Abstract
Introduction
cervical cancer ranks as the fourth most common cancer among females worldwide. Approximately 528,000 new cases of cervical cancer are reported annually, and about 85% of them occur in less-developed countries. The lack of skilled medical staff and pre-screening procedures is the main cause of the high fatality rate in these countries. Cervicography images are the gold standard procedure for the evaluation of cervical cancer; however, the high intra-class inconsistency makes the diagnosis process more challenging for skilled medical specialists.
Method
In this work, we propose a fully automated computer-aided diagnosis (CAD) system for classifying cervical cancer using Cervicography images. Data augmentation is performed in the initial phase to address dataset imbalance. Subsequently, we proposed two novel deep learning modules: the 11-Parallel Inverted Residual Bottleneck Blocks (11-PIRBnet) architecture and the 9-Parallel Inverted Residual blocks with Self-Attention Mechanism (9-PIRSANet). Both modules are fused at the network level via a depth concatenation layer to form a new network, 375NFNet. The proposed network is trained on the selected dataset, whereas the hyperparameters are initialized through Bayesian Optimization (BO). For feature extraction, a depth concatenation layer is used during testing to combine information from both deep learning modules. Finally, the extracted features are classified using a shallow neural network (SNN) to produce the final classification.
Result
To evaluate the model, experiments were conducted on a publicly available cervical screening dataset of Cervicography images, and results demonstrate an accuracy of 95.5%, a precision of 95.4%, and an area under the curve of 0.97. When compared with several pre-trained techniques, the proposed architecture achieved significant improvement in accuracy, precision, and number of trainable parameters.
Conclusion
The proposed 375NFNet architecture demonstrates remarkable accuracy and efficiency in classifying cervical cancer through cervicography images, which shows its potential as a valuable tool in resource-constrained environments.
Keywords
Introduction
The common malignancy, cervical cancer, affects mainly women in developing countries with high rates of morbidity and mortality.1,2 Cells in cervical tissue begin to grow and reproduce uncontrollably, without adhering to the correct mechanisms for cell division, resulting in CrC, a cancerous tumor. 3 In the early stages of cervical cancer, symptoms may not be evident for many years, and the symptoms related to the first stage of the disease are not obvious. 1 The leading causes of cervical cancer include early pregnancy, smoking, having multiple relationships, oral hormonal contraception, premature intimacy, weakened immune systems, and irregular menstrual cycles. 4 The symptoms of cervical cancer related to sexual intercourse include moderate pain, vaginal discharge, and irregular uterine bleeding. 4 It is more common in women over 30. 5 Women who suffer from untreated cervical HPV infections typically take 15–20 years for normal cells to develop into cancer. Still, this process may be accelerated for those who have weakened immune systems.3,6
Cervical cancer primarily occurs due to the integration of high-risk human papillomaviruses (HPV) into the host genome, with nearly 99% of cases linked to sexually transmitted HPV infections. 7 It is estimated that more than one hundred HPV genotypes exist, of which at least fifteen are associated with cervical cancer and other cancerous growths. 8 As invasive cervical cancer is detected early, 92% of females who develop the disease will survive for five years. But just 44% of women with cervical cancer encounter an early diagnosis. As a result of the spread of cervical cancer to other tissues or organs, the 5-year survival rate drops to 58%. 9 Globally, approximately 600,000 women suffer from this disease each year. Nearly 300,000 women die from the disease each year. Therefore, it is strongly recommended that middle-aged women undergo annual cervical screening. According to the World Health Organization (WHO), it is the second most prevalent cancer in developing countries and the fourth most common cancer worldwide in women. 10
In cervical cancer screenings, human papillomavirus (HPV) testing, as well as visual inspection following acetic acid application (VIA), are commonly utilized. 11 Cervical cancer is often diagnosed with cytopathology screening. During a cervical cytopathological examination, a physician uses brushes to collect cells from the patient's cervix, then exfoliates the cells and arranges them on a glass slide. 12 Under a microscope, cytopathologists examine samples to assess whether they may contain malignant tumors. On each plate, there are thousands of cells. As a result, the manual examination is highly problematic and prone to error even for specialists. Thus, a better solution is widely required for this issue, which in turn provides better accuracy. 4 The number of deaths and diseases caused by cervical cancer has prompted many scientists throughout the world to look for ways to reduce it. However, the large volume of photographs increases the pathologists’ and physicians’ workload. Despite the availability of numerous commercial techniques for diagnosing CrC abnormalities, these methods were deemed expensive and required human expertise to operate. 13 When lesions are overanalyzed, low-grade cervical lesions are treated unnecessarily, which increases the risk of infection and economic difficulty. Despite the availability of numerous commercial techniques for diagnosing CrC abnormalities, these methods were deemed expensive and required human expertise to operate. 14
Computer-aided diagnostics (CAD) systems have recently been the subject of multiple research studies for lung malignancies, colorectal cancers (CRC), and various other kinds of cancer.15,16 The use of CAD systems can be very beneficial to physicians and pathologists when it comes to early detection and treatment of cervical cancer and other cancers. 7 In addition to diagnosing and tracking cervical cancer, CAD imaging systems facilitate expert doctors in decision-making, resulting in more precise and prompt treatment. 16 A CAD system is based on a few key steps: image preprocessing, segmentation of cancer or infection regions, feature extraction and reduction, and classification. The preprocessing involves contrast enhancement and noise removal from the input images. A higher-contrast image facilitates accurate segmentation of the infected or cancerous region. Feature extraction is another important step in the CAD system for classifying disease regions into relevant categories such as Type 1, Type 2, and Type 3. A few sample images of these types are shown in Figure 1. 17

