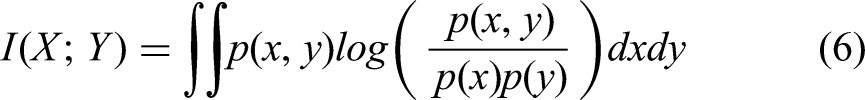Abstract
Background
Deep generative models can improve the generalization of deep learning in medical imaging by enriching limited training data with diverse, realistic synthetic images.
Purpose
To assess whether Denoising Diffusion Probabilistic Models (DDPM) generated synthetic MRI, with and without mutual information (MI) regularization, enhances brain tumor classification across heterogeneous datasets.
Study Type
Retrospective.
Population
A total of 559 patients with low and high grade brain tumors (LGG, HGG) were included from two datasets: public dataset (BraTS, n = 335) and clinical dataset (TASMC, n = 224), used exclusively to evaluate model generalization.
Field Strength/Sequence
1.5 T/3.0T-MR / T1WI, T1WI + C, T2WI, and FLAIR images.
Assessment
DDPM models were trained to generate synthetic MR images of low grade glioma (LGG) and high grade glioma (HGG), with a variant incorporating MI. Image quality was assessed using Pearson-correlation, Frechet-Inception-Distance (FID) and Inception-Score (IS). For classification purposes. For classification, a 2D ResNet-152 was trained under four setups: (1) real images (baseline), (2) +augmentation, (3) +DDPM, and (4) +DDPM + MI. Performance was assessed by accuracy and F1-score. Robustness was tested through cross-dataset evaluation using a 5-fold ensemble.
Results
The DDPM models, with and without MI, generated high-quality synthetic images, achieving FID = 31.47, 45.00, and IS = 1.50, 1.25, respectively. Lower FID and higher IS indicate enhanced realism and diversity, suggesting that MI improved both the quality and variability of the generated images. Cross-dataset evaluation demonstrated that DDPMs with MI achieved superior generalization performance in brain tumor classification task, with accuracies of 0.89 and 0.85 for BraTS-to-TAMSC and TAMSC-to-BraTS evaluations, respectively. These results outperform the baseline model (0.87, 0.80), traditional data augmentation (0.85, 0.78), and the standard DDPM without MI (0.82, 0.83).
Data Conclusion
DDPM + MI with ensemble learning significantly improves brain tumor generalization across diverse datasets, consistently outperforming baseline, traditional augmentation, and standard DDPM. This combination offers a robust solution for cross-institutional clinical applications.
Introduction
Deep learning (DL) algorithms have demonstrated significant efficacy in brain lesion segmentation and classification tasks, particularly when leveraging multi parametric MRI data (T1WI, T1W + C, T2WI, and FLAIR). 1
Generalization is a critical aspect of DL models, especially in medical image based models, where the ability to accurately predict unseen data is essential for clinical applications. Medical imaging datasets frequently encompass only a narrow spectrum of pathological diagnosis. Relatively homogeneous image acquisition protocols, and patient demographics, potentially lead to model overfitting. 2 The efficacy of DL models in classifying unseen data is crucial, yet frequently compromised by the constrained nature of training datasets. 3 This limitation often leads to overfitting, where models perform exceptionally well on familiar data but falter when presented with unseen inputs. The discrepancy between performance on training data and generalization to new, unseen data, poses a significant challenge in deploying these models in real-world scenarios. This is particularly evident in critical domains such as medical imaging. 4
To address these challenges, solutions such as data augmentation, and synthetic data generation, can be employed.5,6 Synthetic data generated images can serve as an effective form of data augmentation by enriching the training dataset with diverse and realistic variations. However, these strategies must be implemented carefully to ensure that they enhance model generalizability without introducing artifacts that could further skew classification results.
