Abstract
Introduction
Pituitary neuroendocrine tumors (PitNETs) are a common type of intracranial benign tumor that can be divided into non-functioning PitNETs (NF-PitNETs) and hormone-secreting PitNETs including prolactinomas, growth hormone-secreting adenomas, ACTH-secreting adenomas, and TSH-secreting adenomas. 1 At present, dopamine agonists, somatostatin analogues and selective estrogen receptor modulators are employed in the treatment of hormone-secreting PitNETs, 2 while clinical trials have been conducted to evaluate the efficacy of medical treatment in NF-PitNETs, such as dopamine agonists.3-5 Our prior research identified DRD2 high expressed in 25%-30% of NF-PitNETs. 6 The efficacy of treatment is largely reliant on the expression of specific receptors on PitNETs cells, namely dopamine receptor D2 (DRD2), somatostatin receptor 2 (SSTR2), and estrogen receptor 1 (ESR1).7-9 Because of the absence of reliable methods to predict molecular expression, current protocols mandate a six-month pharmacological trial period with serial MRI evaluations and serum prolactin (PRL) monitoring, deferring surgical intervention until confirmed drug resistance. Developing a pretreatment radiomic prediction model could substantially shorten ineffective treatment durations, thereby reducing medication-related adverse effects, financial burdens, and risks of tumor progression. For high-surgical-risk NF-PitNETs, noninvasive imaging assessment of DRD2 expression could guide personalized DA therapy selection. These advancements suggest that integrating radiomic biomarker analysis with molecular pathology testing within clinical workflows may enable precise patient stratification and optimized therapeutic decision-making. Such multimodal classification models could shift PitNET management from empirical approaches to precision medicine paradigms. Consequently, prediction of DRD2, SSTR2, and ESR1 expression status represents a critical advancement for personalized PitNET management, potentially transforming clinical practice through targeted treatment allocation and resistance anticipation.
Some studies have been conducted to assess the magnetic resonance imaging (MRI) signal intensity and DA/SSA resistance in prolactinomas and growth hormone (GH)-secreting tumors.10-12 Dogansen et al reported a correlation between the intensity of the baseline T2-weighted signal and drug responsiveness in both prolactinomas and GH adenomas, 10 but Burlacu et al failed to find the same association. 11 Other studies used radionuclides to predict the drug response of DAs or SSAs.13,14 However, the research result value has been reduced due to the high equipment requirements, expensive inspections, and small research scales. Moreover, although ESR1, as the main target of endocrine therapy, correlates with the drug resistance and invasiveness of PitNETs, few studies have constructed a preoperative predictive model to predict its expression level. 15 Above all, a predictive tool with high repeatability, easy implementation, and easy promotion is urgently needed to profile the expression of key targets for drug treatment in PitNETs and help make individual therapeutic decisions and further clinical research.
The area of radiomics analysis has grown tremendously in recent decades with the development of identification tools and the increase in the size of datasets. This is a new area that refers to the process of converting high-throughput medical image information into high-dimensional data and then mining it to provide evidence to improve clinical decision. Radiomics based on high-resolution digital image characterization is more effective than other conventional approaches. 16 Previous studies demonstrated that radiomics has a good predictive ability for the diagnosis and treatment of PitNETs, including the ability to predict whether the tumor invades the cavernous sinus, the extent of tumor consistency, and the proliferative index, furthermore, several studies have reported that radiomics can predict not only the subtype of PitNETs, but also the sensitivity to conventional treatment.17-22 Rui et al developed an ensemble learning model for preoperative prediction of silent cortical adenoma subtypes in nonfunctioning PitNETs, with an area under the curve (AUC) of 0.927. 23 However, there are some shortcomings and drawbacks in analyzing images with traditional radiomics in terms of object segmentation and hard-coded characteristic acquisition. For further analyze image data, the newly developed deep learning radiomics methods can extract quantitative and tremendous information derived from digital images through supervised learning. 24 To date, no studies have used deep learning to predict the expression of drug targets in PitNETs. Therefore, if the radiomics, and deep learning features can be reasonably combined, it will be of great help in the establishment of predictive models.
A dynamic multilayer perceptron (MLP) was developed based on image representation, and finally the MLP interacts with multimodal features in higher and wider dimensions to fully exploit the potential of multimodal information. They performed extensive experiments to demonstrate the effectiveness of such a method. Thus, this study aimed to compare the single machine learning model and the multimodal models of joint deep learning features, prove that adding deep learning features can enhance the predictive performance of the model. Eventually, our study developed a predictive model to predict the expression of DRD2, SSTR2, and ESR1 in PitNETs using dynamic MLP fusing radiomics and deep learning features.
