Abstract
Rising global cancer rates are projected to significantly increase by 2050, highlighting the urgent need for improved scalable prevention, early detection, and personalized therapy tools. Artificial intelligence (AI) has demonstrated significant capabilities in diverse oncology tasks, leveraging high-dimensional data from medical imaging, molecular profiles, and electronic health records for applications in radiology, digital pathology, genomics, prognostication, and treatment selection. Nevertheless, the clinical adoption of most AI systems is still limited by the black box issue, that is, prediction without clear explanation, which, in turn, limits the confidence and accountability of clinicians as well as their ability to communicate with patients. In this review, we searched sources over the years (2015-2025) from PubMed, Scopus, and Web of Science for evidence on explainable AI (XAI) methodologies that may provide greater interpretability and trust in oncologic practice. Local interpretable model-agnostic explanation and Shapley additive explanations (LIME and SHAP) are model-agnostic methods that offer local and global feature attribution and help clinicians to understand the main influential factors behind model predictions. The complementary approaches, such as Gradient-weighted Class Activation Mapping (Grad-CAM), Integrated Gradients and DeepLift, also bring the explainability to image- and genomics-based processes, whereas more recent strategies (eg, Anchors, Prototypical Part Network (ProtoPNet), and contrastive or counterfactual explanations) also focus on enhancing stability and clinical utility. Irrespective of such developments, several issues continue to be experienced, including computational load, inconsistency in explanations, domain transfer, deployment into clinical processes, bias, privacy issues, and changing regulatory requirements. In general, XAI can transform oncology AI to become clinically interpretable, transparent prediction of outcomes, which will make its application safer by adhering to strict validation procedures, human control, and patient-centered communication. By providing a comprehensive and clinically grounded overview, this review aims to support researchers, clinicians, and stakeholders in advancing trustworthy and transparent AI deployment in oncology.
Plain Language Summary Title
Overcoming the Black Box Challenge in Cancer Diagnosis and Care
Plain Language Summary
Cancer cases are expected to rise dramatically in the coming decades, creating an urgent need for better ways to detect and treat the disease. Artificial intelligence (AI) can support doctors by analyzing medical scans, genetic data, and health records to improve early diagnosis and guide treatment decisions. However, many AI tools work like “black boxes”—they provide results without showing how those results were reached. This lack of transparency makes it difficult for doctors and patients to fully trust AI recommendations.
Our review looks at “explainable AI” (XAI), a growing field that focuses on making AI decisions clearer and easier to understand. Methods such as Local Interpretable Model-Agnostic Explanations (LIME) and Shapley Additive Explanations (SHAP) can show which features or risk factors influenced a prediction, while other tools highlight important areas in medical images. These approaches help doctors validate AI findings, communicate more effectively with patients, and build confidence in using AI in real-world cancer care.
We conclude that explainable AI could make cancer treatment safer, more transparent, and more personalized. For this to happen, researchers, clinicians, and policymakers must work together to improve technical methods, set ethical standards, and ensure patients receive clear and understandable explanations.
Introduction
In the forthcoming decades, cancer incidence is anticipated to rise significantly due to causes such as exposure to pollutants, tobacco use, and unhealthy lifestyles. Recently, Bray et al 1 predicted in a demographics-based study that the global cancer incidence will increase to 35 million cases by 2050, representing a roughly 77% increase from the 2022 level. Nonetheless, Artificial intelligence (AI) could mitigate that increase by enhancing cancer prevention, facilitating early diagnosis, and personalizing cancer treatment. 2 AI tools can analyze extensive datasets, such as medical images, genetic profiles, and electronic health records, enhancing early detection rates and accurately identifying at-risk individuals. 3 Furthermore, AI-driven prediction models facilitate the formulation of treatment plans and drug development, yielding more efficacious medicines and improved patient outcomes. 4 Thus, AI has significantly transformed oncology, enabling earlier cancer detection and personalized treatment. However, AI's clinical integration is hindered by the so-called “black-box” problem, wherein the decision-making process behind AI predictions is opaque and difficult for clinicians to interpret and trust. 5 The lack of transparency in this regard is a major problem for clinicians who are required to comprehend, trust, and discuss AI-produced suggestions with patients. 6 For deep integration of AI into oncology, in which decisions have downstream consequences on patient lives, overcoming the black box challenge is crucial. This narrative review considers the implications of these challenges and outlines avenues to increase transparency and trust in the use of AI systems in oncology. This review explores the potential of XAI methods to enhance AI transparency in oncology, improve clinician trust, and enable more effective patient care.
A number of previous studies have examined XAI in healthcare or oncology contexts, but this literature tends to focus on a specific set of commonly used approaches (mostly SHAP, LIME, and Grad-CAM), mainly focuses on quantitative performance metrics, or lacks sufficient attention to clinical trust and rigorous validation as well as practical constraints of real-world applications. This present review attempts to overcome these shortcomings by providing an integrative synthesis of existing, as well as emerging, XAI approaches and methods: intrinsically interpretable modeling, counterfactual and exemplar-based modeling, global interpretability methods, and pragmatic toolkits, and contextualizing the discussion within the context of workflows and decision-making in the oncology field. For example, Mohamed et al 7 conducted a PRISMA-oriented systematic review of XAI use in diagnosis, prognosis, and therapeutic planning of cancer where the majority of research included the existing formats of explanation and highlighted their shortcomings, including insufficient clinician interaction. Conversely, the current review follows a trust-focused, deployment-based viewpoint, which focuses on human-AI cooperation, is more critical of the dangers to validity, and deals with ethical and regulatory readiness, and considers the less-represented methodological choices and toolchains such as Partial Dependence Plot and Individual Conditional Expectation (PDP-ICE), permutation feature importance, Explain Like I’m 5 (ELI5), and QLattice in addition to the traditional SHAP, LIME, and Grad-CAM strategies. Again, Cui et al 8 focused on the interpretability in radiology and radiation oncology but the present scope includes the full extent of oncology and a multimodal paradigm that goes beyond imaging and includes genomic, radiomic, tabular, and combined analytical pipelines. Although Sadeghi et al 7 present an overarching healthcare synthesis of XAI families, the current review will focus on the issues that are unique to oncology deployment specifically the dataset shift across hospitals and scanners, collinearity in features in radiomics and genomics, and the use of calibration, and offers a prospective framework demonstrating how such challenges can be overcome as the only viable path to trustful oncological AI.
To strengthen transparency despite the narrative orientation, the review is based on a PRISMA -ScR methodology and is supported by a modified checklist that is presented as an appendix. Through the combination of methodological breadth with robust clinical and translational outlook, this narrative review provides a differentiated and prospective framework that will be used to develop transparent, reliable, and clinically significant AI systems in the field of cancer.
Although the most popular techniques applied to tabular, general black-box, and vision-based models are SHAP, LIME, and Grad-CAM, respectively, limiting a review to these techniques alone risks diminishing important families of explainability techniques. Based on this, the current study takes a larger taxonomy of XAI methods applicable to healthcare decision-making. Besides feature-attribution and saliency approaches, we also have intrinsically interpretable models, example-based explanations, counterfactual and recourse-oriented explanations, concept based and attention based explanations, surrogate and rule-extraction explanations, uncertainty, calibration-based, and causality-informed explanations that affect the way explanations should be interpreted in practice. Incorporating global interpretability, practitioner toolkits, and interpretable-by-design modeling (QLattice) provide a broader and more implementation-oriented reference. This increased coverage is aimed at making the study useful to researchers, clinicians, and other stakeholders and retain the discussion within the framework of practical clinical implementation. The objectives of this review include to:
Thoroughly review the XAI techniques used in oncology, including current and novel techniques. Categorize and frame XAI techniques based on methodological underpinnings, and to their applicability to oncology-specific modalities of data. Appraise the capabilities, constraints, and clinical utility of the various XAI methods, with special emphasis on reliability, robustness, and interpretability in high-stakes oncologic decision-making. Determine the most important research paucities that impede the safe and effective translation of XAI into the regular practice in oncology. Provide future directions for XAI in oncology, such as collaboration between humans and AI, ethical and regulatory aspects, and both clinical workflow integration.
