Abstract
Introduction
Breast cancer metastasis remains the leading cause of disease-related mortality. While calmodulin-regulated spectrin-associated protein 3 (CAMSAP3) has been implicated as a metastasis suppressor in other cancers, its role in breast cancer remains poorly defined. This study aimed to investigate the prognostic significance and functional impact of CAMSAP3 in breast cancer.
Methods
CAMSAP3 protein expression was evaluated by immunohistochemistry in 137 primary breast cancer specimens and correlated with clinicopathological features and survival outcomes. Kaplan–Meier and Cox regression analyses were performed to assess distant relapse-free survival (DRFS). Functional assays, including wound healing and Transwell invasion assays, were conducted in MDA-MB-231 and HCC1937 cells following CAMSAP3 overexpression or knockdown. External mRNA-level validation was performed using the Kaplan–Meier Plotter database.
Results
High CAMSAP3 expression was significantly associated with favorable features, including lower histological grade, hormone receptor positivity, reduced proliferation index, and fewer triple-negative phenotypes. Patients with high CAMSAP3 expression exhibited significantly improved DRFS, which was independently validated using a public mRNA dataset showing improved distant metastasis-free survival (DMFS). In vitro, CAMSAP3 overexpression inhibited migration and invasion of MDA-MB-231 cells, while CAMSAP3 knockdown enhanced these phenotypes in HCC1937 cells.
Conclusion
CAMSAP3 is a potential metastasis suppressor and independent prognostic marker in breast cancer. Its high expression is associated with less aggressive tumor characteristics and improved survival, possibly through inhibition of cancer cell migration and invasion.
Introduction
Breast cancer is the most prevalent malignancy and the leading cause of cancer-related mortality among women globally. In 2020 alone, it resulted in approximately 2.3 million new cases and 685 000 deaths. 1 Notably, over 90% of breast cancer-related fatalities are attributed to distant metastases, which remain largely incurable despite advancements in diagnosis and treatment. Even among patients diagnosed at an early stage, 20%–30% ultimately develop metastatic disease, underscoring the persistent challenge of recurrence and systemic spread.2,3 These realities highlight the urgent need to identify key regulators of metastasis—particularly metastasis suppressor genes—and to elucidate their mechanisms of action to inform novel therapeutic strategies.4,5
Factors that drive cancer metastasis include the properties of cancer cells and the tumor microenvironment. Among these factors, the cytoskeleton—particularly microtubules—plays a critical role in cancer cell motility. Microtubules are essential components of the cytoskeleton that regulate cell migration, division, and intracellular transport. 6 The precise regulation of microtubule dynamics, including their stability and organization, is crucial for maintaining cell polarity and facilitating directional cell movement, processes fundamental to both normal development and pathological conditions like cancer metastasis. 7 Disruptions in microtubule stability and function, often influenced by altered expression or activity of microtubule-binding proteins, are frequently observed in metastatic cancers, contributing to enhanced invasiveness and dissemination. 8 In breast cancer, microtubule-stabilizing agents such as paclitaxel have been shown to effectively inhibit the progression of metastatic disease, highlighting the potential of microtubule-related pathways as therapeutic targets. 9 Their dynamics are tightly controlled by microtubule-binding proteins, including calmodulin-regulated spectrin-associated protein 3 (CAMSAP3). CAMSAP3 binds to the minus ends of microtubules, stabilizing and orienting them.10,11 In lung cancer, decreased expression of CAMSAP3 has been associated with poor prognosis and increased invasion.12–14 However, its expression profile, functional role, and prognostic significance in breast cancer remain largely unknown.
A previous study reported a potential link between CAMSAP3 and breast cancer metastasis. 15 Given the established critical role of microtubule dynamics and their regulatory proteins in cancer cell motility and the limited understanding of CAMSAP3's specific involvement in breast cancer metastasis, we hypothesized that CAMSAP3 may affect breast cancer metastasis through microtubule-related mechanisms. Therefore, this study aims to evaluate the clinical significance of CAMSAP3 expression in breast cancer tissues, investigate its association with metastasis and patient outcomes, and assess its biological functions in cell migration and invasion in vitro.
Materials and Methods
Population
This study was approved by the Ethics Committee of Affiliated Hospital of Jiaxing University (First Hospital of Jiaxing). The study adhered to the Declaration of Helsinki, and written informed consent was obtained from all patients for their participation in the study and the publication of their anonymized data. The reporting of this study conforms to Reporting recommendations for tumor marker prognostic studies (REMARK) guidelines. 16 This retrospective study involved selecting cases based on specific inclusion and exclusion criteria without stratification or matching. Clinicopathological data were gathered from all consecutive patients meeting these criteria, treated from January 1, 2012, to December 31, 2013, with follow-up extending to December 31, 2023. The median follow-up time was 9.7 years. Clinical and pathological data were obtained from the hospital's electronic medical records system and the institutional pathology archives to ensure completeness and consistency. All patient details were de-identified prior to data analysis to protect patient privacy and ensure anonymity. The inclusion criteria were as follows: 1) primary breast cancer; 2) female sex; 3) age ≥ 18 years; 4) pathological type of invasive breast cancer, nonspecial type; 5) TNM anatomical stage I ∼ III according to the American Joint Committee on Cancer (AJCC) eighth edition; 6) acceptance of radical surgery (including but not limited to modified radical surgery and breast-conserving radical surgery) and reasonable postoperative adjuvant treatment; and 7) complete clinical and pathological data. The exclusion criteria for patients were as follows: 1) ipsilateral supraclavicular lymph node metastasis; 2) preoperative neoadjuvant chemotherapy or neoadjuvant endocrine therapy; 3) dual primary breast cancer; 4) a combined history of other malignant tumors; 5) pregnant or lactating; 6) mental illness or incapacity; and 7) lack of sufficient tumor tissue in the tissue specimen bank for research use.