Materials and Methods
Study Design
This retrospective study was conducted in accordance with the TRIPOD statement for prediction model development and validation. 25 Patients with PitNETs were collected (n = 256) to perform a retrospective study. The diagnosis of these patients was verified by an examination of archived histopathological slides, which were reviewed by experienced neuropathologists to ensure accurate diagnosis of PitNETs. Characteristic data including gender, age, hormone-hypersecreting tumors, Knosp grade and tumor diameter were collected.
The inclusion criteria included (1) patients with a confirmed diagnosis of PitNETs by histologic and immunohistochemical staining; (2) identifiable MRI data, including preoperative and postoperative, as well as those containing T1-weighted (T1WI), T2-weighted (T2WI), and contrast-enhanced T1-weighted (T1CE); and (3) MRI images obtained within one week preoperative. Exclusion criteria: (1) Patients who have received surgical management or radiotherapy, or chemotherapy or for PitNETs; (2) PitNETs with unclear immunohistochemical staining; (3) poor quality of MRI images; (4) poor quality RNA of tissue samples. Finally, there were 186 patients included in our study. The cohort size was determined using the event-per-variable rule (EPV ≥10) and radiomics stability thresholds, ensuring robustness for 15-20 candidate features per receptor. All eligible cases from 2017–2020 were included to maximize generalizability.
Tissue Microarray
In order to confirm the presence of PitNETs tissue all the hematoxylin eosin-stained original sections were reviewed. Typical areas of tumor tissue were obtained for tissue microarrays (TMAs), which were constructed from formalin-fixed paraffin-embedded tissue samples containing replicate 1 mm cores.
Monoclonal antibodies were used to detect DRD2 (Abcam, ab150632), SSTR2(Abcam, ab134152), and ESR1 (Abcam, ab32063) following the previously described protocols. 26 The quantification of the stain intensity of DRD2, SSTR2, and ESR1 was conducted by immunoreactive score (IRS), a score of 0-1 for negative staining, 2-3 for low score, 4-8 for medium score, 9-12 for high score. 27
Quantitative Reverse Transcription Polymerase Chain Reaction (RT-qPCR)
PitNETs specimens were collected immediately after surgical resection and stored in a refrigerator at −80 °C for further examination. Trizol reagent (Invitrogen, Carlsbad, CA, USA) was used to extract the total RNA according to the manufacturer's instructions. The RT-qPCR was performed as described previously, 28 using the SYBR Premix ExTaqTM (TAKARTA) with the Applied Biosystems Prism 7900 system (Applied Biosystems, Life Technologies, USA). Furthermore, the results of RT-qPCR were normalized according to the geometric mean of the reference gene (glyceraldehyde-3-phosphate dehydrogenase). The primers used for RT-qPCR are listed in Supplementary Table 1.
Image Acquisition
MRI examinations were performed using a 3.0 T scanner (Achieva, Philips Healthcare, Amsterdam, The Netherlands) equipped with a dedicated eight-channel head coil. MRI scans were reviewed by two neuroradiologists (Dr Y and Dr Z, 10+ years of experience) to confirm tumor characteristics. ROI delineation was performed by trained researchers under neurosurgeon (21 years of pituitary surgery experience, 10 years of pituitary surgery experience) supervision, with histopathology correlation for validation. The image acquisition protocol included the following sequences: Coronal T1-weighted imaging with the following parameters: repetition time (TR) of 300 ms, echo time (TE) of 16 ms, field of view (FOV) of 150 mm, slice thickness of 3 mm, and matrix size of 232 × 192. T2-weighted imaging with identical parameters to the T1-weighted sequence, except for a matrix size of 232 × 196. Contrast-enhanced T1-weighted images with the same parameters as the T1-weighted sequence, but with a matrix size of 216 × 167. For the contrast-enhanced T1-weighted images, the acquisition time ranged from 2 to 4 min after injection of 0.1 mmol/kg gadolinium-based contrast agent, administered intravenously at a rate of 2 mL/s via the antecubital vein.