Through this focused synthesis, the review aims to inform researchers, clinicians, and stakeholders on best practices and unmet needs in building transparent and reliable AI models for oncology.
Methodology
This study was conducted as a narrative review designed to synthesize and contextualize XAI methods in oncology and did not follow a strict systematic review or meta-analysis protocol. The narrative review methodology was used to allow integrating heterogeneous approaches, data forms and application settings conceptually, which would not readily adapt to formal meta-analysis processes or strict systematic review guidelines. This narrative review prioritizes conceptual clarity, methodological breadth, and clinical relevance over exhaustive enumeration, which is appropriate given the heterogeneity and rapid evolution of XAI methods in oncology.
Ensuring Methodological Clarity, Transparency, and Reproducibility
Although formal systematic procedures (eg, protocol registration, duplicate independent screening, quantitative pooling) were not applied, several measures were implemented to ensure transparency and reproducibility for readers. First, the review process was explicitly structured and documented, including information sources, search terms, inclusion rationale, and synthesis strategy. Second, the review mimicked the PRISMA-ScR framework, which guided clear reporting of literature identification, selection rationale, and thematic organization, without implying full compliance with scoping or systematic review standards. Third, all methodological choices and limitations inherent to a narrative review design are explicitly acknowledged. A brief PRISMA-ScR–informed checklist summarizing the reporting elements addressed in this narrative review is provided as Supplemental Material 1 (Supplementary Table S1).
Literature Sources and Search Strategy
A literature search was carried out across reputable scientific databases related to the research field of oncology, biomedical studies, and artificial intelligence including PubMed, Scopus, and Web of Science for peer-reviewed articles published between 2015 and 2025. Search terms included combinations of “artificial intelligence”, “explainable AI”, “XAI”, “oncology”, “LIME”, “SHAP”, “Grad-CAM”, “interpretability”, and “clinical decision support systems”. Priority was given to studies demonstrating clinical relevance, use-case applications in radiology, pathology, genomics, and treatment decision-making. Emerging methods, ethical implications, and patient-centered approaches were also considered. Additional sources were identified through snowballing from references in key papers. Clinical experts including oncologists were also consulted.
Rationale of Study Selection and Inclusion
Only English-language articles that addressed AI interpretability techniques in oncological contexts were included. The inclusion criteria were as follows: (i) the study used machine-learning or deep-learning on oncology-relevant tasks (ie, diagnosis, prognosis, treatment response, risk stratification); and (ii) at least one XAI or interpretability technique was used in the study. They were both the widely used and the emerging XAI techniques in order to safeguard conceptual comprehensiveness. Non-oncology studies that lacked a discrete explainability element and those that were not described methodologically were eliminated. Because of the narrative review format, formal duplicate screening and quantitative eligibility scoring were not done; relevance was determined iteratively depending on domain applicability, methodology contribution and prominence of citations.
Data Extraction and Thematic Synthesis
Qualitative information was extracted on each of the included studies, and includes the type of cancer, the type of data, model type, XAI method(s), level of explanation (local/global), validation method and the strengths and limitations reported. As an alternative to statistical data synthesis, the results were synthesized thematically to compare the XAI approaches to oncology use-cases in order to reveal typical patterns of methodology and also to reveal the gaps in clinical validation and implementation. The findings of this study were presented and discussed simultaneously using sub-sections to enhance flow and clarity.
Scope and Methodological Limitations
Being a narrative review, this study does not claim to cover all the literature available, and does not follow a formal PRISMA-guided selection pipeline. However, by explicitly documenting search sources, inclusion rationale, and synthesis strategy, the review ensures reproducibility of approach while retaining the flexibility necessary to integrate diverse and rapidly evolving XAI methodologies in oncology.
Explainable AI (XAI) Techniques
XAI encompasses various techniques designed to make machine learning (ML) models more interpretable. They clarify what drives a prediction and how model behavior changes across inputs. The drive towards clinically interpretable AI can also be identified in the related fields, including the application of explainable model-driven solutions in the noninvasive diagnosis of clinically relevant portal hypertension, 9 gestational diabetes prediction, 10 and sickle cell diseases. 11
In the oncology, can serve various purposes: (i) through making predictive outcomes consistent with the known oncologic theory and biological processes; (ii) by making predictive models more auditable and valid; and (iii) by making predictive model output more communicable to the stakeholders. 12 More importantly, explainability can be expressed either locally, in the context of a prediction of a single patient, or on a global scale, across the behavior of the model as a whole. Additionally, explainability can be done post-hoc, by explaining a previously fitted black-box model, or intrinsically, by the use of models which are interpretable in nature. 13 XAI is increasingly used to support interpretability across multimodal pipelines (imaging, radiomics, genomics, and pathology). 7 Here we present these techniques according to their modes of operation.
Local, Model-Agnostic Explanations: LIME and SHAP
LIME offers local interpretability, explaining individual predictions by perturbing inputs and observing the resulting changes in model behavior.
7
LIME is attractive because it can be applied to many model types and provides intuitive feature contribution summaries. However, LIME explanations can be unstable across runs and sensitive to feature correlation and sampling choices; these conditions are common in radiomic and multi-omic oncology data.
14
SHAP, based on cooperative game theory, provides consistent feature attribution, allowing a deeper understanding of how individual features influence a model's output.
15
When evaluating the suitability of XAI techniques for oncology applications, several factors must be considered: effectiveness
Visual Explanations for Imaging Models: Grad-CAM and Saliency Methods
For image-based oncology tasks, Gradient-weighted Class Activation Mapping (Grad-CAM)
Backpropagation-Based Attribution for Deep Models: Integrated Gradients and DeepLIFT
Integrated Gradients (IG) calculate feature relevance by adding up gradients on a path between a reference input and the target input that is of interest to the model in question. Deep Learning Important FeaTures (DeepLIFT) attribute generates a neuron activation that is used to attribute model output to input feature by comparing it to neuron activation resulting from a given reference baseline. 21 These methods have been effectively applied to a variety of deep learning designs, such as but not restricted to image analyses and they have been shown to produce more consistent attribution maps than gradient based methods, which use raw gradients only. Their interpretability depends greatly on baseline/reference selection and input scaling, which can be non-trivial in clinical settings. Integrated Gradients and DeepLIFT focus on computing the contributions of each input feature to the model's predictions, offering transparency in tasks such as genomic data analysis or drug response prediction. 22
Surrogate Models and Rule Extraction (Global Interpretability)
An interpretable model can be trained to provide a high-level view of model behavior and decision boundaries. Global surrogates (eg, training an interpretable model to approximate a black-box model) and rule extraction methods give a high-level view of model behavior and also give a high-level view of decision boundaries. They can be applicable in governance, audit and communication to the stakeholders. Its main shortcoming is that of approximation error: a surrogate might seem interpretable but not faithfully represent the original model, particularly in high-sparsity-density regions or highly complicated interactions.
7
Surrogate models, such as decision trees or linear regression, provide approximations of more complex black-box models, offering clinicians a clearer, albeit simplified, view of AI decision-making.