Sample images of cervical cancer screening. 17
For cervical cancer, traditional features such as color and texture are usually extracted; however, accuracy drops when redundant information is removed. Therefore, a few researchers used feature reduction techniques to resolve this issue, which was later passed to the classifiers for the final classification. The traditional techniques reduced the performance of a CAD system when input data is large in amount, and there is a high redundancy in the features; therefore, the recent trends of deep learning (DL) show a high impact on it by extracting important features using raw images without any segmentation step or reduction methods. A convolutional neural network (CNN) is a type of deep learning model that consists of several layers, including a convolutional layer, a ReLU layer, a pooling layer, a fully connected layer, and a classification layer. The convolutional layer extracts features, which are further refined by the activation and pooling layers. To further link the important features, it is essential to add a mechanism that not only reduces the feature vector length but also improves the CAD performance. For cervical cancer, it is essential to design a model that not only accurately classifies Types 1, 2, and 3, but also runs in minimal time.
Alquran et al 5 offered the first algorithm capable of classifying Pap smear images into seven separate classifications to assist in the diagnosis of cervical abnormalities. The ResNet101 model was utilized to extract automated features, and a Support Vector Machine (SVM) classifier was employed for classification. Using a cascade method with five polynomial SVM models, the system successfully classified high levels of abnormalities and distinguished between mild and moderate dysplasia with roughly 92% accuracy. Habtemariam et al 11 designed a reliable system for automatically categorizing different types of cervixes and cervical cancer. They used 4005 colposcopy images and 915 histopathology images for the experimental process. To identify the transformation zone and extract the region of interest (ROI) from cervix pictures, a lightweight MobileNetv2-YOLOv3 model was trained. The EfficientNetB0 model was then trained on histogram-matched histopathology pictures for the classification of cervical cancer, and these extracted images were subsequently classified using that model for cervix type. For the cervical cancer classification, the presented method achieved an accuracy of 94.5%, whereas the 96.84% accuracy was obtained for cervix type classification. Kalbhor et al 21 employed a deep learning framework for the classification of cervical carcinoma images. The first method utilized various ML methods with pre-trained models as feature extractors to achieve classification accuracy, with ResNet-50 obtaining the highest accuracy of 92.03%. By optimizing pre-trained models, the second method used GooleNet through TL and achieved a maximum classification accuracy of 96.01%.
Bingol et al 8 presented a hybrid deep learning model for the classification of Pap smear pictures to identify cervical cancer. The Gaussian approach was applied to enhance the images in the SIPaKMeD dataset, and feature maps were derived from the original and enhanced datasets. Mobilenetv2 and Darknet53 models were used to process these features. Neighborhood Component Analysis (NCA) was used to reduce the dimensionality of the combined feature maps. Next, various classifiers were used to classify the optimized features; the Support Vector Machines (SVM) classifier yielded the best results, with an accuracy of 98.90%. Deo et al 22 introduced CerviFormer, a cross-attention-based Transformer technique for the classification of cervical cancer in Pap smear images. CerviFormer uses transformers and makes very few assumptions regarding the amount of data it receives. The model is efficient at managing enormous datasets because it uses a cross-attention method to condense large-scale input data into a compact latent Transformer module. The model exhibited 96.67% accuracy for 3-state classification on the SIPaKMeD dataset and 94.57% accuracy for 2-state classification.
S. Nurmaini et al 23 introduced a novel deep learning method to improve the decision-making about cervical precancerous lesions through cervicograms, taken before and after applying acetic acid. The proposed model aims to improve the accuracy and consistency of VIA (Visual Inspection with Acetic Acid) analysis by utilizing the strengths of Slicing Aided Hyper Inference (SAHI), Yolo v8 and cancer treatment guidelines. Experimental findings reveal that the proposed model outperforms existing methods by achieving high accuracy (90.78%), sensitivity (91.67%), and specificity (90.96%) respectively. G. Atteia et al 24 proposed a DL-based support system incorporated with a hybrid feature reduction and optimization module for the identification of cervical cancer in liquid-based cytology smear images. The proposed system is integrated with a sparse Autoencoder with a binary Harris Hawk metaheuristic optimization algorithm to select the most critical features from the overall feature set of input images. Three pretrained CNNs are used to retrieve the overall supplemented feature set, and a Bayesian optimized KNN classifier is used on this improved feature map for the classification of pap smear images. On evaluation, the proposed methodology shows classification accuracy, sensitivity, and specificity of 99.9%, 99.8%, and 99.9%, respectively.
D.D. Himabindu et al 25 presented a deep learning technique that leverages swin transformer and ensemble of deep learning models to detect cervical cancer through colposcopy images. In the proposed technique, a Weiner filtering model is used for pre-processing, a Swin Transformer is used for feature extraction, and an ensemble of three models, namely an autoencoder (AE), a bidirectional gated recurrent unit (BiGRU), and a deep belief network (DBN), is utilized for the detection process. Upon testing, this sophisticated architecture achieves 99.44% accuracy over existing models. A.K. Sharma et al 26 utilized deep learning in 2 ways for cervical cancer detection. First, they extracted features from pretrained models using various ML techniques and then applied transfer learning to cervical cancer images through pretrained methods. By combining ResNet-50 and VGG16 in an effective way, the proposed approach achieves an accuracy and F1-score of 94.53% and 93%, respectively. A. Hunzala et al 27 proposed a hybrid approach that combines transfer learning, deep convolutional neural networks and ensemble models for cervical cancer detection. The authors preprocessed the dataset by performing Error Level Analysis (ELA) and then applied four transfer learning techniques, ie, Efficient-NetB0, ResNet-50, MobileNetV2, and DenseNet201, on the dataset. These techniques are compared with four DCNN architectures (AlexNet, ZfNet, HighwayNet and LeNet-5). Additionally, an ensemble model AZL that leverages AlexNet, ZfNet and LeNet is proposed and results demonstrate that AZL achieves 99.92% accuracy, outperforming individual DCNN architectures and transfer learning techniques.
To have a better understanding of previous work done for cervical cancer classification, all the related studies discussed in this section are summarized in the Table 1 below.
Summary of Literature Review.

The limitations in the above studies, as discussed in Table 1, are needed to be addressed by proposing new classification systems for cervical cancer identification.
Proposed a customized 11-Parallel Inverted Residual Bottleneck Blocks (11-PIRBnet) architecture that consists of a twin parallel layer structure. The designed model is lightweight with a few parameters. Proposed a customized 9-Parallel Inverted Residual blocks with Self-Attention Mechanism (9-PIRSANet) architecture for the classification of cervical cancer. The proposed architecture extracted important information from self-attention mechanism. Proposed a network-level fused CNN architecture called fused 375NFNet that concatenated the depthwise information of 11-PIRBnet and 9-PIRSANet architectures into a single layer. Hyperparameters of this fused model are optimized through Bayesian Optimization instead of manual selection for enhanced training performance. A detailed statistical analysis has been performed to validate the performance of proposed fused architecture on the selected dataset. In addition, several ablations studies are also performed to analyze the generalizability and scalability of proposed fused model.
Proposed Methodology
In this work, a novel network-level fused self-attention architecture is proposed for the classification of cervical cancer. The proposed architecture is based on two steps- training and testing, as shown in Figure 2. In the training step, the dataset has been divided into a training set and a testing set. After that, augmentation has been performed on the training set to increase the number of images and add diversity for efficient training. After that, two CNN models are proposed, named the customized 11-Parallel Inverted Residual Bottleneck Blocks (11-PIRBnet) architecture and the customized 9-Parallel Inverted Residual blocks with Self-Attention Mechanism (9-PIRSANet) architecture. 11-PIRBnet consists of 200 layers with approximately 7.5 million parameters, and 9-PIRSANet consists of almost 175 layers, accounting for approximately 8.0 million parameters. Later on, the output of both models has been fused using a network-level fusion technique, and the final architecture is obtained, named fused 375NFNet, which contains a total of 375 layers with 15.5 million parameters. Hyperparameters of this fused model are optimized through Bayesian Optimization and performed training. In the testing phase, the trained model has been utilized, and features are extracted from the depth concatenation layer that is fed to shallow neural network classifiers for final cervical cancer classification. The technical specification is given in Table 2.
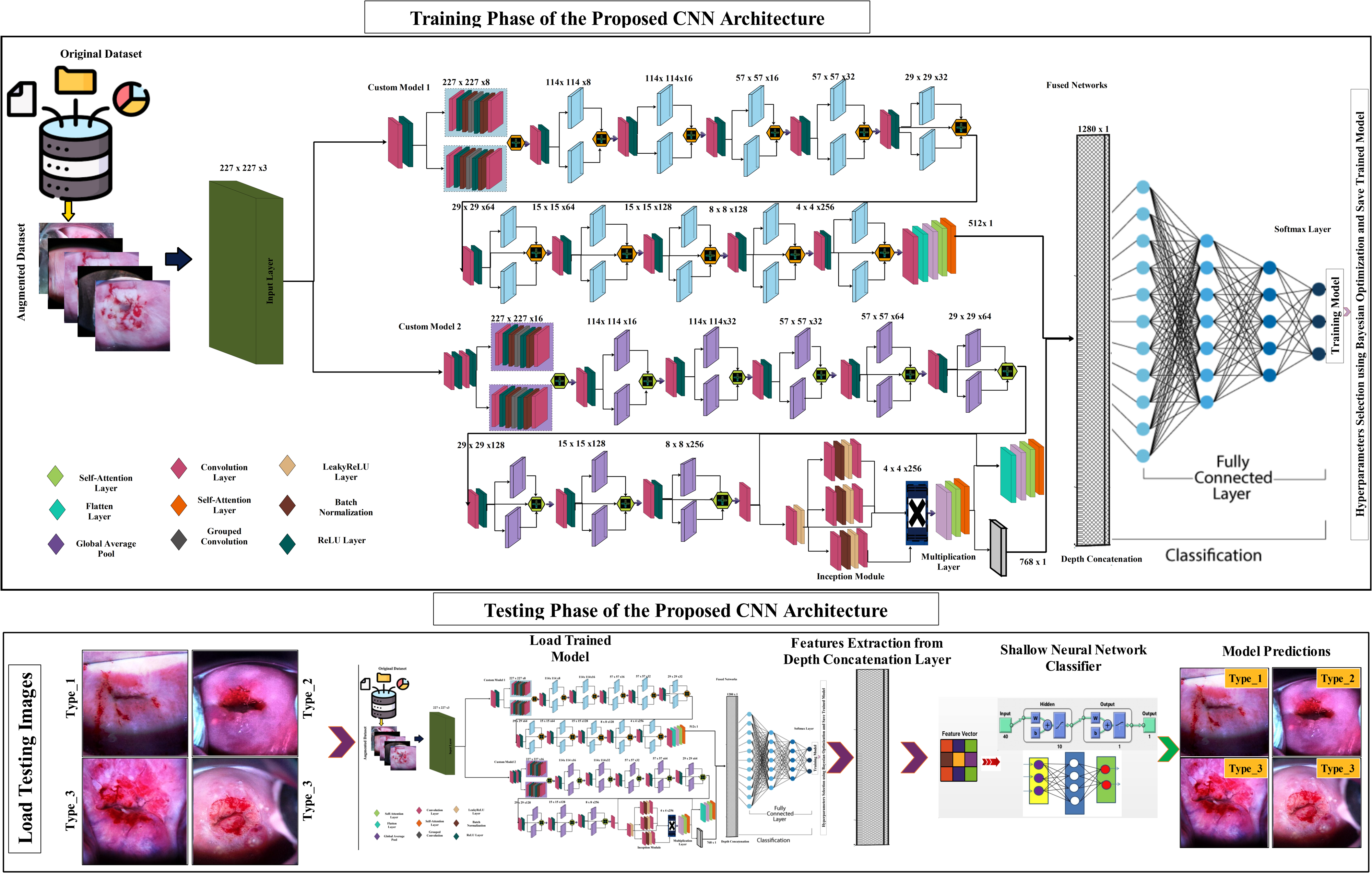
Proposed detailed architecture of 375NFNet for cervical cancer classification.
The Technical Specifications of Proposed Model.