Image Segmentation and Feature Extraction
3.0 T MRI (Achieva, Philips Healthcare, Amsterdam, The Netherlands) scanning of pituitary gland and dynamic 3.0 T MRI with Gd-DTPA contrast enhancement should be performed to identify the presence of PitNETs. Patients were examined in the supine position. The last MRI images within one week before surgery were obtained as the ultimate medical images. Moreover, we confirmed the inclusion of T1WI, T2WI and T1CE sequences in the coronal and sagittal positions in the pre- and post-operative images. All patients’ scanned images were manually drawn for regions of interest (ROIs) by two investigators under the supervision of an experienced neurosurgeon using ITK-SNAP software (University of Pennsylvania, www.itksnap.org). Based on the marked ROI image, all the images are cropped into a three-dimensional lesion area to eliminate other irrelevant information in the image. The T1WI image was used as a benchmark, and the images of the other two modalities were resampled to the T1WI image for subsequent multimodal input to make the voxel difference between images of different modalities as little as possible. Finally, all images were normalized by Z-Score before being input into the network to reduce the pixel intensity variation between different patient images, which made the pixel intensity scale in a certain range.
Then, the Densely Connected Convolutional Network-121 (DenseNet-121) architecture was adapted to develop our deep convolutional neural networks (DCNNs) for deep learning feature extraction. In this study, 1037 radiomics features were extracted from each modality image of each patient's MRI. These features included shape and size features, texture features, first-order and high-order features.
Model Establishment and Validation
The MRI dataset of 186 patients with PitNETs were divided into training sets (148 patients) and validation sets (38 patients) with a ratio of 4:1. Five-fold cross-validation was conducted on the training set to refine the training of the network parameters. Subsequently, the training set and optimized parameters were used to train the final model, and the validation set results were calculated and statistically analyzed according to the indicators. Additionally, multiple groups of comparative experiments were designed, and the effect of various traditional machine learning on this task was reviewed to validate the effectiveness of the methods in our study. To address potential biases from various scanners, we applied z-score normalization to all extracted features. Irrelevant features were removed using the T-test, concentrating on those with p-values less than 0.05. Following this, we utilized the Least Absolute Shrinkage and Selection Operator (LASSO) to discard features with zero coefficients and to highlight the most significant features. And Random Forest, Support Vector Machine, Logistic Regression, and K-Nearest Neighbor models were established separately. The predictive performance of the model was evaluated using a range of metrics, including the area under the receiver operating characteristic curve (AUC) with a 95% confidence interval (CI), accuracy (ACC), sensitivity (SEN), and specificity (SPE).
Network Architecture
The dense dilated inception network comprised the backbone network DenseNet-121, a radiomics branch network, and dynamic MLP. The backbone network DenseNet-121 down sampled the image and extracted features of different sizes, different dimensions, and more abstract. The radiomics branch network comprised two linear layers to find the nonlinear relationship between radiomics and predicted values. The first linear layer reduced the dimension of one-dimensional features to 1024 dimensions, and the second layer reduced the dimension. When it reaches 512 dimensions, the dynamic MLP model finally maps the omics features to the image features to obtain a weighted feature map for output. Each input image feature of the dynamic MLP model was the output of the previous block. First, the one-dimensional radiomics features were reshaped into a two-dimensional matrix to map them to image features. Then, weights were extracted using the Rectified Linear Unit activation function and layer normalization. Finally, the weights were mapped to image features.
Statistical Analysis
The continuous variables (age, tumor volume, and tumor max size) were analyzed using the Kruskal–Wallis rank sum test, 29 and the discrete variables (sex, Knosp score, p53, and ki-67) were analyzed using the chi-square test to determine the differences in the distribution of variables between cohorts. All tests were two-sided with a significance of 0.05.
Results
Cohort Characteristics and Data Partitioning
Following TRIPOD guidelines for prediction model development and validation, the enrolled patients were stratified into training (n = 148) and validation (n = 38) sets using randomized stratified sampling to preserve the distribution of key clinical outcomes (Figure 1). Baseline demographic and clinical characteristics are summarized in Table 1. The average age of these patients was 51.9 years and ranged from 16 to 82 years old. 56 patients (30.1%) were functional PitNETs and 130 patients (69.9%) were non-functional PitNETs. Knosp grading stratification revealed 117 patients (62.9%) with grade 0–2 (non-invasive) and 69 (37.1%) with grade 3–4 (invasive) lesions. Immunohistochemical results identified p53 positive in 69 cases (37.1%) and elevated Ki-67 (≥3%) in 46 (24.7%). No significant differences between training and validation sets in age, sex, or functional status (p > .05). However, the validation set exhibited higher proportions of Knosp grades (p = .02) and p53 positivity (p = .037), reflecting intentional stratification to test model generalizability (Supplementary Tables 2 to 4).