23
Techniques like Surrogate Models provide a simple, interpretable alternative but may fail to capture the full complexity of deep models, especially in high-dimensional domains like radiomics.
24
Table 1 provides a brief comparison of the various XAI techniques, their advantages, and limitations. Despite their popularity, both LIME and SHAP exhibit limitations that can affect their reliability in clinical oncology. LIME is known to produce unstable explanations, particularly in the presence of highly correlated input features
Comparison of XAI Techniques: Advantages & Limitations.

Beyond SHAP, LIME, and Grad-CAM: A Broader Taxonomy of XAI Methods for Healthcare
Within this broader taxonomy, SHAP and LIME remain central for local feature-attribution in tabular and general black-box settings, while Grad-CAM and related saliency approaches remain prominent in deep vision models. Nevertheless, the bigger picture is that no one approach is adequate across modalities and clinical operations; the triangulation of explanations, that is, a combination of feature attributions, counterfactuals and example-based evidence is frequently more justifiable in clinical practice. Although some of these methodologies are yet to be broadly validated or routinely applied to the oncology practice, including them provides a complete landscape of references and provides insight into the future directions as to which the further methodological integration could be made. Importantly, the low adoption is not synonymous to the lack of relevance. In oncology, where clinical decisions are high-stakes, longitudinal, and multimodal, explainability requirements extend beyond model accuracy to include biological plausibility, robustness across cohorts, interpretability across disease stages, and compatibility with clinical workflows. As a result, emerging or underdeveloped XAI tools can take on a central role in addressing the lack of satisfaction in the demands relating to trust calibration, treatment planning, and regulatory approval, despite the early stage of the available evidence.10,11
Intrinsically Interpretable Models (Built-in Transparency)
Intrinsically interpretable models can be completely understood without examining the underlying model. Another major substitute to post hoc explainability is the use of models whose design can be interpreted directly. Some examples are the sparse linear or logistic regression, the generalized additive models (including interpretable GAM variants), decision trees, rule lists, and scoring systems. Such models are usually provided with transparent relations of features to outcomes and can eliminate the need to provide retrospective layers of explanation. 34 In clinical practice, the benefits of intrinsically interpretable models can also be seen when clinical accountability and auditability are of primary importance. However, these models can be compromised in terms of predictive performance in very complex tasks and their interpretability can be lost when feature engineering, interactions and high-dimensional inputs are non-trivial. 35
Case-Based Reasoning and Prototypes (Example-Based Explanation)
The example-based methods explain a prediction with references to similar situations (nearest neighbors), prototype representatives, or most influential training examples. Analogies with previous patients may be easier to understand than abstract weights of features, which, in turn, would help clinicians with face-validity and reviewing cases. 36 Nevertheless, example-based explanations are sensitive to the data quality, their representativeness, and the selected metric of similarity; they can also reveal the privacy threats unintentionally unless they are managed. 37
Counterfactual Explanations and Algorithmic Recourse (Actionability)
Counterfactual explanations explain what would happen to a model with minimal perturbations to the input. Counterfactuals in clinical decision support can be used to increase actionability, making predictions connected to the possible intervention or monitoring plan. 38 However, counterfactual validity can be jeopardized when the proposed changes are clinically impossible to implement (eg, fixed features), they do not respect causal restrictions, or they are not consistent with established care pathways. 39 In line with that, clinically based constraints and consequential logic are necessitated when giving counterfactuals in the health context.
Concept-Based Explanation and High-Level Interpretability
Concept based approaches seek to align model reasoning to match human interpretable clinical concepts (such as edema patterns, some radiographic appearance, laboratory syndromes) as opposed to raw pixels or individual features. Such correspondence can fill the semantic disparity between model internals and clinician reasoning. 40 Concept definition and the need to ensure that the model really makes use of the concepts and not just spurious correlates is a continuous challenge, especially when dealing with different populations and imaging equipment.
Attention Mechanisms: Useful (Non-Explanatory) Mechanisms
Attention weights and attention maps are commonly used as natural-language and imaging tasks often used as an intuitive attribute of attention. Regions or tokens related to model processing can be emphasized by attention and qualitative review could be aided by it. Attention, however, does not imply fidelity to the importance of causality, and its interpretability is determined by the architecture used and the validation methods. 41 Attention based visualizations must therefore be introduced with care and where possible with faithfulness checks or additional techniques (eg, perturbation tests, counterfactuals).
Uncertainty, Calibration, and Causality: Interpretability in Context
Explainability is most relevant when it is combined with a trusted uncertainty quantification and calibration since clinicians should be able to know not only the reasoning behind a prediction but also the confidence in a prediction and what conditions may cause a prediction to fail. 42 Furthermore, several of the outputs of explanation are meant to be associational, not causal, lack of causal support causes explanations to strengthen confounding or data artifacts. Thus, XAI results are to be discussed together with calibration measures, external validation, sub group results and clinical plausibility checks. 43
The Permutation Feature Importance (PFI)
PFI is used to determine feature relevance through the quantification of the predictive performance loss after feature value permutation. Due to its model-agnostic character and computational simplicity, PFI is commonly used in fast global interpretability and debugging of a model. However, PFI can give biased estimates when predictors are correlated and can miss the significance of variables that share information with others. 44 Therefore, careful interpretation and sensitivity analysis are required, particularly when working with clinical data that often has collinearity.
Partial Dependence Plot (PDP) and Individual Conditional Expectation (ICE)
PDPs represent the average connection between a predictor and model result whereas ICE graphs illustrate instance-specific response curves. The PDP–ICE approach is especially beneficial in oncological settings with either risk scores, dose response curves, or continuous biomarkers because it is capable of revealing non-linear effects and interactions that can be obscure in purely local attribution approaches. 45 However, PDPs are misleading in correlated feature space or extrapolation in sparsely populated feature space, ICE plots can be useful in detecting heterogeneity but can still be used to show association as opposed to causality.
Explain Like I’m 5 (ELI5)
ELI5 is an interpretability pragmatic toolkit, which offers various explanatory results, such as model weight inspection of linear models, feature contribution elucidations of chosen models, and feature importance through permutation. Its major strength is its usability, which allows applied researchers to produce transparent summaries during the process of model development and reporting. 46 ELI5 should, however, be seen as an interface tool, not as a complete XAI methodology, the quality of its explanations depends upon the underlying estimator, and its assumptions.
QLattice
QLattice is a symbolic modeling approach wherein the target of the modeling is small functional representations, such as concise equations or graphs, predicting the relationships thus giving intrinsically interpretable models. The symbolic models may be used to supplement the opaque predictors in situations where the decision logic required by stakeholders is straightforward and auditable (like in clinical risk stratification), and provides transparent alternatives or high-level hypotheses about the association between biomarkers. 47 Just like any other model-search method, QLattice is prone to overfitting, cohort-specific instability, and poor performance in complex tasks, hence external validation and clinical plausibility tests are inseparable.
Historical Development of XAI: A Timeline Perspective
The evolution of XAI has been instrumental in addressing the interpretability challenge posed by black-box AI models. These milestones highlight how the field has progressed from foundational techniques to domain-specific adaptations that enhance clinical trust and usability, as outlined below.
2016 – Introduction of LIME: Ribeiro et al
27
introduced LIME, a model-agnostic technique that explains individual predictions by locally approximating the black-box model. It marked the first widely adopted effort to offer interpretability across model types in healthcare. 2017 – Emergence of SHAP: Lundberg and Lee
28
introduced SHAP, which uses game theory to fairly distribute feature importance. Its ability to offer both global and local explanations made it a favored tool in oncology applications, such as genomic mutation analysis. 2017 – Grad-CAM Developed for Visual Explanations: Grad-CAM became a key method for producing visual heatmaps in CNNs, revolutionizing explainability in medical imaging—particularly in radiology and pathology.