Dataset Collection and Augmentation
The Cervical Cancer Screening Dataset on Kaggle was released to the public in 2017. 42 The selected dataset images are in RGB in nature. 43 The cervical images in the collection are categorized into three types: Type 1 (625), Type 2 (3042), and Type 3 (1698). 17 Type 1- The majority of the cervix's exterior is visible; Type 2- both the internal and external cervical parts are visible, and Type 3- only the internal cervical portion is visible. These classifications are used to help assess the risk of cervical cancer, with each type representing different anatomical variations of the cervix that can influence cancer risk assessment and screening strategies. The labeling process was completed by three skilled gynecologists, including senior medical doctors and an emergency surgical officer. Their combined expertise ensured the accurate classification of cervical images, which is critical for assessing the risk of cervical cancer and improving the reliability of the diagnostic process. 11 Figure 3 illustrates a few sample images of this dataset.

A visual illustration of data augmentation process for cervical cancer images.
The initial dataset of images was insufficient to effectively train a deep CNN model. To enhance the training performance of the proposed 375NFA, additional images were incorporated through data augmentation techniques. However, before this, dataset was split 50-50 into training and testing data by utilizing a stratified random sampling approach. Instead of conventional splits like 80-20 or 70-30, we adopted 50-50 split to ensure balanced evaluation on this limited dataset. To ensure reproducibility and maintain class relationship, all the images in the respective class are shuffled randomly with a fixed seed (42) and then split into 2 equal halves. Class proportions in both sets are verified after splitting. Now, to ensure class balance, data augmentation is applied on the training set. This approach increased the diversity of the training set, leading to more accurate and robust model results. Three operations were applied for data augmentation: flipping left-right, flipping up-down, and rotating the images by 90 degrees. These transformations increased the diversity of the dataset, improving the model's ability to generalize. Figure 3 visually demonstrates these augmentation techniques, and a detailed overview of the entire dataset is provided in Table 3. In this table, it is noted that the original Kaggle dataset consists of 5365 RGB cervicography images (categorized into three classes: Type I (625), Type II (3042) and Type III (1698)). These images are resized to 227×227 and normalized on the scale of (0-1). This dataset is then divided into training and testing sets using a ratio of 50-50. After that, data augmentation process was performed on training data and generated 2000 images of each type. Summary is given under Table 3.
Brief Description of Cervical Cancer Screening Dataset.

Proposed 375NFA CNN Architecture
In this work, we proposed a novel network-level fused CNN architecture named 375NFA. The proposed architecture consists of two different modules i-e 11-PIRBnet and 9-PIRSANet that are fused based on network-level in the final phase of fully connected layer. The design behind this architecture is motivated by the challenges observed in the existing methods. Traditional residual blocks follow wide-narrow-wide pattern that results in high computational cost. The proposed inverted residual bottleneck blocks follow narrow-wide-narrow approach that leads to computational efficiency and feature preservation. Instead of sequential stacking, we employed a parallel network with asymmetric configuration (11 vs 9) for multi-scale feature extraction and complementary feature learning while maintaining computational complexity. Additionally, Self-attention mechanism is incorporated to model long-range dependencies and handle intra class variability. Network-level Fusion via depth concatenation employed in this model further improves classification performance by allowing end-to end joint training and optimization. The purpose of this network is to get the maximum information of both designed CNN modules and obtained the improved cervical cancer classification accuracy. The proposed 375NFA architecture contains 15.5 million total trained parameters with 375 layers.