The patients’ clinicopathological data included age, menstrual status, diameter of the primary breast cancer tumor, axillary lymph node status, histological grade, and estrogen receptor (ER), progesterone receptor (PR), human epidermal growth factor receptor 2 (HER2), Ki67, cytokeratin 5/6 (CK5/6), and epidermal growth factor receptor (EGFR) status. ER and PR were detected using immunohistochemistry (IHC), and a positivity threshold of ≥1% of cells was considered positive. 17 HER2 was tested using IHC and fluorescence in situ hybridization (FISH). The negative criteria were as follows: negative or 1 + by IHC or patients with 2 + by IHC who underwent further testing for the HER2 gene by FISH, and the result was negative (HER2/CSP17 < 2.0 and average HER2 copy number per cell < 4.0). 18 Ki67 was detected using IHC, and the proportion of Ki67-positive cells among all tumor cells was visually estimated. A 30% cutoff was considered the optimal threshold for Ki67 at our center. 19 CK5/6 and EGFR were detected using IHC. The expression levels of CK5/6 and EGFR were considered positive if nuclear staining was 1% or more. 20 Triple-negative breast cancer (TNBC) was defined as being negative for ER, PR, and HER2. Postoperative adjuvant treatments, including chemotherapy, radiotherapy, endocrine therapy, and targeted therapy, were administered to patients based on established national and institutional clinical practice guidelines prevalent during the study period and the multidisciplinary team's recommendations. Specific treatment regimens varied by individual patient and tumor characteristics but were not randomized.
Prognostic indicators included distant relapse-free survival (DRFS) and breast cancer-specific survival (BCSS). The DRFS was defined as the period from the date of radical surgery to the date of distant metastasis of breast cancer or death. The BCSS was defined as the period from the date of radical surgery to the date of death caused by breast cancer recurrence and metastasis. 21
Tissue Sample Collection and Processing
This study collected primary breast cancer and corresponding adjacent normal breast tissue specimens (n = 137) from patients who underwent therapeutic surgery at our center between January 1, 2012, and December 31, 2013. All tumor tissue samples were immediately fixed in 10% neutral buffered formalin upon collection, routinely processed, and embedded in paraffin blocks. These formalin-fixed, paraffin-embedded (FFPE) tissue blocks were then stored at room temperature in the institutional pathology archives until use.
Immunohistochemistry
Paraffin-embedded breast cancer sections were deparaffinized, rehydrated, and treated with 3% H2O2 to block endogenous peroxidase activity. Antigen retrieval was performed by boiling in sodium citrate buffer for 3 min. After cooling, 10% goat serum was applied to block nonspecific binding. Sections were incubated overnight at 4 °C with the primary antibody (Invitrogen, PA5-60528, 1:100, Waltham, USA), followed by the HRP-conjugated secondary antibody (Proteintech, SA00001-1, 1:10000, Beijing, China). A chromogenic reagent was added for 1–2 min, then slides were counterstained with hematoxylin, dehydrated through graded alcohol, cleared in xylene, and sealed. All sections were then observed and imaged using a Leica DM3000 microscope.
Five non-overlapping high-power fields were randomly selected. Staining intensity was scored as 0 (none), 1 (weak), 2 (moderate), or 3 (strong); the percentage of positive cells was scored as 0 (0%), 1 (≤25%), 2 (26-50%), 3 (51-75%), or 4 (76-100%). 22 The two scores were summed to yield a composite IHC score. All immunohistochemical staining was evaluated by two independent experienced pathologists who were blinded to the patients’ clinicopathological characteristics and study outcomes.
Patients were categorized into low- and high-CAMSAP3 expression groups using a 5-point cutoff value, determined by receiver operating characteristic (ROC) curve analysis to maximize sensitivity and specificity for predicting DRFS based on Youden's J statistic (sensitivity + specificity – 1). This threshold also aligned with the median IHC score for CAMSAP3 in this cohort (n = 137).
Cell Culture
The breast cancer cell MDA-MB-231 (National Collection of Authenticated Cell Cultures, SCSP-5043, RRID: CVCL_0062, Shanghai, China) were derived from an adenocarcinoma of the breast (pleural effusion), and their identity was confirmed by the vendor via short tandem repeat (STR) profiling. They were cultured in a mixed solution of 10% fetal bovine serum (FBS) (SERANA, S-FBS-SA-015, Waltham, USA) and 1% penicillin‒streptomycin (Biosharp, BL505A, Hefei, China) in DMEM (BDBIO, L100-500, Hangzhou, China). HCC1937 cells (Procell Life Science & Technology Co., Ltd, CL-0093, RRID: CVCL_0290, Wuhan, China) were derived from a primary breast ductal carcinoma, and their identity was confirmed by the vendor via STR profiling. They were cultured in RPMI-1640 medium (Gibco, 31800-105, Waltham, USA) supplemented with 10% FBS and 1% penicillin‒streptomycin. All cells were cultured in a humidified cell culture incubator at 37 °C with 5% CO2. When the cells reached 80%-90% confluence, they were passaged at a ratio of 1:2 or 1:4. The cells were trypsinized and passaged once every 2-3 days. Cells in the logarithmic growth phase were used for subsequent experiments.
