Abstract
Background
Lung adenocarcinomas manifesting as part-solid nodules (PSNs) represent a distinct clinical subtype where accurate preoperative determination of pathological invasiveness critically influences both prognosis and surgical decision-making. This multicenter study aims to develop an ensemble machine learning classifier that integrates computed tomography (CT) radiomic signatures with clinical-radiological features to enhance the preoperative prediction of invasive status.
Methods
We retrospectively analyzed 344 patients with pathologically confirmed lung adenocarcinoma presenting as PSNs across three medical centers. Following random allocation into training (n = 240) and validation (n = 104) sets (7:3 ratio), we extracted 1239 quantitative radiomic features from preoperative thin-section CT scans. Through rigorous feature engineering, we constructed a radiomic score using least absolute shrinkage and selection operator regression. We systematically evaluated both single-algorithm classifiers and ensemble approaches (including hard/soft voting and stacking), incorporating both the radiomic score and clinical-radiological features.
Results
Among the various evaluated machine learning models, the stacking classifier, which combines radiomic scores and clinical-radiological features, performed the best, achieving an AUC of 0.84, an accuracy of 0.817, an F1 score of 0.869, a precision of 0.818, and a recall of 0.926.
Conclusion
Our stacking ensemble learning classifier, which synergistically combines CT radiomics signatures with clinical-radiological features, provides a clinically actionable tool for the preoperative prediction of pathological invasiveness in PSN-type lung adenocarcinoma, thereby enhancing individualized surgical planning.
Keywords
Introduction
The global prevalence of lung cancer makes it one of the most lethal malignancies, with adenocarcinoma having the highest incidence among lung cancers.1–3 According to the 2017 guidelines issued by the Fleischner Society, lung cancer observed in computed tomography (CT) imaging is categorized into three distinct types based on the proportion of solid to ground-glass components. These categories include pure ground-glass nodules (pGGNs), characterized exclusively by ground-glass opacity; solid nodules (SNs), which consist entirely of solid tissue; and part-solid nodules (PSNs), which comprise both ground-glass and solid components. 4 Among these, PSNs represent a distinct clinical subtype characterized by a lower prevalence of lymphatic metastasis, visceral pleural invasion, and spread through air spaces compared to SNs. Furthermore, patients with PSNs tend to have better recurrence-free survival (RFS) and overall survival rates than those with SNs, although these rates are worse than in patients with pGGNs.5–7
Lung adenocarcinomas presenting as PSNs can be classified pathologically as adenocarcinoma in situ (AIS), minimally invasive adenocarcinoma (MIA), or invasive adenocarcinoma (IAC), depending on their invasive characteristics. The 5-year recurrence-free survival (RFS) after complete resection approaches 100% for AIS and MIA, whereas it drops to approximately 74.1% for IAC.8,9 Therefore, wedge resection or segmentectomy is generally recommended for AIS and MIA because these procedures can preserve lung tissue and significantly reduce surgery-related complications. In contrast, lobectomy is typically necessary for IAC to achieve a lower tumor recurrence rate, despite its higher risk of adverse surgical outcomes.10,11 Consequently, preoperative prediction of IAC in lung adenocarcinomas manifesting as PSNs is vital for guiding treatment decisions, especially regarding the choice of surgical extent.
Radiomics has shown considerable potential in elucidating the biological characteristics of lung cancer through quantitative imaging analysis. Recent studies highlight its clinical effectiveness in predicting high-grade patterns in stage I lung adenocarcinoma, 12 enhancing preoperative tumor grading accuracy, 13 and differentiating immune-related pneumonitis from tumor progression in stage IV non-small cell lung cancer during immunotherapy, 14 thereby impacting clinical decisions. Furthermore, hybrid models that combine radiomics with clinical and radiological features demonstrate promising potential for preoperative evaluation of the pathological invasiveness of lung adenocarcinoma presenting as PSNs.15–18 However, in practical applications, selecting the most appropriate statistical method to integrate radiomics with clinical-radiological data can be challenging. Traditional methods relying on single-model frameworks, such as logistic regression with fixed parameters, often fail to capture the complex patterns and relationships within the data.15–18 Machine learning algorithms, on the other hand, can better model nonlinear relationships and high-dimensional interactions, thereby reducing the risks of overfitting and underfitting while enhancing generalizability. Future research in this field would particularly benefit from leveraging machine learning approaches. 19
To address this, we used both single-algorithm classifiers and ensemble machine learning (including hard/soft voting and stacking) to combine multiple models for final predictions,19–22 integrating radiomics with clinical-radiological features. Ensemble classifiers merge different models by combining various algorithms or data perspectives, which help balance their strengths and weaknesses. Consequently, they enhance overall performance in complex medical decision-making, leading to more accurate and reliable predictions than any single model alone. This is the first study to systematically compare and combine these techniques for predicting lung adenocarcinoma invasiveness in PSNs. Our goal is to develop robust prediction models by reducing complexity and maximizing data utilization, thereby guiding future efforts in selecting effective diagnostic tools for personalized treatment planning.
