Abstract
The management of early-stage colon cancer involves surgical resection of the primary tumor with or without chemotherapy, depending on pathological staging. The benefit of adjuvant chemotherapy for stage II and III colon cancer is approximately 5% and 15%, indicating the need for optimization for risk stratification and patient selection. Several studies have revealed that current clinicopathological factors lack precision. Circulating tumor DNA (ctDNA) is cell-free DNA originating from cancer cells and can be detected even in the absence of radiologically detectable disease among patients with colon cancer. Recent cohort studies revealed that ctDNA is one of the most significant prognostic factors for patients with early-stage colon cancer, surpassing pathological and clinical risk factors. Prospective cohort studies also suggest there may be a predictive role for ctDNA on the decision for consideration of adjuvant therapy. Currently, randomized clinical trials are enrolling to better define this role. In this review article, we review recent literature on ctDNA and its role in patients with colon cancer. We also elaborate on the future clinical utility of ctDNA in clinical practice and the unmet need for research to optimize currently available ctDNA assays.
Introduction
Colorectal cancer remains prevalent worldwide, with over 150,000 new cases anticipated in the US alone in 2024. 1 The gold standard of treatment in limited-stage colon cancer remains surgery; however, the role of adjuvant therapy is determined by pathological clinical risk factors. Although patients with stage III colon cancer are recommended to receive adjuvant therapy, the benefit of chemotherapy is limited to 10–15% in this population, and it is only ∼5% for patients with stage II colon cancer.2,3 This indicates that the majority of the patients who receive adjuvant chemotherapy do not derive benefit, and clinicopathological risk factors carry significant limitations for appropriate patient selection for adjuvant treatment of colon cancer.
Circulating tumor DNA (ctDNA) is increasingly becoming a reliable, non-invasive technique to capture cell-free DNA deriving from cancer cells and examine the underlying genetic characteristics of cancers (Figure 1). 4 While typically isolated from the peripheral blood, ctDNA can also be obtained from other sources such as cerebrospinal fluid, stool, urine, peritoneal fluid, and pleural fluid. 5 Collection often requires EDTA or cell-stabilizing tubes. The short half-life of ctDNA (approximately 2 h) with rapid blood clearance creates an opportunity to use ctDNA as a biomarker of persistent microscopic residual disease after curative surgical resection. 6 A broad range of assays are used to analyze ctDNA; these can largely be grouped into tumor-informed and tumor-agnostic platforms (Table 1). 7 Current technologies utilized for the detection of ctDNA and its origin include polymerase chain reaction (PCR), next-generation sequencing (NGS), and methylomics. 7 PCR utilizes a series of DNA primer sequences that are targeted to specific known mutations in primary tumors obtained from surgical samples and are, therefore, only able to detect specific mutations in the ctDNA, and this approach is commonly used for tumor-informed platforms. By contrast, NGS techniques are able to broadly sequence cell-free DNA, and this technology has been combined with blood-based methylation profiling of cell-free DNA to estimate the origin of ctDNA, and these are considered tumor agnostic assays. 8 These technical advances in ctDNA testing have uncovered new opportunities for the detection of minimal residual disease for patients with colon cancer.
