Abstract
Introduction
Preoperative differentiation among adenocarcinoma in situ (AIS), minimally invasive adenocarcinoma (MIA), and invasive adenocarcinoma (IAC) is crucial for guiding ground-glass nodule (GGN) management. This multicenter study evaluated the comparative utility of deep learning (DL), radiomics, and conventional machine learning (cML)-based clinicoradiographic models for this ternary classification.
Methods
We developed four DL models (DenseNet-121, ResNet-10, ResNet-18, and VGG-13) for the ternary classification of AIS, MIA, and IAC using multicenter CT datasets. For comparative analysis, we constructed two additional classification models: (1) a radiomics model employing feature engineering through analysis of variance, recursive feature elimination with cross-validation, and least absolute shrinkage and selection operator, and (2) the cML-based clinicoradiographic model utilizing 12 different classifiers. The performance of all models was evaluated using the macro area under the curve (Macro-AUC) metric.
Results
847 GGNs postoperatively confirmed as lung adenocarcinoma were included in this multicenter study, which were randomly split into a training set (70%, n=592) and a validation set (30%, n=255). The DL model ResNet-10 demonstrated superior performance, achieving a Macro-AUC of 0.8055 (95% CI: 0.7723-0.8387), an accuracy of 0.6300 (95% CI: 0.5541-0.6764), and an F1-score of 0.4206 (95% CI: 0.3821-0.4598). This performance surpassed that of the radiomics model, which had a Macro-AUC of 0.7801 (95% CI: 0.7432-0.8170), an accuracy of 0.6100 (95% CI: 0.5276-0.6204), and an F1-score of 0.5505 (95% CI: 0.4983-0.6017), and the cML-based clinicoradiographic model, which achieved a Macro-AUC of 0.7770 (95% CI: 0.708-0.846), an accuracy of 0.6000 (95% CI: 0.5376-0.6604), and an F1-score of 0.4438 (95% CI: 0.3925-0.4961).
Conclusion
The ResNet-10 network established a novel ternary classification model for predicting the invasiveness of GGNs. This approach provides clinically actionable insights that support surgical planning and facilitate risk-adapted management.
Introduction
Lung adenocarcinoma, the dominant histological subtype of lung cancer, remains a primary contributor to global cancer-related mortality. 1 The expanded use of low-dose computed tomography (CT) screening has significantly increased the detection of early-stage lung adenocarcinomas, which often present as pulmonary ground-glass nodules (GGNs). 2 Current CT-based classification categorizes GGNs into two types: (1) pure GGNs, characterized by homogeneous ground-glass attenuation without any solid component on a mediastinal window, and (2) part-solid nodules, which contain homogeneous ground-glass attenuation along with a solid component on a mediastinal window. While pure GGNs are generally associated with preinvasive lesions, such as adenocarcinoma in situ (AIS) or minimally invasive adenocarcinoma (MIA), pathological examinations reveal invasive adenocarcinoma (IAC) in up to 27% of cases. 3 This discrepancy between radiological appearance and pathological invasiveness underscores the diagnostic challenges associated with GGNs. Consequently, optimizing CT-based diagnostic criteria and refining surgical management strategies for GGNs have become critical priorities in early-stage lung cancer research.4–6
Current challenges in predicting invasiveness primarily stem from overlapping clinicoradiographic features (eg, age, lobulation, and spiculation) and the limited representativeness and high subjectivity among GGN subtypes. 7 Radiomics models, while proficient at extracting quantitative imaging features, often rely on handcrafted feature sets (eg, shape, texture, and wavelet features) that are constrained by mathematical formulations. This can lead to the oversight of biologically relevant patterns not captured by existing radiomics analysis.8–10 In contrast, deep learning (DL) leverages convolutional networks to automatically extract discriminative features from chest CT scans. This method supports end-to-end training from pixels to diagnostic outcomes, optimizing feature extraction and classification jointly. Consequently, it enables accurate preoperative stratification of invasiveness in GGNs by capturing subtle spatial patterns beyond human visual perception.11,12
However, existing DL models have largely focused on computer vision methods, and most studies have limited their outcomes to binary classifications (eg, AAH, AIS, and MIA vs IAC13,14 or AAH and AIS vs MIA and IAC15,16). Only a small number of studies have developed ternary classification models,17,18 highlighting the critical need for accurate preoperative stratification. While regular follow-up is recommended for AIS, sublobar resection is often adequate for cases of MIA; in contrast, IAC typically necessitates lobectomy with lymph node dissection.4–6 Moreover, existing models predominantly rely on single-center cohorts with limited sample sizes and lack comparative validation against radiomics or clinicoradiographic benchmarks.13–18
To address these gaps, this multicenter study develops four mainstream DL architectures (DenseNet-121, ResNet-10, ResNet-18, VGG-13) for the ternary classification of AIS, MIA, and IAC, and rigorously benchmarks their performance against radiomics and clinicoradiographic models. Our work aims to establish an optimal stratification framework to guide personalized management strategy and surgical intervention for early lung cancer patients.