Materials and Methods
Study Population
In this retrospective three-center study, we collected CT images and pathological diagnoses from patients who underwent surgical resection for lung adenocarcinoma between January 2021 and June 2023. The inclusion criteria were as follows: (a) PSNs with a maximum diameter of 5–30 mm on preoperative CT images; (b) postoperative pathological examination confirming lung adenocarcinoma; and (c) a chest CT scan performed within 14 days prior to surgery. The exclusion criteria included: (a) patients who had received preoperative chemoradiotherapy; (b) patients with multiple primary lung cancers or other malignancies; and (c) unqualified images containing artifacts, atelectasis, or pneumonia, which complicated segmentation. The screening flowchart of enrolled patients is provided in Supplement Figure 1. This retrospective study enrolled 344 patients with pathologically confirmed lung adenocarcinoma presenting as PSNs. Among these patients, 235 (68.3%) were diagnosed with IAC, while 109 (31.7%) were classified as non-invasive adenocarcinoma. Within the non-IAC cases, 98 (28.5%) were classified as MIA, and 11 (3.2%) as AIS. Of the 344 enrolled lung adenocarcinoma patients, 223 (64.8%) were male, and 121 (35.2%) were female, with a median age of 59 years (interquartile range [IQR]: 52-66). The participants were divided into training (n = 240, 69.8%) and validation (n = 104, 30.2%) sets in a 7:3 ratio without stratification. The training set included 156 males (65.0%) with a median age of 59 years (IQR: 52-66.2), while the validation set comprised 67 males (64.4%) with a median age of 59.5 years (IQR: 52-66).
The institutional review boards of all participating institutions approved the study (2021-07-009 from Xiangtan Central Hospital, KY2020147 from the Affiliated Hospital of Southwest Medical University, and 2023YJSLL-121 from the Affiliated Hospital of Guilin Medical University). Due to the retrospective nature of the study and the use of anonymized data, the boards waived the requirement to obtain written informed consent.
CT Radiological Feature Evaluation
The CT scanning parameters used in this study were consistent with those from our previous research.12,13 For further details on the acquisition of CT images, please refer to Supplemental Digital Content 1.
Eligible CT images collected from the three centers were co-registered and stored in DICOM format. These images were subsequently imported into the Picture Archiving and Communication System to visualize all enrolled PSNs. The CT radiological features of these PSNs on high-resolution CT images were interpreted by senior cardiothoracic radiologists, who were not informed of the pathological diagnoses. Tumor size was defined as the maximum diameter, specifically the longest perpendicular diameter in the axial plane, according to the eighth edition of the TNM classification of lung cancer. 23
Additionally, the consolidation-to-tumor ratio (CTR) was defined as the ratio of the maximum size of the solid component to the maximum tumor size, as illustrated in the schematic diagram (Supplement Figure 2). CTR measurements were taken for the maximum diameter of both the entire tumor and the solid component within the lung window, which has a width of 1500 Hounsfield units (HU) and a level of −700 HU.24,25 The solid component was characterized as a region of heightened opacification that fully concealed the vascular markings beneath it. In contrast, ground-glass opacity is defined as an area with a minor, homogeneous increase in density that does not obscure the vascular markings below. Finally, the other CT radiological features included location, margin, shape, lobulation sign, spiculation sign, vascular convergence sign, vacuole sign, and pleural indentation sign. Any disagreements in the CT image evaluation were discussed collaboratively until a consensus diagnosis was reached.
Lesion Delineation and Radiomics Workflow
All CT images underwent semiautomatic segmentation of PSNs using 3D Slicer software (version 5.2.2; www.slicer.org). The segmentation process began with an initial contouring protocol executed by a radiologist-in-training who specialized in thoracic oncology imaging and had five years of relevant experience. This initial contouring was subsequently refined through an iterative process by an attending thoracic radiologist with 15 years of subspecialty expertise, ensuring the accuracy of the volumetric measurements.
The radiomic analysis followed a structured pipeline consisting of four main phases: image processing, feature quantification, dimensionality optimization, and predictive modeling. First, we performed image resampling using the PyRadiomics package within the Python programming environment to standardize voxel spacing to 1 × 1 × 1 mm3 (x, y, z). The resampling algorithm was based on the B-spline curve, with the interpolator set to sitkBSpline (degree = 3), ensuring smooth and accurate interpolation of the image data. Next, we adjusted the bin width to partition the pixel intensity range of 0–255 into five intervals. This binning approach effectively discretized the intensity values, reducing the dimensionality of the feature space and facilitating subsequent analysis. Second, we extracted 1239 quantitative radiomics features per PSN, including shape, texture, and wavelet-filtered attributes. To standardize these features, normalization was performed using a z-score transformation (mean μ=0, standard deviation σ=1). Following feature extraction, univariate statistical testing (two-tailed t-test, α=0.05) was applied to exclude non-discriminative features. A pairwise collinearity assessment was then conducted to identify redundant variables using Pearson's correlation coefficient (|r| ≥ 0.9). In cases where correlation was detected, one feature from each cluster of correlated features was iteratively eliminated through a stochastic selection process.