48
2019 – Integration of XAI in Cancer Classification Models: Researchers began integrating LIME and SHAP into neural networks for breast cancer classification and tumor grading from histopathology images, providing visual justifications for model decisions.
49
2021–2023 – Deep Integration of XAI in Multi-modal Oncology Models: With the surge in multi-modal data (genomic, imaging, and clinical records), hybrid and ensemble methods leveraging SHAP and Grad-CAM started appearing in radiogenomics and treatment response prediction studies.
50
2024 – Advances in Real-Time XAI and Federated Learning: New developments, such as instance-specific SHAP (InstanceSHAP) and federated XAI strategies, are emerging to meet the growing demand for interpretable models trained across decentralized oncology datasets.
13
The Black Box Phenomenon
Black boxes are AI systems whose internal decision logic is difficult to understand. This lack of explainability in oncology can raise questions about AI recommendations, particularly when a life-altering treatment choice is at stake. 7 This lack of transparency in the oncological field is not an unimportant inconvenience; instead, it has significant clinical confidence, patient safety, responsibility, and regulatory acceptability implications. Even a model that has a high predictive accuracy can fail to explain the reason behind its findings (such as the reason to categorize a lesion as a malignancy, place a patient in a high-risk category, or to predict a response to treatment). Without clear thinking clinicians might not be in a position to determine whether a prediction is biologically plausible, is because of spurious association, or it is due to latent confounding. 51
There are various elements that cause the black-box phenomenon in oncology AI. The first is the complexity of models: deep neural networks are trained on high-dimensional non-linear representations, which are hard to represent as rules or relationships understandable by humans. 52 Second, heterogeneity and high dimensionality of the data: oncology processes actively combine imaging (CT, magnetic resonance imaging MRI, histopathology) variables, radiomics, genomics and electronic health record (EHR) variables, thus forming feature space in which correlated predictors and latent structure make interpretation difficult. 53 Third, dataset artifacts, shortcut learning: models can learn based on non-clinical signals (scanner specific patterns, staining variability, site specific practices, or documentation habits) that are correlated with outcomes but not biological aspects of the disease. 54 Fourth, distribution shift: performance and explanations may differ between institutions due to scanner differences, protocols, patient population or change in guidelines and the behavior would be unpredictable at deployment. 55
The clinical risks of oncology AI black-box are well known. The absence of interpretability may lead to automation bias where clinicians trust model outputs more to an extent of uncertainty or being incorrect, or may use potentially useful tools less, due to distrust. It may also hinder the analysis of errors and quality assurance, making it hard to diagnose failure modes (such as poor performance in particular subgroups), and it makes medico-legal accountability hard, since the decisions that affect diagnosis and treatment need justifiable reasons. This means that overcoming the black box requires not only correct models, but also information that the models are correct, stable and interpretable clinically in a manner that facilitates safe decision-making.
Evidence abound suggesting that without a clear understanding of how AI reaches its conclusions, healthcare professions may be reluctant to trust tools driven by these technologies, conversely undermining their clinical utility. 5 Oncology in particular tends to have a high stake for such decisions because they are inherently complex and require multidisciplinary teams to evaluate large volumes of patient data. The inability to interpret AI output can limit collaborative decision-making by prompting team members to question the AI recommendations. 56 Patients may also exhibit skepticism towards technology they are unfamiliar with. The growing degree of complexity of medical data has rendered AI a compelling method for analysis and decision-making in oncology. 57 The “Black Box” dilemma, characterized by restricted interpretability and explanatory capacity, impedes the acceptability of AI in clinical practice. 57 In response to this issue, researchers have established XAI frameworks like LIME and SHAP to clarify complex models and facilitate their integration into medical procedures. 58 Figure 1 is a simple illustration of black box and XAI processes. Many oncology applications, including breast cancer classification 59 and Alzheimer's disease detection, 60 have made use of LIME and SHAP. These approaches provide advantages, including the assessment of feature significance, the visualization of correlations, and the evaluation of individual predictions. 58 Some researchers opine that opaque decisions violate physicians’ ethical obligations, while others maintain that such decisions are prevalent in medicine and that empirical validation of accuracy may outweigh the necessity for explanations. 61 Notwithstanding obstacles in research, regulation, and privacy, black-box medicine presents significant advantages in personalized healthcare by utilizing advanced algorithms to analyze extensive health statistics. 62

A simple illustration of black box and XAI processes.
The Need for Explainable AI (XAI) in Oncology
AI algorithms often produce highly accurate but nonintuitive outputs, particularly in deep learning models due to their complex nature. However, the decision-making pathways in these models are not readily accessible, and despite their ability to analyze millions of data points, including medical images and genomic data, they are limited to providing answers to specific questions. 63 This opacity presents a challenge in oncology, as it necessitates the creation of individual treatment plans that require clear rationales for each decision. To tackle this issue, a promising approach is XAI, which intends to make AI models more interpretable and transparent. This helps oncologists comprehend the rationale behind a model's conclusions, builds trust, and lays the foundation for adopting AI-driven insights. 64
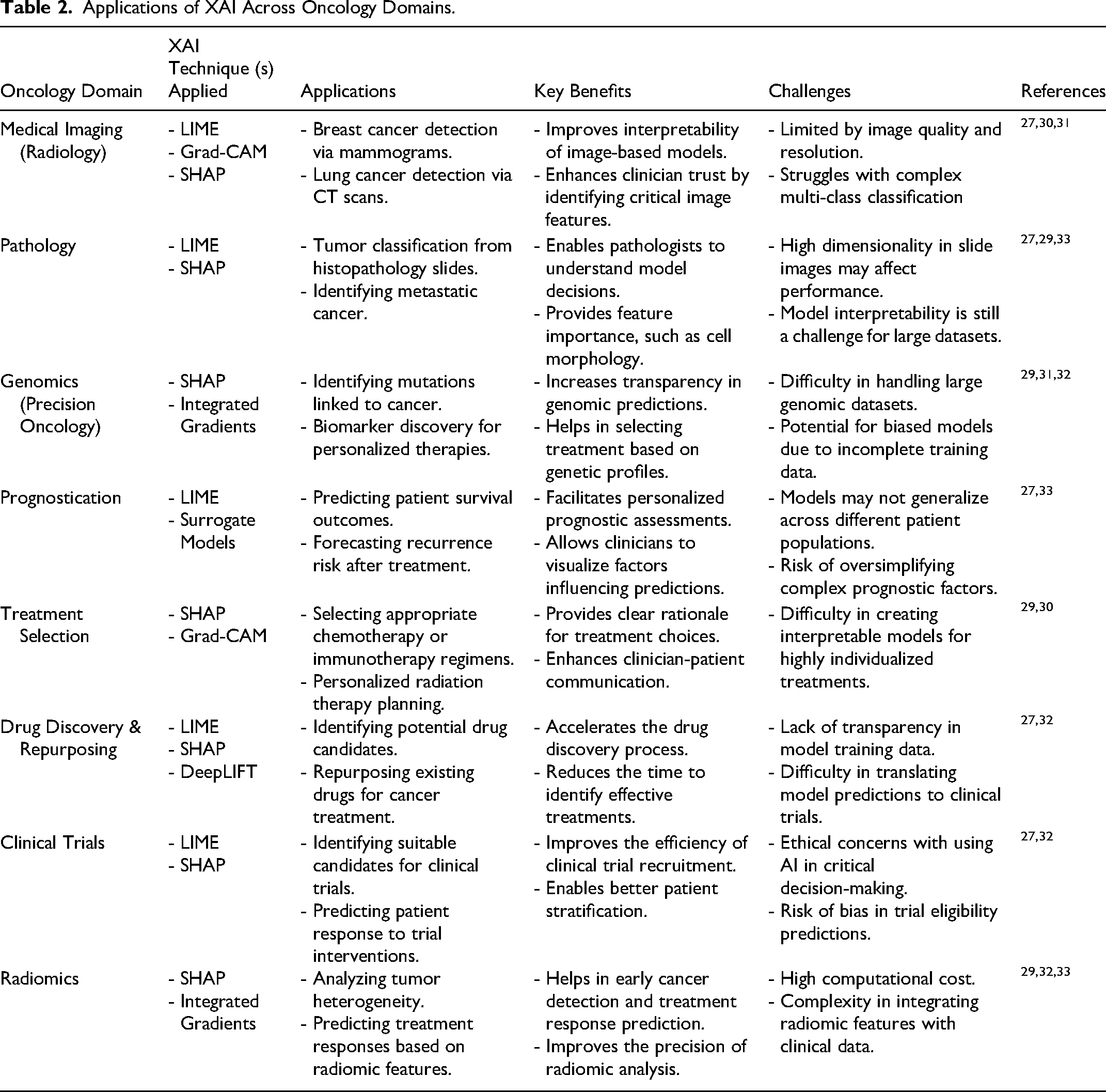
The application of XAI in oncology spans several domains, improving both the interpretability and the clinical utility of AI-driven decision-making systems. Table 2 summarizes the applications of XAI across diverse oncology domains. In medical imaging
Applications of XAI Across Oncology Domains.
