Abstract
Introduction
Lung cancer is one of the leading causes of cancer-related mortality among both men and women worldwide, 1 which is classified as 2 main types: small cell lung cancer and nonsmall cell lung cancer (NSCLC), accounting for about 15% and 85%, respectively. Among NSCLC, lung adenocarcinoma (LUAD) is the most common subtype accounting for more than 50% and has an increasing frequency. 2 Over the past decades, the discoveries of oncogenic driver genes and mutations for lung cancer allows the development of personalized targeted therapy, especially EGFR-TKIs. Gefitinib has been widely used as front-line EGFR-TKIs therapy in LUAD patients harboring sensitive EGFR mutations and confers favorable disease control. However, acquired resistance is eventually inevitable after an average of 9 to 11 months of gefitinib use, and the underlying mechanism remains a biological challenge. 3 Although it is believed that second mutations of EGFR, genetic amplifications or mutations of other driver genes, and histological transformation involving in EGFR-TKIs resistance may serve as predictive factors, most of these signatures need tumor biopsy for detection which largely limits its clinical practice. Therefore, the discovery of predictive biomarkers for acquired resistance to EGFR-TKIs resistance by different molecular signatures on different types of biopsies is of great clinical significance. The circulating RNAs, such as miRNAs, long noncoding RNAs (lncRNAs), and circRNAs, detected in plasma are emerging as a novel class of candidate biomarkers for molecular monitoring during targeted therapy.
Circular RNAs (circRNAs) are a newly defined class of noncoding RNAs originated from back-splicing events. 4 Based on its closed circular structure, circRNA specifically exhibits extraordinary stability, due to the lack of accessible ends exposed to nucleolytic degradation. 4 Recently, a line of studies has shown that circRNAs contain target sites for miRNAs and act as competitive endogenous RNA (ceRNA) to regulate miRNAs and other noncoding RNAs-related signaling axis, thus exerting various biological functions in cancer development. 5 With the development of circRNAs microarray and high-throughput second-generation sequencing, the identification of circRNAs expressed in various cancers is gradually coming to light. Studies have shown that circRNAs play an indispensable role in the occurrence, development, metastasis, and drug resistance of cancer. For example, hsa_circ_0004015 is reported to regulate the proliferation, invasion, and TKI resistance of NSCLC 6 ; Hsa_circ_0007798 was shown to enhance the gefitinib sensitivity of LUAD cells via activating ASK1-dependent apoptosis. 7 In supporting of its intrinsic features and based on these research evidences, circRNA is emerging as a novel potential biomarker for cancer prognosis and treatment.
In this present study, we did a comprehensive analysis of circRNAs expression profiles in gefitinib-treated lung adenocarcinoma patients, and the analysis strategy of this study is shown in a flow chart (Figure 1). A total of 8 peripheral plasma samples from 2 groups of LUAD patients (4 gefitinib-sensitive and 4 gefitinib-resistant) harboring EGFR-sensitive mutations (L858R or 19Del) were collected to perform circRNAs microarray assay, The results showed that there were 2571 differentially expressed circRNAs (DECs) between 2 groups. Based on the bioinformatics analysis for these DECs involving in diverse biological processes, we constructed a circRNA-miRNA network including 10 circRNAs and their corresponding miRNAs. And we validated that hsa_circ_0030591 and hsa_circ_0040348 were indeed upregulated in gefitinib-resistant patients by qRT-PCR. Our study provides data on plasma circRNAs expression profiles for NSCLC patients with acquired gefitinib resistance, and we anticipate that hsa_circ_0030591 and hsa_circ_0040348 may serve as a potential biomarker for NSCLC patients treated with gefitinib.

Flow chart of this study design.
Methods
Patients and Samples Collection
A total of 16 LUAD patients with EGFR Exon 19 deletion or Exon 21 L858R mutations treated with gefitinib in Ningbo Hwamei Hospital of the University of Chinese Academy of Sciences between March 2017 and May 2020 were included in this study. In the beginning, each patient is sensitive to gefitinib. After 5 to 24 months of treatment, 8 patients remained sensitive to gefitinib and were designated as the gefitinib-sensitive group, while 8 patients developed resistance and were designated as the gefitinib-resistant group. 8 patients’ peripheral plasma samples (4 gefitinib-resistant and 4 gefitinib-sensitive patients) were used for microarray assay, and all 16 patients’ plasma samples were used for validation by RT-PCR. 2.5 mL of peripheral plasma was collected from each patient and was stored in the PAXgene plasma RNA tubes (Becton, Dickinson and Company, American). The plasma samples were collected at a room temperature of 18–25°C and stored in −80°C refrigerator. This study was approved by the Medical Ethics Committee of Ningbo Hwamei Hospital, and all patients signed the informed consent before they were enrolled.