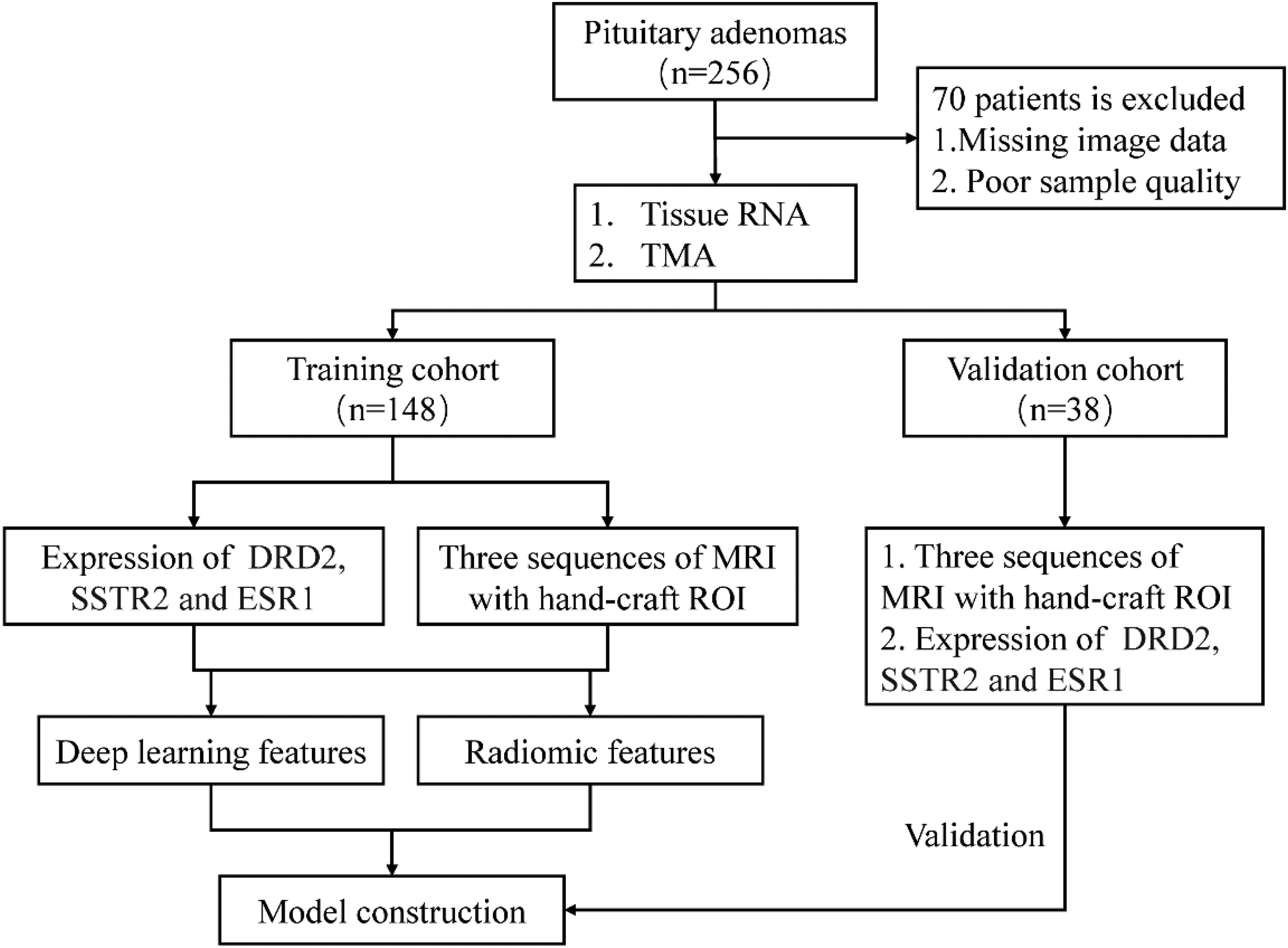
Flow Diagram of Patient Enrolment.
Clinical Characteristics of Patients in the Training and Validation Sets.