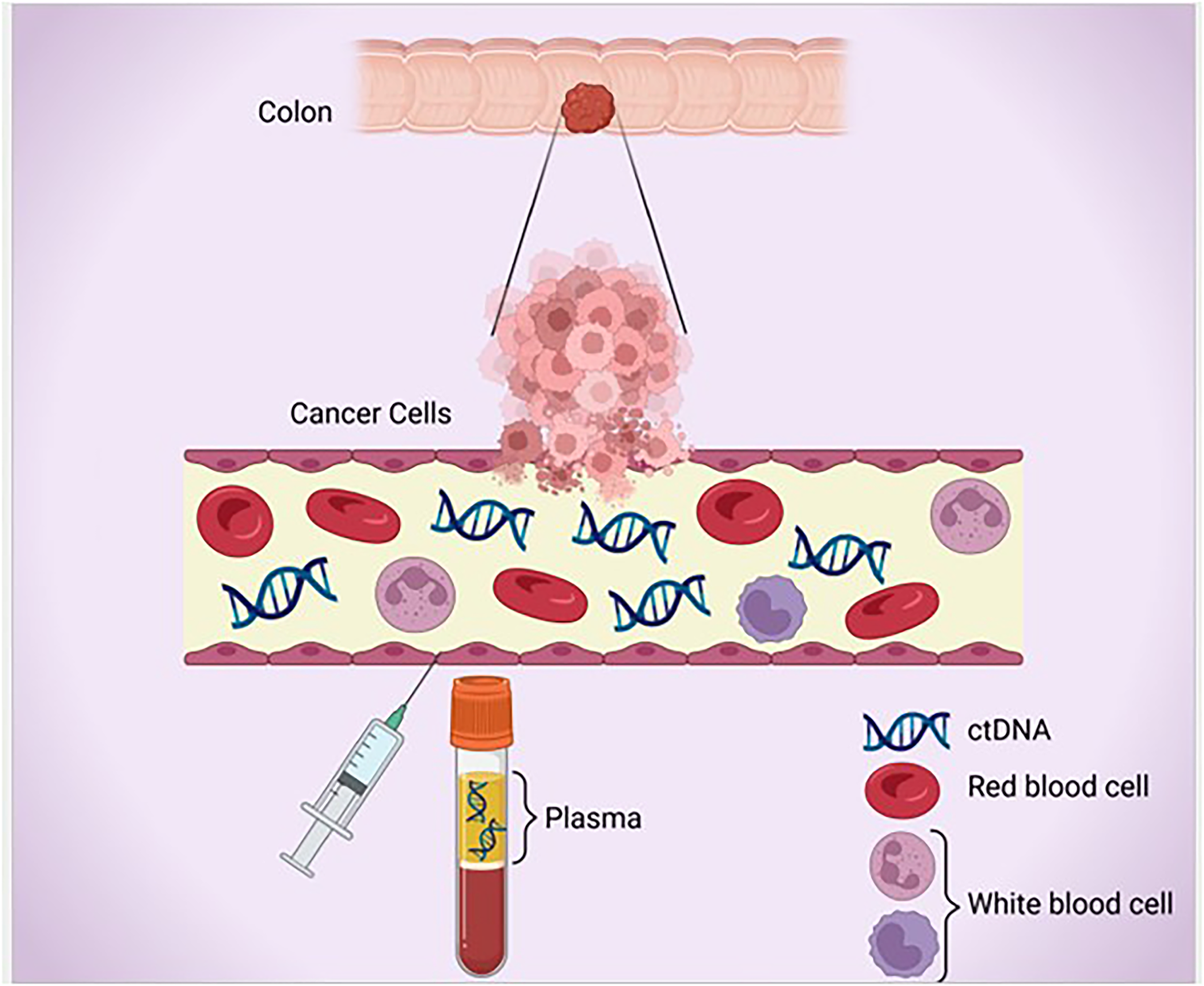
Circulating tumor DNA and its detection in plasma.
Circulating Tumor DNA Assays.

Abbreviations: NGS, Next Generation Sequencing; PCR, Polymerase Chain Reaction; WES, Whole Exome Sequencing.
The use of ctDNA has the potential to significantly enhance the ability of clinicians to risk stratify patients with colon cancer and determine appropriate adjuvant treatment based on the presence of minimal residual disease. This review will examine the current literature surrounding the use of ctDNA as a prognostic marker and its biomarker role for more precise determination of adjuvant treatment, as well as detection of targetable early micrometastatic disease following adjuvant therapy.