Proposed 11-Parallel Residual Blocks based Self-Attention architecture for cervical cancer classification.
At the start of the network, there is a 3×3 Convolutional layer with ReLU activation, a stride of 1, and a depth of 8. The first parallel IBRBs were added. Each block contains seven layers. The first convolutional layer has a 16-depth size with a stride of one and a 1×1 filter size. Then, a ReLU is added with a normalization layer. Then, a grouped Convolutional layer was added that has a 3×3 filter size with a normalization layer. The first residual block concludes by adding the last convolutional layer with 8 depths, a stride value of 1, and a filter size of 1×1. Then, these IBRBs are connected to other layers by adding the first additional layer. After that, a Convolutional layer along with a ReLU has been added with a kernel size of 3×3 with a stride value of 2 and a depth size of 8.
The second parallel residual block has been added with the same layer pattern in which convolutional layers have dimensions 16 and 8 depth size with the same kernel size of 1×1 and stride 1. The second additional layer has been added to add the weights of this parallel block. With transitional layers, a Convolutional with ReLU is additionally used, which has a 3×3 kernel size with a stride of 1 and a depth of 16. Third, parallel residual blocks are added to the same layer pattern in which convolutional layers have dimensions 32 and 16 depth size, having the same kernel size 1×1 with strides 1. The output of this block has been added to an additional layer. After that, the transitional layers have been added that included a Convolutional of 3×3 kernel size, stride 2, and a depth size 16. The ReLU has been attached after the convolutional layer.
In fourth parallel block, the depth dimension has been increased as 32 and 16 with the same kernel of 1×1 and strides 1. The output of both parallel paths of this block has been added into a single additional layer that followed by the transitional layers such as Convolutional layer and ReLU having 3×3 kernel size with stride value is 1 and depth size 32. In these four blocks, no skip connections were added to maintain the dimensions of the resultant layers.
In the fifth and sixth parallel residual blocks, the depth size of both convolutional layers has been set as 64 and 32 with 1×1 kernel size and strides 1. In between these blocks, transitional layers such as Convolutional with ReLU activation have been added, with a filter size of 3×3, depth values of 32 and 64, and a stride value of 2. The change of the stride value is to reduce the weight matrix. Seventh and eighth, parallel residual blocks having the exact dimensions 128 and 64 depth size with the same kernel 1×1 and a single stride value. In between these blocks, transitional layers of depth size 64 and 128, respectively, were added.
Ninth and tenth, parallel residual blocks having same dimensions 256 and 128 depth size with the same filter size 1×1 and stride value of one. In between these blocks, transitional layers such as convolutional layer with ReLU has been added, which has the same kernel 3×3 with strides 2 and a depth size is 128 and 256, respectively. Eleventh, parallel residual block has been added and both convolutional layers depth size of 512 and 256, respectively. In addition, the kernel size 1×1 and strides 1 has been selected. An additional layer has been added that connects the skip connection and inside layers weights. After that, few more layers have been added such as convolutional layer having 3×3 filter size, stride value of one and depth size is 512. Later on, a Global Average Pool (GAP) connected to control the size of the network that further connected with a flattened layer. The output of this layer is passed to self-attention layer that computed more inside features of the input image.
The feature maps generated from the previous convolution layer were used to derive the self-attention maps within each convolution block. By paying attention to every position in the same image with varying weights, as a result, it was calculated and expressed how much attention a position attracts: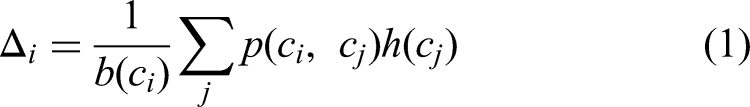
Where, i represents the point at which the response was calculated, j enumerated all positions in the same image, the scalar value is computed through function p and it is computed between i and 



Here 
Here

Proposed 9-Parallel Residual blocks architecture with Inception and self-attention modules for cervical cancer classification.
The second parallel residual blocks has the same layers pattern in which convolutional layers has 32 and 16 depth size with the same 1×1 kernel size and strides one. Then, second additional layer has been added that follows by the transitional layers such as convolutional and ReLU activation of filter size 3×3, stride 1, and a depth size of 32. Third and fourth, parallel residual blocks having the same dimensions 64 and 32 depth size with the same 1×1 kernel and strides one. In between these blocks, transitional layers are added, which has the same 3×3 filter size with strides 2 and 1 and a depth size is 32 and 64, respectively.
In the fifth and sixth parallel blocks, same depth size of 128 and 64 has been selected for the convolutional layers along with 1×1 kernel size and strides 1. In between these blocks, transitional layers with 3×3 kernel and strides 2 and 1, respectively. The transitional layers are standard 3×3 conv layers followed by ReLU activation that are integrated to adjust the feature dimensions before entering the next block. Moreover, the depth size is 64 and 128 has been selected. In the seventh and eighth parallel blocks, the depth size has been increased such as 256 and 128, whereas the kernel and strides remains same. In between these blocks, transitional layers are added of depth size is 128 and 256, respectively. In the ninth parallel block, depth size is further increased to 512 and 256; however, the filter size and stride value remain same as 1×1. Each block extracted information is combined through additional layer along with a skip connection.
The transitional layers are added after this block of 3×3 kernel and depth size 512. To control the dimension of extracted weights of the network, we added a global average pool layer that follows a flattened layer. The output of this layer is passed to self-attention layer for the extraction of more informative features. Self-attention process is described under first module (Eq. (1)- (6)). The output of self-attention layer is passed to fully connected layer. In parallel with these layers, one convolutional layer was added having depth size of 256 and 3×3 kernel. A LeakyReLU is further added for the non-linearity. After that, a single inception module was added in which the first three parallel convolutional added have the same depth size of 512, 3×3 kernel size and strides 1. Then, three grouped convolutions were added with three Leaky ReLU layers, and then again three convolutional layers added having the same dimensions of 256, 3×3 kernel size and strides 1. The outputs of this module are connected through a multiplication layer. After that, a global average pool layer is added that follows the flattened and self-attention layer. Both self-attention layers are added to each other through depth-concatenation layer for enricher the features information of each image for more accurate classification.
Proposed Network-Level Fusion and Training
In the final stage, features from both modules extracted from the self-attention and depth concatenation layers are passed to another depth concatenation layer, as illustrated in Figure 5. The depth concatenation layer features are fed to softmax layer for the classification. The complete fused architecture is shown in Figure 6. In this figure, the cross entropy is utilized as a loss function for softmax classifier. The softmax function is defined as follows:

Proposed network-level fused self-attention architecture for cervical cancer classification.
In this work, we utilize BO for hyperparameters tuning of proposed CNN architecture. The tuned hyperparameters are initial learning rate and momentum, whereas the batch size is selected 64 and SGDM is an optimizer.
Bayesian optimization is defined as a Gaussian-based optimization method that utilize a probabilistic model to select hyperparameters while balancing exploration and exploitation. It is designed to maximize an unknown objective function
Equations (8) and (9) represents the acquisition function such as improvement based:

The Equation (10) represents the method possible to evaluate the probability of improvement using normal distribution: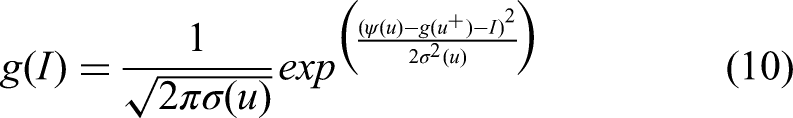
The integral of the density function estimates the expected improvement (EI) that defined in equations (11) and (12).

Where
Algorithm of Bayesian Optimization.

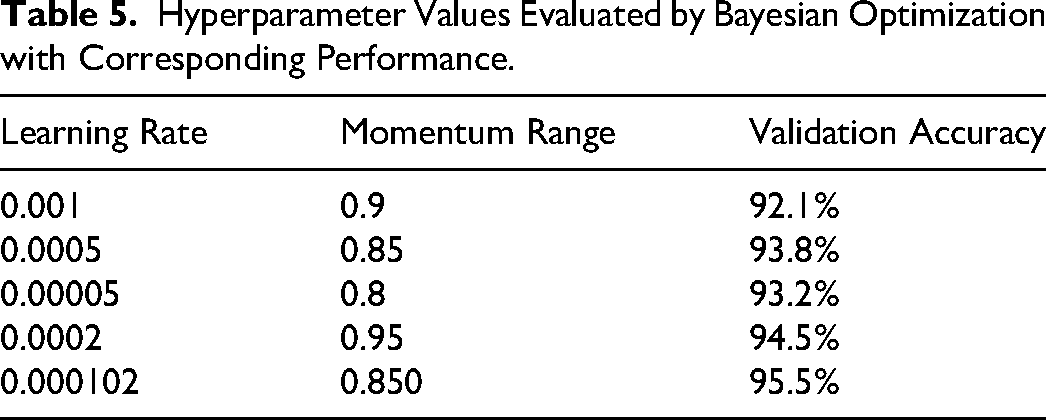
Using this formulation of BO, the initial learning rate and momentum range is passed as input and obtained optimized values such as 0.000102 and 0.850, respectively. However, different values of hyperparameters evaluated by the BO are shown in the Table 5.
Hyperparameter Values Evaluated by Bayesian Optimization with Corresponding Performance.