Strategies for Enhancing AI Transparency and Explainability
There is much promise in using artificial intelligence (AI) to improve oncology to enhance cancer diagnosis, prognosis, and treatment personalization. However, we often criticize AI models, including deep learning approaches, for being “black boxes,” meaning we are unable to understand how these models make decisions. Again, transparency and explainability issues associated with AI recommendations pose challenges for clinicians and patients to trust them. To encourage the adoption of AI in oncology, it is crucial to go beyond merely improving predictive accuracy and instead focus on developing algorithms that are easy to understand. To reach this goal, we can employ the following strategies:
Integrating Feature Importance Analysis: Feature importance ranking, saliency maps, and attention mechanisms, among other techniques, can also be used to identify which inputs (eg, imaging features or genetic markers) matter most in an AI model's decision. Identifying the features or data points that significantly influence an AI model's decision can enhance its interpretability.
7
For example, in the fields of radiomics, saliency maps can identify salient regions whose presence in the medical image helped the AI make the original diagnosis.
73
There are also novel architectures that enable researchers to make not only predictions but yield explanations for their decisions as well. A prime example of this is a recent study by Hassan et al
74
that showed how a deep learning model can simultaneously identify retinal disease biomarkers and present a “map” to explain its diagnostic reasoning. This transparency can potentially empower clinicians to validate their findings against their clinical judgment. Employing Model-Agnostic (Post-hoc Explanation) Methods: Post-hoc approaches employ AI models to produce explanations that align with the decisions made by the AI model. Some techniques, such as LIME and SHAP, can explain AI systems after they have been built without changing the structure of the models themselves. These methods make it easier to understand complicated models, which makes predictions easier for oncologists to understand. Eventually, they understand why certain treatment suggestions or risk assessments are made.
75
Developing Hybrid Models: Machine learning models can combine with traditional statistical methods to form accurate and explainable systems. Hybrid models combine the strengths of deep learning in pattern recognition with the transparency of conventional approaches like logistic regression.
63
Sometimes, choosing inherently interpretable models, like decision trees or logistic regression, over complex deep neural networks is more beneficial. Even though these simpler models may not always achieve the highest accuracy, they serve as more straightforward explanations of decisions, thereby enhancing transparency in clinical applications.
7
Integrating Human Expertise: Including clinicians in the AI decision-making process ensures the pairing of AI outputs with expert knowledge. Using AI predictions gives clinicians a glimpse into why the AI is making the recommendations it does, as well as assessing whether the recommendations would actually agree with clinical intuition.
76
Researchers from Stanford and the University of Washington have developed an auditing framework that uses human expertise and generative AI to evaluate classifier algorithms. This synergy can be useful to spot for biases or spurious correlations in AI output.
77
Model Validation and Regulatory Standards: First and foremost, we must delineate clear expectations and procedures for the development and deployment of AI in healthcare settings. Rigorous validation of AI models on large, diverse datasets is thus required to ensure their reliability across different patient populations.
78
Furthermore, by adhering to emerging regulatory frameworks and standards for AI in healthcare, we can develop explainable algorithms for the oncology field. This would aid in fostering trust between staff and patients, while also upholding the importance of patient safety.
7
Fostering a Culture of Collaboration: Interdisciplinary collaboration among oncologists, ethicists, and data scientists is crucial for an inclusive understanding of AI systems. Clinicians may gain more confidence in employing these technologies through consistent workshops and training sessions focused on AI tools.
79
Building Trust Through Validation and Clinical Trials: Although transparency is crucial for fostering confidence, it is insufficient on its own to guarantee the dependability of AI algorithms in practical cancer contexts; comprehensive validation and clinical trials are essential to confirm the validity of these algorithms.
80
External validation of AI models using robust and diverse datasets helps confirm the generalizability of these models.
81
Furthermore, employing AI as a decision-support mechanism in clinical trials rather than as a substitute for human discernment can enhance oncologists’ trust. This enables doctors to participate in the creation of AI models, ensuring that the emerging AI addresses the practical needs and concerns of the oncology community.
82
Addressing Ethical Challenges: The ethics of artificial intelligence in cancer must not be disregarded. Artificial intelligence algorithms cause significant concern because training datasets may not adequately represent all patient attributes. To eliminate inequities in cancer care, it is essential to overcome these biases by utilizing diverse and representative datasets.
83
Regulatory organizations must develop clear criteria for AI use in healthcare settings by prioritizing transparency, patient safety, and full accountability.
84
Enhancing Patient Trust and Communication in XAI-Assisted Oncology
While most XAI research focuses on clinician interpretability, patient trust in AI-assisted care is equally critical. In oncology, where treatment decisions carry profound emotional and physical consequences, patients must understand and feel confident in the tools guiding their care. However, current XAI outputs, such as SHAP plots or Grad-CAM heatmaps, are often too technical for non-expert audiences. As AI-supported decisions increasingly influence diagnosis and treatment options, patients must be able to comprehend the rationale behind these recommendations to make informed choices and maintain confidence in their care.
85
Patient-Friendly Explanations: Traditional XAI outputs, such as SHAP plots or Grad-CAM heatmaps, are often too technical for patients to interpret. To bridge this gap, AI systems must generate simplified, layperson-level explanations tailored to patient literacy.
86
For example, rather than presenting a heatmap, an AI system could summarize its output as: “The model recommends treatment A because your imaging results and tumor markers resemble those of patients who responded well to this therapy.” Clinicians Communication Strategies: Clinicians serve as intermediaries between patients and AI-driven systems. Therefore, effective communication training is essential. Clinicians should be trained to translate XAI outputs into understandable terms, using analogies (eg, “like a second opinion that shows its reasoning”) and visual aids.
87
Shared decision-making tools could incorporate interactive XAI visualizations with “what-if” scenarios to demonstrate how changes in patient features might affect predictions.
88
Tailoring the level of explanation based on the patient's health literacy and preferences is very crucial.
