Abstract
Introduction
Lung adenocarcinoma (LUAD), the most prevalent histological subtype of non-small cell lung cancer (NSCLC), remains a major contributor to cancer-related mortality worldwide. 1 Despite significant advancements in diagnostic and therapeutic strategies, including targeted therapies and immunotherapies, the five-year survival rate for individuals with advanced-stage lung adenocarcinoma remains alarmingly low. 2 The progression and metastasis of tumors, driven by intricate interactions between cancer cells and their surrounding microenvironment, pose significant challenges to improving patient outcomes. 3 The poor clinical outcomes are largely attributed to tumor heterogeneity, therapeutic resistance, and the aggressive nature of LUAD characterized by rapid progression and high metastatic potential. 4
A growing body of evidence suggests that tumor progression is not solely governed by cancer-intrinsic factors but is critically influenced by the tumor microenvironment (TME). 5 Among the key components of the TME, the extracellular matrix (ECM) has emerged as a dynamic structure that not only provides mechanical support but also actively regulates tumor behavior, including proliferation, migration, invasion, and response to therapy.6,7 Dysregulated ECM remodeling, driven by matrix metalloproteinases (MMPs), fibroblasts, and cytokine signaling, contributes to a pro-tumorigenic niche and promotes EMT, a critical step in cancer metastasis. 8 Therefore, targeting the ECM and its regulators has garnered increasing attention as a potential strategy to inhibit LUAD progression and improve treatment efficacy.
Neuregulin 4 (NRG4), 9 a lesser-studied member of the epidermal growth factor (EGF) family, is primarily known for its physiological role in metabolic homeostasis,10,11 NRG4 binds to ErbB4, a member of the ErbB receptor tyrosine kinase family, and has been implicated in regulating glucose metabolism, thermogenesis, and lipid homeostasis in adipose tissue and liver.12,13 In the context of cancer, most research on neuregulins has focused on NRG1 and NRG2 due to their well-established roles in ErbB receptor activation and tumorigenesis. 14 In contrast, the biological function of NRG4 in cancer remains largely unexplored. Emerging studies have suggested that NRG4 may exert tumor-suppressive effects in certain malignancies such as hepatocellular carcinoma and prostate cancer, potentially through modulation of cellular proliferation and inflammatory signaling.15,16 However, its relevance in LUAD has not yet been systematically investigated.
In this study, we sought to elucidate the role of NRG4 in the progression of lung adenocarcinoma. Through integrative analyses of LUAD patient datasets and in vitro/in vivo experiments, we found that NRG4 expression is significantly reduced in LUAD tissues and is inversely correlated with the expression of key genes involved in ECM remodeling and EMT. Functional assays demonstrated that treatment with recombinant NRG4 (rNRG4) suppresses LUAD cell proliferation, migration, and impairs tumor growth and metastasis. Transcriptomic profiling further revealed that rNRG4 attenuates ECM-related gene expression, highlighting a previously unrecognized mechanism by which NRG4 may exert anti-tumor activity. Notably, high NRG4 expression was associated with favorable clinical outcomes in LUAD patients, supporting its potential as a prognostic biomarker and therapeutic target. Collectively, these findings provide novel insights into the tumor-suppressive role of NRG4 in LUAD and underscore its therapeutic relevance in disrupting ECM-mediated tumor progression.
Materials and Methods
Cell Culture and Reagents
Human lung adenocarcinoma cell lines A549 (ATCC:CCL-185; RRID:CVCL_0023) and HCC827 (ATCC:CRL-2868; RRID:CVCL_2063) were cultured in 1640 and mouse lung adenocarcinoma cell LLC (ATCC:CRL-1642; RRID:CVCL_5653) were cultured in DMEM, all the cell medium were supplemented with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin at 37 °C in a humidified incubator with 5% CO₂. rNRG4 was purchased from Sinobiological and prepared according to the supplier's instructions.
Cell Proliferation Assay
Cell proliferation was assessed using the Enhanced Cell Counting Kit-8 (CCK8, Beyotime) according to the manufacturer's instructions. Cells were seeded in 96-well plates and treated with various concentrations of rNRG4 (0, 25, 50, 100 ng/mL). At 24 h, CCK-8 solution was added, and absorbance at 450 nm was measured.