Materials and Methods
Patient Enrollment
Ethics Approval Statement This multicenter retrospective study was reviewed and formally approved by the institutional review boards (IRBs) of Xiangtan Central Hospital (Approval No. 2021-07-009), Liuyang Hospital of Traditional Chinese Medicine (Approval No. 2022-016), and the Affiliated Hospital of Southwest Medical University (Approval No. KY2020147). The study was conducted in accordance with the ethical standards laid down in the Declaration of Helsinki and its later amendments. Due to the retrospective nature of the study and the use of fully de-identified data—where all personal identifiers were permanently stripped prior to analysis—the requirement for informed consent was waived by the respective IRBs.
Eligible cases were selected based on the following criteria: (1) histologically verified lung adenocarcinoma presenting as GGNs on preprocedural thin-section CT imaging; (2) nodule diameters ranging from 5 to 30 mm; and (3) absence of prior biopsy or anticancer therapies. Exclusion criteria included: (1) insufficient quality of radiological or pathological records; (2) an interval exceeding 6 months between CT acquisition and surgical resection; and (3) the presence of multifocal adenocarcinomas. After rigorous screening based on the above criteria, a total of 847 GGNs were ultimately enrolled in this multicenter study and randomly divided into a training set (70%, n = 592) and a validation set (30%, n = 255) for model development and performance evaluation. A graphical abstract for this study is presented in Figure 1.

A graphical abstract summarizing the key findings and methodologies of this study.
Four Deep Learning Models
The CT acquisition protocols used in this multicenter study are detailed in Supplemental Digital Content 1. The nodule segmentation and image preprocessing methods for modeling are provided in Supplemental Digital Content 2 and Supplemental Digital Content 3, respectively.
To ensure a fair and controlled comparison of intrinsic architectural advantages, we implemented a strictly standardized training and optimization protocol across all four DL models (DenseNet-121, ResNet-10, ResNet-18, and VGG-13). All experimental implementations were conducted on an NVIDIA A6000 GPU platform. Each model was trained for a fixed budget of 200 epochs using the AdamW optimizer with identical hyperparameters: an initial learning rate of 1 × 10−4, a batch size of 16, and a weight decay of 1 × 10−4.
To guarantee adequate convergence and avoid local minima for all architectures, we employed a consistent learning rate consisting of a warm-up phase (10% of total epochs) followed by cosine annealing to adaptively tune parameters. Furthermore, to rigorously address class imbalance within the multicenter dataset, a weighted cross-entropy loss function was applied uniformly across all models, assigning class-specific weights of [2.0, 3.0, 5.0] to the AIS, MIA, and IAC categories, respectively.
The image processing pipeline for all architectures followed a consistent structure: a backbone consisting of convolutional layers for local feature extraction and pooling layers for dimensionality reduction. This sequential process transforms low-level image features into high-level semantic feature maps. A subsequent flattening operation converts the 3D feature tensor into a 1D vector, preserving inter-channel spatial relationships. Finally, the flattened vector passes through fully connected layers for feature fusion, culminating in a Softmax activation function to generate the probability distribution for the ternary classification task. To further validate the robustness of our results, 95% confidence intervals (CIs) were calculated for key performance metrics using bootstrapping.19–21
Conventional Machine Learning-Based Clinicoradiographic Models
The clinicoradiographic features assessed in this study include age, sex, nodule type, nodule size, location, shape, margin, lobulation sign, vascular convergence sign, vacuole sign, spiculation sign, and pleural indentation sign. Details regarding these features can be found in Supplemental Digital Content 4.