89
Building Trust Through Transparency: Studies show that patients are more likely to trust AI decisions when they are accompanied by transparent rationales and endorsed by their healthcare provider. Thus, clinicians play a key role in mediating the AI-human trust relationship
Critical Appraisal of XAI in Oncology: from Plausible Explanations to Clinically Trustworthy Evidence
Even though XAI is frequently introduced as a solution to the problem of the black box, having an explanation are not essentially tantamount to truth, causality, or clinical usefulness. In oncology, in which decision-making involves high stakes and extends over time, that is, screening, diagnosis, choice of therapy, and response monitoring, an explanation must satisfy at least three conditions to be associated with clinical relevance: (i) faithfulness, which means that the interpretation of the underlying decision logic of the model is accurately represented; (ii) stability, which means that the interpretation remains consistent under minor perturbations; and (iii) clinical utility, which means congruence with oncologic reasoning and support of safe clinical actions. 92 Many published oncology researches give graphic or visually stimulating descriptions, eg, heatmaps or ranked-feature visualizations, but do not formally certify these properties, and thus run the risk of deploying a model that is interpretable, but also insecure or misleading.12,93
Method-Specific Limitations and Failure Modes in Oncology
Two of the most popular model-agnostic feature attribution methods, SHAP and LIME, susceptible to dependance structures and are unstable. 94 Oncology datasets often have correlated predictors, eg, radiomic texture families, multi-omic signatures, and correlated laboratory panels. In these correlated environments, feature attribution might be non-unique, that is, several correlated variables can share predictive credit and the resulting importance scores can change significantly across resampling plans, cross-validation folds, or pre-processing choices. 95 LIME additionally depends on local perturbation schemes, kernel bandwidth and surrogate model choice; these design options may produce different explanations of the same patient, thus undermining clinician trust. 14 SHAP may give local and global attributions, but its empirical accuracy is dependent upon the calibration of the model, preprocessing of the data and the feature dependance, which are often not reported in clinical AI studies. 96
Grad-CAM and saliency mapping methods tend to generate highly appealing visualizations, but their empirical validity is low. In the field of oncologic imaging (such as histopathology, CT/MRI, and mammography), saliency maps are often viewed as reflecting the spatial areas that were used in model decision-making. Such emphasized areas, however, can be due to non-representative artifacts, like scanner markers, staining artifacts, or compression patterns, and thus reflect shortcut learning and not true pathological signals.97,98 Furthermore, saliency representations are prone to the difference caused by different model architectures or preprocessing pipelines. Without quantitative measures of faithfulness, (such as deletion and insertion curves, occlusions sensitivity, and perturbation based measurements), saliency maps are to be considered as hypothesis-generating devices but not as conclusive proof of biologically based interpretations.
Global approaches like PFI and PDP-ICE can provide inaccurate results under correlation and extrapolation of features. Practically, PFI can underestimate the relative role of features which share information and overestimate role in case there is information leakage or proxy variables. 99 PDP assumes covariates to be marginally independent and may give spurious effects where not empirically supported; ICE, in contrast to being useful in the revelation of heterogeneity, is purely associational. 100 These methodological limitations are presumed to be especially relevant to the oncology field, where clinical decision-making can be directly influenced by interpretation of the importance of biomarkers. Based on this, diagnostic checks should be used to supplement effect plots with sufficient data support (data density), sensitivity to collinearity, and consistency across clinical meaningful subgroups.
The inherent characteristics of surrogates and rule extraction are the exchange of interpretability with approximation error. Surrogate models can provide useful global summaries that can be used to govern and communicate with stakeholders, but create a false impression of understanding in cases where surrogate fidelity is not measured. 101 In the oncology, surrogates should be evaluated on fidelity at local clinical areas (like borderline-risk situations, treatment-threshold situations) rather than using aggregate measures of agreement.
Why interpretability alone does not guarantee trust: calibration, uncertainty, and dataset shift
Clinical trust is undermined less by the absence of explanations than by unreliable performance under real-world conditions. Datasheet shift including scanner variation, staining protocols, site-specific case mix and changing guidelines are regular occurrences in oncology predictive models, and can change both the predictions and the explanations attached to them. Therefore, explanations should be viewed together with estimates of calibration and uncertainty. A model with a good explanation but poor calibration can do more harm than one with a poor explanation but good calibration. The most justifiable deployment strategy in oncology is consequently one that simultaneously provides explanation, confidence and external validation (such as the factors that motivated the prediction, the confidence of the model, and the situations where the model is known to fail). 102
Clinical Utility and Human Factors: Avoiding Automation Bias
Even explanations of high quality may be harmful when they lead to automation bias, that is, overreliance on the output of the model when clinicians defer conclusion. Where necessary, explanations in oncology tumor boards and workflow-based environments must be crafted to promote the calibrated trust and not persuasiveness. This requires a more human-based evaluative method: exploring the ways in which clinicians in different positions make sense of explanations, evaluating the effect of explanations on diagnostic accuracy and time-to-decision, and establishing whether explanations reduce or increase inequalities in patient subgroups. 103 The explanations must also be practical: where possible, they should also connect the model outputs with alternatives that are clinically viable (such as further imaging or confirmatory testing) but avoid limitations such as immutable properties and causal plausibility.
Threats to Validity in Oncology XAI Studies
Threats to internal validity: Leakage of outcome-proximal variables (like treatment codes), confounding due to site-specific clinical practices, and failure to report preprocessing procedures, can produce inflated performance and misleading explanations.
Threats to construct validity: Explanatory outputs are often interpreted as a ground truth, despite the inherently associational quality of their outcomes; without an interpretative framework that is conscious of the concept of causality, the results of such explanations may serve to strengthen spurious correlations or proxy variables.
Threats of external validity: Lack of external, temporal validation together with ineffective assessment of subgroup generalizability and domain drift between different scanners or institutional contexts can interfere with predictive performance and the accuracy of the related explanations.
Threats of interpretability: Explanation uncertainty, with the addition of perturbations, such as resampling variation, additive noise, feature correlation, and lack of faithfulness testing can yield explanations that are plausible but not faithful to model reasoning.
Threats of human factors: Unsafe deployment due to automation bias, false interpretation of heatmaps, and lack of clinician training can lead to unsafe use even when technical explanations are available.
Ethical, Legal, and Social Implications of XAI in Oncology
While XAI improves transparency in clinical decision-making, its application in oncology raises significant ethical, legal, and social challenges that must be addressed to ensure responsible deployment and equitable patient outcomes.
Bias and fairness: AI models, particularly those trained on non-representative datasets, may inadvertently perpetuate healthcare disparities by providing suboptimal predictions for underrepresented patient populations. For instance, skin lesion classifiers trained primarily on images of light-skinned individuals have shown decreased accuracy in detecting melanoma in darker skin tones, leading to diagnostic bias.
104
In oncology, similar risks arise if tumor histology models are developed on data from a single ethnicity or gender.
105
To mitigate this, XAI techniques can identify biases by visualizing how specific patient attributes influence predictions, ensuring equitable healthcare delivery.
106
However, unless datasets are balanced and diverse, even interpretable models can reinforce systemic biases. Data privacy and Security: AI in oncology typically involves processing highly sensitive data, including imaging, genomic, and electronic health records. To safeguard patient confidentiality, XAI approaches must integrate robust data anonymization techniques, ensuring that personal data is protected without sacrificing model performance.
107
Techniques such as differential privacy Regulatory and governance frameworks: The deployment of XAI in clinical oncology must adhere to emerging regulatory and governance frameworks Human Expertise hurdles: Clinicians face information overload, and complex visual outputs from methods like SHAP or Grad-CAM can be difficult to interpret within time-limited settings.