The trained model is finally employed in the testing phase for the feature extraction and final classification.
Proposed Architecture Testing Phase
In this section, the testing process of the proposed architecture has been presented with visual output, whereas the results are discussed under Section 3. The trained model is employed in this work and utilized the testing images set for features extraction. Features are extracted from the depth concatenation layer from the trained model using equation (13).
Where,
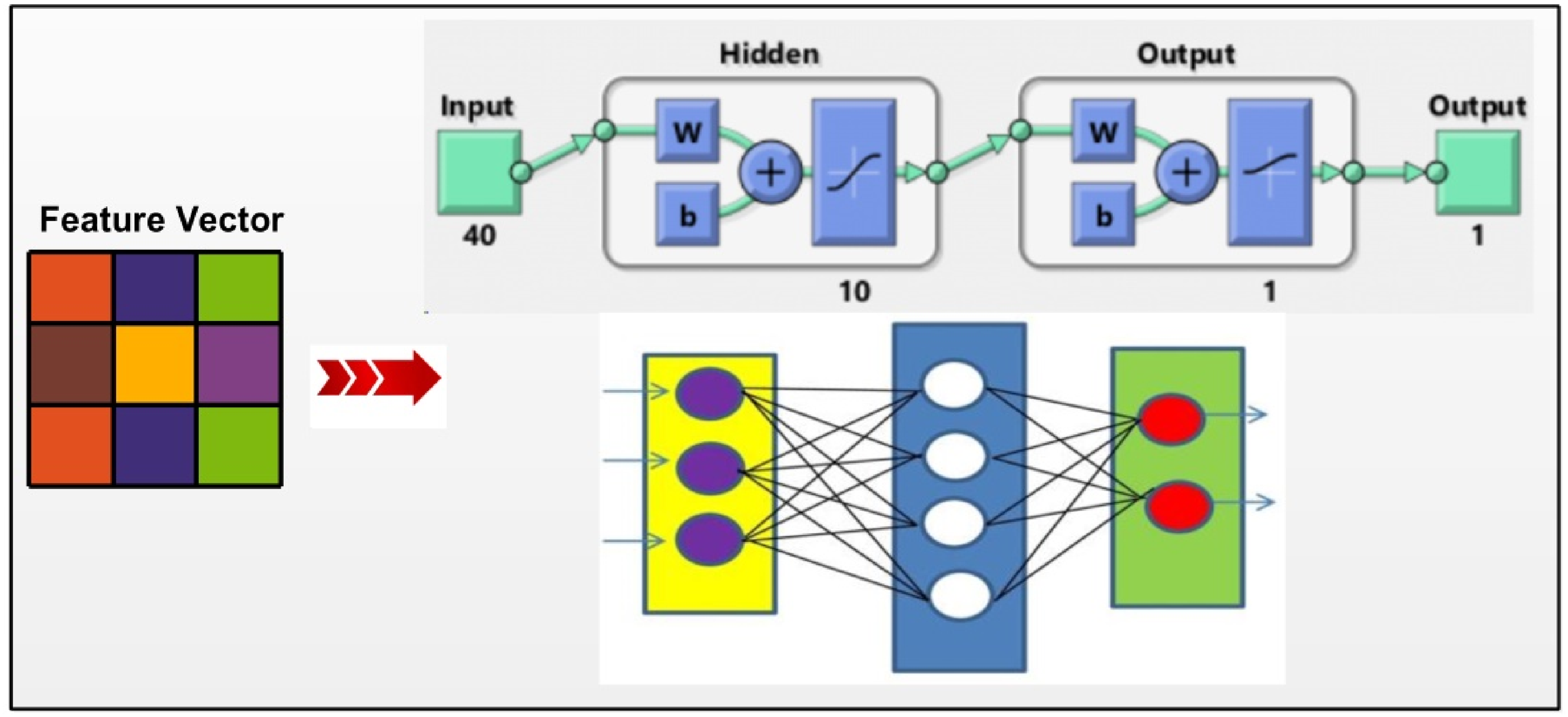
Proposed shallow neural network for the classification of cervical cancer.
Results and Discussion
In this section, the proposed architecture for the cervical cancer classification is interpreted through qualitative and quantitative measures. The dataset of this work is discussed under section 2.1 and sample images are shown in Figure 1. The dataset has been divided into a 50:50 approach and then performed data augmentation on training data. Using training data, the proposed architecture is trained and later utilized in the testing phase. In the testing phase, we utilized a 10 Fold Cross validation for the classification results, whereas the classification accuracy is compared with several well-known neural network and machine learning classifiers such as Cubic SVM, Fine Gaussian SVM, narrow neural network, medium neural network, Bi-layered Neural Network, and Tri-layered Neural Network, respectively. Each classifier performance is analyzed through accuracy, precision, recall, F1-Score (F1-Sc), FNR, and AUC. The entire architecture is implemented on MATLAB 2024a using a Personal Desktop Computer with 128GB of RAM, 20GB Graphics card Tesla V100.
Conducted Experiments
For the evaluation of the proposed architecture, we performed three different experiments:
In the first experiment, proposed 11-PIRBnet architecture is employed and trained on the selected dataset that later used in the feature extraction and classification. In the second experiment, proposed 9-PIRSANet architecture is utilized for the feature extraction and classification. In third experiment, the entire proposed fused 375NFNet architecture is employed for the feature extraction and classification.
Experiment 1
The results of this experiment are presented in Table 6. The proposed 11-PIRBnet architecture features are classified using machine learning classifiers and obtained the highest accuracy of 94.9% by Shallow NN classifier. Other measures such as recall is 94.8, precision is 95.0, F1-Sc is 94.89, FNR is 5.2, and AUC is 97.1, respectively. A confusion matrix shown in Figure 8 can further verify these measures. In this figure, it is observed that the correct prediction rate of Type 1, Type 2, and Type 3 is 94.6%, 95.9%, and 94.1%, respectively. The FNR of Type 3 is higher than Type 1 and Type 2 classes. PRC curves in Figure 8 also show strong performance with AUPRC values of 96.63% (Type 1), 97.53% (Type 2), and 96.81% (Type 3). Similarly, these measures are computed for other listed classifiers and obtained precision rate for each classifier is 92.1 (cubic SVM), 95.4 (FGSVM), 93.4 (Narrow NN), 94.9 (Medium NN), 90.0 (Bi-layered NN), and 89.5 (Trilayered NN), respectively. Overall, the achieved 94.9% accuracy demonstrates that this module is suitable for initial screening; however, the high false-negative rate suggests the need for an additional review mechanism. However, the balanced precision-recall curves across all types indicate that this module can serve as a classifier in low-resource settings.
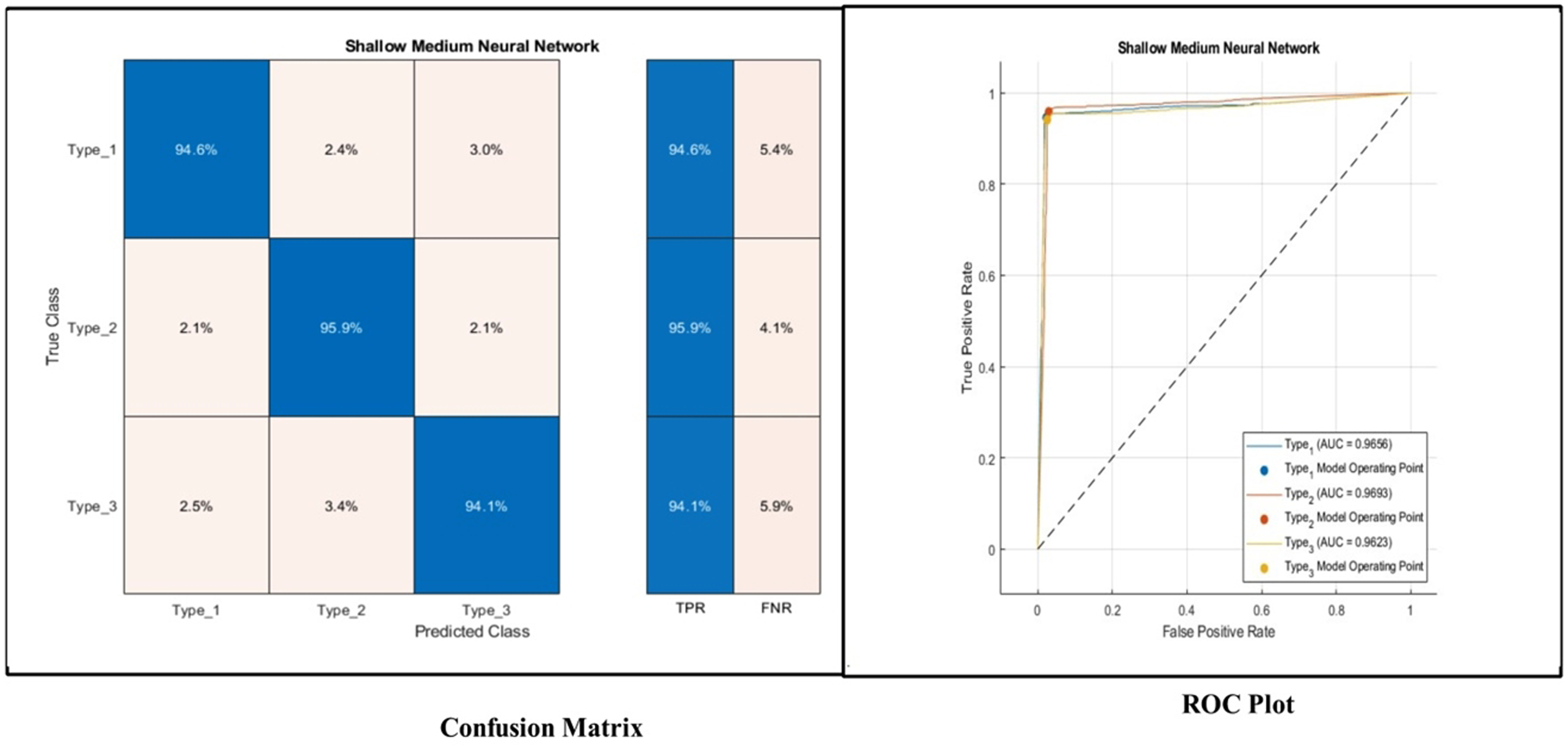
Confusion matrix, ROC plot and PRC plot of shallow neural network for proposed 11-parallel inverted residual blocks architecture.
Cervical Cancer Classification Results Using Proposed 11-Parallel Inverted Residual Blocks (11-PIRBNet) Architecture.