For all analyses, we utilized scikit-learn, a Python library for conventional machine learning (cML). 22 The curated clinicoradiographic features served as input variables to predict the ternary histological subtypes (AIS, MIA, and IAC) as classification targets. Twelve cML classifiers were implemented: logistic regression, random forest, support vector machine (SVM), gradient boosting, k-nearest neighbors, Naive Bayes (NB), decision tree, extreme gradient boosting, light gradient boosting machine, adaptive boosting, extremely randomized trees, and categorical boosting. In this study, GridSearch with five-fold cross-validation was systematically employed to evaluate various parameter combinations and select the optimal hyperparameters based on the cross-validation results from the training set.23,24 The process of hyperparameter optimization across various cML models is detailed in Supplemental Digital Content 5.
Radiomics Model
The radiomics analysis was executed in three sequential phases: feature extraction, dimensionality optimization, and predictive modeling. In the initial phase, 1239 quantitative imaging features for each GGN were derived using PyRadiomics (v3.0.1), followed by standardization via z-scoring to mitigate batch effects. This high-dimensional radiomics profiling of preoperative thin-section CT scans established a candidate feature pool for each GGN.
To predict the pathological invasiveness of lung adenocarcinoma presenting as GGNs, we employed a multi-stage dimensionality reduction framework adapted from the methods proposed by Zuo et al and Pan et al17,18 To strictly prevent data leakage, all feature selection and model construction steps were performed exclusively on the training set. First, univariate analysis of variance (ANOVA) with a significance threshold of p < 0.05 was applied to exclude features lacking discriminatory power. Next, recursive feature elimination with cross-validation (RFECV) was conducted using 5-fold cross-validation to iteratively refine the feature set by removing the least informative variables until optimal predictive performance was achieved. 25 Finally, L1-regularized LASSO regression was used to further enforce feature sparsity by shrinking non-informative coefficients to zero, resulting in a compact yet highly discriminative feature signature.
Model Selection Protocol
The predictive performance of the DL, clinicoradiographic, and radiomics models was rigorously benchmarked. The primary assessment utilized multi-class receiver operating characteristic (ROC) analysis, with the macro-averaged area under the curve (Macro-AUC) serving as the principal metric. Complementary discriminative metrics, including diagnostic accuracy, precision, recall, and F1-score, were quantified to evaluate classification consistency across pathological invasiveness. The final model selection was based on the maximal Macro-AUC stability across validation sets for the ternary classification of invasiveness in GGNs. 26 The reporting of this study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines for cohort studies. 27
Results
Patient Characteristics
A total of 847 patients with GGNs who had a pathological diagnosis of lung adenocarcinoma were included in this study. Among these patients, 155 (18.3%) were diagnosed with AIS, 256 (30.2%) with MIA, and 436 (51.5%) with IAC. Of the total, 592 patients were included in the training set, among whom 104 (17.6%) were diagnosed with AIS, 190 (32.1%) with MIA, and 298 (50.3%) with IAC. In the validation set, 255 patients were included, with 51 (20%) diagnosed with AIS, 66 (25.9%) with MIA, and 138 (54.1%) with IAC.
The baseline characteristics of all patients are summarized in Table 1. A comparative analysis of baseline characteristics showed no statistically significant differences (p > 0.05 for all variables) between the training and validation sets, indicating that the groups are comparable.
Comparative Analysis of Clinicoradiographic Features Between Training and Validation Sets.

Performance of Four Deep Learning Models
Detailed performance metrics for four DL models, including Macro-AUC, accuracy, recall, precision, and F1-score, are presented in Figure 2. For the ternary classification of adenocarcinoma into AIS, MIA, and IAC, ResNet-10 achieved the highest Macro-AUC of 0.8055, followed by DenseNet-121 with 0.792, VGG-13 with 0.7805, and ResNet-18 with 0.7795. The architecture of the developed ResNet10 is illustrated in Figure 3.

Comprehensive performance metrics for the four deep learning models, including Macro-AUC, accuracy, recall, precision, and F1-score.

Architectural representation of the developed ResNet-10 model.
Performance of cML-Based Clinicoradiographic Models
Detailed performance metrics for the 12 cML models, including Macro-AUC, accuracy, recall, precision, and F1-score, are presented in Table 2 and Figure 4. For the ternary classification of AIS, MIA, and IAC, the SVM-based clinicoradiographic model achieved the highest performance with a Macro-AUC of 0.7770 (0.708-0.846) and an accuracy of 0.6000, demonstrating that rigorous hyperparameter optimization via GridSearch allowed the model to maximize the utility of the available clinicoradiographic features. Conversely, the NB model recorded the lowest Macro-AUC of 0.6768 (0.600-0.754).