RNA Extraction, Amplification and Labeling
We used Trizol reagent (Invitrogen) to extract total RNA containing small RNA from whole plasma samples and purified it with miVana miRNA isolation kit (Ambion, Austin, TX, USA) according to manufacturer's protocol. The OD260/280 reading was measured using a spectrophotometer (Nanodrop ND-1000) to determine the purity and concentration of RNA. RNA integrity was detected by 1% formaldehyde denaturing gel electrophoresis. RNA was digested, amplified, and labeled by the cRNA Amplification and Labeling Kit (CapitalBio, Beijing, China) according to the manufacturer's product instruction.
CircRNA Microarray Assay
The purified RNA was hybridized with the CapitalBio Technology Human CircRNA Array v2 which was designed with 4 identical arrays per slide (4 × 180 K format) and each array containing probes interrogating about 170,340 human circRNAs.
The circRNA array data were analyzed by the GeneSpring software V13.0 (Agilent) and R software package (R version 3.1.2). Box plot was used to visualize the distribution of the intensities of all circRNAs in the samples before and after normalization. Principal component analysis (PCA) of circRNA expression profiles was completed by the Stats package. Scatter plot was used to assess the differences in circRNA expression between the 2 groups. Volcano plot was used to display the DECs. Heatmap was used to analyze the DECs in different samples completed by pheatmap package. Threshold values of fold change (FC) ≥2 or ≤0.5 with P < 0.05 were considered significantly differentially expressed.
Construction of the circRNA-miRNA Network
The miRanda-3.3 software was used to select the DECs and predict their corresponding target miRNA. Narrow down by fold change > 1.5, P < 0.05 and each original signal value > 100, 5 circRNAs associated with the ATP-binding cassette (ABC) transporter pathway, and 5 circRNAs associated with the PI3K-Akt signaling pathway were selected togethering with their predictive target miRNAs to construct the circRNA-miRNA RNA network.
Validation of circRNAs Expression by qRT-PCR
Quantitative real-time PCR (qRT-PCR) was performed by SYBR-Green Premix Ex Taq Kit (Takara Bio, Japan), and the remix was applied by ABI PRISM 7500 sequence detection system (Life Technologies, USA) at 95 °C for 5 min and amplified by 40 cycles of denaturing at 95 °C for 15 s, 60 °C for 20 s, and 72 °C for 40 s. The CT value was the fractional cycle number at which the fluorescence exceeded the given threshold. GAPDH was used to normalize the RNA preparation. The relative expression levels of circRNAs were calculated using 2−△△Ct method. The primers were designed using Primer 6.0. The primers sequences of hsa_circ_0030591 were as follows: 5′-GTGCTCACTGGATTGTCTTCAT-3′ (forward) and 5′-GACTTTTCCCAGGCGTACATTT-3′ (reverse); The primers sequences of hsa_circ_0040348 were: 5′-CATGTGCAAAACCTTCCAACAC-3′ (forward) and 5′-TAACACGAGAGATCCACGGTAC-3′ (reverse). The primers sequences of GAPDH were: 5′-CAAATTCCATGGCACCGTCA-3′(forward) and 5′-AGCATCGCCCCACTTGATTT-3′ (reverse).
Statistical Analysis
SPSS v19.0 (SPSS, Inc., Chicago, IL, USA) was used for all statistical analyses, and data were expressed as the mean ± SD. Student's two-tailed unpaired t-test was used to compare the circRNAs expression levels between 2 groups. P < 0.05 was considered statistically significant.