Feature Selection and Model Development
Previous studies showed that the mRNA expression of ribonucleic acid of ESR1, DRD2, and SSTR2 strongly correlated with the expression of protein levels.30,31 Following the Image Biomarker Standardisation Initiative (IBSI) guidelines, a radiomics workflow was implemented to extract 1284 quantitative features from preoperative T1/T2-weighted MRI sequences. Next, univariate t-test filtering (p < .05) was used to eliminate non-discriminative features, followed by LASSO regression (λ optimized via 10-fold cross-validation) to address multicollinearity, identifying 23 robust radiomic features predictive of molecular targets (ESR1, DRD2, SSTR2). These selected features were then used to train four traditional machine learning models: Random Forest, Support Vector Machine, Logistic Regression, and K-Nearest Neighbor (Supplementary Figure 1). In contrast to conventional multimodal fusion approaches, dynamic Multilayer Perceptron (dMLP) architecture was developed to synergistically integrate handcrafted radiomic features with deep learning-derived convolutional features (Figure 2). The dMLP employed adaptive attention gates to dynamically weight contributions from radiomics and neural network outputs, enabling context-aware fusion of multimodal information. This fused representation was further processed through two fully connected layers with dropout regularization (rate = 0.3) to enhance generalization, thereby addressing overfitting risks while improving the model's capacity to map complex relationships between imaging signatures and molecular expression. This model ensured both interpretability and high-dimensional pattern capture, establishing a robust framework for non-invasive prediction of therapeutic targets in PitNETs.

The Analysis Process of Radiomics. The Radiomics and Deep Learning Feature Extraction and Dimension Reduction Process Shown in the Figure is from the Preoperative Coronal MRI.
Model Performance and Clinical Utility
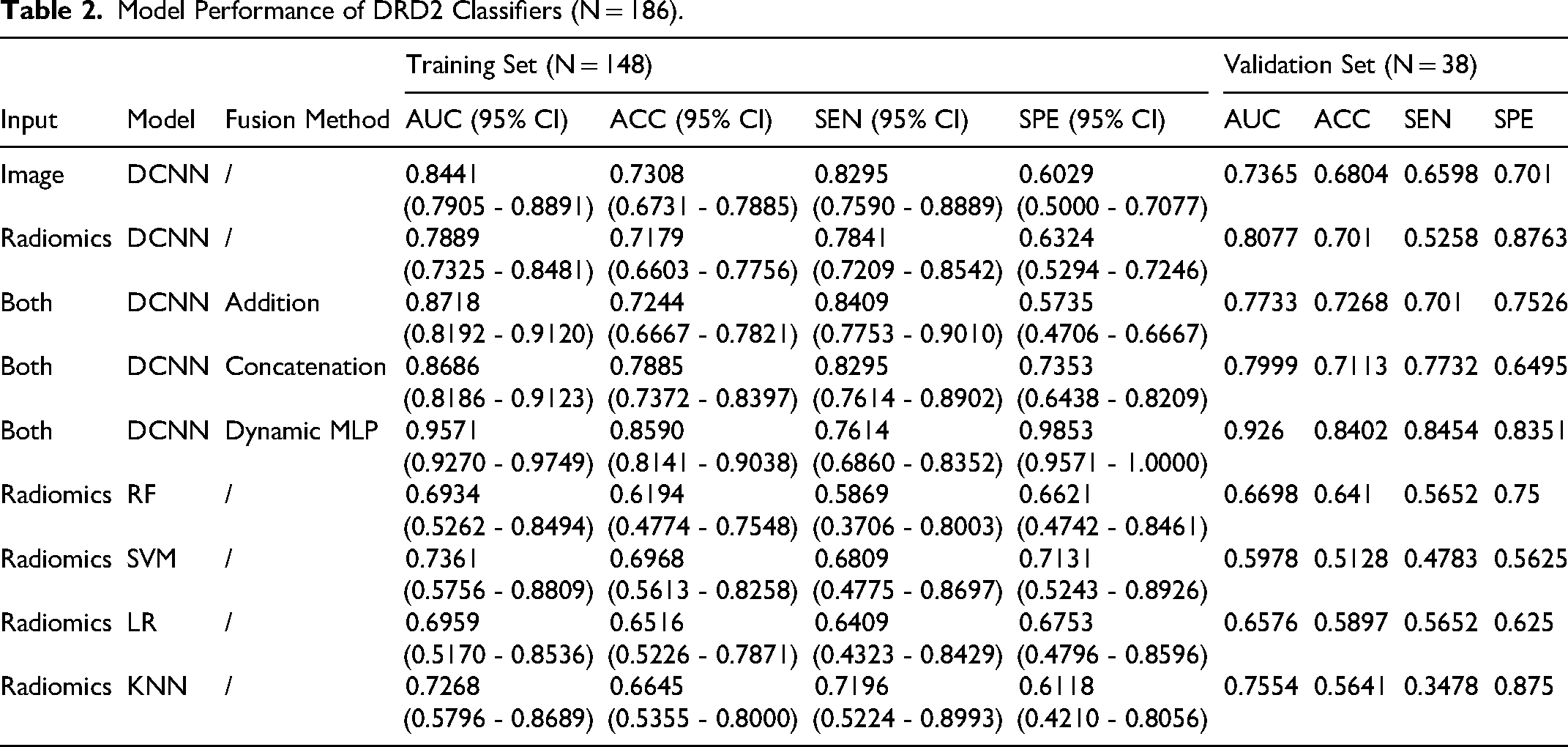