Detailed performance metrics for various cML models, encompassing Macro-AUC, accuracy, recall, precision, and F1-score.
Comparative Performance Analysis of Machine Learning Models Leveraging Clinicoradiographic Features for Ternary Classification of Pulmonary Ground-Glass Nodule Invasiveness.

Performance of Radiomics Model
Quantitative radiomics profiling yielded 1239 features per GGN. Sequential dimensionality reduction was implemented through a three-tiered protocol: first, ANOVA with a significance threshold of α = 0.05 retained 1024 features with discriminative capacity. Next, RFECV iteratively pruned redundant features, culminating in 354 non-collinear radiomics descriptors (Figure 5). Finally, L1-penalized LASSO regression identified 25 features with non-zero coefficients, establishing the final radiomics signature. This radiomics model achieved a Macro-AUC of 0.7801 for the ternary classification of adenocarcinoma in AIS, MIA, and IAC.

RFECV utilized for radiomic feature reduction.
Optimal Model Selection for Ternary Classification Diagnostic Tasks
The comparative effectiveness analysis of DL, clinicoradiographic, and radiomics models is presented in Table 3 and Figure 6. Among these, the DL model ResNet-10 demonstrated superior performance with a Macro-AUC of 0.8055, surpassing the radiomics model, which achieved a Macro-AUC of 0.7801, followed closely by the SVM-based clinicoradiographic model with a Macro-AUC of 0.7770.
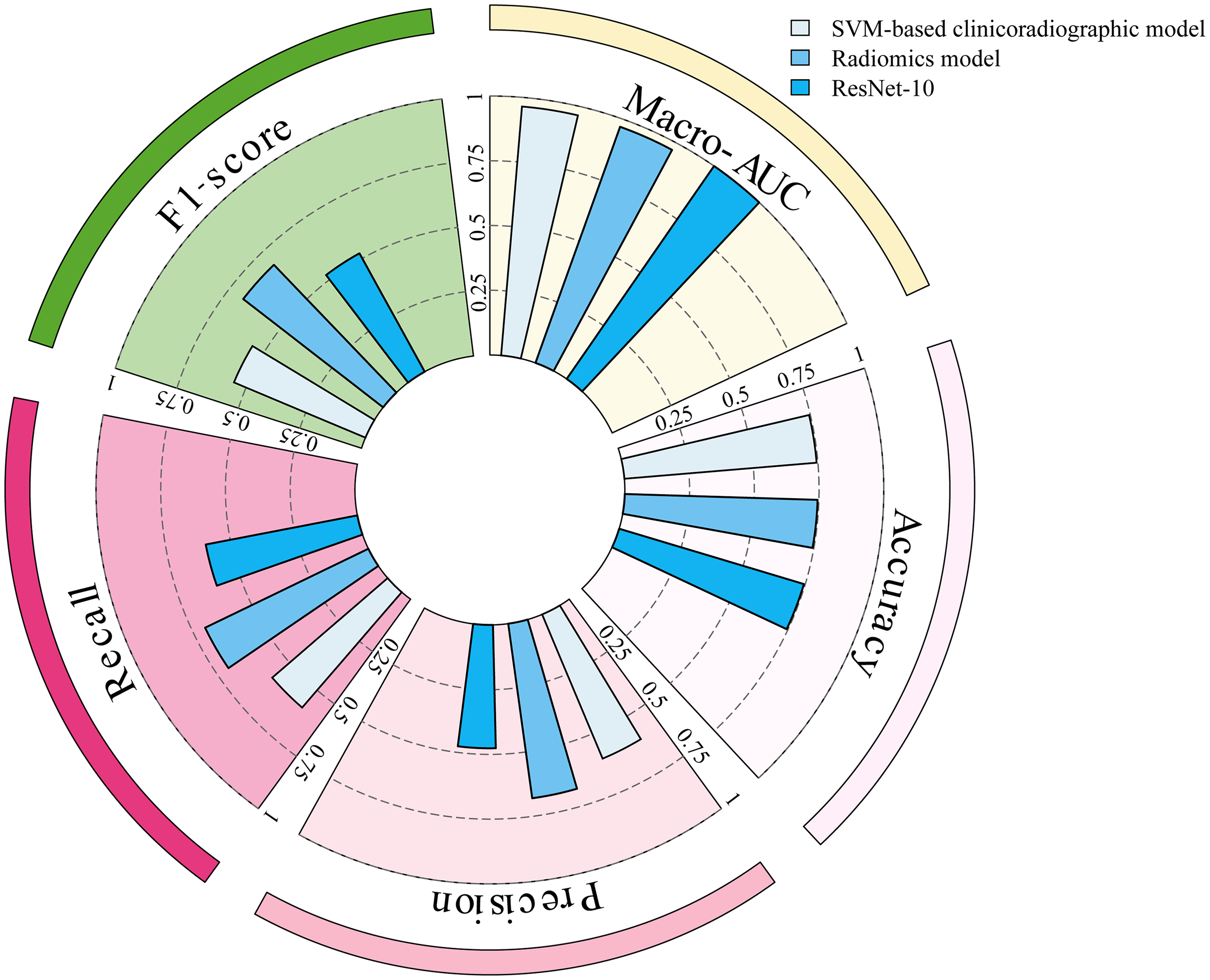
Comparative analysis of effectiveness among deep learning, clinicoradiographic, and radiomic models.