Results
Expression Profiles of circRNAs in Gefitinib-Sensitive and Gefitinib-Resistant Groups
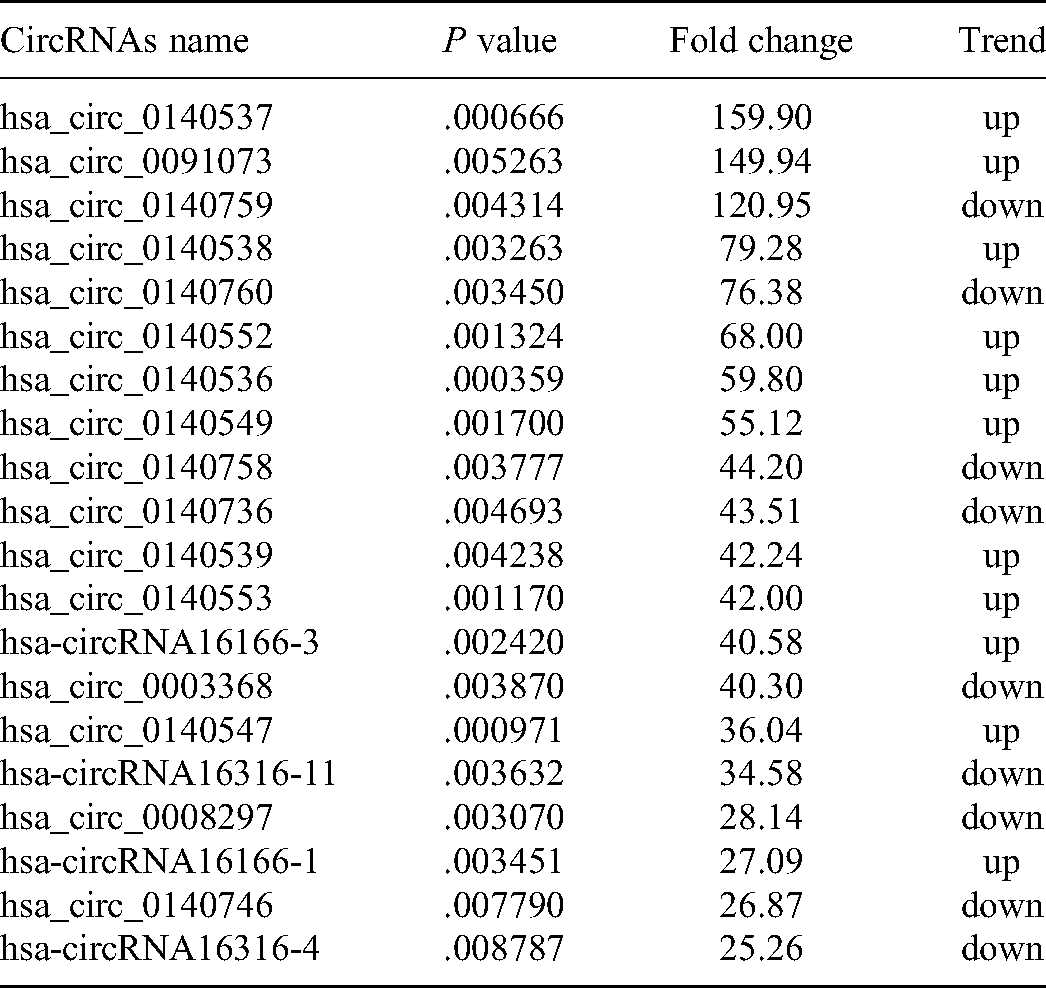
To explore the potential circRNAs related to EGFR-TKI resistance, the Human circRNA Array V2 chip detection was performed in 8 peripheral plasma samples from 4 gefitinib-sensitive and 4 gefitinib-resistant patients; the clinical features of all 8 patients are listed in Table 1. Quality control for microarray was performed on the raw and normalized data. The box plots showed the raw probe intensity and the normalized intensity after log2 RNA signal transformation (Figure 2A and B). PCA analysis showed a clear segregation between 2 groups, indicating distinguished circRNA expression patterns (Figures 2C and S1A). In total, 2571 circRNAs were identified to be differentially expressed between these 2 groups, including 1011 upregulated and 1560 downregulated circRNAs (P < 0.05; FC≥2 or ≤0.5), as shown in the scatter plots (Figure S1B) and the volcano plots (Figure 2D). The distribution of the parent genes of these candidate DECs was completed as circos plot, showing a wide distribution almost on all human chromosomes (Figure 3). Then the DECs were included in the hierarchical cluster heatmaps among gefitinib-sensitive and gefitinib-resistant groups and were also shown good discrimination between these 2 groups (Figure 4). The top 20 DECs with the rank of significant fold changes and their detailed information are listed in Table 2.