Cell Migration Assays
Cell migration was analyzed using transwell assays and performed according to previous reports. 17 2 × or 5 × 10^5 cells were seeded in the upper chamber of transwell inserts and treated with rNRG4. After 16 h, migrated or invaded cells were fixed, stained, and counted under a microscope.
Quantitative Real-Time PCR (qPCR)
Total RNA was extracted from cells/tissues using TRIzol RNA isolation kit (Takara) following the manufacturer's instructions. 18 RNA was reverse-transcribed into complementary DNA (cDNA) using HiScriptTM II Q RT SuperMix for qPCR (+gDNA wiper) (Vazyme). Quantitative PCR was performed using ChamQ Blue Universal SYBR qPCR Master Mix (Vazyme) on an ABI QuantStudio. The primers used for target genes are listed in Supplementary Table S1. Relative gene expression was calculated using the 2^−ΔΔCt method, with Gapdh as the internal control.
Hematoxylin and Eosin (HE) Staining and Immunohistochemistry (IHC)
Tissues were fixed in 10% formalin for 12-24 h, embedded in paraffin, and sectioned. Paraffin-embedded tissue sections were deparaffinized, rehydrated, stained with HE (Beyotime) according to the manufacturer's instructions.For IHC assay, sections were incubated with a primary antibody against MMP9 and E-cadherin at 4 °C overnight. After washing, sections were incubated with a biotinylated secondary antibody and streptavidin-HRP, followed by DAB substrate for visualization. Nuclei were counterstained with hematoxylin. Images were obtained using an optical microscope (Nikon).
Immunofluorescence (IF)
The cells were fixed in 10% formalin, followed by blocking and staining in antibody diluent containing 10% goat serum. Subsequently, the cells were incubated overnight at 4 °C with mouse anti-Vimentin antibodies. Afterward, the cells were treated with the corresponding secondary antibodies. Finally, the cells were mounted with 4′,6-diamidino-2-phenylindole (DAPI) and imaged using fluorescence microscopy.
Immunoblotting
Cells were lysed in RIPA lysis buffer (Beyotime) supplemented with PMSF and a protease inhibitor cocktail (Beyotime). Total protein extracts were denatured by boiling in Laemmli sample buffer containing SDS. Denatured proteins were resolved using SDS-PAGE and subsequently transferred onto nitrocellulose (NC) membranes via wet electroblotting. Membranes were blocked with 5% (w/v) non-fat milk dissolved in TBST for 1 h at room temperature, followed by three 5-min TBST washes. Membranes were probed with primary antibodies (listed in Supplementary Table S2) overnight at 4 °C with gentle agitation. Following primary antibody incubation, membranes were washed three times with TBST (5 min per wash) and incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies for 1.5 h at room temperature with gentle agitation. Membranes were then washed three times in TBST (5 min per wash). Protein bands were visualized using an enhanced chemiluminescence (ECL) substrate kit (Beyotime) using a Servicebio Chemiluminescence Imager (SCG-W3000) with exposure times optimized between 10 s to 5 min.
RNA Sequencing and Bioinformatics Analysis
RNAseq was performed in shanghai applied protein technology. Total RNA from rNRG4-treated samples and primary tumors was extracted using the TRIzol reagent. RNAseq was conducted by Applied Protein Technology, utilizing the NovaSeq platform for data processing. Differentially expressed genes (DEGs) were identified, and pathway enrichment analysis was performed to evaluate the effects of rNRG4 treatment. DEG analysis was carried out with a significance threshold of P < .05 and |fold change| ≥ 2. For KEGG enrichment analysis, the clusterProfiler package (v3.12.0) was used, with a cutoff of P < .05 to identify significantly enriched pathways.
Lung Cancer Model
Twelve-week-old C57BL/6 mice were subcutaneously injected with 5 × 10⁵ LLC cells to establish lung cancer models. Following tumor formation, the mice received paratumoral injections of 200 ng/tumor/day rNRG4 or vehicle (PBS) control. 12 Tumor volume was measured weekly, and metastatic nodules in the lungs were quantified after four weeks. The reporting of this study conforms to the ARRIVE 2.0 guidelines. 19 All procedures involving animals were conducted in accordance with the Guide for the Care and Use of Laboratory Animals, eighth Edition. 20
Online Data Mining
Clinical data of Nrg4 expression analysis for lung patients were downloaded from the Kaplan–Meier Plotter (https://kmplot.com/analysis). In the analysis process, gene expression data and overall survival information are downloaded from TCGA and the GTEx projects, using a standard processing pipeline. The database is handled by a PostgreSQL server.