Comparative Performance of Different Models for Preoperative Prediction of Pathological Invasiveness in Pulmonary Ground-Glass Nodules.

Figure 7 depicts the multi-class ROC curves for the derived binary classification tasks. ResNet-10 exhibited superior discriminative capacity across all histological subtypes. For AIS identification, the model achieved a leading AUC of 0.8498, representing a relative improvement of 4.1% over the radiomics model (AUC = 0.8164) and 8.2% over the SVM-based clinicoradiographic model (AUC = 0.7854). Similarly, in the diagnosis of MIA, ResNet-10 demonstrated robust performance with an AUC of 0.7076, outperforming the radiomics (AUC = 0.6817) and SVM models (AUC = 0.7001) by relative margins of 3.8% and 1.1%, respectively. Notably, for IAC detection, ResNet-10 attained the highest AUC of 0.8500, surpassing both the radiomics (AUC = 0.8331; +2.0% relative improvement) and SVM models (AUC = 0.8457; +0.5% relative improvement). These findings underscore the consistent superiority of ResNet-10 in stratifying GGN invasiveness, with the most substantial performance gain observed in the identification of AIS.

Multi-class ROC curves for deep learning, clinicoradiographic, and radiomic models.
Discussion
In this multicenter study, we developed four mainstream DL architectures (DenseNet-121, ResNet-10, ResNet-18, and VGG-13), alongside a cML -based clinicoradiographic model and a radiomics model for the ternary classification of AIS, MIA, and IAC related to GGNs. Among these models, ResNet-10 demonstrated robust performance within our multicenter cohort, showing consistent advantages across binary classification tasks, particularly in identifying AIS.
ResNet-10 outperformed other DL architectures in the ternary classification of AIS, MIA, and IAC associated with GGNs. The reasons for its superior performance are as follows19–21: 1) ResNet-10 reduces overfitting through its simplified architecture. Its shallow design, which features fewer layers and parameters, mitigates overfitting, especially in small medical datasets, which typically consist of ≤1000 samples. While deeper networks (such as ResNet-18 and DenseNet-121) risk memorizing noise, ResNet-10 maintains residual connections that promote stable gradient flow, effectively balancing simplicity and expressiveness. This design enables robust generalization across multicenter data, as evidenced by its comparable or superior performance relative to deeper models. 2) ResNet-10 captures critical shallow features relevant to clinical cues. The classification of GGNs relies heavily on low-level features, such as nodule size and type, which are effectively captured by the initial layers of ResNet-10. In contrast, deeper models often engage in redundant hierarchical abstractions that are irrelevant to the task. This not only wastes computational resources but also increases the risk of noise negatively impacting the model's performance. The architecture of ResNet-10 restricts feature abstraction to clinically meaningful depths, ensuring a focus on task-critical signatures while avoiding a decrease in clinical relevance.
ResNet-10 outperformed the radiomics model and followed the SVM-based clinicoradiographic model. ResNet-10's end-to-end feature learning capability offered distinct advantages in this study over traditional radiomics and clinicoradiographic models. Radiomics relies on handcrafted features (such as texture and shape) that may fail to capture complex, nonlinear relationships present in medical images.28,29 In contrast, ResNet-10 automatically learns hierarchical features from raw pixel data, including microstructural patterns that are invisible to the human eye. This capability allows it to detect early signs of invasiveness, such as subtle architectural distortions or contrast enhancements that radiomics pipelines might overlook.