Several models have been proposed for synthetic MRI data generation. GAN models have been effectively employed to generate synthetic T1WI and FLAIR MRI, which are then utilized for brain lesion segmentation on the BraTS dataset.7,8 Conte et al accomplished this task using an image-to-image translation GAN model. 9 Moshe et al successfully used a GAN model to generate missing T1WI, T2WI, FLAIR, or both, for classification tasks. 10 Guan and Loew demonstrates that incorporating GAN-generated images into the training dataset can mitigate classifier overfitting when distinguishing between normal and abnormal tissue. However, their results indicate that traditional data augmentation techniques yielded marginally superior performance compared to the GAN-based approach. 11
It has been shown that images, including medical images12–14 generated by Denoising Diffusion Probabilistic Models (DDPMs) are of higher quality to those generated by GANs.15,16 DDPMs have emerged as an alternative to GANs in generative modeling, offering advantages in training stability, sample quality, and flexibility. They have been shown to generate higher-quality medical images, resulting in improved diversity and fewer artifacts within the outputs.17,18 In medical imaging tasks, Yuan et al and Salehinejad et al used DDPMs to manage missing data and address class imbalances in both Alzheimer's and chest x-ray classification tasks.19,20 Yi et al investigated the generalization capabilities of DDPMs, defining generalization as the correlation between the training set and the quality of generated images. They proposed that a lower correlation and lower mutual information value, indicates better generalization, helping distinguish models that truly learn data distributions from those that memorize. 21
The aim of this study was to evaluate the impact of DDPM-generated synthetic MRI data, with and without mutual information regularization. This impact enhancing brain tumor classification and generalization across heterogeneous datasets.
Material and Methods
The network was trained and tested on Linux system, using a single Graphical Processing Unit (GPU), CUDA 9.1 device, Nvidia RTX2080 Ti. Analysis was performed with Python software (Python 3.7.7) along with Fastai software (1.0.61).
Datasets
The Brain Tumor Segmentation Challenge (BraTS, 201922–24) dataset, available for download at https://www.med.upenn.edu/cbica/brats2019/data.html, used for model training and evaluation. BraTS dataset included 335 patients, scanned between 2012 and 2018 (estimation, based on24,25), 76 cases with low grade glioma (LGG), and 259 cases with high grade glioma (HGG), respectively. Each scan contains T1WI, T1WI + C, T2WI and FLAIR images and mask of the enhancing tumor.
BraTS dataset was acquired with different clinical protocols and various scanners from multiple (n = 19) institutions. For each subject, the central tumor slice (based on the provided segmentation of the enhancing lesion) was selected.
TAMSC dataset: A local dataset from the Tel Aviv Sourasky Medical Center (TAMSC), used exclusively for evaluation assessed, the model's generalization capabilities. The TAMSC dataset included 224 patients scanned between 2008–2021. Of them, 37 cases with LGG, and 187 cases with HGG were studied. The MRI scans were part of routine clinical assessments. 3D U-net,26,27 a validated model specifically trained for brain tumor segmentation, was employed to automatically segment the enhancing lesion area. All resulting segmentations were reviewed and manually refined, as needed.
The study was approved by the TAMSC institutional review board (IRB) which waived informed consent (IRB approval number 0200-10).
Denoising Diffusion Probability Model (DDPM)
DDPM works by incrementally corrupting the data
When the noise schedule
Where
In the reverse process, the backward transition
Where the mean
Where
During training, Smooth L1 loss (Huber loss) was used to train the denoising network
Denoising Diffusion Probability Model with Mutual Information (DDPM + MI)
In order to improve diversity and reduce overfitting in synthetic image generation, we incorporated Mutual Information (MI) into the DDPM loss function. While DDPMs were generating high-quality images, they produced outputs highly correlated with the training data.28–30 Additionally, MI as a regularization term encouraged the model to generate more diverse samples by reducing dependency on the input. MI defined as equation (6):
The total loss combined the standard denoising loss and the MI term as equation (7):
This equation has been measured as informing the generated image and sharing with the input. This approach helps the model produce realistic images while avoiding memorization, leading to better generalization in clinical classification tasks.