111
Most clinical systems lack seamless integration with XAI tools, and oncology professionals often lack formal training to use them effectively. Interpretability needs also vary by specialty, complicating tool design. Moreover, AI explanations must align with clinical priorities to be useful.
112
Addressing these hurdles requires clinician-centered design, adaptable explanation formats, and integration with existing health IT systems to ensure practical adoption and trust in XAI.
Challenges in Designing Clinical Trials for XAI in Oncology
While validation and clinical trials are critical to the clinical integration of AI in oncology, XAI-specific systems introduce novel and complex challenges that differ from traditional AI trials. Designing robust trials for explainable models requires not only assessing predictive performance but also evaluating interpretability, clinician acceptance, and patient safety in real-world workflows. Addressing these challenges will require interdisciplinary coordination between trialists, oncologists, AI developers, human factors experts, and bioethicists. The creation of consensus guidelines for XAI evaluation in oncology which are analogous to CONSORT-AI or SPIRIT-AI for general AI is a critical next step toward trustworthy clinical validation.
Defining Evaluation Metrics for Interpretability: Standard clinical trial outcomes such as sensitivity, specificity, or progression-free survival do not adequately capture the value of model explanations. Trials involving XAI systems must include new metrics, such as explanation fidelity (how well explanations align with model logic), usability scores by clinicians, and trust indices. However, these measures remain non-standardized, limiting their regulatory acceptance.
113
Complexity of Multi-Stakeholder Endpoints: XAI systems are expected to satisfy multiple stakeholders: clinicians (interpretability), patients (understandability), and regulators (safety and accountability). Designing trials that measure outcomes for all groups adds complexity, requiring qualitative surveys, cognitive task analysis, and patient satisfaction assessments alongside traditional clinical endpoints.
114
Risk of Cognitive Overload and Misuse: In trials, exposing clinicians to detailed explanations may backfire, leading to over-reliance on AI, misinterpretation of heatmaps, or distraction from critical clinical reasoning.
115
Determining the appropriate level of detail and the best interface for presenting XAI outputs must be rigorously tested in simulated and live settings. Trial Design Heterogeneity: There is a lack of consensus on how to structure XAI trials. Should they be randomized controlled trials (RCTs), observational evaluations, or implementation science studies
116
? Hybrid designs may be required, such as cluster-RCTs with human-AI interaction arms or crossover trials comparing XAI versus black-box AI systems with human feedback loops. Regulatory Ambiguity and Ethics: Existing regulatory frameworks (eg, FDA SaMD or EU AI Act) do not yet explicitly address how to assess the interpretability of AI models. Moreover, ethical questions arise in trials where AI influences high-stakes decisions like cancer prognosis, especially if XAI explanations are later found misleading.
117
Pre-registration protocols must clearly define how explanations will be used, disclosed, and evaluated. Integration with Electronic Health Records (EHRs): To mimic real-world clinical environments, XAI systems in trials must be embedded within oncology information systems and EHRs. However, interoperability issues, delays in model output rendering, and lack of clinician training in XAI interfaces can compromise trial integrity and data reliability.
86
Research Gaps in XAI for Oncology
Despite the fast development of XAI, its introduction in the existing literature has a number of significant gaps that slow down its reliable introduction into the daily routine of oncological practice. These gaps highlight that progress in oncology-focused XAI must extend beyond algorithmic transparency toward clinically validated, human-centered, and ethically governed systems. To begin with, there is a gap in the form of lack of standardized evaluation measures of explainability. Literature uses heterogeneous, generally subjective, measures of interpretability, making cross-model, cross-dataset, and cross-institution comparisons problematic. 7 There is a necessity to have validated and clinically based benchmarks that assess faithfulness, stability, and usefulness of explanations simultaneously.
Second, little prospective and real-world validation is a significant gap. The vast majority of XAI methods in the oncology field are retrospectively evaluated or in controlled settings and have limited information on their performance, strength, or impact on clinical decision-making in realistic workflows of various patient groups. 97
Third, lack of sufficient integration of clinical context and domain knowledge is also a factor. Many of the explanations are feature-based or graphical and not patterned to oncologic logic, disease biology or cures. There is a need to conduct research to formulate concept-based and context-appropriate explanations that can be projected onto clinically meaningful constructs. 92
Fourth, there is limited research on the gaps around human- AI interaction and usability. There is little empirical information on how clinicians process, believe or take action on XAI outputs, especially in high-stakes oncologic decision making. 118 End-user systematic studies are needed to maximize the explanation design, reduce automation bias, and enable successful human-AI interaction.
Fifth, there are still ethical, fairness, and regulatory vacuity. The detection of bias, the subgroup-specific reliability of the explanations, auditability, and the adherence to the regulatory rules are not covered consistently in the extant literature. Bringing the XAI development at the parity of developing emergent regulatory systems in medical AI systems of high risk is also a need that has not been met.
Lastly, causal and action gaps are still present. The majority of explanations are associative and not causal thus limiting their application in the treatment planning or intervention strategy. Causal inference, counterfactual reasoning, and quantification of uncertainty need to be combined in future studies that can help to improve the clinical utility of XAI in the medical field of oncology. 119
Future Directions
The evenness between model complexity and interpretability is crucial for the advancement of AI applications in oncology. While advances in XAI have increased the transparency of machine learning models, several critical gaps must be addressed to ensure clinical relevance and widespread adoption.
Research Gaps in Real-World XAI Benchmarking: One of the most pressing limitations is the lack of publicly available, high-quality, real-world datasets that include both clinical outcomes and ground-truth explanations. Most XAI evaluations are based on synthetic or retrospective datasets that do not reflect the dynamic nature of clinical decision-making. This limits the generalizability and practical validation of XAI systems in oncology.
120
There is a growing need for curated benchmark datasets that incorporate imaging, genomic, and clinical records, along with clinician-annotated explanation labels for comparative evaluation of XAI methods. Emerging XAI Methods for Complex Clinical Contexts: Current XAI techniques like LIME and SHAP offer valuable insights, but their scope remains limited when applied to highly individualized, multi-modal oncology models. Novel approaches such as contrastive explanations, which highlight why a prediction was made instead of an alternative diagnosis or treatment, and counterfactual explanations, which show what minimal changes would lead to a different outcome, are increasingly being explored.
121
These patient-specific explanation methods offer more actionable insights and align closely with how clinicians and patients reason about treatment options. Redefining Evaluation Metrics for XAI in Oncology: Traditional performance metrics like accuracy, precision, and recall are insufficient for evaluating the utility of XAI in clinical practice. Future research should develop and standardize XAI-specific evaluation frameworks, including explanation fidelity, clinical usefulness, user satisfaction and trust, and comprehension and actionability, to assess the model's effectiveness in real-time decision-making. Integrating these metrics into AI development cycles will help ensure that explanations are not only technically correct but also practically valuable. Multimodal and Federated Future Models: The future of oncology AI lies in multimodal data integration, where imaging, genomics, and structured clinical records are combined to generate holistic models. This will require more sophisticated XAI tools capable of disentangling and visualizing contributions from heterogeneous data sources.