Clinicoradiographic models, which integrate clinical variables (such as age and nodule size) with radiological findings, face similar limitations. These models require manual feature selection, which can introduce bias and diminish generalizability. In contrast, ResNet-10 bypasses these issues by learning data-driven representations directly from images, eliminating the need for priori feature engineering. Its performance in multicenter settings further underscores its robustness to inter-institutional variability in imaging protocols and patient demographics. 7
Regarding the cML-based clinicoradiographic models, despite applying rigorous GridSearchCV to identify optimal hyperparameters, the classification accuracy plateaued at approximately 60%. This suggests a performance ceiling inherent to handcrafted semantic features (such as spiculation or lobulation). 28 While interpretable, these features lack the high-dimensional resolution required to fully differentiate the subtle, continuous morphological transitions between AIS, MIA, and IAC compared to the end-to-end feature learning of ResNet-10.
It is worth noting that while the Radiomics model yielded a higher F1-score compared to ResNet-10 (0.5505 vs 0.4206), we prioritized ResNet-10 as the optimal architecture based on its superior Macro-AUC (0.8055) and overall accuracy (0.6300). The F1-score is a threshold-dependent metric that can be disproportionately influenced by class imbalance and the specific operating point derived from the validation set. In contrast, the Macro-AUC provides a robust, threshold-independent assessment of the model's global discriminative power. More importantly, from a clinical perspective, ResNet-10 demonstrated higher efficacy in the critical binary tasks—specifically in distinguishing AIS (AUC = 0.8498) and IAC (AUC = 0.8500)—which are the primary determinants for stratifying patients into conservative management versus surgical intervention.
When reconfigured for the binary classification of IAC versus non-IAC (comprising AIS and MIA), ResNet-10's performance gains align remarkably well with clinical management priorities. While MIA and IAC typically necessitate surgical intervention, AIS is increasingly considered for conservative management.4–6 By focusing on this most clinically actionable distinction, ResNet-10 simplifies diagnostic workflows and facilitates the rapid triage of high-risk patients. This capability is particularly valuable in large-scale screening programs, where minimizing false negatives for IAC is critical for optimizing patient outcomes.
There are several limitations to our work. First, our study exclusively included pathologically proven lung adenocarcinomas from post-surgical specimens, thereby excluding indeterminate GGNs managed with surveillance. This exclusion may overestimate the model's performance in screening scenarios where benign or pre-invasive nodules are more prevalent. To enhance our findings, further prospective validation in screening cohorts is necessary to assess real-world generalizability. Second, although this study utilized a multicenter cohort, the validation strategy was based on a stratified hold-out split of pooled data rather than a geographically independent external validation (eg, training on two centers and testing on a third). While we utilized stratified sampling to ensure balanced representation of centers and classes, the lack of a fully independent external test set means that the generalizability of our models to unseen institutions requires further verification in future prospective studies. Third, the absence of longitudinal postoperative surveillance data limits our ability to assess the model's prognostic validity regarding clinical outcomes, especially those related to different treatment programs. Collecting such data in future studies will be crucial for enhancing the robustness of our findings. Finally, the exclusion of multifocal adenocarcinomas creates a critical gap in clinical applicability. Future iterations of our research will integrate whole-lung graph neural network correlation studies to improve multifocal classification.
In conclusion, this multicenter study demonstrates that ResNet-10 shows promise as an effective DL architecture for the ternary classification of GGNs into AIS, MIA, and IAC. ResNet-10 competitively outperforms radiomics and cML-based clinicoradiographic models in terms of global discriminative ability (Macro-AUC), despite trade-offs in other metrics. Additionally, in subsequent binary classification tasks, ResNet-10 demonstrates superior discriminative ability for IAC detection. While further prospective validation is required, the diagnostic consistency observed across multicenter data provides a preliminary foundation for clinical decision support, particularly for optimizing GGN stratification and precision therapy in early lung cancer.