Prognostic Role of ctDNA After Curative Surgery for Patients with Colon Cancer
Cell-free DNA shedding dynamics vary significantly based on the origin of the cancer, and the site of metastasis in colon cancer is known to have higher rates of circulating tumor allele fractions compared to other solid tumors. 9 Due to this fact, within the past decade, prospective cohort studies were conducted to test the prognostic role of ctDNA as a surrogate of minimal residual disease (MRD) for patients with colon cancer. In one of the earlier studies, Tie et al 10 investigated the ctDNA for patients with early-stage colon cancer. In this prospective cohort study of 230 patients with resected stage II colon cancer, patients underwent tumor-informed ctDNA testing at 4 −10 weeks post-operatively and received adjuvant chemotherapy (ACT) at the clinician's discretion. Investigators identified a 7.9% post-operative (post-op) ctDNA positivity rate. Among ctDNA-positive patients, 79% experienced recurrence at a median follow-up of 27 months. By contrast, only 9.8% of patients with negative post-operative ctDNA experienced recurrence (hazard ratio (HR), 18; 95% confidence interval, 7.9 to 40; P < .001). They further demonstrated an association between positive ctDNA after completion of ACT and inferior recurrence-free survival (HR, 11; 95% CI, 1.8 to 68; P = .001), revealing a highly potent prognostic value of ctDNA (Table 2).
Overview of Studies Evaluating the Prognostic Role of ctDNA in Limited Stage CRC.

A follow-up multi-center cohort study by Tie et al 12 examined 96 patients with stage III colon cancer with the aim of examining the impact of post-operative ctDNA status on recurrence-free survival. In this study, blood samples were collected 4 −10 weeks after surgery and after completion of ACT. The baseline post-op ctDNA positivity rate was 21% (20/96), which was higher than patients with stage II colon cancer. The results demonstrated that positive post-op ctDNA was associated with inferior recurrence-free survival (HR, 3.8; 95% CI, 2.4-21.0; P < .001). The authors also investigated the prognostic value of ctDNA status after completion of ACT on the recurrence-free interval (RFI). Results showed a marked association between ctDNA positivity and decreased 3-year RFI. Patients with post-ACT positive ctDNA had 30% 3-year RFI, whereas this was 77% for patients with negative ctDNA (HR, 6.8; 95% CI, 11.0-157.0; P < .001). In the multivariate analysis, including clinicopathologic risk factors, post-op ctDNA remained the single most independent prognostic variable (HR, 7.5; 95% CI, 3.5-16.1; P < .001). The study by Diehn et al. 11 utilized NGS-based ctDNA to examine post-operative ctDNA status in a cohort of patients with stage II and III colon cancer. Uniquely, post-operative blood samples were taken very close to the time of resection with a mean duration of 10 days. The results demonstrated a significantly inferior 2-year relapse-free survival among patients who tested positive for ctDNA (17% vs 88%; HR 10.3; 95% CI 2.3-46.9; P < .00001). A similar association was noted for time to recurrence (HR 20.6; 95% CI 3.1-139.0; P < .00001) and overall survival (HR 3.4; 95% CI 0.5-25.8; P = .041).
A landmark paper by Kotani et al (The CIRCULATE-Japan-GALAXY study) 15 investigated the molecular dynamics of a tumor-informed ctDNA assay among 1039 patients with stage II-IV colon cancer who underwent complete surgical resection. CtDNA status was measured 4 weeks after surgery, which is considered as the MRD window. Among all stage patients, the ctDNA positivity rate was 18%, and this was 2%, 7.6%, and 23% among patients with stage I, II, and III colon cancer, respectively. Notably, 43% of patients who underwent surgical resection of stage IV cancer had ctDNA positivity. In the same study, the investigators identified a significant association between ctDNA positivity and RAS, BRAF, and MSI status. For example, among patients with MSI-H colon cancer, ctDNA positivity was 5%, and this was 19% among patients with MSS colon cancer. The investigators identified that 61.4% of patients with positive post-operative ctDNA experienced recurrence versus only 9.5% of those who were ctDNA negative (HR 10.0, 95% CI 7.7-14.0, P < .0001) and 18-month disease-free survival (DFS) was 38.4% versus 90.5%, respectively. This association was noted across all-stage diseases with an HR of 37, 18, 9.6, and 5.9 for stages I, II, III, and IV, respectively. In the multivariate analysis that included clinical, pathological, and molecular risk factors status, post-operative ctDNA positivity was the single most important prognostic factor (HR 10.82, 95% CI 7.07-16.6, P < .001). By contrast, more traditional risk factors such as microsatellite status, pathological T and N stage, and RAS status were not significantly associated with risk of recurrence after adjustment with ctDNA status. Most recently, the updated analyses of this study published by the investigators validated the findings of the original report, and ctDNA was found to be the most powerful prognostic marker in multivariate analysis (HR = 12.08, P < .001). 16 Interestingly, persistent ctDNA negativity during the surveillance period was an even more powerful predictor of better survival outcomes, with an HR of 44.24 (P < .0001) compared to one-time testing. This indicates that ctDNA is a dynamic biomarker, and the prognostic value and sensitivity of ctDNA testing significantly improves with repeated testing. This also indicates there is an unmet need for research to improve the rate of false negative results, which is likely due to low-level DNA shedding, by advancing the technology to capture ultra-low levels of ctDNA.