Transfection of siRNAs or Plasmid
Transfection was performed when the cell density reached 70%∼80%. Two centrifuge tubes were prepared. For transfection, 2 μL of Lipofectamine 2000 (Invitrogen, 11668-019, Waltham, USA) was mixed with 50 μL (for siRNAs) or 100 μL (for plasmid) of Opti-MEM (Gibco, 31985-070, Waltham, USA) in tube A. Simultaneously, siRNA (Beijing Tsingke Biotech Co., Ltd, Beijing, China) or 4/8 μg of CAMSAP3 overexpression plasmid (YouBio, Changsha, China) was combined with 50 μL (for siRNAs) or 100 μL (for plasmid) of Opti-MEM in tube B. The siRNA sequences used were as follows: siR-CAMSAP3-1 sense (CGACGGAUUGAGGCCAUAUUC) and antisense (GAAUAUGGCCUCAAUCCGUCG); siR-CAMSAP3-2 sense (GCUGAUGGACGACCUCGAUAA) and antisense (UUAUCGAGGUCGUCCAUCAGC); siR-CAMSAP3-3 sense (AUCGCAUUCUGGAGGAAAUUG) and antisense (CAAUUUCCUCCAGAAUGCGAU). The preparation solutions were thoroughly mixed in tubes A and B. The culture medium was aspirated, the cells were washed twice with 1× phosphate-buffered saline (PBS), and 3 mL of fresh culture medium was added. The prepared mixture was added dropwise to the cells. Follow-up experiments were performed 48 h after transfection. We designed three siRNAs targeting different sequences and selected the one exhibiting the most effective knockdown efficiency, as validated by Western Blot (WB), for subsequent experiments.
Real-Time Quantitative PCR (RT-qPCR)
Total RNA was extracted using RNAex (500 μL; Accurate Biology, AG21101, Changsha, China) and phase-separated with 1-Bromo-3-chloropropane (BCP) (100 μL; Sigma-Aldrich, B9673, Taufkirchen, Germany). After centrifugation (12 000 rpm, 4 °C, 15 min), the aqueous phase was collected, precipitated with isopropyl alcohol (500 μL; Acros 327270010, Waltham, USA), and centrifuged (12 000 rpm, 4 °C, 10 min). The RNA pellet was washed twice with 75% ethanol (Sinopharm, 10009218, Shanghai, China), air-dried, and resuspended in DEPC-H₂O (Biosharp, BL510B, Hefei, China). Purity was assessed via OD260/280 and OD230/260 ratios. For reverse transcription, 1.5 μg RNA was mixed with Oligo dT (1 μL; Accurate Biology, AG11728, Changsha, China) in 13 μL DEPC-H₂O, heated at 70 °C for 5 min, then chilled. The reaction (20 μL) included 4 μL RT Buffer (5×), 1 μL dNTP (10 mM), and 1 μL M-MlV RTase (Thermo, K1622, Waltham, USA), and was incubated at 37 °C for 60 min and 70 °C for 15 min. qPCR was performed with SYBR Green (ABclonal, RK21204, Wuhan, China) in a 10 μL system containing 4 μL cDNA (25 ng), 0.5 μL each primer (forward: CCCCGTCAGGTCTCATGTCC; reverse: CGATTCTTCTGCGGTTCGTTCA). Cycling conditions were: 95 °C for 5 min; 40 cycles of 95 °C for 5 s and 60 °C for 4 s.
Western Blot
Cells were washed three times with PBS and lysed in ice-cold, in-house lysis buffer for 10 min. After centrifugation at 14 000 rpm, 4 °C for 10 min, the supernatant was collected and protein concentration was determined using a commercial assay (Thermo, 23228, Waltham, USA). Equal amounts of protein (20 μg per lane) were separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) (NCM Biotech, P2011, Suzhou, China) at 200 V for ∼49 min and transferred onto polyvinylidene difluoride (PVDF) membranes (Millipore, IPVH00010, Taufkirchen, Germany) using a constant current of 1.3 A for 10 min. Membranes were blocked with 5% bovine serum albumin (BSA) (Solarbio, A8020, Beijing, China) for 2 h at room temperature, then incubated overnight at 4 °C with a primary antibody (Biorbyt, orb165365, 1:1000, Cambridge, United Kingdom), followed by a 2 h room temperature incubation with secondary antibody (Sigma-Aldrich, F1804, 1:1000, Taufkirchen, Germany). After washing, membranes were developed using enhanced chemiluminescence (Biosharp, BL523A, Hefei, China), and signals were captured with a 15-s exposure.
Migration and Invasion Assays
For the wound healing assay, MDA-MB-231 (4 × 10⁵) and HCC1937 (5 × 10⁵) cells were seeded into 6-well plates. Once confluent, a linear scratch was created using a 10 μL pipette tip. Detached cells were removed by PBS washing, followed by the addition of 1 mL serum-free medium. Images were captured at 0 h and subsequent time points from 9 predefined fields per well using an inverted microscope (Olympus IX73). The migration rate was calculated as:
(scratch width at 0 h – scratch width at N h) / scratch width at 0 h × 100%.
For the Transwell invasion assay, cells were resuspended in serum-free medium and 1 × 10⁴ cells were seeded into upper chambers coated with Matrigel (ABW, 082704, Xiamen, China) at a concentration of 50 µL/cm², solidified at 37 °C for 30 min. The lower chambers were filled with 0.5 mL complete medium containing FBS. After 24 h incubation, non-invading cells were removed with cotton swabs. Cells that invaded to the underside were fixed with ice-cold 4% paraformaldehyde for 30 min, stained with crystal violet (Aladdin, C110703, Shanghai, China) for 15 min, and imaged. Cell numbers were counted from five random high-power fields using ImageJ software (NIH, USA).