Supplemental Material
sj-docx-1-tct-10.1177_15330338261423265 - Supplemental material for Preoperative Ternary Classification of Pulmonary Ground-Glass Nodules (AIS/MIA/IAC): ResNet-10 Outperforms Radiomics and Clinicoradiographic Models in Multicenter Study
Supplemental material, sj-docx-1-tct-10.1177_15330338261423265 for Preoperative Ternary Classification of Pulmonary Ground-Glass Nodules (AIS/MIA/IAC): ResNet-10 Outperforms Radiomics and Clinicoradiographic Models in Multicenter Study by Dan Long, Zhichao Zuo, Huchuan Zhou, Wanyin Qi, Sanhong Zhang, Jinqiu Deng and Ziqiang Yang in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-2-tct-10.1177_15330338261423265 - Supplemental material for Preoperative Ternary Classification of Pulmonary Ground-Glass Nodules (AIS/MIA/IAC): ResNet-10 Outperforms Radiomics and Clinicoradiographic Models in Multicenter Study
Supplemental material, sj-docx-2-tct-10.1177_15330338261423265 for Preoperative Ternary Classification of Pulmonary Ground-Glass Nodules (AIS/MIA/IAC): ResNet-10 Outperforms Radiomics and Clinicoradiographic Models in Multicenter Study by Dan Long, Zhichao Zuo, Huchuan Zhou, Wanyin Qi, Sanhong Zhang, Jinqiu Deng and Ziqiang Yang in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-3-tct-10.1177_15330338261423265 - Supplemental material for Preoperative Ternary Classification of Pulmonary Ground-Glass Nodules (AIS/MIA/IAC): ResNet-10 Outperforms Radiomics and Clinicoradiographic Models in Multicenter Study
Supplemental material, sj-docx-3-tct-10.1177_15330338261423265 for Preoperative Ternary Classification of Pulmonary Ground-Glass Nodules (AIS/MIA/IAC): ResNet-10 Outperforms Radiomics and Clinicoradiographic Models in Multicenter Study by Dan Long, Zhichao Zuo, Huchuan Zhou, Wanyin Qi, Sanhong Zhang, Jinqiu Deng and Ziqiang Yang in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-4-tct-10.1177_15330338261423265 - Supplemental material for Preoperative Ternary Classification of Pulmonary Ground-Glass Nodules (AIS/MIA/IAC): ResNet-10 Outperforms Radiomics and Clinicoradiographic Models in Multicenter Study
Supplemental material, sj-docx-4-tct-10.1177_15330338261423265 for Preoperative Ternary Classification of Pulmonary Ground-Glass Nodules (AIS/MIA/IAC): ResNet-10 Outperforms Radiomics and Clinicoradiographic Models in Multicenter Study by Dan Long, Zhichao Zuo, Huchuan Zhou, Wanyin Qi, Sanhong Zhang, Jinqiu Deng and Ziqiang Yang in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-5-tct-10.1177_15330338261423265 - Supplemental material for Preoperative Ternary Classification of Pulmonary Ground-Glass Nodules (AIS/MIA/IAC): ResNet-10 Outperforms Radiomics and Clinicoradiographic Models in Multicenter Study
Supplemental material, sj-docx-5-tct-10.1177_15330338261423265 for Preoperative Ternary Classification of Pulmonary Ground-Glass Nodules (AIS/MIA/IAC): ResNet-10 Outperforms Radiomics and Clinicoradiographic Models in Multicenter Study by Dan Long, Zhichao Zuo, Huchuan Zhou, Wanyin Qi, Sanhong Zhang, Jinqiu Deng and Ziqiang Yang in Technology in Cancer Research & Treatment
Footnotes
Abbreviation
Ethics Approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committees of the participating centers.
Informed Consent Statement
Due to the retrospective nature of the study and the use of fully de-identified data—where all personal identifiers were permanently stripped prior to analysis—the requirement for informed consent was waived by the respective IRBs.
Author Contributions
Sanhong Zhang and Huchuan Zhou contributed to the study conception and design. Data collection was performed by Sanhong Zhang, Jinqiu Deng, and Ziqiang Yang. The first draft of the manuscript was written by Dan Long, Zhichao Zuo, and Wanyin Qi. Zhichao Zuo provided supervision and contributed to the revision of the manuscript. All authors read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (grant no. 82403308), Hunan provincial 14th Five-Year Plan AppliedCharacteristic Discipline – Clinical Medicine ( No. Xiang Jiao Tong[2022] 351) and the Aid Program for Science and Technology InnovativeResearch Team in Higher Educational Institutions of Hunan Province, China.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All data included in this study are available upon request by contact with the corresponding author.
Use of Artificial Intelligence
Generative artificial intelligence (AI) tools were used solely for language editing and grammatical refinement of this manuscript to improve readability. No AI tools were used to generate or modify scientific data or content.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