DDPM Model Training
The algorithm that is used with the adamW optimization, along with smooth L1 loss, and in combination with 100 epochs and 0.0002 learning rate was used in this research. The DDPM model was implemented with and without MI regularization, and separated for the different tumor types. This generated HGG and LGG based upon original images (Figure 1B). Training used a linear noise schedule over 1000 diffusion steps in this process. The batch size was determined by extracting the number of images or samples in the current batch from the shape of the pixel data. The model based on the lowest validation loss was chosen over all the other models.

Input data. A. Extraction of lesion patches, B. Experimental design for data classification involves four classification model setups: (1) real images, (2) real images with augmentation, (3) real images with images generated by Denoising Diffusion Probabilistic Models (DDPM), and (4) real images with images generated by DDPM with Mutual Information (MI) optimization.
The DDPM with MI are incorporated into the loss function as regularization. This is trained under the same conditions (The standard DDPM). Along with the MI term, weighted at 0.1 and the L1 smooth loss weighted at 0.9.
DDPM Model Evaluation
1. Fréchet Inception Distance (FID) and Inception Score (IS)
Evaluation of the DDPM model results were performed by comparing the generated images to the input images based on FID and IS.
31
The FID metric mirrors the resemblance between the distribution of real images and generated images in a feature space. A lower FID score indicated more similarity between real and generated image distribution. The highest IS metric pinpointed quality and diversity of the generated images.
32
In order to ensure consistency with standard 3-channel datasets and methodologies, we needed to reduce the 4-channel data to 3 channels using an encoder to preserve the structural information.
2. Pearson Correlation - Image Similarity to Training Data
To assess the similarity between synthetic images and the training data, we used the Pearson correlation coefficient, which computed the highest correlation with any training image. We focused on the average of the highest correlations between synthetic and training images across different datasets. This approach highlighted how similar the synthetic images were to the training set - and whether they reflected excessive similarity (ie, memorization) or appropriate diversity. The high correlations between values may indicate memorization or data leakage, while lower connections suggest greater diversity (a desirable trait in generative tasks).
Application for Tumor Classification
The segmented enhancing tumor was used as the target area for classification. Each image was first cropped around the center of the enhanced lesion area, and then resized to a 96 × 96 image size (Figure 1A). This size was chosen as a practical compromise—large enough to capture the tumor and surrounding tissue, yet efficient for DDPM training. This design is in line with previous studies using DDPMs on localized regions. 33
The BraTS data was split, at the subject level, into datafiles of 80% training and 20% testing. The training dataset was split into 80% training and 20% validation. For cross-cataloging evaluation, models instructed on the BraTS training dataset were tested on the TAMSC dataset (BraTS-to- TAMSC). The model was adapted to the full TAMSC dataset where it was tested on the BraTS dataset (TAMSC-to-BraTS).
A 2D Resnet152 convolutional neural network was employed for model training. Following this optimization, the training was conducted with batch size = 32, utilizing Focal Loss as a loss function with α = 0.25 and γ = 2. The initial learning rate was set to 0.01, and the training spanned 200 epochs. Additionally, the network structures were modified to accommodate 4 channels of input data (accounting for the different MRI contrasts).
The classification was performed under four setups (Figure 1B): (1)
Integrating Ensemble Modeling for Enhanced Robustness
To enhance prediction accuracy and robustness, we employed an ensemble model using 5-fold cross-validation within each dataset. This approach combined the outputs of five models by averaging their weights. Overall, ensemble models provided a more robust and accurate classification by mitigating weaknesses and amplifying strengths. 34
Evaluation of Classification Results
Classification performance across the four experimental setups was evaluated using 5-fold cross-validation, within each dataset and cross-dataset evaluation between the BraTS and TAMSC datasets, measuring accuracy and F1 scoring as performance metrics.
