hepatocellular carcinoma is currently the fifth most common solid tumor worldwide and the second leading cause of cancer-related deaths in China. Surgery is one of the curative treatments to HCC, 17 but the rate of 5-year survival is below 12%. 18 Recurrence is the risk factor for low survival rate, and hence it is necessary to find the biomarker of prognosis.
In this study, we found that LUZP mRNA was overexpressed in HCC tissues. Luzp is a regulatory protein encoded by LUZP. It contains a segment of 20 amino acids that is rich in basic residues at the amino (N)-terminal flanking region of the ZIP motif. 19 The segment that comprises the basic region and the periodic arrays of leucine residues is referred to as “basic region leucine zipper” or bZIP motif. This basic region has been thought to facilitate DNA binding by interacting with the negatively charged phosphate groups in DNA. Krebs et al found that luzp is a bona fide subunit of the meta-coactivator complex, which enhances formation of the preinitiation complex, leading to transcription initiation. 20 Hsu et al found that knockout of LUZP in mice causes perinatal death, and the LUZP (−/−) embryos displayed a cranial neural tube closure defect. 10 Those studies indicate that LUZP may play a key role in the nervous system development. Overexpression of LUZP or mutation in a vital area of LUZP may cause cancer. 14 Dysfunction of LUZP was found in cervical cancer, breast cancer, and pancreatic carcinoma. 11,13,14 In the current study, we found that LUZP was overexpressed in HCC.
Hepatocarcinogenesis is a complex process that involves accumulation of genetic and epigenetic changes. 6 Major risk factors for HCC include HBV or HCV infection and alcoholic liver disease. 21 In China, HBV is the major risk factor for HCC. The HBV–DNA integration is frequently detected in HBV-positive HCC cells. 22 . Murakami et al demonstrated that HBV integration could lead to aberrant target gene transcription and cause hepatocarcinogenesis. 23 The LUZP can regulate the expression of DNA. Upregulation of LUZP may facilitate the development of HCC. But the current study failed to show significant difference in the HBV infection rate between the high- and low-expression groups. The role of LUZP in hepatocarcinogenesis needs further investigation.
Overexpression of LUZP is associated with poor prognosis in patients with breast cancer and pancreatic carcinoma. 13,14 Our results showed an association of high LUZP expression with more aggressive tumor behavior, including higher α-FP level, advanced TNM stage, and higher recurrence incidence. High α-FP level and advanced TNM stage are clinicopathologic markers of invasiveness and unfavorable prognosis in patients with HCC. 24 –26 Consistently, the results of this study showed an association of high LUZP expression with shorter overall survival.
However, our study failed to identify a link between expression of LUZP and vascular invasion or lymph node metastasis. We also failed to show an association of expression of LUZP with gender, age, or alcohol intake. Such findings may reflect the limited sample size, and further studies are needed.
In summary, the current study established a correlation between high LUZP expression in the tumor tissue and poor prognosis in patients with HCC. Further studies are required to investigate the role of LUZP in the progression of HCC.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article: The work was supported by the National Natural Science Foundation of China (No. 81373172).