Statistical Analysis
Data are presented as mean ± SEM. Statistical comparisons were performed using Student's t-test or one-way ANOVA. Kaplan-Meier survival analysis was used for survival data. The statistically significant was tagged in each figure.
Results
NRG4 is Closely Associated with the Epithelial-Mesenchymal Transition (EMT) in Lung Adenocarcinoma
To explore the relationship between NRG4 and EMT in lung adenocarcinoma, we analyzed TCGA database correlations between NRG4 expression and EMT-related genes. Our analysis revealed that NRG4 is negatively correlated with the EMT inducer TGF-B1 (Figure 1A) and mesenchymal-specific markers Vimentin (VIM) and Fibronectin (FN1) (Figure 1B, C). Conversely, NRG4 showed positive correlations with tight junction-associated genes (TJP1, TJP2, TJP3) (Figure 1D-F). These results suggest that NRG4 plays a role in suppressing EMT progression.

NRG4 is Closely Associated with the Epithelial-Mesenchymal Transition (EMT) in Lung Adenocarcinoma. (A-F) TCGA Data Revealed the Corelation Between NRG4 and TGF-B1 (A), FN1 (B), VIM (C), TJP1 (D), TJP2 (E), TJP3 (F). Significance Defined as P < .05.
rNRG4 Inhibits Lung Adenocarcinoma Cell Migration and EMT in Vitro
To evaluate the effect of rNRG4 on cell migration, transwell assays demonstrated that rNRG4 significantly reduced the migratory ability of lung adenocarcinoma cells (Figure 2A, B). Further analysis showed that rNRG4 inhibited mesenchymal-specific genes (Cdh2, Vimentin, Fibronectin) while upregulating epithelial-specific genes (Cdh1, Zo-1) (Figure 2C-E). At the protein level, rNRG4 also suppressed mesenchymal-specific proteins (CDH2, Vimentin, Fibronectin) while upregulating epithelial-specific proteins (CDH1, ZO-1) (Figure 2F-K), confirming its inhibitory effect on EMT and migration.