Experiment 2
Table 7 describes the results of this experiment. In this experiment, proposed 9-PIRSANet model features are extracted and passed to the classifiers for the classification. The Shallow NN classifier attained the utmost accuracy 94.6%, whereas the recall 94.8%, precision 95%, F1-Sc is 94.89, FNR value is 5.2, and AUC is 0.98, respectively. A confusion matrix shown in Figure 9 can further verify these measures. The correct prediction rate of each type in this figure is 95.5, 96.2, and 92.7%, respectively. Compared these prediction rates with Experiment 1, it is noted that a minor reduction is occurred for Type 3, whereas the improvement is noted in Type 1 and Type 2 class. Similarly, the precision rate for the other listed classifiers is 92.4 (Cubic SVM), 93.2 (FGSVM), 95.2 (Narrow NN), 90.7 (Medium NN), 90.1 (Bi-layered NN), and 77.6 (Tri-layered NN), respectively. Overall, it is noted that the performance of few classifiers has been improved and for the few of them has been minor reduced. The enhanced Type 1 accuracy demonstrates the importance of self-attention for rare anatomical presentations; however, the decline in accuracy of Type 3 suggests that this module may underperform on internally focused cervical views. To get the advantage of these two proposed architectures, we proposed a fused network that combine model and presented in the next experiment.

Confusion matrix and ROC plot for shallow neural network using proposed 9-PIRSANet.
Classification Results of Proposed 9-PIRSANet for Cervical Cancer Classification.

Experiment 3
In this experiment, the entire proposed architecture has been employed and performed classification. An analysis of deep features is performed from fused 375NFA network and passed to the classifiers for the final classification. Shallow NN obtained highest accuracy of 95.5% that is improved than the reported accuracy of experiment 1 (94.9) and experiment 2 (94.6), as presented in Table 8. The recall rate of Shallow NN is 95.4, precision rate is 95.4, F1-Sc is 95.4, and AUC is 0.97, respectively. These measures can be further verified through a confusion matrix, illustrated in Figure 10. This figure illustrates that type 1 class correct prediction rate is 96.9%, whereas the type 2 and type 3 correct prediction rates are 96.2 and 94.0%, respectively. Compared with experiment 1 and 2, it is observed that the correct prediction rate has been improved after the proposed networks fusion. Similarly, the precision rate of other classifiers is significantly improved such as 95.5 (FGSVM), 94.9 (Narrow NN), 95.7 (Wide NN), 94.8 (Bi-layered NN), 94.6 (Trilayered NN), respectively. The enhanced performance across all three types ensures the reliable identification of different anatomical representations while the balanced PRC curves ensure confident predictions. Hence, the proposed 375NFA architecture improved the accuracy and precision rates compared to the individual CNN architectures.

Confusion matrix of shallow neural network using proposed 375NFNet architecture.
Proposed Fused 375NFNet Architecture Results for Cervical Cancer Classification Using Cervical Screening Dataset.