The final derivation of the radiomic signature was accomplished using least absolute shrinkage and selection operator (LASSO), incorporating nested 10-fold cross-validation to minimize the risk of overfitting (Supplement Figure 3). Features that retained non-zero coefficients following the regression analysis were integrated into a composite radiomic score (Radscore) through a weighted linear combination of the selected features.
Machine Learning Framework
The Radscore was synergistically integrated with clinical-radiological features to act as composite predictors in our computational modeling pipeline. We employed dual-layer analytical protocols that encompass both foundational and sophisticated statistical constructs, facilitating the preoperative discrimination of invasive adenocarcinoma manifesting as PSNs.
Single-Algorithm Classifiers
Fundamental predictive architectures implemented monolithic machine intelligence frameworks, including random forest (RF), adaptive boosting (AdaBoost), light gradient boosting machine (LightGBM), gradient boosting decision tree (GBDT), extreme gradient boosting (XGBoost), and categorical boosting (CatBoost). These methodologies, which employ arboreal integration through stratified data subspace allocation, wherein individual decision trees undergo parallel optimization across partitioned feature-phenotype cohorts, yield promising results in the analysis of biomedical data, particularly in the diagnosis of lung cancer.26–29
Ensemble Machine Learning Classifiers
Ensemble machine learning classifiers utilize advanced integration strategies, including stacking classifiers and voting mechanisms. The stacking classifier architecture adopts a hierarchical two-level structure: the base level integrates predictions from six primary classifiers (RF, GBDT, XGBoost, LightGBM, AdaBoost, CatBoost), while the meta-level employs a logistic regression meta-classifier to optimally combine these outputs through cross-validated probability calibration, as depicted in Figure 1. Parallel voting strategies operate through two distinct consensus protocols: hard voting, which determines class membership by majority rule across base classifier decisions, and soft voting, which calculates class probabilities via the weighted average of individual classifier confidence scores. Additionally, stacking classifiers—built upon a dual integration framework—enhance predictive robustness by simultaneously mitigating model bias through stacked generalization and minimizing variance via consensus-based decision aggregation. This framework maintains strict separation between base learner training spaces and meta-learner optimization domains, ensuring modular architecture and preventing information leakage. 30

Architecture of the Stacking Classifier.
Model Selection Protocol
The experimental design schematic outlines our multi-criteria optimization framework (Figure 2). Diagnostic performance quantification utilizes the area under the receiver operating characteristic (ROC) curve (AUC) as the principal criterion, supplemented by an extensive evaluation matrix that encompasses accuracy, precision, recall, and F1 score. Stratified partitioning allowed the development cohort to facilitate model parameterization, while the independent validation cohort enabled unbiased performance benchmarking. The classifier demonstrating maximal AUC robustness during validation analyses was designated as the optimal configuration. 31

Schematic Representation of the Experimental Design Utilized in this Study.
Interpretability Analysis Framework
Model explainability was enhanced through the implementation of Shapley Additive Explanations (SHAP) to quantify feature contribution hierarchies.32,33 A sequential optimization protocol was initiated through ordered feature integration based on SHAP-derived importance rankings. The algorithmic workflow involved cyclical retraining of classifier architectures with progressive feature inclusion, coupled with validation cohort AUC recalibration at each augmentation stage. This dynamic feature integration paradigm enabled simultaneous monitoring of incremental predictive gain and the evolution of SHAP value trajectories. Through incremental learning frameworks, the parametric impact of individual predictors on global model performance was visualized via multivariate gradient mapping, establishing explicit correlations between feature-space perturbations and performance metric fluctuations across sequential learning iterations.
Statistical Analyses
All statistical analyses were performed using R software (version 3.6.3). Categorical variables were expressed as frequencies and percentages (%), whereas continuous variables were summarized as means with standard deviations for normally distributed data or as medians with interquartile ranges for non-normally distributed data. To compare groups, chi-square tests were used for categorical variables, and either independent samples t-tests or Mann-Whitney U tests were applied for continuous variables, depending on their distribution. A two-tailed p-value of less than 0.05 was considered statistically significant.
Result
Baseline Characteristics
This retrospective study enrolled 344 patients with pathologically confirmed lung adenocarcinoma presenting as PSNs. As shown in Table 1, a comparative analysis of the demographic and radiological characteristics between the training and validation sets revealed no statistically significant differences (all p > 0.05). This lack of significant disparities indicates a high level of homogeneity, ensuring balanced feature distributions for subsequent model development and evaluation.