Bioinformatics Validation
External validation was conducted using the Kaplan–Meier Plotter based on probe 1568617_a_at, which represents the CAMSAP3 gene. 23 The JetSet algorithm was employed to identify the optimal probe set. 24 Following the application of quality control filters, which included the removal of redundant samples and the exclusion of biased arrays, survival analyses were performed using auto-selected optimal cutoffs. The dataset comprised a sample size of 910 breast cancer patients, as determined by the query. Breast cancer subtype distribution data were not available from the Kaplan–Meier Plotter. Both distant metastasis-free survival (DMFS) and overall survival (OS) were assessed.
Statistical Analysis
Statistical analysis was conducted using SPSS 22.0 software. GraphPad 8.0 software was used to create the graphs. Normally distributed measurement data were typically presented as the mean ± standard deviation (SD), and an independent sample t test was used for hypothesis testing. Enumeration data were analyzed using Pearson's chi-square test. Survival analysis utilized the Kaplan‒Meier method and log-rank test to assess differences in survival curves between groups. Univariate and multivariate survival analyses utilized Cox regression analysis. Variables with a P value <0.10 in the univariate survival analysis were included in the multivariate survival analysis. The covariates included age, menstrual status, tumor diameter, lymph node status, histological grade, and the levels of ER, PR, HER2, Ki67, CK5/6, EGFR, and CAMSAP3. These variables were selected based on clinical relevance and univariate screening. Hazard ratios (HR) were provided with 95% CI. To determine the optimal CAMSAP3 IHC score cutoff for predicting DRFS, ROC curve analysis was performed. The cutoff was selected to maximize sensitivity and specificity based on Youden's J statistic (sensitivity + specificity – 1), with the area under the curve (AUC) and its 95% CI calculated to assess discriminatory ability. All the statistical tests were two-sided, and a P value less than 0.05 indicated a statistically significant difference. For in vitro experiments, including RT-qPCR, wound healing, and Transwell invasion assays, three biological replicates (n = 3) were performed, each with two technical replicates, which were averaged to yield one data point per biological replicate for statistical analysis. Western blot experiments were conducted with three biological replicates (n = 3) without technical replicates. The representative data are presented.
Results
Patient Characteristics
A total of 137 patients were included in the study (Figure 1). Among these, 54 patients (39.4%) were classified into the low CAMSAP3 expression group, while 83 patients (60.6%) were categorized into the high expression group based on a 5-point cutoff. The median age of the participants was 55 years (range: 29-80), and the median follow-up duration was 9.7 years. The baseline clinical and pathological characteristics, including ER, PR, HER2, and Ki-67, exhibited significant differences between the two groups, as summarized in Table 1. During the follow-up period, 18 patients developed distant metastasis (8 to bone, 7 to lung, 2 to liver, and 1 to brain), and 6 patients succumbed to breast cancer.

Flowchart of Patient Selection. Of 209 Initially Enrolled Patients, 72 were Excluded Based on Predefined Criteria, Resulting in a Final Cohort of 137 Breast Cancer Cases.
Clinicopathological Characteristics of Patients and Their Associations with CAMSAP3 Expression.

Bold values indicate statistical significance (P ≤ 0.05).
CAMSAP3, calmodulin-regulated spectrin-associated protein 3; ER, estrogen receptor; PR, progesterone receptor; HER2, human epidermal growth factor receptor 2; TNBC, triple-negative breast cancer; CK5/6, cytokeratin 5/6; EGFR, epidermal growth factor receptor.
Breast Cancers with Elevated CAMSAP3 Expression Exhibit Features of Lower Malignancy
CAMSAP3 expression was significantly reduced in breast cancer tissues compared to adjacent normal tissues (Figure 2A). The ROC curve analysis was performed to determine the optimal CAMSAP3 IHC score cutoff for predicting DRFS events, yielding an AUC of 0.65 (95% CI: 0.52-0.78), indicating moderate discriminatory ability. The optimal cutoff of ≥5, based on Youden's J statistic (sensitivity + specificity – 1), achieved a sensitivity of 70% and specificity of 60% for predicting DRFS events. This threshold also corresponded to the median IHC score in our cohort (n = 137). Using this cutoff, patients were stratified into a low-expression group (<5 points, n = 54) and a high-expression group (≥5 points, n = 83) (Figure 2B). Compared with the low-expression group, patients with high CAMSAP3 expression exhibited a lower histological grade, higher ER and PR positivity, fewer TNBC cases, and reduced expression of Ki67, CK5/6, and EGFR (Table 1). These findings suggest that high CAMSAP3 expression is associated with tumor features indicative of reduced malignancy.