86
Similarly, federated learning which enables training across decentralized datasets while preserving privacy will demand new XAI solutions that work consistently across sites, despite data heterogeneity. Additionally, continuous model validation will become essential as AI systems are deployed in clinical settings, requiring ongoing assessments to ensure that models remain accurate and relevant over time. Human–AI collaboration: One of the key future direction is the creation of human-AI collaborations where clarifications will enable effective communication between clinicians and models. The XAI has to become a complement to clinical judgment instead of its replacement, and facilitate shared decision-making, allow clinicians to question or challenge model outputs, and foster calibrated trust. Human-AI interaction also requires high-quality usability testing, formal training of the clinicians, and customized adaptive explanation interfaces based on the particular role of the user (such as oncologists, pathologists, and tumor board members). Ethical and regulatory systems: The implementation of XAI in oncology should be informed by clear ethical values and regulatory frameworks that cover transparency, accountability, fairness, patient safety and data management. The proposed research directions should coincide with the clarification of the explainability approaches with new regulatory demands on high-risk AI systems, whereby explanations are auditable, reproducible, and medico-legally scrutinized. From regulatory standpoint, medical AI systems have dynamic oversight structures. The European Union propsed Artificial Intelligence Act defines AI systems used in the healthcare sector as a high-risk area and thus require strict adherence to transparency, human control, risk, and post-market surveillance.
122
Similarly, the regulatory authorities such as the U.S. Food and Drug Administration (FDA) emphasize the significance of good ML practice (GMLP), transparency of the models, performance monitoring, and change management of AI-based medical devices.
123
Although implicitly these regulatory frameworks require AI outputs to be sufficiently interpretable to support clinical accountability, enable auditability and guide the processes of decision-making, these regulatory frameworks do not dictate specific XAI methods. In the future, the development of XAI in the field of oncology needs to be more aligned with the changing regulatory and clinical demands. This alignment entails the development of explanation methodologies that are technically sound as well as documentable, reproducible and auditable across the model iterations and deployment environments. Moreover, potential validation, ongoing performance checkup, and clear reporting of lack of certainty and failure modes will become the essential components of regulatory compliance. Moreover, explainability must be made to assist human-in-the-loop decision-making. The design makes clinicians be able to override, contextualize or query model outputs, thus avoiding passive adoption of algorithmic recommendations. All these factors reaffirm the need to have XAI solutions that are technically advanced and at the same time, more concrete in terms of regulatory and clinical practicality.
Limitations of the Study
While this study synthesizes the current state and potential of explainable AI in oncology, it primarily reflects perspectives from computer science, informatics, and biochemistry. This review has a number of limitations that should be considered keenly in drawing the implications of XAI to oncology. The absence of direct contributions from practicing clinical oncologists or pathologists is a notable limitation. To begin with, the research is a narrative synthesis, not a systematic review or meta-analysis; therefore, search strategy, study selection, risk-of-bias assessment were not performed according to the standardized guidelines, which could be the source of selection and publication bias. Second, the XAI literature is also not homogeneous in terms of algorithms, methods of explanation, datasets, endpoints, and reporting practices, thus making cross-study comparison challenging and preventing the possibility of making quantitative conclusions regarding clinical benefit. Third, much of the evidence is done on a retrospective basis or in a simulated environment with much less guiding prospective, multi-centered studies; therefore, actual performance, calibration, workflow integration, and patient-centered outcomes (eg, trust, shared decision-making, equity) are not fully characterized. Fourth, the explanations are frequently not validated with reference to clinician reasoning or plausible biological processes, and assessment is often based on proxy measures that are not necessarily relevant in practice in terms of utility or safety. Fifth, there is limited generalizability due to dataset shift, lack of representation of different populations and tumor subtypes, and confounding by imaging protocols and site effects, which can increase bias despite apparent accuracy. Lastly, explainability, privacy, and accountability policies put forward by the regulatory, legal, and ethical authorities are constantly changing, which means that some of the recommendations might require revision with the advancement of the standards. Future studies must focus on transparent reporting and reproducible evaluation both of model performance and of model explanations, do pre-registered prospective studies, multi-institutional validation and human-centered assessment of model explanations with clinicians and patients. These measures will be necessary to guarantee that XAI will improve interpretability and at the same time improve safety, fairness, and clinically meaningful outcomes in oncology.
Conclusion
XAI is a critical aspect of overcoming the long-standing black-box challenge that stands in the way of the reliable clinical application of AI in oncology. XAI methods are essential for enhancing the transparency, trust, and usability of AI systems in oncology. While widely used methods such as SHAP, LIME, and Grad-CAM can improve transparency, their clinical value depends on explanation faithfulness, stability, and usability, particularly in heterogeneous oncology settings spanning imaging, radiomics, genomics, and multimodal decision-making. This review attests that interpretability should be taken as a sociotechnical imperative, integrating technical elucidations with clinical workflow integration, human supervision and governance controls, and not be thought of as an algorithmic adjunct. To reach a broad coverage, this review goes beyond the analysis of SHAP, LIME, and Grad-CAM with intrinsically interpretable models; counterfactual and example-based explanations; concept-based and attention-based strategies, and surrogate and global interpretability strategies. The discussion highlights the fact that the usefulness of explanations in healthcare settings requires strict calibration, solid validation, and compliance with the clinical feasibility limits. By providing clear and interpretable model predictions, XAI techniques facilitate better decision-making, improve patient outcomes, and ensure that AI-driven treatments align with clinical expertise. However, overcoming challenges related to bias, data privacy, and regulatory compliance will require concerted efforts from researchers, clinicians, and policymakers. Addressing these gaps through prospective multi-center studies, uncertainty- and calibration-aware explanation interfaces, human-centered evaluation, and regulatory-ready reporting will be essential for advancing transparent, reliable, and clinically meaningful AI systems in oncology. As the number of people getting cancer is expected to rise around the world by 2050, solving the “black box” problem in oncology with XAI could lessen the bad effects of cancer by improving patient outcomes, should this terrible prediction come true.
Supplemental Material
sj-docx-1-tct-10.1177_15330338261434649 - Supplemental material for Overcoming the Black Box Challenge: Building Trust in Artificial Intelligence Algorithms in Oncology
Supplemental material, sj-docx-1-tct-10.1177_15330338261434649 for Overcoming the Black Box Challenge: Building Trust in Artificial Intelligence Algorithms in Oncology by Esther Ugo Alum, Chukwuoyims Kevin Egwu, Vaithiyalingam Subramanian Manjula, Patience Owere Ekpang, Joseph Enyia Ekpang II, Darlington Arinze Echegu, Benedict Nnachi Alum and Daniel Ejim Uti in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Acknowledgements
None
Ethics Approval
This study does not require an Ethics statement as it is based solely on publicly available data and does not involve direct interaction with human subjects.
Consent to Participate
Not applicable.
Consent to Publish Declaration
Not applicable.
Credit Authorship Contribution Statement
Conceptualization: EUA, DAE, BNA
Methodology: EUA, CKE, DAE, POE, VSM, JEE, BNA, DEU
Investigation: EUA, CKE, DAE, POE, VSM, JEE, BNA, DEU
Data Interpretation: EUA, CKE, DAE, POE, VSM, JEE, BNA, DEU
Resources: EUA, CKE, DAE, POE, VSM, JEE, BNA, DEU
Supervision: VSM, JEE, CKE
Validation: POE, BNA
Visualization: EUA, VSM
Writing – original draft: EUA, DEU
Writing – review & editing: EUA, CKE, DAE, POE, VSM, JEE, BNA, DEU
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Competing Interest Declaration
We declare no competing interests.
Availability Statement
All data are utilized in the manuscript.
Clinical Trial Date of Registration
Not applicable.
Clinical Trial Registration Number
Not applicable.
Clinical Trial Registry
Not applicable.
AI Disclosure Statement
During the preparation of this manuscript, the authors used Quilbot and Grammarly tools for language editing, and grammar correction improvement. However, after its use, the authors thoroughly reviewed, verified, and revised all assisted content to ensure accuracy and originality. The authors, therefore, take full responsibility for the integrity and final content of the published article.
Supplemental Material
Supplemental material for this article is available online.