Comparative Analysis of Clinical-Radiological Features Between the Training and Validation Sets.
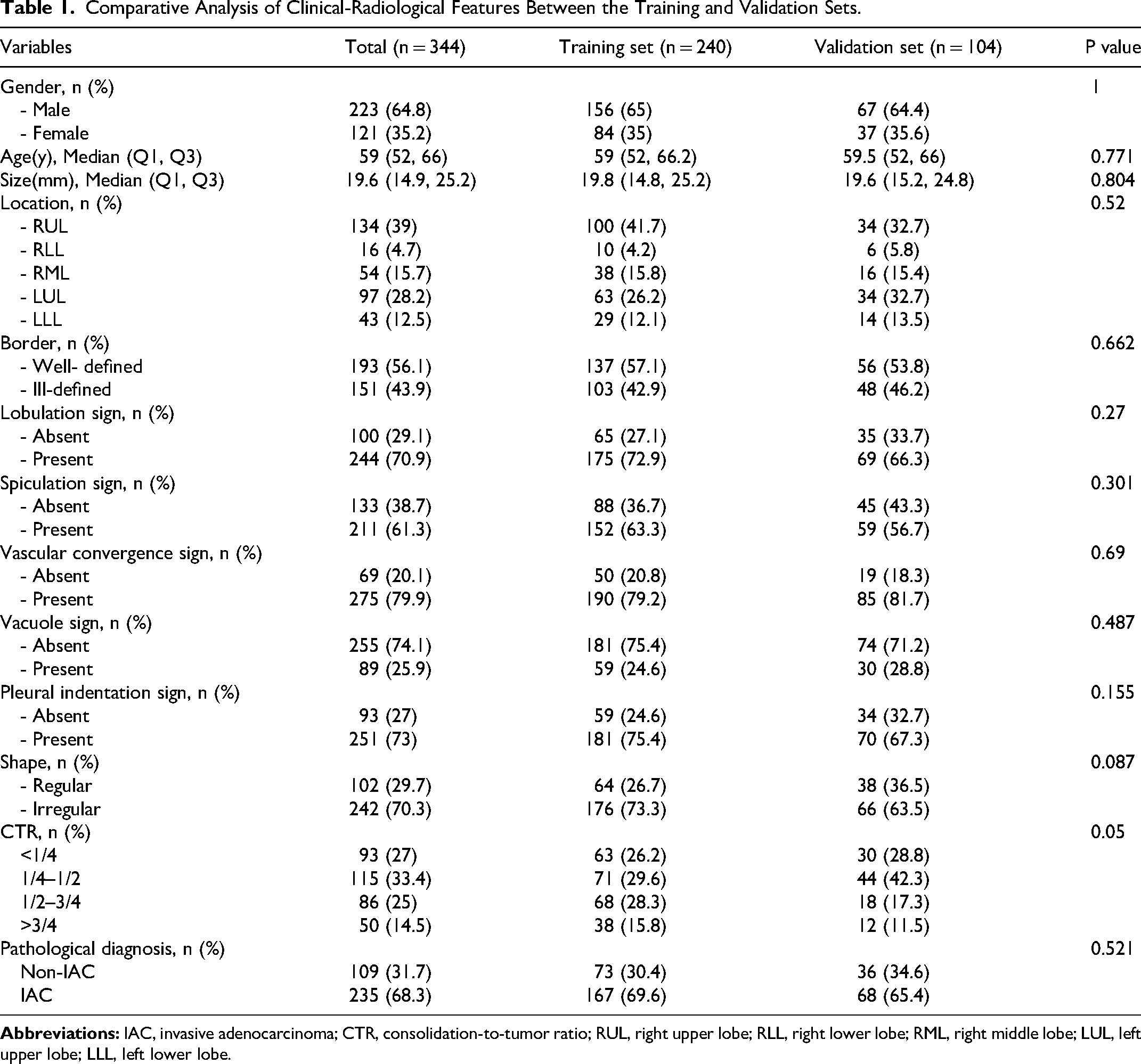
The diagnosis of lung adenocarcinoma presenting as PSNs highlighted that pathological invasiveness was more common among male patients. This invasiveness was associated with advanced age, larger tumor sizes, the presence of vacuole signs, and higher CTR (see Supplement Table 1).
Radiomic Analysis
Quantitative radiomic profiling of preoperative thin-section CT scans resulted in the identification of 1239 candidate radiomics features for each PSN. A rigorous dimensionality reduction protocol was employed to systematically refine these features in two phases. First, 639 radiomic features were selected using batch t-tests. This was followed by the retention of 188 radiomic features through a pairwise collinearity assessment, where Pearson's correlation coefficient (|r|) was set at ≥0.9. Subsequently, this optimized set of radiomic features underwent LASSO regression with 10-fold cross-validation. This process led to the identification of 17 non-collinear features that maintained predictive significance (λ=0.027; see Supplemental Figure 4). The details of these features are provided in Supplemental Digital Content 2. Additionally, inter-feature dependency analysis using Spearman's rank correlation confirmed minimal multicollinearity across the final signature components (see Supplement Figure 5), with absolute coefficient magnitudes consistently maintained below critical multicollinearity thresholds.