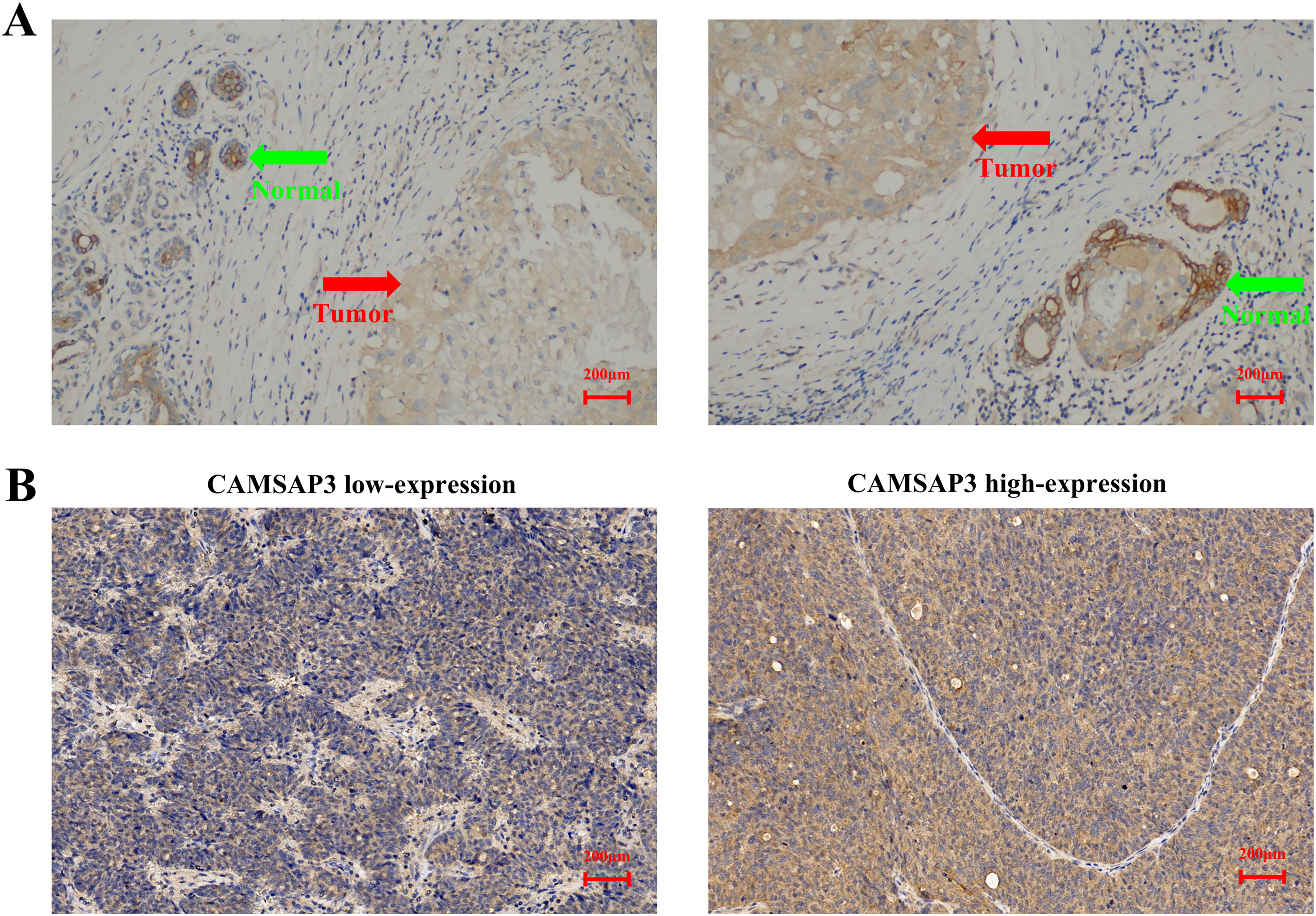
Immunohistochemical Detection of CAMSAP3 Expression in Clinical Breast Cancer Specimens. (A) Representative Image Illustrating Lower CAMSAP3 Expression in Primary Breast Cancer Tissue (Red Arrow) Compared to the Adjacent Normal Breast Tissue (Green Arrow) (×200 Magnification). (B) Representative Images of Low and High CAMSAP3 Expression in Breast Cancer Tissues (×200 Magnification). CAMSAP3, Calmodulin-Regulated Spectrin-Associated Protein 3.
High CAMSAP3 Expression is Associated with Improved Distant Recurrence Outcomes
Kaplan-Meier survival curves demonstrated that high CAMSAP3 expression was associated with significantly improved DRFS (P = 0.001, Figure 3A). BCSS analysis revealed no significant difference between the groups (P = 0.140, Figure 3B). Univariate Cox analysis identified lymph node status and CAMSAP3 as significant prognostic factors for DRFS. In the multivariate analysis, both factors remained independent predictors (Table 2), reinforcing the role of CAMSAP3 as a prognostic biomarker. Stratified analysis further demonstrated that the positive effect of high CAMSAP3 expression on DRFS persisted irrespective of lymph node status (Figure 3C and D).

High CAMSAP3 Expression is Associated with Improved Prognosis. (A-D) Kaplan-Meier Survival Analyses from Our Single-Center Cohort (n = 137) Showing that High CAMSAP3 Expression Correlates with Improved DRFS in the Total Cohort (A; P = 0.001) and in Stratified Subgroups of Lymph Node-Negative (C; n = 94, P = 0.005) and Lymph Node-Positive Patients (D; n = 43, P = 0.022). No Significant Correlation was Observed with BCSS (B; P = 0.140). (E-F) External Validation Using the Kaplan-Meier Plotter (n = 910). High CAMSAP3 mRNA Expression is Significantly Associated with Longer DMFS (E; HR = 0.70, 95% CI: 0.53−0.92; Log-Rank P = 0.010), while a Favorable but Non-Significant Trend is Observed for OS (F; HR = 0.75, 95% CI: 0.55−1.01; Log-Rank P = 0.059). CAMSAP3, Calmodulin-Regulated Spectrin-Associated Protein 3; DRFS, Distant Relapse-Free Survival; BCSS, Breast Cancer-Specific Survival; DMFS, Distant Metastasis-Free Survival; HR, Hazard Ratio; CI, Confidence Interval; OS, Overall Survival.
Univariate and Multivariate Analyses of DRFS.