Expression profiles of circRNAs in gefitinib-sensitive and gefitinib-resistant groups. (A-B) Box plots are showing the distribution of circRNA expression in the 8 samples (red box: the sensitive group, blue box: the resistant group). (A) Is showing the raw probe signal, and (B) is showing the normalized probe signal. (C) PCA analysis (2D) of circRNA expression profiles. (D) The volcano plot. The abscissa value is −log10(P-values) and the ordinate value is log2(fold change). Both in (B) and (C), the upregulated genes are marked in red, the downregulated genes are marked in green, and the nonsignificantly different genes are marked in black.

The circos plots. The outer colored bands indicate different chromosomes, and the inner circles indicate the degree of difference of different genes in the chromosomal position. Red represents upregulated differential genes, green represents downregulated differential genes, and the length of the column represents the fold change of the differential genes.
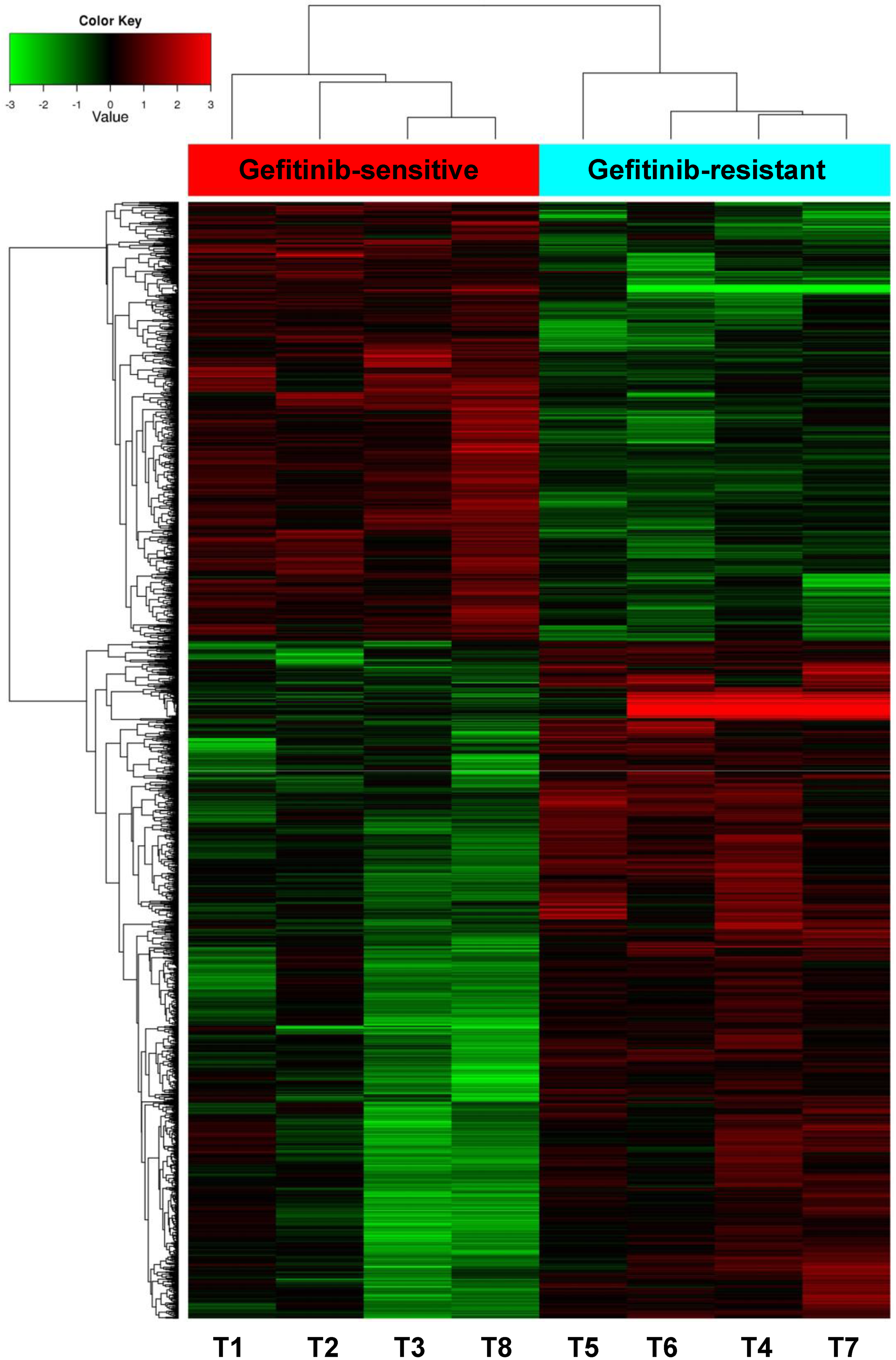
The heatmaps. Cluster analysis diagram of the sample. T1, T2, T3, and T8 are samples from patients in the sensitive group, while T4, T5, T6, and T7 are samples from patients in the resistant group.