The Radscore is used for building radiomics models. In this binary classification task, we aim to distinguish IAC from non-IAC in lung adenocarcinoma presenting as PSNs. The radiomics model achieved an AUC of 0.799, an accuracy of 0.798, an F1 score of 0.840, a precision of 0.873, and a recall of 0.809.
Machine Learning Performance Comparison
Within our predictive modeling framework, radiomics-derived Radscore and clinical-radiological features served as dual-channel inputs for evaluating pathological invasiveness in lung adenocarcinomas presenting as PSNs. Based on ROC analysis (Figure 3), a comparative evaluation of diverse algorithmic architectures revealed substantial performance disparities among the models, with clear stratification in AUC values. Notably, the XGBoost implementation demonstrated the optimal discriminative capacity with an AUC of 0.831 when evaluated as a base learner. In contrast, ensemble strategies exhibited varying levels of efficacy: conventional voting architectures, comprising hard voting (AUC = 0.770) and soft voting (AUC = 0.830), displayed suboptimal performance relative to single-algorithm benchmarks. However, the stacking classifier achieved the peak diagnostic accuracy of AUC = 0.840 through meta-learner optimization (Table 2). Furthermore, a multidimensional performance evaluation (Figure 4) quantified classifier effectiveness across key metrics, revealing that the stacking classifier attained a balanced performance with an accuracy of 0.817, an F1 score of 0.869, a precision of 0.818, and a recall of 0.926.