Taieb et al 17 investigated the prognostic role of ctDNA among patients enrolled in the PRODIGE-GERCOR trial. In this post-hoc analysis, 1345 patients with stage III colon cancer had ctDNA testing, and the investigator used tumor-agnostic ctDNA testing. Notably, the ctDNA positivity rate was 13.8%- and 3-year DFS was 66.3% versus 76.7% for patients with positive and negative ctDNA, respectively (HR = 1.55, P = .006). In this study, both the ctDNA positivity rate and risk of recurrence were highly different from those of other studies, indicating that the sensitivity and specificity of assays can significantly vary, leading to highly distinct outcomes.
Most recently, a US-based study, the BESPOKE CRC 19 reported results from a prospective cohort study investigating a tumor-informed ctDNA platform among patients with stage II and III colorectal cancer. CtDNA positivity rate was 6.9% and 22.4% among patients with stage II and III colorectal cancer, which is consistent with the findings of the Japanese GALAXY study. CtDNA positivity was associated with significantly inferior disease-free survival (DFS) with an HR of 25.7 and 18.1 for patients with stage II and III colorectal cancer, respectively. The strength of the association found in this study calls into question whether the historical prognostic factors that we use for adjuvant treatment decisions are precise enough for the selection of the right population for adjuvant therapy.
Several other studies also investigated the role of ctDNA for patients with colorectal cancer with different assays (Table 1) and confirmed the prognostic role of ctDNA.8,13,14,18,20,21 However, it is important to note that ctDNA dynamics for patients with colon and rectal cancer significantly differs, 22 and higher incidence of ctDNA false negative rates can be seen among patients with early-stage rectal cancer with ctDNA positivity range of 75%–85% while this is ∼95% for patients with early-stage colon cancer. 22 It is also important to note that the treatment paradigm of rectal cancer has also evolved into total neoadjuvant approaches where the use of ctDNA is relatively limited for treatment decisions, and its applicability to the clinical practice of rectal cancer appears to be limited to surveillance. Therefore, studies should separately examine colon and rectal cancers to better define the role of this novel technology.
Predictive Role of ctDNA Benefit from Adjuvant Chemotherapy
The utility of ctDNA as a prognostic biomarker has, in turn, led researchers to ask whether ctDNA can also be used to guide treatment decisions in CRC. In fact, some prospective cohort studies also investigated the predictive value of ctDNA for patients who underwent ACT. In their study, Tie et al. demonstrated that among 16 patients with stage III colon cancer patients with positive post-op ctDNA, 9 out of 16 patients (56%) cleared the ctDNA following adjuvant therapy. 12 The GALAXY trial investigators reported a significant benefit of ACT among patients with stage II/III colon cancer and with positive ctDNA with HR of 6.59 (P < .0001). 15 In this study, 18-month DFS rates were 22% versus 61% in patients with and without ACT, respectively. Notably, this benefit was not seen among patients with negative ctDNA with HR of 1.7 (95% CI 0.8-3.7, P-0.16), indicating beyond prognostic value, ctDNA may also be a predictive value for adjuvant chemotherapy. Most recently, the investigator reported updated analyses of this study, and notably, the benefit of chemotherapy was not seen among patients with negative ctDNA (HR 1.4 P = .09), while significant benefit was seen among patients with positive ctDNA (HR = 4.3, P < .0001). 16
The BESPOKE study also showed a significant association with ctDNA positivity and benefit from ACT. 19 In this study, HR for the benefit of ACT was 3.06 (P = .0025) versus 1.47 (P = .23) for patients with and without a positive ctDNA test result, respectively. Although studies suggest there is a potential predictive role of ctDNA for ACT, it is important to note that these were non-interventional, uncontrolled studies with relatively early results, and long-term outcomes may further shed light on the predictive value of ACT, particularly for patients who test ctDNA negative.