Discussion
A detailed discussion has been conducted for the proposed architecture in terms of qualitative and quantitative measures. The proposed architecture is shown in Figure 2 that trained on Cervical Cancer images (see Figure 1). The data augmentation was performed at the initial phase to improve the training process. Later on, the each single proposed module is evaluated on testing images to check the strength of proposed model. Results of these modules are discussed in Tables 6 and 7, whereas the fused model results are presented in Table 8. From these results, this illustration shows that the fused structure significantly improved the correct prediction performance. To further validate the proposed model, we performed several ablation studies.
Ablation Study 1: Data Augmentation Versus Without Augmentation
In the first ablation study, we analyzed the performance of the proposed architecture after and before data augmentation. The main purpose of this ablation study is to analyze the effect of data augmentation step in the learning of CNN architecture. Figure 11 shows the accuracy and time analysis. It is observed in this visualization that the accuracy before augmentation was 84.6, 85.3, and 90.1% for individual CNN modules and proposed fused model. However, after the fusion process, the accuracy is improved 4%-9% that is a huge improvement. Similarly, the time comparison is also conducted just for the training process and it is observed that the time after the augmentation process was increased due to more number of images. Hence, this ablation study show that the augmentation process stretch the training time but on the other side, this step improved the testing accuracy of the proposed architecture.

Ablation study 1- accuracy and time comparison after and before augmentation using proposed architecture.
Ablation Study 2: Initial Learning Rate Selection
In the second ablation study, we analyzed our proposed architecture with different learning rate such as learning rate selection through BO and manual initialization. We initialized six manual initial learning rates such as M1 (0.001), M2 (0.005), M3 (0.012), M4 (0.00016), M5 (0.0002), and M6 (0.00001). Figure 12 illustrated training accuracy of this ablation study and it is noted that the selection through BO obtained higher training accuracy. Also, M4 and M5 obtained better performance on the selected dataset. Hence, it is concluded that the BO based initialization returned better training accuracy.

Ablation study 2- initial learning rate based training accuracy analysis on selected augmented dataset.
Ablation Study 3: Pre-Trained Models
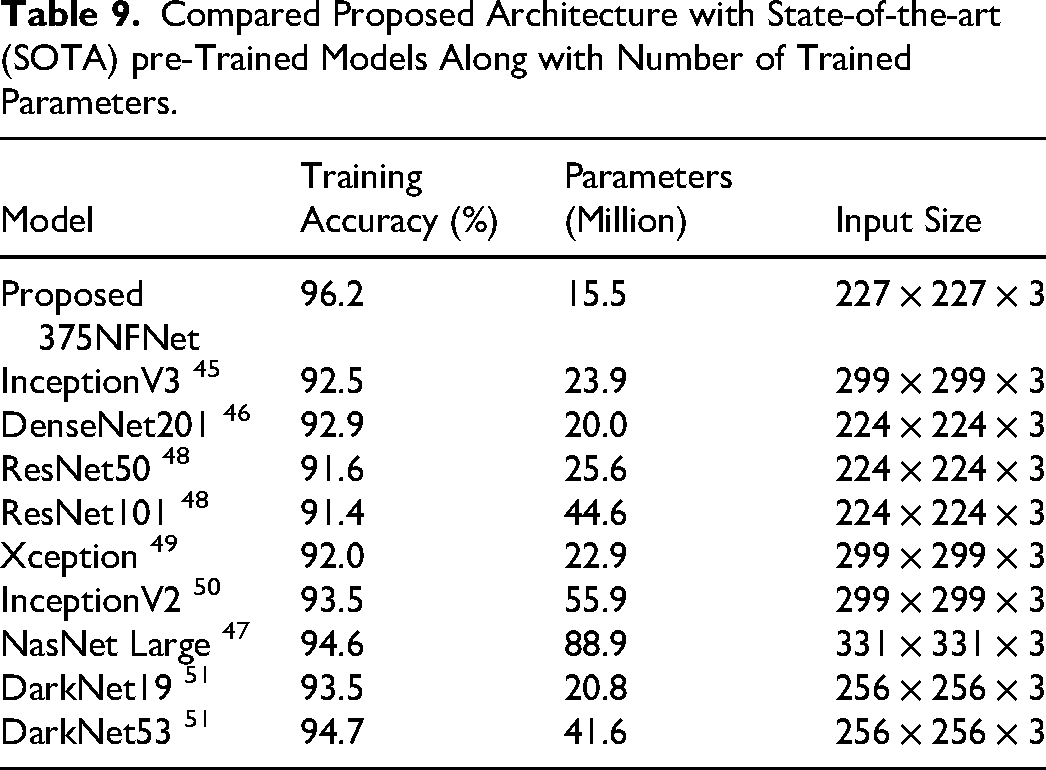
In this ablation study, we compared the proposed model with several pre-trained models, as listed in Table 9. In this table, accuracy, total trained parameters, and input size is given for each selected model. These models are trained on augmented cervical cancer dataset and training accuracy is noted for each. The hyperparameters are selected through BO, similar to proposed 375NFNet. The InceptionV3 45 obtained an accuracy of 92.5%, whereas the number of trainable parameters are 23.9(M). For the DenseNet201, 46 the obtained accuracy is 92.9%, whereas the trainable parameters are 20.0 (M). For the rest of the models, the training accuracy and total trainable parameters are (91.6, 25.6(M)), (91.4, 44.6 (M)), (92.0, 22.9 (M)), (93.5, 55.9(M)), (94.6, 88.9 (M)), (93.5, 20.8(M)), and (94.7, 41.6(M)), respectively. The NasNet Large 47 the model demonstrates that the fused framework has the highest number of trainable parameters, whereas the proposed 375NFNet achieves the lowest number of trainable parameters at 15.5 million (M) while maintaining a training accuracy of 96.2%.Hence, by leveraging a reduced number of trainable parameters, the proposed 375NFNet model achieved improved training accuracy and decreased computational time.
Compared Proposed Architecture with State-of-the-art (SOTA) pre-Trained Models Along with Number of Trained Parameters.
In the last, we also compared the proposed architecture testing accuracy with few pre-trained models while testing all of them on the same Cervical screening dataset. As listed in Table 10, it is noted that the EfficientNetB0 52 attained the utmost accuracy of 92.9%, and precision is 93.6, recall rate is 92.8, and F1-Score value is 93.19, respectively. The second best accuracy is achieved by SqueezeNet Pre-Trained53,54 model of 92.7%, whereas the precision rate is 93.5 and F1-Score is 93.09%, respectively. The Mobilenetv2 Pre-Trained 55 architecture is also obtained the third-best accuracy of 92.4% on Cervical Screening Dataset. When compared these models with proposed architecture, it is observed that the obtained accuracy is 95.5 and precision rate is 95.4% that is improved than the pre-trained models. In order to verify that the observed performance improvements of proposed 375NFNet over pretrained models are statistically significant rather than random variation, we conducted 10-fold stratified cross validation and paired t-tests. The 95% CI represents the range in which true accuracy difference lies with 95% confidence, t-statistic measures how many standard deviations the observed performance is from zero and p-value shows the probability that such improvement occurs randomly. As shown in Table 8, narrow range of 95% CI, high t-statistic values and low p-values suggest that proposed model is robust and statistically significant over pretrained architectures. However, this improved classification performance comes with a computational trade-off as the proposed 375NFNet requires moderate computational resources as compared to light-weight architectures like MobileNet-V2 and SqueezeNet, which makes it less practical for resource-constrained mobile devices. Figure 13 shows the final prediction results of the proposed architecture. In this figure, the original images features are extracted compared with trained model that in returned a label output such as Type 1, Type 2, and Type 3.
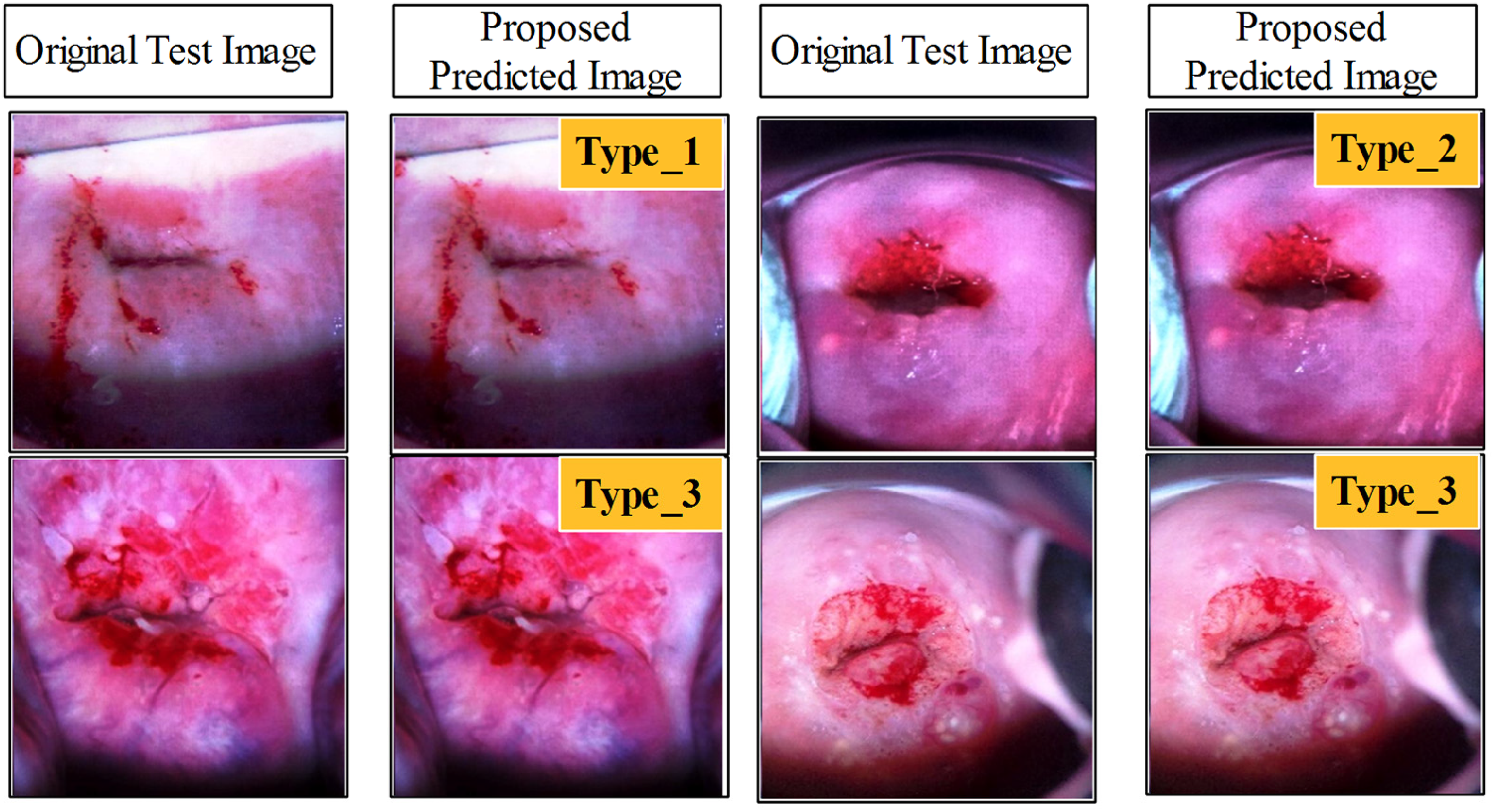
Proposed 375NFNet model prediction results with shallow neural network classifier.
Comparison of Proposed Fused Architecture Performance with Several State of the art pre-Trained Models Using Selected Cervical Screening Dataset.