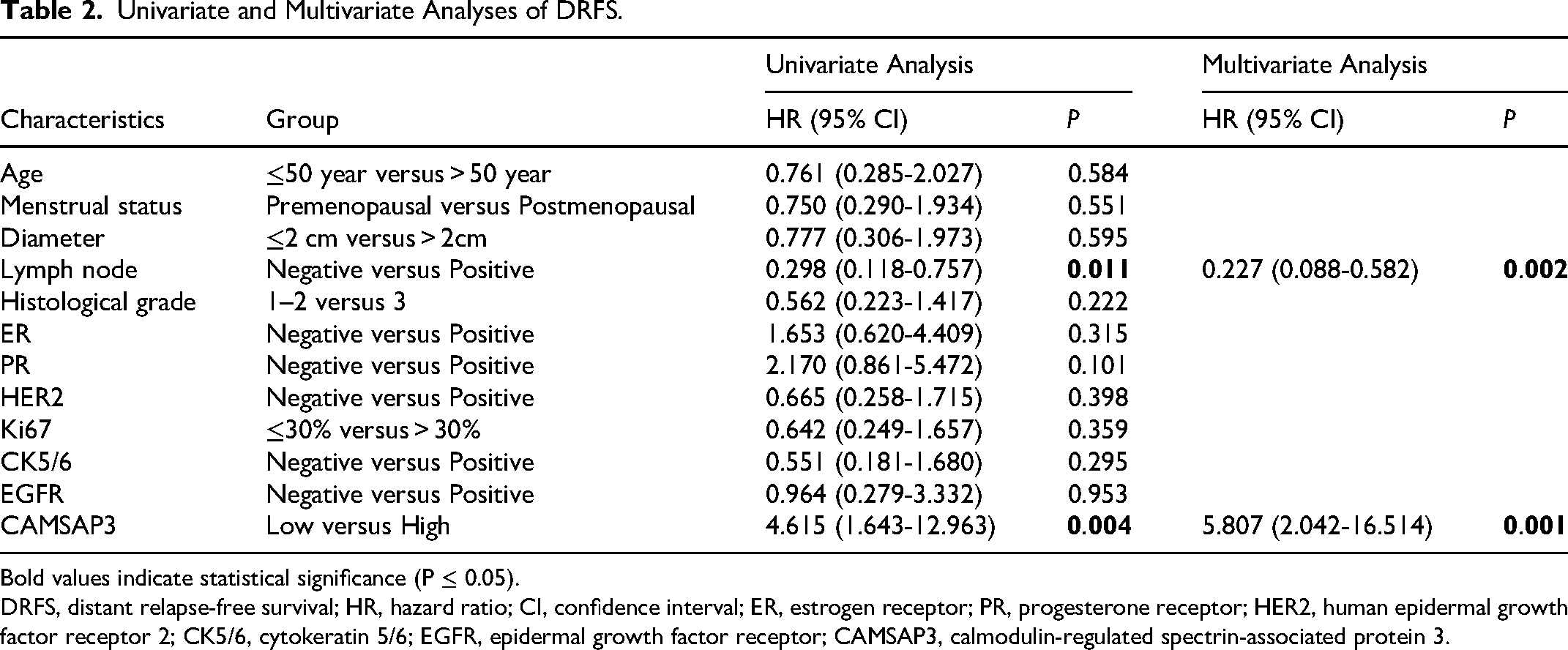
Bold values indicate statistical significance (P ≤ 0.05).
DRFS, distant relapse-free survival; HR, hazard ratio; CI, confidence interval; ER, estrogen receptor; PR, progesterone receptor; HER2, human epidermal growth factor receptor 2; CK5/6, cytokeratin 5/6; EGFR, epidermal growth factor receptor; CAMSAP3, calmodulin-regulated spectrin-associated protein 3.
External mRNA-based validation using the Kaplan–Meier Plotter demonstrated that high CAMSAP3 expression was significantly associated with improved DMFS (HR = 0.70, 95% CI: 0.53-0.92, log-rank P = 0.010, Figure 3E) in a cohort of 910 breast cancer patients. A favorable trend was also observed for overall survival (OS) (HR = 0.75, 95% CI: 0.55-1.01, P = 0.059, Figure 3F), although this did not reach statistical significance. These findings are consistent with our IHC-based results and further support the prognostic role of CAMSAP3 in breast cancer.
CAMSAP3 Inhibits Cell Migration and Invasion in Vitro
RT-qPCR analysis confirmed low endogenous expression of CAMSAP3 in MDA-MB-231 cells and high expression in HCC1937 cells (Figure 4A). CAMSAP3 overexpression in MDA-MB-231 cells significantly reduced both migration and invasion, as demonstrated by wound healing and Transwell assays (Figure 4B–D). To investigate the effects of CAMSAP3 silencing, three distinct siRNAs (siR-CAMSAP3-1, siR-CAMSAP3-2, and siR-CAMSAP3-3) were synthesized and evaluated for their knockdown efficiency. Analysis by WB showed that all three siRNAs effectively reduced CAMSAP3 protein levels in HCC1937 cells to varying degrees, with siR-CAMSAP3-3 exhibiting the most potent knockdown (Figure 5A). Therefore, siR-CAMSAP3-3 was selected for subsequent experiments. Conversely, the siRNA-mediated knockdown of CAMSAP3 in HCC1937 cells significantly enhanced these behaviors (Figure 5B–D). These in vitro findings indicate that CAMSAP3 suppresses the migratory and invasive properties of breast cancer cells.