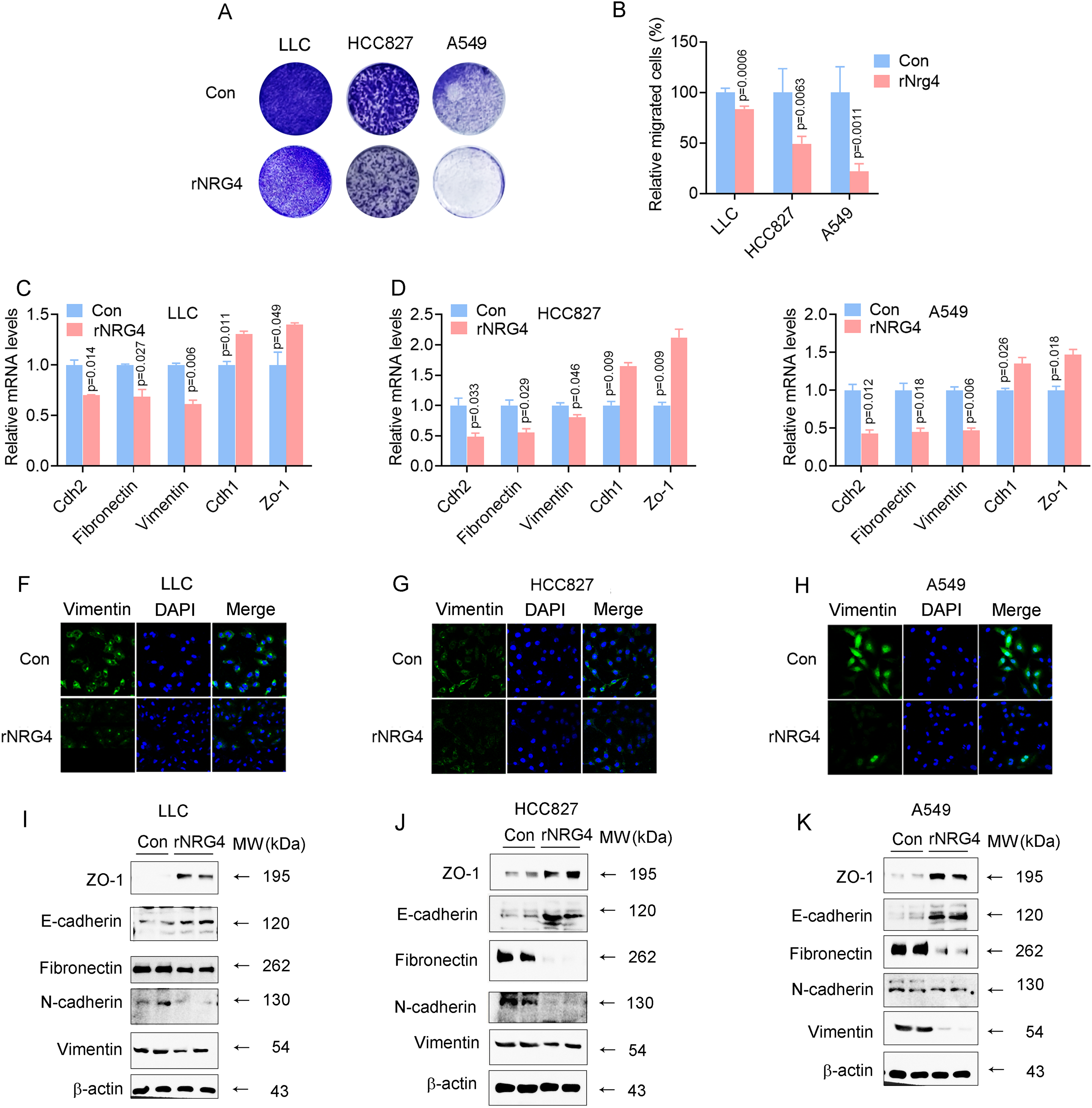
rNRG4 Inhibits Lung Adenocarcinoma Cell Migration and EMT in Vitro. (A) Representative Images from Transwell Assays show Migration of 2 × 105 LLC Cells, 5 × 105 HCC827 Cells, and 2 × 105 A549 Cells After 16 h of culture. (B) Quantification of Migrated Cells Per Field. (C-E) qRT-PCR Analysis of Mesenchymal Markers (Cdh2, Vimentin, Fibronectin) and Epithelial Markers (Cdh1, Zo-1) in LLC, HCC827, and A549 Cells Treated with 100 ng/mL rNRG4 for 24 h. (F-H) Representative Immunofluorescence Images Showing Vimentin Expression in LLC (F), HCC827 (G), and A549 (H) Cells Treated with 100 ng/mL rNRG4 for 24 h. (I-K) Immunobloting Images Showing the Expression of EMT-Related Protein Markers in LLC (I), HCC827 (J), and A549 (K) Cells Treated with 100 ng/mL rNRG4 for 48 h. Three Independent Experiments were Performed in Each Case. Data were Presented as mean ± SEM, Compared to the Control (Con, PBS) Group, with Significance Defined as P < .05.
rNRG4 Inhibits Cell Proliferation
Given the importance of cell proliferation in lung cancer progression, we assessed the effect of rNRG4 on the proliferation of lung adenocarcinoma cell lines. rNRG4 treatment significantly inhibited proliferation of LLC, HCC827, and A549 cells in a dose-dependent manner (P < .01), with the strongest effect observed at 100 ng/mL(Figure 3A-C). rNRG4 at a concentration of 100 ng/mL significantly suppresses the expression of proliferation-associated markers Ki67 and Pcna at both the mRNA level (Figure 3D-F) and protein level (Figure 3G-1). These findings highlight rNRG4 as a potent inhibitor of cell proliferation.