Results
The mean tumor volume was 2.54 ± 1.41 mL (range: 0.3-10.5 mL) for HGG and 2.72 ± 3.01 mL (range: 0.34-14.47 mL) for LGG. Statistical analysis using the Mann–Whitney U test showed no significant difference in tumor volumes between HGG and LGG (P = .14), supporting the suitability of the selected patch size across tumor types.
DDPM Model Evaluation
The FID and IS values presented in Table 1 indicates that the generated images achieved a notable level of quality and diversity. The results aligned with other studies, 31 confirming the high image quality and diversity of the generated images. The Pearson correlation between the generated and real images, reached a mean value of 0.55 ± 0.11 for both data sets. When mutual information was incorporated into the DDPM model, the correlation decreased to a mean value of 0.47 ± 0.11, suggesting reduced memorization and increased diversity. Nonetheless, image quality remained comparable, as reflected by similar IS values and only a moderate impact on FID scores.
Evaluation of Generated Images Model on BraTS and TAMSC Datasets.

HGG, high grade glioma; LGG, low grade glioma; FID, Fréchet Inception Distance; IS, Inception Score.
Tumor Classification
BraTS Dataset (Baseline)
This model reached a mean 5-fold accuracy level of 0.90 ± 0.03, and F1 score of 0.91 ± 0.03 for the validation dataset. The mean accuracy of 0.86 ± 0.03 and F1 score of 0.86 ± 0.03 for the test dataset shows the output of the base model. Combining Ensemble model, by averaging the output of 5-fold weight, has improved the overall test dataset accuracy and F1 score to 0.97. Based on those results, an Ensemble model was employed for all subsequent inference results.
Model Generalization - Cross-Dataset Evaluation