One such landmark interventional study, the DYNAMIC trial, 23 in which 455 patients with stage II colon cancer were randomly assigned to have treatment decisions (ACT) guided by either ctDNA or conventional management. CtDNA was measured at 4- and 7-weeks post-surgery; in the ctDNA group, a positive result at either time point prompted the initiation of ACT, while ctDNA-negative patients were not treated. The primary endpoint of this study was 2-year DFS outcomes. After a median follow-up period of 37 months, 15% of patients in the ctDNA group received ACT versus 28% in the standard group (RR, 1.82; 95% CI, 1.25 to 2.65). Notably, patients in the interventional arm received more oxaliplatin-based chemotherapy than those in the standard arm (62% vs 10%). In a comparison of two-year recurrence-free survival, ctDNA management was found to be non-inferior to standard management (95% CI, −4.1 to 6.2, noninferiority margin, −8.5 percentage points). Two-year DFS rates were 93.5% versus 92.4% in ctDNA-guided and standard management arms, respectively. However, among patients with negative post-op ctDNA, patients with T4N0 disease had 3-year DFS of 81.3% while this was 94.2% for those with T3N0 disease. This indicates that patients with T4 disease and negative ctDNA are at high risk of recurrence (HR 2.6 95% CI 1.01 to 6.71). Although this study demonstrated that the use of ctDNA-guided clinical decision-making was able to reduce exposure to ACT without reducing recurrence-free survival for patients with T3 disease, relatively inferior DFS outcomes seen among patients with T4 disease indicate that single time point ctDNA testing may not be sufficient for decision making for this high-risk group.
The COBRA trial 24 also investigated the role of ctDNA for adjuvant chemotherapy for patients with T3N0 colon cancer. This study utilized an earlier form of tumor agnostic ctDNA assay. A total of 635 patients with low-risk colon cancer were randomized to receive either standard of care versus ctDNA-guided management. Patients with positive post-operative ctDNA received 6 months of ACT, and ctDNA was re-measured at 6 months to assess for clearance. However, among the 16 total patients with positive ctDNA, 43% in the surveillance arm experienced ctDNA clearance after 6 months versus 11% in the chemotherapy arm. The study has since been terminated, given these negative results with a higher incidence of ctDNA clearance with observation only compared to ACT, which was probably driven by false positive ctDNA results in the context of the tumor agnostic testing in which unrelated genetic variants can be captured if testing is not optimized. Ongoing studies such as the CIRCULATE-US trial6,25 are currently enrolling patients to investigate the predictive role of ctDNA for patients with high-risk stage II and stage III colon cancer. In this study, patients will assigned into two cohorts based on ctDNA status. Patients who test negative for ctDNA (cohort A) will undergo randomization to standard of care ACT versus ctNDA surveillance without ACT. Patients in the cohort with positive post-op ctDNA (cohort B) will be randomized to receive either standard of care ACT versus FOLFIRINOX (5-fluorouracil and folinic acid plus oxaliplatin plus irinotecan). Patients who initially test ctDNA negative and develop ctDNA positivity later in the course of surveillance will be re-randomized in cohort B to receive either the standard-of-care ACT or FOLFIRINOX (delayed adjuvant approach). Currently, several other trials are investigating the predictive role of ctDNA for adjuvant chemotherapy decisions, including the CIRCULATE-Japan and Dynamic III trials (CCTG C029).