rNRG4 Inhibits Cell Proliferation. (A-C) CCK8 Assay of LLC (A), HCC827 (B), and A549 (C) Cells Treated with Different Doses of rNRG4 for 24 h. D-E. qRT-PCR Analysis of Cell Proliferative Genes (Ki67 and Pcna) of LLC (D), HCC827 (E), and A549 (F) Cells Treated with 100 ng/mL rNRG4 for 24 h. (G-I) Immunobloting Images Showing the Expression of Cell Proliferation and Apoptosis-Related Proteins in LLC (G), HCC827 (H), and A549 (I) Cells Treated with 100 ng/mL rNRG4 for 48 h. Three Independent Experiments Were Performed in Each Case. Data Were Presented as Mean ± SEM, Compared to the Control (Con, PBS) Group, with Significance Defined as P < .05.
rNRG4 Inhibits Lung Adenocarcinoma Progression
To evaluate the therapeutic potential of rNRG4, we treated a lung adenocarcinoma mouse model with rNRG4 (200 ng/tumor/day) for 20 days. rNRG4 significantly reduced primary tumor growth, as evidenced by decreased tumor volume and weight (Figure 4A-D). Moreover, rNRG4 effectively suppressed lung cancer metastasis (Figure 4E, F), demonstrating its robust anti-tumor effects.

rNRG4 Inhibits Lung Adenocarcinoma Progression. (A) The Mouse Model of rNRG4 Treated LLC Bearing Tumors. (B) The Tumor Volume of LLC Tumors in the Presence of rNRG4. (C.D) The Morphology (C) and Tumor Weight (D) of Primary Tumors. (E, F) The Morphology (E) and Lung Metastases (F) of the rNRG4 Treated LLC Bearing Mice. Data were Presented as Mean ± SEM, Compared to the Control (Con, PBS) Group, with Significance Defined as P < .05.
rNRG4 Partially Inhibits EMT in Vivo
Building on our previous findings that rNRG4 inhibits EMT and cell migration in vitro, we assessed its effect on EMT markers in vivo. Protein analysis revealed that rNRG4 significantly increased E-cadherin expression in both primary tumors (Figure 5A) and lung tissues (Figure 5B). Further analysis demonstrated that rNRG4 inhibited EMT in primary tumors (Figure 5C) and lung tissues (Figure 5D). These results suggest that rNRG4 effectively inhibits EMT in vivo.

rNRG4 Partially Inhibits EMT in Vivo. (A-D) the Representive Images of HE Staining and IHC of MMP9 and E-Cadherin of Primary Tumors (A) or Lung Metastasis (B). qRT-PCR Analysis of Mesenchymal Markers (Cdh2, Vimentin, Fibronectin) and Epithelial Markers (Cdh1, Zo-1) in Primary Tumors or Lung Tissues from Mice Treated with rNRG4. Data Were Presented as Mean ± SEM, Compared to the Control (Con, PBS) Group, with Significance Defined as P < .05.
rNRG4 Inhibits Extracellular Matrix Organization and Component
To further explore the functional mechanism of rNRG4 in cancer progression, we performed RNAseq analysis on primary tumors. The results revealed that rNRG4 regulated numerous immune-related events in the primary tumors (Figure 6A). Beyond immune-related genes, ECM organization and collagen-containing ECM emerged as key pathways influenced by rNRG4 in inhibiting cancer progression (Figure 6B). To validate the role of ECM remodeling, we analyzed the correlation between NRG4 and MMPs, finding that NRG4 was negatively correlated with MMP2, MMP9, and MMP12 (Figure 6C-H). These findings suggest that rNRG4 inhibits extracellular matrix organization and its components, contributing to its anti-cancer effects.

rNRG4 Inhibits Extracellular Matrix Organization And Components. (A) GO and KEGG of all DEG from RNAseq Data of Primary Tumors Treated with rNRG4. (B) GO and KEGG of DEG, Besides Immune Related Genes, from the Above RNAseq Data. (C-E) TCGA Data Revealed the Corelation Between NRG4 and MMP2(C), MMP9(D), MMP12(E). (F-H) Immunobloting Images Showing the Expression of MMPs in LLC (F), HCC827 (G), and A549 (H) Cells Treated with 100 ng/mL rNRG4 for 48 h. Two Independent Experiments were Performed in Each Case. Data were Presented as mean ± SEM, Compared to the Control (Con, PBS) Group, with Significance Defined as P < .05.