Figure 2 shows classification performance heatmaps comparing four different scenarios (Baseline, +Augmentation, +DDPM, +DDPM + MI) across two transfer directions: (BraTS→TAMSC and TAMSC→BraTS). The integration of mutual information into the DDPM model, combined with ensemble learning, resulted in improvements in generalization across datasets. Combining DDPM with MI has significantly outperformed all other methods, by achieving the highest accuracy and F1-score of 0.89 and 0.90 for BraTS-to-TAMSC, along with 0.85 and 0.86 for TAMSC-to-BraTS. Such results highlight the effectiveness of combining MI with ensemble learning, in order to enhance model robustness, reduce overfitting, and improve transferability to unseen datasets in cross-dataset evaluations.
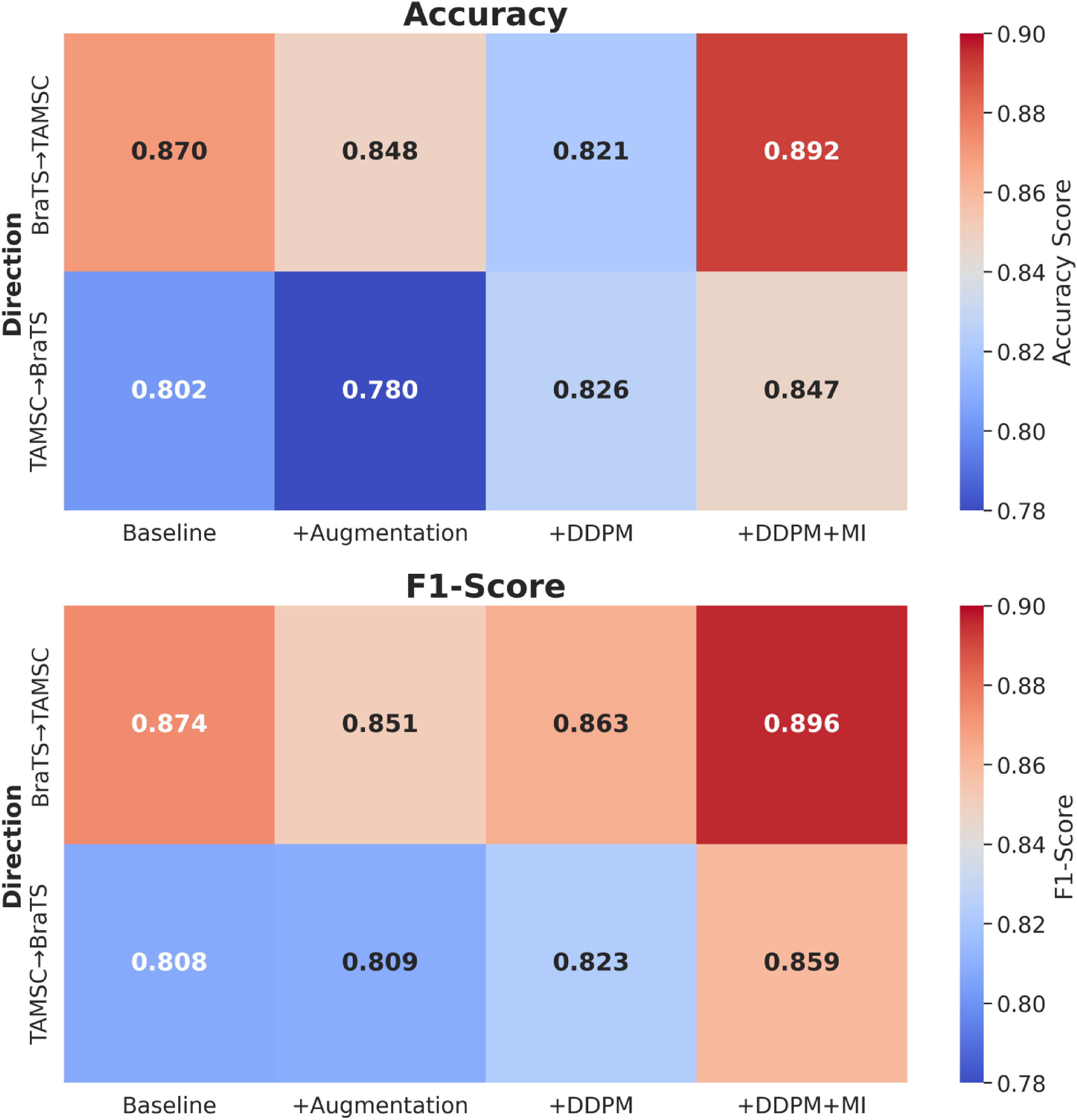
Classification performance heatmaps comparing four different scenarios.
Discussion
Our findings demonstrate that DDPM with mutual information regularization and ensemble learning, substantially improved cross-dataset generalization in brain tumor classification. By consistently outperforming baseline models, traditional augmentation, and standard DDPM created approaches across heterogeneous clinical datasets.
Model generalization presents a critical challenge in deploying DL models for clinical applications. Models trained on one dataset often struggle to perform well on unseen data from different institutions.20,21 In clinical data, this is particularly important, due to the inherent variability in patient populations, imaging devices, and acquisition protocols across varied hospitals or regions. Ensuring robust generalization is key for achieving reliable, real-world performance in clinical settings. 35
It was suggested that integrating synthetic images into model training can significantly enhance DL model generalization. By increasing the diversity of the training set, we seamlessly adapt models to better integrate with unseen conditions and external datasets.