The use of ctDNA as a Surveillance Strategy for Colorectal Cancer After Completion of Curative Therapy
Given the utility of ctDNA as a prognostic biomarker, the potential for its use as a surveillance strategy to monitor for recurrence is also being explored. It is important to note that ctDNA is a dynamic marker, and repeated tests can have advantages for surveillance. The study of Tie et al identified that among 62 patients with negative post-op ctDNA, 6 (10%) patients seroconverted to ctDNA positive during the surveillance. Kotani et al 15 compared 12-week post-operative ctDNA status with carcinoembryonic antigen (CEA) levels, which is typically utilized in the clinical setting to monitor for recurrence. Their results demonstrated a concordance of positivity/negativity between ctDNA and CEA status in 81.3% of patients and a discordance in 18.7% of patients. When analyzing discordant cases, 70% of the ctDNA positive/CEA negative patients experienced relapse as compared to 12% of the ctDNA negative/CEA positive patients. In the same study, patients whose test converted to ctDNA positive had significantly worse DFS outcomes compared to patients who persistently had negative tests (18 months DFS of 33.8% vs 92.1%). These results suggest that ctDNA positivity has the potential to be a strong predictor of relapse and can be used for surveillance. Similarly, in the BESPOKE study, 19 350 patients with stage II-III CRC were included in the initial analysis, of which 232 received ACT, and 118 were observed following curative resection. Interestingly, 44.4% of patients with ctDNA clearance experienced disease recurrence. In all of these patients, ctDNA turned positive prior to radiological evidence of relapse. Furthermore, serially ctDNA positive patients were found to have significantly worse DFS as compared to serially ctDNA negative patients (HR = 124.3, 95% CI: 29.8-518.7, P < .0001), and it was a more powerful predictor of recurrence than single time point ctDNA testing (HR = 20.8). In this study, the investigators also showed that approximately 40% were able to undergo oligometastatic-directed therapy, and 55% of those patients remained disease-free. Notably, the study by Dasari et al 26 showed that 86 patients out of 1049 developed ctDNA positivity, and among those, 46 (53%) had measurable metastatic disease at the time of ctDNA measurement, and only 47% had true MRD positivity. Although overall, these findings are encouraging for the surveillance use of ctDNA for patients with early-stage colon cancer, there is a clear unmet need for improved assays, while more mature data with long-term outcomes is needed on whether these interventions would alter the overall survival of the patients.
The use of ctDNA for the Detection and Elimination of Micrometastatic Disease After Adjuvant Therapy
The use of ctDNA for the elimination of micro-metastatic disease is also being actively explored in several studies. The ongoing PEGASUS study investigated the role of FOLFIRI in the elimination of micrometastatic disease. In this study, 35 patients with post-op ctDNA positivity were treated with 3 months of CAPEOX, and among those, 11 patients (31%) experienced ctDNA clearance. Most notably, the investigators reported that 48% of patients with post-ACT (CAPEOX) positive ctDNA cleared ctDNA after receiving FOLFIRI, indicating a potential role of FOLFIRI in micrometastatic disease.27,28 The study by Pappas et al. 29 is currently investigating the eradication of occult metastatic disease defined by positive ctDNA in the blood using a tumor agnostic assay. In this study, ctDNA screening is obtained 3-6 weeks after ACT completion, and patients with negative initial ctDNA were surveilled every 3 months with imaging and ctDNA testing. The study investigates several different approaches, including FOLFIRI and precision medicine therapeutics after ACT for patients with stage III colon cancer. Patients with positive initial ctDNA and actionable biomarkers (V600E, HER2, MSI-H) received 6 months of biomarker-directed therapy. The primary outcome is ctDNA clearance one month after completion of additional therapy or surveillance. Another stage II/III study by Yaeger et al 30 aims to evaluate whether Encorafenib and Cetuximab improve DFS in patients with BRAF V600E stage II or stage III colon cancer after standard adjuvant therapy. Patients will be enrolled in ctDNA detectable and undetectable cohorts; in the ctDNA detectable cohort, the primary endpoint will be ctDNA clearance rate at 6 months, whereas this will be 6 months of ctDNA recurrence-free survival in the non-detectable cohort. Collectively, there are increasing efforts to utilize ctDNA as a surrogate marker of persistent occult metastatic disease with the intention of eradicating micrometastatic disease.