NRG4 is Positively Correlated with Overall Survival of Lung Adenocarcinoma Patient
We analyzed the relationship between NRG4 expression and overall survival in lung adenocarcinoma patients and found that higher NRG4 levels were positively correlated with improved overall survival (Figure 7A). Together, these findings demonstrate that NRG4, particularly in its recombinant form, inhibits lung adenocarcinoma progression by targeting key processes such as cell proliferation, migration, EMT, and ECM remodeling. Its positive association with patient survival further highlights its potential as a therapeutic target for lung adenocarcinoma (Figure 7B).

NRG4 is Positively Correlated with Overall Survival of Lung Adenocarcinoma Patient. (A) TCGA Data Demonstrate a Significant Correlation Between NRG4 Expression and Overall Survival (Auto select best cutoff) in Lung Adenocarcinoma. (B) Graphical Abstract Illustrating the Role of NRG4 in Lung Adenocarcinoma Progression, Highlighting its Inhibitory Effects on Cell Proliferation, Cell Migration, and ECM Components.
Discussion
In this study, we provide compelling evidence that NRG4 serves as a multifaceted suppressor of lung adenocarcinoma progression. Through a combination of transcriptomic analysis, functional assays, and in vivo xenograft models, we demonstrate that NRG4 inhibits key oncogenic processes, including cell proliferation, migration, EMT, and ECM remodeling. These findings not only uncover a novel tumor-suppressive function for NRG4 in LUAD, but also support its potential as a candidate for further investigation in combating this aggressive malignancy.
One of the most notable findings of our study is the inverse correlation between NRG4 expression and mesenchymal markers such as Vimentin and Fibronectin, both of which are hallmarks of EMT and poor prognosis in LUAD patients.21-23 EMT is a critical driver of cancer progression, enabling epithelial tumor cells to acquire mesenchymal features, increasing their migratory and invasive capabilities, and promoting metastasis.24-26 Our data reveal that NRG4 not only downregulates mesenchymal markers, but concurrently upregulates epithelial markers such as E-cadherin. This dual modulation suggests that NRG4 may function as a gatekeeper of the epithelial phenotype, thereby impeding EMT-driven dissemination. These observations are consistent with recent studies showing that modulation of EMT status can profoundly influence therapeutic resistance and immune evasion,27-29 further underscoring the clinical relevance of targeting this process.
In addition to its impact on EMT, NRG4 significantly modulates ECM dynamics. The ECM is not merely a structural scaffold; it is a biologically active component of the tumor microenvironment that governs a range of cancer cell behaviors, including proliferation, migration, and resistance to therapy.30,31 The extracellular matrix is a vital component of the tumor microenvironment, 22 providing structural support 23 and influencing cell behavior. 32 Our transcriptome data and subsequent validation experiments demonstrate that NRG4 negatively regulates several ECM-associated genes, particularly MMP2, MMP9, and MMP12, which are known mediators of ECM degradation and metastatic dissemination.33-35 This suppressive effect on ECM remodeling implies that NRG4 may also modulate tumor-stromal interactions, a concept increasingly recognized as pivotal in cancer biology. 36 By altering the biophysical and biochemical properties of the tumor niche, NRG4 may limit cancer cell escape from the primary site and inhibit secondary site colonization.
The therapeutic implications of our findings are further strengthened by the use of rNRG4 in both in vitro and in vivo settings. rNRG4 consistently inhibited tumor growth, reduced metastatic burden, and restored epithelial architecture, thereby confirming its tumor-suppressive capabilities. Notably, survival analysis based on publicly available datasets revealed a positive correlation between high NRG4 expression and overall survival in LUAD patients, suggesting potential utility of NRG4 not only as a therapeutic agent but also as a prognostic biomarker.
Nevertheless, there are limitations in the current study that warrant consideration. Our in vivo experiments were conducted using a subcutaneous xenograft model, which does not fully recapitulate the native lung microenvironment or metastatic routes of LUAD. 37 While this model offers a tractable system for rapid evaluation of tumor growth and molecular modulation, it lacks the spatial and mechanical cues inherent to the orthotopic lung milieu. Orthotopic models, involving direct implantation of tumor cells into the lung parenchyma, offer superior physiological relevance are critical for studying metastatic spread via natural anatomical routes such as the vasculature or pleural cavity. 38 Although logistical and ethical constraints precluded the use of such models in the current study, future investigations will prioritize the adoption of orthotopic systems to validate the clinical translatability of our findings.