In this study we used DDPMs for synthetic MRI image generation. Studies have demonstrated that DDPMs improve classification tasks by generating realistic synthetic MRI data and addressing class imbalances. However, DDPMs alone tend to produce samples with high correlation to the training data, which can limit their effectiveness in promoting generalization. To overcome this, incorporation of mutual information (MI) into the loss function,21,28,36 aims to reduce connection with the training set and enhance data diversity. 37 This offers a promising solutions to the overfitting problem, commonly encountered in small, imbalanced datasets.13,17,18 In the current study, we evaluated the contribution of synthetic data generation with MI-augmented DDPMs for improving model generalization in brain tumor classification tasks. We achieved high image quality, as demonstrated by FID and IS scoring. 32
2D classification was selected due to computational constraints and limitations of current DDPM architectures, which are optimized for 2D slice generation given memory and stability challenges. This approach enabled efficient training with MI regularization, ensemble learning, and cross-dataset evaluation. The 2D strategy also allowed for the creation of larger training sets by extracting tumor-centered slices, thereby enhancing DDPM stability. Central slices were chosen to support lesion segmentation and to capture the most diagnostically relevant features.
To further evaluate the impact of synthetic data on model robustness, we conducted cross-dataset generalization tests (training the classification model on the BraTS dataset and testing it on an external cohort (TAMSC dataset), and vice versa). These evaluations reflect real-world clinical scenarios, where models are expected to generalize across institutions with differing imaging protocols and patient populations. Four inference settings were assessed: (1) baseline, (2) baseline with traditional data augmentation, (3) baseline with standard DDPM-generated images, and (4) baseline with DDPM-generated images incorporating MI.
While traditional augmentation is well-established for improving classification performance, the use of DDPM-based synthetic data revealed more nuanced outcomes.38,39 Notably, the DDPM model with MI, consistently achieved superior performance across datasets. These findings underscored the limitations of traditional augmentation techniques and standard DDPMs, in capturing sufficient variability and highlighting the critically low correlation with in the training data.
To further enhance stability and performance, we incorporated ensemble learning during training. Specifically, we employed a 5-fold cross-validation strategy, where the model was trained independently on five different subsets of training data. The final prediction was obtained by averaging the outputs of these five models. This ensemble approach helped reduce prediction variance and mitigated the risk of overfitting to specific data distributions. Ultimately this leads to more robust and reliable performance across diverse test sets.
These findings indicate that synthetic data plays a crucial role in improving model generalization across diverse test datasets. It creates entirely new, diverse samples, simulating scenarios absent in the original dataset. Added to this findings, our models are able to generalize more effectively adjusting for unseen conditions or rare edge cases.40,41
Limitations
This study is limited to conventional MRI (T1WI, T1WI + C, T2WI, and FLAIR sequences), which includes other MRI modalities or advanced imaging techniques, that might reveal further benefits or limitations of such proposed. In addition, the use of DDPMs and large models like ResNet152, demands substantial computational resources, potentially limiting their accessibility in resource-constrained settings. It is also acknowledged that 3D classification could offer richer volumetric context and is proposed as a direction for future work, particularly to enhance anatomical coherence and capture spatial dependencies across slices.
Conclusion
This study has demonstrated that the combination of DDPM-generated synthetic data, with mutual information as regularization and ensemble learning, substantially improves brain tumor classification performance across diverse datasets. This approach consistently surpassed conventional methods including baseline models, traditional augmentation, and standard DDPM implementations, offering an effective strategy for addressing cross-institutional variability in clinical imaging.
Footnotes
Acknowledgments
Ethics Approval
Approved by the Tel Aviv Sourasky Medical Center IRB (No. 0200-10). The study was conducted in accordance with the Declaration of Helsinki.
Informed Consent
Patient consent was waived by the IRB due to the retrospective and anonymized nature of the data.
Author Contributions
Yael H. Moshe – Conceptualization, Methodology, Analysis, Writing.
Mina Teicher – Supervision, Theory, Review & Editing.
Moran Artzi – Conceptualization, Methodology, Data Curation, Clinical Insight, Supervision,
Review & Editing.
All authors approved the final manuscript.
Funding
Israel science foundation 205501.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability
Use of Artificial Intelligence
No generative AI was used for data or content creation. ChatGPT was used only for minor language editing; Language editing was performed by a professional English editor.