ROC Analysis Comparative Evaluation Showcasing Significant Performance Discrepancies among Various Algorithmic Architectures.

Multidimensional Performance Evaluation Quantitatively Assessing Classifier Efficacy Across Key Metrics, Including Accuracy, Precision, Recall, and F1 Score.
Assessment of Diagnostic Efficacy Across Diverse Machine Learning Approaches.

Model Interpretation
When assessing the contribution of the first layer to the stacking classifier, the RF algorithm exerted the most significant influence, followed by CatBoost, LightGBM, XGBoost, AdaBoost, and GBDT, as shown in Figure 5. A global interpretation of the stacking classifier for the second layer, based on SHAP values, is presented in Figure 6. Among the features analyzed, Radscore had the most significant contribution, followed by CTR, size, age, border, pleural indentation sign, vacuole sign, gender, location, shape, spiculation sign, lobulation sign, and vascular convergence sign. Additionally, Figure 7 illustrates the decoding of gender-specific feature contributions via the mean absolute SHAP values to address potential differences between male and female patients regarding both radiomic and clinical-radiological features. In this analysis, males contributed 53.7%, while females contributed 46.3%.

Bee Diagram Illustrating the Contribution of the First Layer Within the Stacking Classifier Framework.
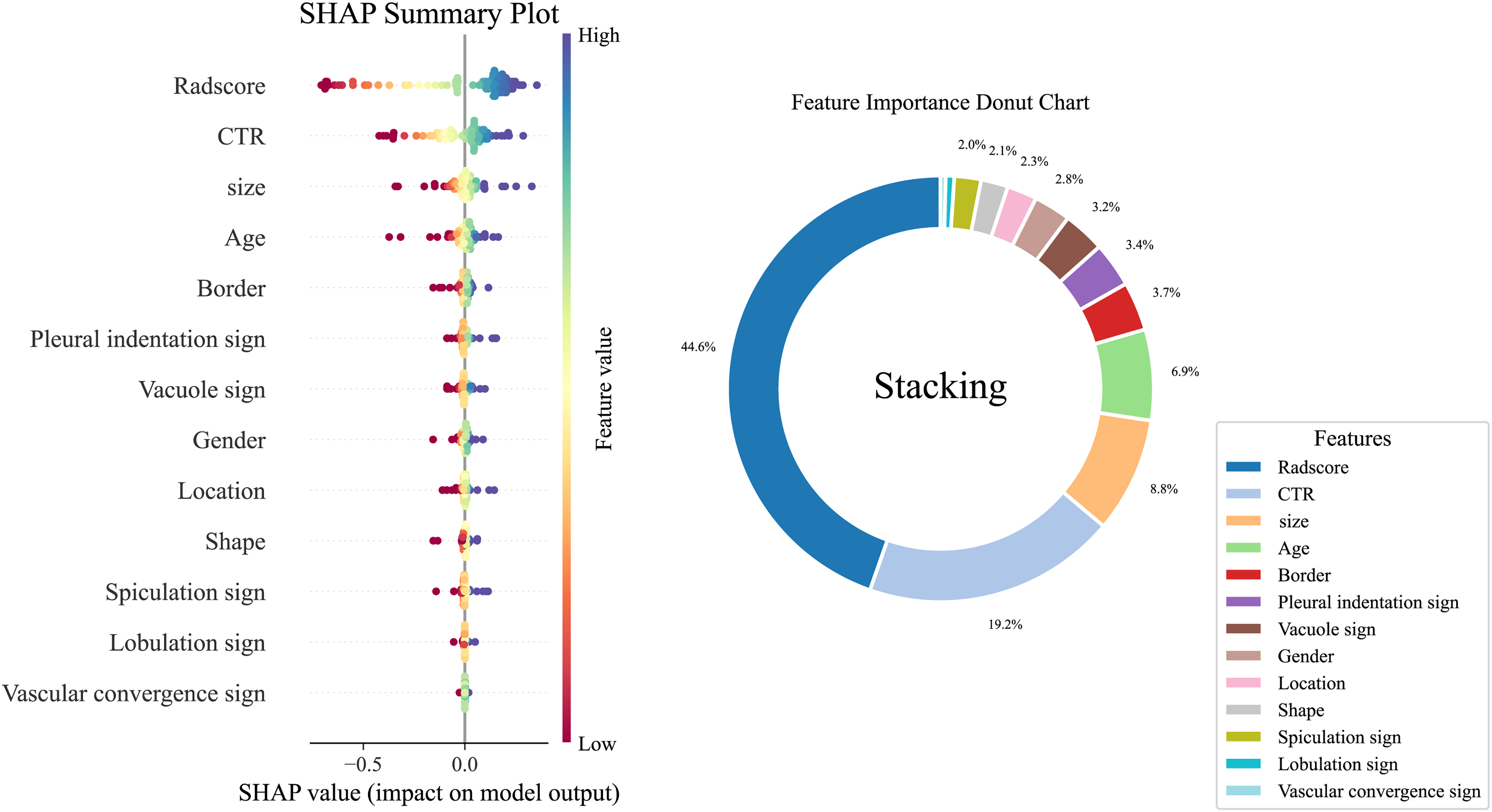
Global Interpretation of the Stacking Classifier for the Second Layer, Informed by SHAP Values.
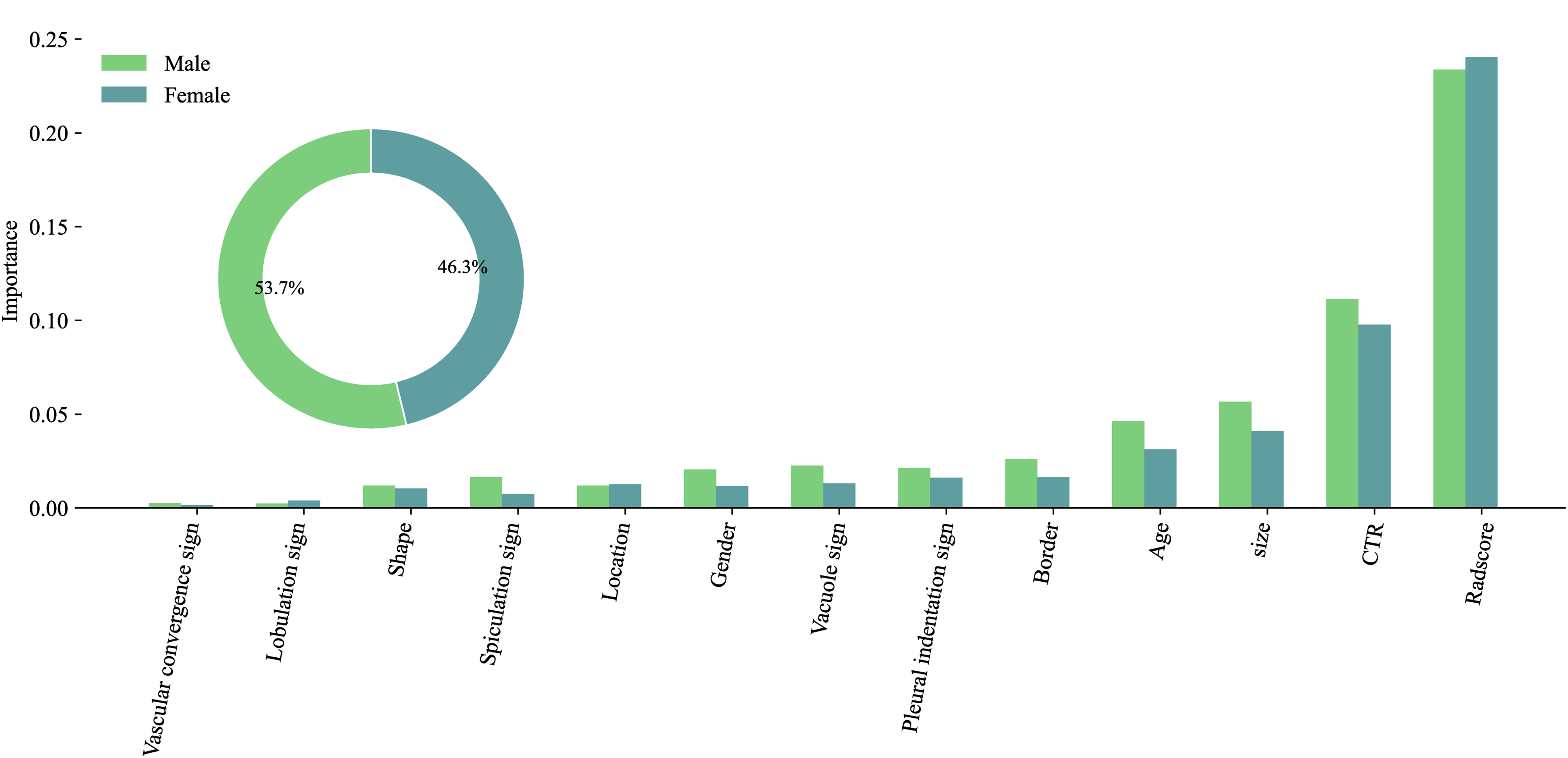
Male & Female Feature Contributions: Decoding with Mean Absolute SHAP.