CAMSAP3 Overexpression Suppresses Migration and Invasion of MDA-MB-231 Breast Cancer Cells. (A) Relative CAMSAP3 mRNA Expression in MDA-MB-231 and HCC1937 Cells, as Determined by RT-qPCR. (B) WB Showing Increased CAMSAP3 Protein Expression in MDA-MB-231 Cells Following Plasmid-Mediated Overexpression. (C) Wound Healing Assay Indicating Reduced Migratory Capacity Upon CAMSAP3 overexpression. (D) Transwell invasion assay demonstrating suppressed invasion in CAMSAP3- overexpressing cells. All in vitro Experiments were Independently Repeated Three Times. Data are Presented as Mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001. CAMSAP3, Calmodulin-Regulated Spectrin-Associated Protein 3; RT-qPCR, Real-Time Quantitative PCR; WB, Western Blot.
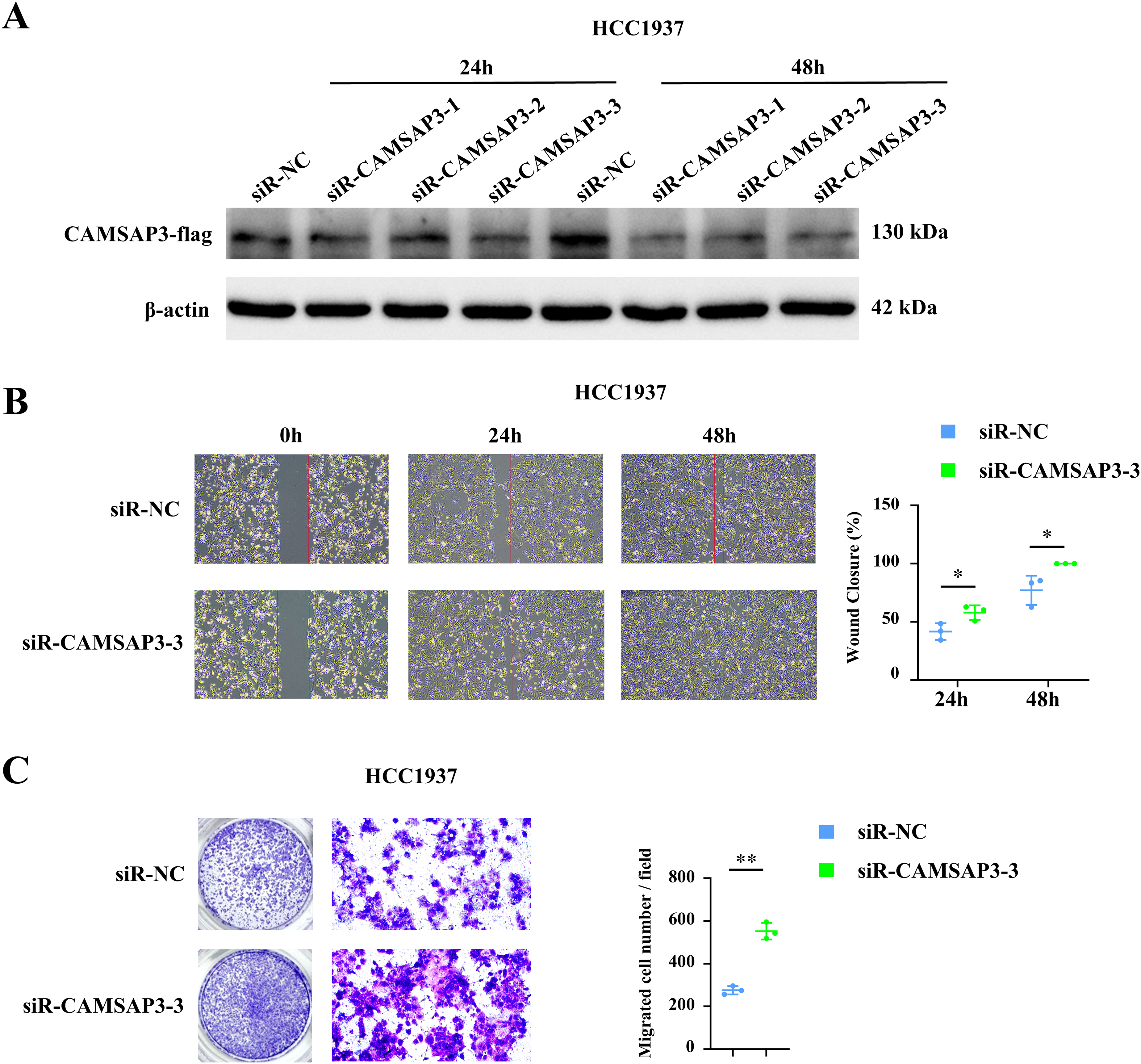
Knockdown of CAMSAP3 Enhances Migration and Invasion in HCC1937 Breast Cancer Cells. (A) WB Confirming Reduced CAMSAP3 Protein Expression Following siRNA-Mediated Knockdown. (B) Wound Healing Assay Showing Increased Migratory Capacity after CAMSAP3 Knockdown. (C) Transwell Invasion Assay Indicating Enhanced Invasion Upon CAMSAP3 Depletion. All In Vitro Experiments were Independently Repeated Three Times. Data are Presented as Mean ± SD. *P < 0.05, **P < 0.01. CAMSAP3, Calmodulin-Regulated Spectrin-Associated Protein 3; WB, Western Blot.