Another area for future exploration is the identification of signaling pathways downstream of NRG4. Given that NRG4 primarily signals through the ErbB4 receptor,12,13 it is conceivable that it modulates key downstream effectors such as PI3K/AKT, STAT5, or ERK/MAPK pathways, all of which are implicated in cell survival, migration, and differentiation.39,40 However, whether these pathways are involved in the anti-EMT and anti-ECM effects observed in LUAD remains to be determined. Moreover, our RNA-seq data revealed potential alterations in immune-related pathways following rNRG4 treatment. This raises the intriguing possibility that NRG4 may also exert immunomodulatory functions within the tumor microenvironment, either by directly influencing immune cell recruitment or by reshaping the ECM in a way that facilitates anti-tumor immunity. Considering the increasing success of immunotherapies in lung cancer, combinatorial strategies involving NRG4 and immune checkpoint inhibitors may offer enhanced therapeutic efficacy.
Lastly, the translational potential of NRG4 warrants investigation in the context of combination therapy. As monotherapies often fail to produce durable responses in advanced-stage LUAD due to compensatory signaling and heterogeneity, combining NRG4 with conventional chemotherapy, tyrosine kinase inhibitors, or immunotherapeutic agents may synergize to achieve greater clinical benefit. Future preclinical and clinical studies should aim to define the optimal therapeutic windows and dosing strategies for such combinations.
Conclusion
In conclusion, our study identifies NRG4 as a critical suppressor of lung adenocarcinoma progression via coordinated inhibition of EMT, ECM remodeling, and cellular proliferation and migration. These findings expand our understanding of NRG4 beyond its classical metabolic functions and establish a foundation for its future development as a diagnostic biomarker and therapeutic agent. By targeting fundamental processes that drive LUAD aggressiveness, NRG4 represents a promising candidate for the next generation of anti-cancer therapies.
Supplemental Material
sj-docx-1-tct-10.1177_15330338251344424 - Supplemental material for Neuregulin 4: A Key Regulator in Suppressing Lung Adenocarcinoma Progression
Supplemental material, sj-docx-1-tct-10.1177_15330338251344424 for Neuregulin 4: A Key Regulator in Suppressing Lung Adenocarcinoma Progression by Shufan Zhang, BMed, Tianhan Xu, BMed, Simeng Li, BMed, Liming Tang, MD, and Dongmei Wang, PhD in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-2-tct-10.1177_15330338251344424 - Supplemental material for Neuregulin 4: A Key Regulator in Suppressing Lung Adenocarcinoma Progression
Supplemental material, sj-docx-2-tct-10.1177_15330338251344424 for Neuregulin 4: A Key Regulator in Suppressing Lung Adenocarcinoma Progression by Shufan Zhang, BMed, Tianhan Xu, BMed, Simeng Li, BMed, Liming Tang, MD, and Dongmei Wang, PhD in Technology in Cancer Research & Treatment
Footnotes
Acknowledgments
We extend our gratitude to the Large Instrument Platform of Changzhou Medical Center for providing instrumental and technical support.
Ethical Considerations
Animal experiments were approved by the Ethics Committee of Animal Experiments of Nanjing Medical University (Approval No. IACUC-2308061, 2501040).
Author Contributions
Conceptualization, Dongmei Wang; Data curation, Shufan Zhang, Tianhan Xu, Simeng Li; Funding acquisition, Dongmei Wang, Liming Tang; Investigation, Dongmei Wang, Shufan Zhang; Methodology, Shufan Zhang, Tianhan Xu; Project administration, Shufan Zhang, Simeng Li; Resources, Dongmei Wang; Writing -original draft, Shufan Zhang, Dongmei Wang; Writing -review & editing, Shufan Zhang, Simeng Li, Tianhan Xu, Dongmei Wang.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from National Natural Science Foundation of China (82473051, 82273232), Changzhou Sci&Tech Program (LCQYBS202309, 2024CZBJ020, LC2024LCQYBS202309), Changzhou Medical Center of Nanjing Medical University Program (CZKY102RC202301, CMC2024PY06, CMCM202403).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Derived raw data supporting the results of this research can be obtained from corresponding author on request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