To enhance the clinical translatability of our SHAP analysis, we have included a real-world case example in the force plot (Figure 8), which illustrates the prediction explanation for a specific instance. In this plot, positive impacts are shown in red, while negative impacts are displayed in blue. The length of the arrows signifies the magnitude of each feature's impact on the output.

In this Force Plot, it is Evident that the Highest Rad Score (0.17), Followed by CTR (1/4–1/2) and Size (19.72 mm), Positively Influences the Risk of IAC in Lung Adenocarcinomas Presenting as PSNs.
Figure 9 demonstrates the relative contribution of different features to the model's predictive performance, along with their corresponding AUC scores. The top four features (Radscore, CTR, size, age) exhibit significantly higher contribution values compared to the other variables. Notably, CTR shows the strongest contribution, indicating its substantial discriminative power in classification tasks. The next three highlighted features reveal a gradual increase in both contribution magnitude and associated AUC values. However, after incorporating additional features, the AUC began to decline slowly. Despite this, the AUC metric remains consistently above 0.7 across all top features, suggesting that the model maintains adequate discrimination capability.

Relative Contribution of Distinct Features to the Model's Predictive Performance, Accompanied by Their Respective AUC Scores.
Discussion
This multicenter study demonstrates that ensemble machine learning classifiers, which integrate CT radiomic signatures with clinical-radiological features, achieve robust performance in the preoperative prediction of pathological invasiveness for lung adenocarcinoma presenting as PSNs. The stacking classifier, leveraging meta-learning to harmonize heterogeneous base models, outperformed both conventional voting strategies and single-algorithm approaches.
The stacking classifier effectively combines diverse features from base models like RF, CatBoost, LightGBM, XGBoost, AdaBoost, and GBDT by utilizing a meta-learner. Unlike fixed voting methods, stacking classifier employs a trainable meta-learner, such as logistic regression, to capture nonlinear interactions between base model outputs. This approach is particularly valuable in medical imaging tasks, where interpretability and robustness are crucial. By leveraging the strengths of individual base models while mitigating their weaknesses, the stacking classifier provides a robust framework for integrating multimodal biomarkers. Consequently, stacked ensemble methods are especially advantageous in this specific context.20–22,30
To further explore the interpretability analysis framework using SHAP,32,33 the top four features (Radscore, CTR, size, age) exhibit significantly higher contribution values compared to other variables. The Radscore contributes the most, aligning with its validated role in quantifying tumor heterogeneity, which underlies biological processes such as chaotic angiogenesis and heterogeneous cellular proliferation patterns within solid components.15–18 CTR serves as a surrogate marker for tumor invasiveness in lung adenocarcinoma presenting as PSNs,15–18,34–36 with higher CTR values reflecting aggressive pathological features. Furthermore, the semiquantitative assessment of CTR, divided into four equal parts, is regarded as a valuable tool for the prognostic evaluation of PSNs. 36
Tumor size, defined as the maximum diameter according to the eighth edition of the TNM classification of lung cancer, 23 reflects an infiltrative invasive growth pattern. Additionally, advanced age is associated with tumor invasiveness in lung adenocarcinoma presenting as PSNs, consistent with previous studies.15–18 Notably, earlier research has shown that tumor size becomes a more critical determinant of outcomes.37,38 This emphasizes the importance of integrating clinical-radiological features (CTR, tumor size, age) with quantifications of tumor heterogeneity for precise risk stratification, highlighting comprehensive integration within the current stacking classifier.