Information of Patients Subjected to circRNAs Microarray Assay.
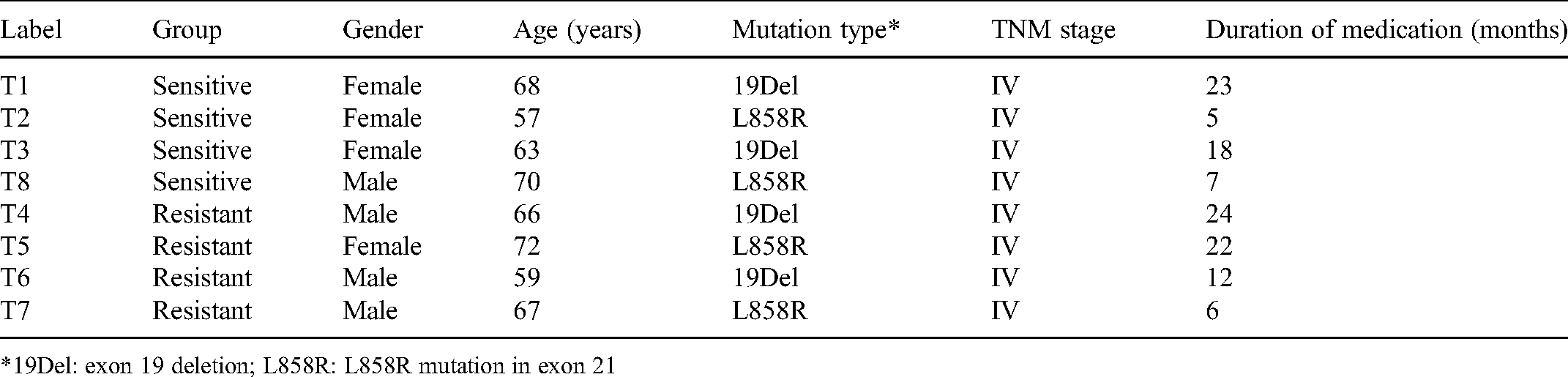

*19Del: exon 19 deletion; L858R: L858R mutation in exon 21
Top 20 Most Significant Aberrant Expressed circRNAs Based on Fold Change.
Functional Annotation of DECs
Next, we performed GO and KEGG pathway enrichment analyses to further explore the functions of these DECs. GO-enrichment items contained 6645 biological processes, 891 cellular components, and 1396 molecular functions, and the top 30 terms are shown in Figure 5A and B, and their hierarchies are shown in Figure S2A to C. The most enriched 3 GO terms of the biological process were chromosome segregation, cell cycle process, and single-organism organelle organization. KEGG pathway enrichment items included 100 BioCyc, 270 KEGG pathway, 91 Panther, and 1130 Reactome, and the top 30 terms are shown in Figure 5C and D, and the KEGG classification is shown in Figure S3. Specifically, DECs were most enriched in the ECM–receptor interaction, the cell cycle, and the Ribosome biogenesis pathways. More importantly, we found that ABC transporters (P = 0.01), which had been well established to involve in multidrug resistance,8,9 and PI3K-Akt signaling pathway (P = 0.02), which are associated with EGFR-TKI resistance,10,11 were both ranked in top 10 pathways enriched.

GO and KEGG pathway enrichment plot. (A-B) Statistics of GO-enrichment plot and bar graph (molecular function, biological process, and cellular component. (C-D) Statistics of pathway enrichment plot and bar graph. Enrichment items are obtained from databases of KEGG, Reactome, Panther, and BioCyc.