Discussion
Breast cancer metastasis remains the leading cause of disease-related mortality, and therapeutic options are limited once distant dissemination occurs. This highlights the urgent need to identify novel molecular regulators that drive metastatic progression and may serve as prognostic or therapeutic targets.25,26 In this study, we demonstrated within a single-center cohort that high expression of CAMSAP3 was associated with favorable clinicopathological features and improved DRFS in breast cancer patients. Importantly, this association was externally validated using a quality-controlled public dataset, which confirmed the correlation between high CAMSAP3 expression and improved DMFS. Additionally, it revealed a favorable—though not statistically significant—trend toward better OS. These findings reinforce the robustness and potential clinical relevance of CAMSAP3 as a prognostic marker. Furthermore, overexpression of CAMSAP3 significantly suppressed breast cancer cell migration and invasion in vitro, supporting its functional role as a metastasis suppressor. While a single-center design ensured standardized data collection and leveraged a unique regional cohort with a 9.7-year median follow-up, it may introduce selection bias and limit generalizability compared to multicenter studies; however, external validation with a larger dataset mitigates these concerns and supports broader applicability.
CAMSAP3 is a microtubule minus-end binding protein involved in cytoskeletal organization and cell polarity. Microtubules play essential roles in structural support and intracellular transport. In cancer cells, dysregulated microtubule dynamics are closely linked to enhanced motility and metastatic potential. 27 For instance, knockout of microtubule-stabilizing genes such as STMN1 and TALIN2 has been shown to suppress tumor cell migration.28,29 Previous studies have suggested that CAMSAP3 has tumor-suppressive functions in lung cancer. However, its role in breast cancer has been less understood. Our findings provide the first evidence that CAMSAP3 may also function as a suppressor of breast cancer metastasis.
Our study revealed that high CAMSAP3 expression was inversely correlated with multiple unfavorable tumor characteristics. Specifically, tumors with elevated CAMSAP3 levels exhibited lower expression of CK5/6 and EGFR—markers frequently associated with basal-like and aggressive breast cancer phenotypes.30,31 In addition, high CAMSAP3 expression was significantly associated with a lower histologic grade, positive hormone receptor status, and reduced prevalence of triple-negative breast cancer. These findings collectively suggest that CAMSAP3 may help maintain epithelial differentiation and suppress tumor aggressiveness. Consistently, functional assays demonstrated that CAMSAP3 overexpression inhibited migration and invasion in both ER-negative and triple-negative breast cancer cell lines, indicating its potential role in limiting metastatic behavior in aggressive subtypes.
Currently, few molecular markers specifically predict metastatic potential in breast cancer. Our findings suggest that CAMSAP3 may serve as a clinically valuable biomarker for stratifying metastatic risk. Given its role in regulating microtubule stability, CAMSAP3 may intersect with cytoskeletal remodeling pathways such as the epithelial–mesenchymal transition (EMT), which warrants further mechanistic investigation.
The identification of CAMSAP3 as a metastasis suppressor opens avenues for clinical translation, particularly through strategies that enhance CAMSAP3 expression or stabilize microtubules to inhibit metastatic progression. Pharmacological agents that promote microtubule stability, such as taxanes (eg, paclitaxel), could be explored in combination with CAMSAP3 upregulation to synergistically suppress cancer cell motility and invasion. Additionally, developing small-molecule enhancers of CAMSAP3 activity could offer a targeted approach to bolster its tumor-suppressive effects, especially in aggressive breast cancer subtypes like triple-negative breast cancer. However, challenges remain, including the need for in vivo validation to confirm these effects and the development of delivery systems to ensure targeted and efficient drug administration. Novel delivery technologies could significantly enhance the feasibility of these therapeutic strategies. Emerging biomimetic nanocarriers, such as extracellular vesicles derived from chimeric antigen receptor monocytes, have demonstrated efficient tumor-targeted delivery via non-invasive routes and may offer a promising platform to potentiate microtubule-stabilizing or CAMSAP3-enhancing therapeutics in metastatic breast cancer. 32
Several limitations should be acknowledged in this study. First, while in vitro assays demonstrated CAMSAP3's suppressive role, in vivo validation and exploration of its specific molecular mechanisms, such as EMT regulation, were not conducted. Second, the IHC-based expression cutoff, though supported by external data, requires further validation in larger, multicenter cohorts. Third, despite designing and validating three siRNAs, functional assays primarily used a single representative siRNA. Although all three siRNAs effectively suppressed CAMSAP3, the reciprocal findings from our overexpression experiments internally validate that observed effects are specific to CAMSAP3 knockdown, mitigating concerns about off-target effects. Finally, using only two cell lines limits the representation of breast cancer heterogeneity, suggesting future investigations with diverse models (eg, EGFR- or HER2-positive) are warranted to fully elucidate CAMSAP3's role.
Conclusion
In conclusion, our study identifies CAMSAP3 as a potential metastasis suppressor and independent prognostic marker in breast cancer. These findings provide a basis for future mechanistic exploration and clinical validation.
Footnotes
Abbreviations
Acknowledgements
Not applicable.
Ethics Approval
This study was approved by the Ethics Committee of the Affiliated Hospital of Jiaxing University (the First Hospital of Jiaxing) (third Floor, Administration Building, Jiaxing First Hospital, No. 1882 Zhonghuan South Road, Jiaxing, Zhejiang, China) under approval number 2023-LY-083 (March 16, 2023). The study adhered to the Declaration of Helsinki.
Consent to Participate
All participants provided written informed consent to participate in this study.
Consent to Publish
Not applicable.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Jiaxing Science and Technology Plan Project (No. 2022AY30019), the Research Fund of the First Hospital of Jiaxing (No. 2023-YB-036), the Jiaxing Key Discipline of Medicine (Mastropathy, Innovation Subject) (No. 2023-FC-001), the Breast Cancer Precision Diagnosis and Treatment Center of the First Hospital of Jiaxing (No. 2021-ZZZX-06), and the Jiaxing Provinces and Cities Jointly Cultivated Discipline—General Surgery (Minimally Invasive) (No. 2023-PYXK-001).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