The present study has some limitations. First, the retrospective nature of the study introduced variability in CT scanners and imaging parameters across and within centers, which may have influenced texture feature extraction and the generalizability of the model. Second, while this stacking classifier effectively aids in the preoperative prediction of pathological invasiveness for lung adenocarcinoma presenting as PSNs, training each layer required separate datasets. This increases computational demands and algorithm complexity, and it did not incorporate molecular or genetic biomarkers known to influence tumor invasiveness, thereby limiting its biological comprehensiveness. Finally, the inclusion of only surgically resected cases may introduce selection bias, as nodules managed non-surgically were excluded, and there was a lack of data on longer postoperative follow-up times, which affects the applicability of results to broader clinical populations.
Conclusion
In conclusion, the stacking ensemble model establishes a new benchmark for preoperative risk assessment in PSNs by synergizing multi-dimensional data streams. Radscore, tumor size, CTR, and age collectively provide complementary insights into morphological features, demographic factors, and tumor heterogeneity. These advances hold significant potential for personalizing therapeutic strategies, particularly in guiding decisions between limited resection and lobectomy for early-stage lung cancer.
Supplemental Material
sj-docx-1-tct-10.1177_15330338251351365 - Supplemental material for Ensemble Machine Learning Classifiers Combining CT Radiomics and Clinical-Radiological Features for Preoperative Prediction of Pathological Invasiveness in Lung Adenocarcinoma Presenting as Part-Solid Nodules: A Multicenter Retrospective Study
Supplemental material, sj-docx-1-tct-10.1177_15330338251351365 for Ensemble Machine Learning Classifiers Combining CT Radiomics and Clinical-Radiological Features for Preoperative Prediction of Pathological Invasiveness in Lung Adenocarcinoma Presenting as Part-Solid Nodules: A Multicenter Retrospective Study by Yunhua Li, Jianbang Ding, Kun Wu, Wanyin Qi, Shanyue Lin, Gangwen Chen and Zhichao Zuo in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-2-tct-10.1177_15330338251351365 - Supplemental material for Ensemble Machine Learning Classifiers Combining CT Radiomics and Clinical-Radiological Features for Preoperative Prediction of Pathological Invasiveness in Lung Adenocarcinoma Presenting as Part-Solid Nodules: A Multicenter Retrospective Study
Supplemental material, sj-docx-2-tct-10.1177_15330338251351365 for Ensemble Machine Learning Classifiers Combining CT Radiomics and Clinical-Radiological Features for Preoperative Prediction of Pathological Invasiveness in Lung Adenocarcinoma Presenting as Part-Solid Nodules: A Multicenter Retrospective Study by Yunhua Li, Jianbang Ding, Kun Wu, Wanyin Qi, Shanyue Lin, Gangwen Chen and Zhichao Zuo in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-3-tct-10.1177_15330338251351365 - Supplemental material for Ensemble Machine Learning Classifiers Combining CT Radiomics and Clinical-Radiological Features for Preoperative Prediction of Pathological Invasiveness in Lung Adenocarcinoma Presenting as Part-Solid Nodules: A Multicenter Retrospective Study
Supplemental material, sj-docx-3-tct-10.1177_15330338251351365 for Ensemble Machine Learning Classifiers Combining CT Radiomics and Clinical-Radiological Features for Preoperative Prediction of Pathological Invasiveness in Lung Adenocarcinoma Presenting as Part-Solid Nodules: A Multicenter Retrospective Study by Yunhua Li, Jianbang Ding, Kun Wu, Wanyin Qi, Shanyue Lin, Gangwen Chen and Zhichao Zuo in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-4-tct-10.1177_15330338251351365 - Supplemental material for Ensemble Machine Learning Classifiers Combining CT Radiomics and Clinical-Radiological Features for Preoperative Prediction of Pathological Invasiveness in Lung Adenocarcinoma Presenting as Part-Solid Nodules: A Multicenter Retrospective Study
Supplemental material, sj-docx-4-tct-10.1177_15330338251351365 for Ensemble Machine Learning Classifiers Combining CT Radiomics and Clinical-Radiological Features for Preoperative Prediction of Pathological Invasiveness in Lung Adenocarcinoma Presenting as Part-Solid Nodules: A Multicenter Retrospective Study by Yunhua Li, Jianbang Ding, Kun Wu, Wanyin Qi, Shanyue Lin, Gangwen Chen and Zhichao Zuo in Technology in Cancer Research & Treatment
Footnotes
Acknowledgments
We would like to extend our sincere appreciation to Editage for their meticulous language editing services, which have significantly enhanced the quality and readability of this manuscript.
Compliance with Ethical Requirements
The institutional review boards of all participating institutions approved the study (2021-07-009 from Xiangtan Central Hospital, KY2020147 from the Affiliated Hospital of Southwest Medical University, and 2023YJSLL-121 from the Affiliated Hospital of Guilin Medical University).
Informed Consent Statement
The study protocol was approved by the institutional review board and due to the retrospective nature of the study, the informed consent requirement was waived.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data for this study are available by contacting the corresponding author upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