All models were evaluated using AUC, sensitivity, specificity, and balanced accuracy (Tables 2 to 4). The performance of the single machine learning classifiers ranged from 0.6322 to 0.9037 in the training set, but it ranged from 0.5322 to 0.7927 in the validation set, indicating overfitting. In contrast, the dMLP model demonstrated superior generalizability, achieving 27%-34% improvements over best-performing traditional models (Figure 3). The receiver operating characteristic curves of DRD2 classifiers, SSTR2 classifiers, and ESR1 classifiers was shown in Figure 4A-1. The accuracy and robustness of the deep learning model were superior to that of traditional machine learning model. Moreover, compared with pure deep learning models, combining machine learning features improved the performance of the model significantly. Results showed that the dynamic MLP model could increase the AUC of DRD2, SSTR2, and ESR1 by about 30%. The AUC of the three classifiers of the validation set using the dynamic MLP model was 0.9260 (DRD2), 0.9084 (SSTR2), and 0.9409 (ESR1). Using this model to guide individual therapeutic decisions (Figure 4J to L), the decision curves indicated that, most times, the dynamic MLP model could add more benefit to patients.

Construction of Densely Dynamic Integration Net.
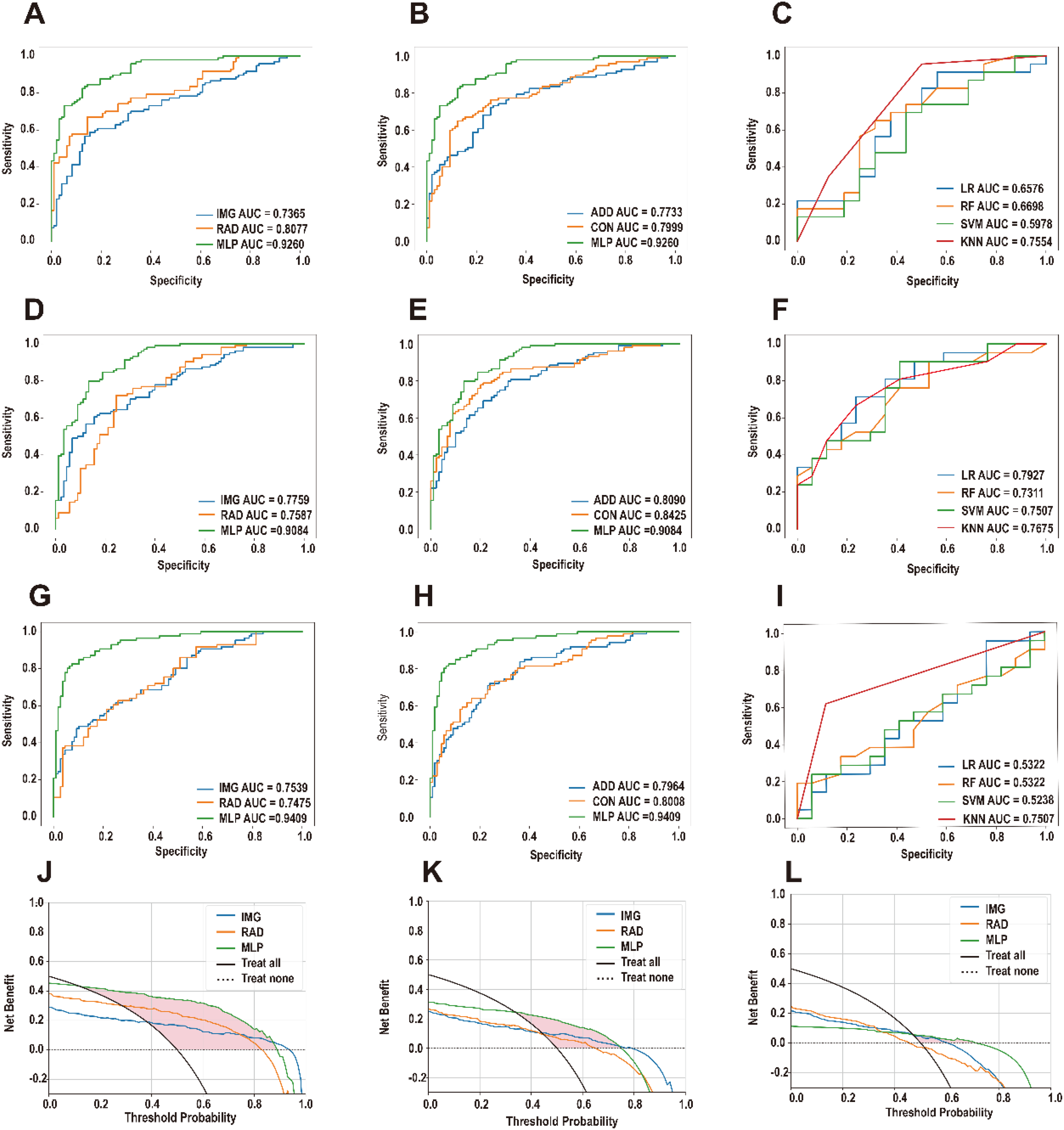
The Model Performance In Validation Set. The Roc Curve And Auc Value of Simple Radiomics Feature Model, Simple Deep Learning Feature Model, Radiomics And Deep Learning Feature Addition Model, Radiomics and Deep Learning Feature Concatenation Model, Mlp Fusion Model Developed By Dcnn, and the Machine Learning Model Classifiers Developed for DRD2 (A-C), SSTR2 (D-F) and ESR1 (G-I). Decision Curve Analysis of DRD2 (J), SSTR2 (K) and ESR1 (L) Models. The Results Showed that the Model Using Mlp Fusion Had Greater Benefitto Patients in Most Cases.
Model Performance of DRD2 Classifiers (N = 186).
Model Performance of SSTR2 Classifiers (N = 186).

Model Performance of ESR1 Classifiers (N = 186).

Discussion
There are many subtypes of PitNETs and medication is a very important treatment for these patients. It is the first line therapy for prolactinomas and an important adjuvant treatment for other types of adenomas, and even patients with refractory PitNETs require a combination of drugs. Many patients experience anxiety at the start of treatment due to concerns about the efficacy of the drugs, especially younger patients with functional adenomas and those who have not been able to control tumor size and hormone levels well after other therapeutic modalities. Therefore, the development of a method to predict the effect of drugs in patients with PitNETs is essential and may help patients to make an optimal choice.