Discussion
The management of colorectal cancer with advanced-stage disease is evolving to precision medicine approaches using several molecular biomarkers, including but not limited to: KRAS, BRAF, HER2, MSI-H, and POLE/POLD1. However, the use of adjuvant chemotherapy is solely based on clinicopathological factors with limited progress over the past decades, and it lacks precision; for this reason, a significant portion of patients who may not benefit from chemotherapy receive adjuvant therapy regardless. CtDNA has been of interest in colon cancer research as shedding tumor-derived DNA appears to be reasonably good for this disease. So far, several prospective cohort studies15,19 uncovered the highly prognostic value of ctDNA, which consistently appears to be more reliable than historical clinicopathological risk factors, leading to prospective interventional studies, some of which are currently enrolling patients globally (CIRCULATE-NA, CIRCULATE-Japan, DYNAMIC III). So far, studies discussed above clearly demonstrate that patients with ctDNA positivity in the MRD window are at the highest risk of recurrence with worse survival outcomes compared to those patients with negative ctDNA. Notably, some studies suggest the level of ctDNA may also be linked to the risk of recurrence and a higher ctDNA level can be seen among patients who eventually developed recurrent disease 31 Nonetheless, spontaneous clearance was reported in only 1.9% of patients in the GALAXY-Japan trial after positive ctDNA in the MRD window, indicating that positivity of ctDNA is the most important determinant for the risk of recurrence.
Although prospective data from the GALAXY 15 and BESPOKE 19 trials are suggestive of the predictive value of ctDNA, it is important to recognize these are not interventional trials; thus, the evidence is purely based on observational outcomes from patients enrolled in these cohort studies. Moreover, most of the analysis reports either 18-month or 2-year DFS, and more mature data is needed to better understand the predictive value of ctDNA, given that 3-year-DFS outcomes are a well-established surrogate of overall survival and well-studied and described in large adjuvant trials.32,33 Perhaps, it is also important to recognize that the shedding pattern of cancer is not independent of the clinical and biological features of colon cancer, as demonstrated by Kotani et al in the GALAXY trial. Similarly, the DYNAMIC trial also proved that ctDNA negativity for patients with T3N0 versus T4N0 colon cancer may be associated with a distinct risk of recurrence (94.2% vs 81.3%). 23 At this time, the evidence indicates that not only the biology of the primary tumor and clinical characteristics, but also the site of the micrometastatic disease may have an impact on ctDNA dynamics, and it may impact its reliability for decision making.22,33,34 For example, one of the limitations of ctDNA appears to be peritoneal metastasis of colon cancer where rate of ctDNA positivity can be as low as 20% depending on the assay utilized. 35 Perhaps, the false negative ctDNA seen with T4N0 colon cancer could be mainly due to the fact that T4 disease increases the risk of peritoneal recurrence where ctDNA has significant limitations. 36 Therefore, studies should factor in the clinical, pathological, and perhaps molecular markers for the analysis of ctDNA-related survival outcomes, which may define distinct approaches for a more precise determination of the therapeutic algorithm of adjuvant therapy for colon cancer. Whether some of the challenges can be overcome with the optimization of assays used in ctDNA detection remains unclear. Currently, the detection limit for most of the commercially available ctDNA testing assays is around 0.01%, 37 and the optimization of these assays may partially improve the challenges we face with ctDNA. It is important to note that additional sources of ctDNA testing, such as peritoneal fluid, may provide additional insight for the use of ctDNA testing in this challenging pattern of recurrence of colon cancer. 38 Nonetheless, the invasiveness of the procedure remains a caveat for the development of this approach for the use of ctDNA for adjuvant decisions.