CircRNA-miRNA Networks
As mentioned earlier, ABC transporters and PI3K-Akt signaling were 2 enriched pathways related to drug resistance; we then narrowed down the top 5 circRNAs (hsa_circ_0030591, hsa_circ_0030574, hsa_circ_0030589, hsa_circ_0030598, and hsa_circ_0030600) associated with ABC transporters pathway and top 5 (hsa_circ_0040355, hsa_circ_0040341, hsa_circ_0105896, hsa_circ_0040348, and hsa_circ_00403681and) associated with PI3K-Akt pathway by significant fold change, respectively. All these 10 circRNAs were subject to miRanda-3.3 and predicted to have 83 target miRNAs (more than one binding site). The RNA networks of circRNA-miRNA are presented in Figure 6. In particular, we found that hsa_circ_0030591 (associated with the ABC transporters pathway) has 12 predictive target miRNAs related to lung cancer with one or more than one binding site, including hsa-let-7, 12 hsa-miR-103b, 13 hsa-miR-125a-5p, 14 hsa-miR-23, 15 and hsa-miR-449c-5p. 16 Similarly, hsa_circ_0040348 (associated with the PI3K-Akt signaling pathway) has 5 corresponding miRNAs with more than one binding site, and was predicted to bind to many lung cancer-related miRNAs with binding site, such as hsa-let-7, 12 hsa-miR-27a-3p, 17 hsa-miR-432-5p, 18 hsa-miR-513c-5p, 19 hsa-miR-622, 20 and hsa-miR-548. 21 More importantly, both of these 2 circRNAs met the selection criteria: (1) original processed signal of the microarray>100; (2) fold change>2; (3) P < .05; (4) upregulated in the gefitinib-resistant group; (5) predicted with more than 5 target miRNAs involving in lung cancer development or drug resistance signals. Based on this, hsa_circ_0030591 and hsa_circ_0040348 were 2 circRNAs selected for further validation, which may potentially involve in resistance of gefitinib treatment of LUAD patients.

CircRNA/miRNA network. Top 10 circRNAs associated with ABC transporters and PI3K-Akt signaling pathways togethering with their 83 target miRNAs for construction of the RNA networks of circRNA-miRNA.
Validation of the Selected DECs by qRT-PCR
To validate the narrow downed DECs from circRNA microarray assay, we used qRT-qPCR to detect the expressions of hsa_circ_0030591 and hsa_circ_0040348 in peripheral plasma of all 16 LUAD patients harboring sensitive EGFR mutation, including 8 gefitinib-sensitive and 8 gefitinib-resistant patients. As shown in Figure 6, both hsa_circ_0030591 and hsa_circ_0040348 were indeed upregulated in the gefitinib-resistant group, compared with the gefitinib-sensitive group. The FCs of hsa_circ_0030591 and hsa_circ_0040348 were 3.41 (P = .011) and 2.54 (P = .039), respectively (Figure 7), which were basically consistent with the results obtained from microarray assay (FC = 2.14 and FC = 2.27).

qRT-PCR validation. The relative expression levels of hsa_circ_0030591 and hsa_circ_0040348 in peripheral plasma of all 16 LUAD patients harboring sensitive EGFR mutation.
Discussion
Gefitinib, as a widely used first-generation of EGFR-TKI, has been successfully applied in the treatment of patients with advanced NSCLC, especially LUAD harboring sensitive EGFR mutations. However, acquired resistance to EGFR-TKI is almost inevitable and most patients suffer disease recurrence after 1 to 2 years. CircRNAs, as an intriguing class of RNA with high stability and complicated functions in gene regulation, have been found to be involved in both cancer onset and progression through diverse biological mechanisms of action.5,22 More importantly, recent studies showed that circRNAs may also play critical roles in drug resistance. 23 Further, circRNA expression can be tissue specific and detected in exosomes, cell-free saliva, and plasma.4,24 Therefore, emerging interests focus on identifying and developing circRNAs as potential biomarkers for diagnostic and therapeutic utilities. 23
In the present study, we collected a total of 16 peripheral plasma samples from LUAD patients harboring EGFR-sensitive mutations (L858R or 19Del) who were treated with gefitinib, including 8 cases of them were considered as gefitinib-sensitive and 8 were gefitinib-resistant. 4 gefitinib-sensitive and 4 gefitinib-resistant samples were subject to performing circRNAs microarray assay. It was found that there were 2571 DECs in the patients with acquired resistance to gefitinib compared with that of sensitive patients. The identified DECs were used for bioinformatics analysis including GO and KEGG analyses, the results showed that a line of items of biological process, cellular component, and molecular function were enriched, and chromosome segregation, cell cycle process, and single-organism organelle organization were the top 3 most significant biological processes. Furthermore, the hierarchy of enriched biological processes suggested mitotic cell cycle process, sister chromatid segregation, and nuclear division, which have been proven to be related with drug resistance in tumors,25,26 might be involved in the mechanism of EGFR-TKI resistance in LUAD. Importantly, we found that ABC transporters and PI3K-Akt signaling pathway were ranked in the top 10 items with the most significant differences in the KEGG pathway. As a well-established multidrug resistance-related signaling and one of the classic pathways of EGFR-TKI in lung cancer, of which dysregulation may lead to rational resistance events consequently. Therefore, we selected the top 10 circRNAs in ABC transporters and PI3K-Akt signaling pathway and their corresponding miRNAs for constructing of circRNA-miRNA RNA network. In particular, many miRNAs associated with lung cancer growth, such as hsa-miR-128, let-7, and hsa-miR-27a,6,12 were predicted to have binding sites within hsa_circ_0030591 and hsa_circ_0040348. We then validated that hsa_circ_0030591 and hsa_circ_0040348 were indeed upregulated in gefitinib-resistant patients by qRT-PCR, which is consistent with the results of microarray.