Previous studies have proved that the expression level of PitNETs key receptors strongly correlates with responsiveness to drug treatment.1,7,9 However, there is currently a lack of a strong noninvasive predictive model of key receptors in PitNETs. The study findings revealed that the three deep learning models based on the combination of the MRI radiomics features and deep learning features predicted the expression levels of three key receptors (DRD2, SSTR2, and ESR1) in patients with PitNETs with high diagnostic accuracy, with high AUC of 0.9260 (DRD2), 0.9084 (SSTR2), and 0.9409 (ESR1), respectively. Since our model is based on routinely acquired structural MRIs without extra cost and invasive manipulations, and it can be applied completely automatically on a conventional computer in a short time, the approach appears to be helpful in guiding the clinical management of PitNETs.
For noninvasive testing of PitNETs receptors, radionuclide testing has been demonstrated in some studies. Tang et al reported that 18F-fallypride positron emission tomography (PET)/MRI revealed the expression of DRD2 in prolactinomas, which were confirmed in 7 patients. 32 For noninvasive profiling of the expression of SSTRs, the 68Ga-DOTATATE PET/computed tomography is of considerable value in the detection of receptors in somatotropin adenomas. 13 However, this detection method or research has considerable limitations, such as high costs, low resolution for microadenomas and small number of research cases. Therefore, target detection through radiomics has obvious advantages over it. Yae et al reported the usefulness of radiomics for predicting drug response in patients with PitNETs. 33 They developed a model to predict the response to DA, which achieved an AUC of 0.78. Similarly, in this study, we performed noninvasive predictive modeling of drug target receptor expression in PitNETs and achieved the expected performance of the model. Therefore, this noninvasive predictive model is more scientific and valuable than traditional drug target prediction.