Comparative Analysis with SOTA
Table 11 presents a comparative analysis of the proposed method with several state-of-the-art (SOTA) approaches for cervical cancer screening across different datasets and imaging modalities. The proposed approach achieved the highest accuracy of 95.5% on the Cervical Cancer Screening Datasets, slightly surpassing the 95.1% reported by Kalbhor et al 19 on Pap smear images and showing a clear improvement over Alquran et al, 5 who obtained 92% accuracy on a similar dataset. Compared to S. Nurmaini et al, 23 who achieved 90.78% using cervicograms, and Saini et al, 18 who reported 81.35% using National Cancer Institute colposcopy images, the proposed method demonstrates a substantial performance gain. Cai et al 20 attained 82.93% on the large-scale HiCervix dataset containing 29 classes, indicating the challenge posed by highly diverse and complex classification tasks. While dataset variability, imaging technique, and class complexity influence reported accuracies, the proposed method consistently outperforms or matches the best results across modalities. This performance advantage highlights the robustness and adaptability of the proposed model, suggesting strong potential for integration into real-world cervical cancer screening workflows and supporting its clinical applicability.
Comparative Analysis of the Proposed Method with State-of-the-art Approaches in Cervical Cancer Screening.

Limitations and Future Work
Despite achieving high classification accuracy, several limitations remain that need to be addressed. First, the proposed model is trained and validated on a single-source dataset from Kaggle, which limits its generalization and robustness. Evaluation of other datasets may affect the model's performance. Also, this model only detects different classes of cervical cancer rather than the presence of cancer itself. Moreover, while a 50-50 train-test split provides balanced evaluation, the limited training data are insufficient to capture the natural variability of clinical practice.
In the future, multicenter validation studies across diverse populations should be conducted to assess the model's generalization. Some explainable AI techniques should also be integrated to make this model reliable and practical for clinical use. The use of vision transformers and attention-based architecture can also be explored to further reduce the model's computational complexity.
Conclusion
Cervical cancer ranks as the fourth most prevalent cancer among women worldwide. The severity of cervical cancer complications underscores the need for effective solutions. This paper proposes a novel network-level fused deep learning architecture for cervical cancer classification. Publicly available cervical screening datasets from Kaggle were utilized to train and validate the proposed framework. Data augmentation was performed at the initial phase to handle the imbalance issue. Two novel deep learning modules are proposed: 11-PIRBnet and 9-PIRSANet. Both modules are fused at the network level via a depth concatenation layer, forming a new network called 375NFNet. After that, the proposed model is trained, and feature extraction is performed in the validation phase. Shallow neural network classifiers are used to classify the extracted features, achieving an accuracy of 95.5%. Based on these results, we conclude the following points:
Data augmentation resolves the problem of imbalance, resulting in better training accuracy. However, it has increased the training time compared to the original extracted dataset. Fusion of two CNN modules in a single layer, such as depth concatenation, improves the learning capability of a network, which in turn increases the precision and recall rates. In addition, the fused network preserves the total number of learnable parameters. Hyperparameter selection using BO improved the training accuracy compared to manual initialization, such as the learning rate. The higher training accuracy led to better testing performance.
Hence, these results show that the system could serve as a valuable decision-support tool for physicians, particularly in resource-constrained settings where both expertise and technology are limited. In the future, a vision transformer- and attention-blocks-based architecture will be designed to further reduce the number of trainable parameters.
Footnotes
Acknowledgments
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. RS-2023-00218176) and the Soonchunhyang University Research Fund. Authors like to thanks Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R440), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Ethics,Consent to Participate,and Consent to Publish Declarations
Not applicable.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Funding
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. RS-2023-00218176) and the Soonchunhyang University Research Fund. Authors like to thanks Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R440), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