For the limitations of our study, due to the nature of this study, we did not strictly calculate the sample size for performing circRNAs microarray assay (a small sample size with 4 gefitinib-sensitive and 4 gefitinib-resistant samples were included), which may result in selection bias. And for the biological validation step, the sample size was also limited to 8 gefitinib-sensitive and 8 gefitinib-resistant blood samples for qRT-PCR. A study with a larger sample size is warranted in the future to further confirm these results. Besides, we respectively collected the samples from matched gefitinib-sensitive and gefitinib-resistant patients; it may be unable to exhibit the dynamic change before and after gefitinib treatment; therefore, it would be a different good angle in the future studies to prospectively collect plasma samples of the same case before and after gefitinib treatment, which may specifically exhibit the dynamic change during the development acquired gefitinib resistance.
In summary, our study provides data and comprehensive analysis on plasma circRNAs expression profiles for NSCLC patients with acquired gefitinib resistance and showed that circRNAs may play key roles in gefitinib resistance. In particular, hsa_circ_0030591 and hsa_circ_0040348 may be candidates for further validation as potential biomarkers for therapeutic monitoring EGFR-TKIs resistance in NSCLC patients.
Supplemental Material
sj-docx-1-tct-10.1177_15330338221139167 - Supplemental material for Comprehensive Analysis of Circular RNA Expression Profiles in Gefitinib-Resistant Lung Adenocarcinoma Patients
Supplemental material, sj-docx-1-tct-10.1177_15330338221139167 for Comprehensive Analysis of Circular RNA Expression Profiles in Gefitinib-Resistant Lung Adenocarcinoma Patients by Junyong Zou, MD, Huiyin Lan, MD, Wei Li, MD, Shuanshuan Xie, MD, Zhongkai Tong, MD, Xiaolian Song, MD, and Changhui Wang, MD in Technology in Cancer Research & Treatment
Footnotes
Author Contributions
CHW and HYL had the idea for the article and provided the final approval of the version to be published. JYZ performed the literature search and data analysis and drafted the manuscript. WL, SSX, ZKT, and XLS were involved in revising the manuscript critically for the important scientific content. All authors listed have made a substantial, direct, and intellectual contribution to the work and approved it for publication.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplemental material. Further inquiries can be directed to the corresponding authors.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics Statement
The studies involving human participants were reviewed and approved by the Medical Ethics Committee of Ningbo Hwamei Hospital of the University of Chinese Academy of Sciences (Approval number: PJ-NBEY-KY-2020-095-01; Approval date: 04/02/2020; Address of the review board: Room 402, Building 3, No. 41, Northwest Street, Haishu District, Ningbo City, Zhejiang, ZIP Code 315010). The patients/participants provided their written informed consent to participate in this study. Written informed consent was obtained from the individual(s) for the publication of any potentially identifiable images or data included in this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Ningbo Health Branding Subject Fund (Grant No. PPXK2018-05), Ningbo Clinical Research Center for Respiratory System Diseases Fund (Grant No. 2022L004), Medical Scientific Research Foundation of Zhejiang Province, China (Grant No. 2019KY594).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