Moreover, our study applied the latest deep learning multimodal fusion method, which included the dynamic MLP model that promoted the image feature's representation ability guided by the multimodal features and improved robustness and generalizability compared to a single predictor. In this study, the combination of radiomics features and deep learning features resulted in almost 30% higher AUC than radiomics features alone. Rui et al verified that the ensemble learning model outperformed the individual models. 23 Our research combined radiomics and deep learning features, similar to the research by Rui et al Furthermore, we constructed, and verified the superiority of the integrated learning model for tumor classification.
Meanwhile, for the drug treatment of PitNETs, the prediction of a single receptor does not always meet the individualized treatment needs of all patients. A prospective trial of 24 patients with acromegaly showed that the combination of cabergoline and a somatostatin receptor ligand (SRL) resulted in a normalized insulin-like growth factor 1 level in 42%-60% of patients, whereas SRL alone did not normalize GH levels, and 21%-71% of patients had GH levels above 2.5 g/L. 34 Additionally, Souteiro et al reported that SSAs and SERMs could be complementary treatments for drug-resistant prolactinomas. 35 Therefore, multitarget prediction is more meaningful for drug treatment. In our study, we simultaneously predicted the three main targets with clinical application value at the same time to improve the quality of individual treatment. When patients with PitNETs are faced with treatment choices, it is possible to predict the potential effect of drugs and provide them with more accurate treatment information to help them make the optimal choice.
However, there were some limitations in this study. Since the poor MRI image quality and sample freshness, the number of patients with PitNETs included in our study was limited. The cohort included only 56 functional PitNETs, limiting statistical power for hormone-specific analyses. This imbalance may reduce the generalizability of the model to rare functional subtypes. While our cohort meets radiomics sample size guidelines, larger multicenter studies and multicenter cohorts are needed to confirm generalizability. In addition, although the fusion of qPCR and TMA data has been proved,36,37 and the coincidence degree is good in our results, further experiments are still needed for verification.
Conclusion
In conclusion, we trained a dynamic MLP DCNN model to differentiate patients with high and low expression of three key PitNETs receptors using traditional radiomics features combined with deep learning features. Considering the reported heterogeneity between the PitNETs receptor profiles, this precise, and practical radiomics model could provide valuable information for personalized treatment decision in PitNETs.
Supplemental Material
sj-docx-1-tct-10.1177_15330338251353305 - Supplemental material for Multimodal Model for Non-Invasive Detection of DRD2, SSTR2 and ESR1 Receptor Profiling in Pituitary Neuroendocrine Tumors: A Retrospective Study
Supplemental material, sj-docx-1-tct-10.1177_15330338251353305 for Multimodal Model for Non-Invasive Detection of DRD2, SSTR2 and ESR1 Receptor Profiling in Pituitary Neuroendocrine Tumors: A Retrospective Study by Jianglong Lu, Xianpeng Wang, Jinghao Jin, Fanjie Xu, Runhua Tang, Cheng Han, Zerui Wu, Zhipeng Su and Yuhang Guo in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Ethical Considerations
Given the retrospective nature of the study, patient informed consent was waived. This study was approved by the Clinical Research Ethics Committee of the First Affiliated Hospital of Wenzhou Medical University (permission number: 2025-R061).
Author Contributions
All authors contributed to the study conception and design. All authors read and approved the final manuscript. All authors contributed as follows: Xianpeng Wang: Conceptualization, Investigation, Data curation, Writing - original draft. Jinghao Jin: Investigation, Methodology, Software, Writing - original draft. Fanjie Xu: Formal analysis, Methodology, Software. Jianglong Lu: Funding acquisition, Methodology, Investigation. Runhua Tang: Methodology, Investigation. Cheng Han: Methodology, Investigation. Zerui Wu: Methodology, Investigation. Zhipeng Su: Funding acquisition, Supervision, Writing - review & editing. Yuhang Guo: Supervision, Writing - review & editing.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from - Basic scientific research project of Wenzhou City of China (grant number Y2023150), Key project jointly constructed by the Science and Technology Department of the State Administration of Traditional Chinese Medicine and the Zhejiang Provincial Administration of Traditional Chinese Medicine (grant number GZY-ZJ-KJ-24087).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Due to patient confidentiality, the MRI con not be provided in the manuscript. If the reader wants to get more information about the MRI, they can contact the corresponding author for further information.
Clinical Trial Number
MR-33-25-025051.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