Currently, ctDNA-guided adjuvant trials are enrolling patients to investigate how reliably and safely ctDNA-guided management can be applied to our daily practice. Perhaps, findings of the DYNAMIC trial23,39 provided insight that patients with low-risk, early stage colon cancer, excluding those with T4 disease, can be observed after careful discussion around risk and benefit with patients given the benefit of adjuvant chemotherapy in stage II colon cancer is limited to approximately 5%. 40 More data is needed for those high-risk stage II colon cancer patients, and perhaps surveillance with recurrent testing of ctDNA in this high-risk group may provide more ground for the use of ctDNA in high-risk colon cancer. Notably, both the GALAXY and BESPOKE trials showed recurrent testing results are more powerful predictors of prognosis than single time point ctDNA testing. Along these lines, the CIRCULATE-NA trial currently tests ctDNA every 3 months for all patients, including those who test negative for ctDNA after resection of tumor. Patients randomized to the observational arm with subsequent seroconversion of ctDNA from negative to positive will be re-randomized into the ctDNA positive cohort in which treatment escalation with FOLFIRINOX is being investigated. Whether this delayed adjuvant chemotherapy can salvage the need for adjuvant treatment for patients with false-negative postoperative ctDNA will be defined in the CIRCULATE-NA. Most recently, the results of the COBRA trial created significant concerns about the use of ctDNA, with possible false positive results and increased spontaneous clearance of ctDNA in non-interventional arm. 24 However, it is important to acknowledge that the ctDNA assay used in this study was an earlier version of ctDNA testing (The guardant LUNAR, a tumor agnostic assay), and testing technology has significantly improved since then.
Finally, ctDNA is of interest in research for occult metastatic disease in the post-ACT setting, and there is growing interest in targeting occult metastatic disease using ctDNA as a surrogate. In the GALAXY-JAPAN study, the median time from ctDNA positivity to radiological recurrence was noted to be 5.9 months. 16 In another, the lead time was reported to be 11.5 months. 18 Notably, the Intercept study by Dasari et al reported only 47% of the patients had ctDNA positivity without measurable disease. Therefore, although ctDNA can be utilized for the detection of micrometastatic disease within a reasonable time period before measurable disease, it appears to be helpful for only a subgroup of patients 26 This indicates an unmet need to advance the technology of ctDNA assays to increase the sensitivity and lead time to expand its applicability to larger cohorts. Nonetheless, there is significant interest and efforts toward the early elimination of MRD for patients with detectable MRD during surveillance. It is important to note that the microenvironment of occult micrometastatic disease has not been well defined, and there is hope that it may be associated with a more permissive microenvironment, leading to better efficacy for immunotherapeutic and precision medicine agents compared to radiologically detectable metastatic disease. Therefore, there is an unmet need for research where we can better define the targetability of positive post-ACT ctDNA (occult metastasis), which will provide more reasoning for ctDNA surveillance in clinical practice. Importantly, the BESPOKE trial showed the feasibility of capturing oligometastatic recurrence with ctDNA testing and related therapeutic intervention, and the impact of these early therapeutic interventions on limited disease on the survival outcomes of patients remains to be seen. It is important to note that most clinical trials utilize ctDNA clearance as the primary endpoint for phase II trials, and it remains unclear whether ctDNA clearance is a reliable surrogate for long-term survival outcomes. At this time, it is important to note that post-ACT positive ctDNA, outside of clinical trials, may result in patient anxiety as there is no established algorithm for the detection of radiological recurrence, and targeting ctDNA with early systemic therapy is investigational only. Therefore, testing ctDNA for surveillance outside of clinical trials should involve extensive discussion with patients and providers before consideration.
Conclusion
Collectively, growing evidence indicates that ctDNA will likely have significant roles in the future management of patients with early-stage colon cancer. While the prognostic impact of ctDNA is well established based on several prospective studies, interventional trials will be more likely to provide a better understanding of the predictive role of ctDNA in patients considered for adjuvant treatment. In fact, if the predictive value of ctDNA is proven in ongoing interventional trials, it will create more opportunities for clinicians to offer adjuvant therapies more precisely than our current clinicopathological risk stratification, which has significant limitations. Testing ctDNA may also provide more insight into the biology of occult metastatic disease and its targetability with novel therapeutics, including precision medicine and immunotherapeutics. Further studies will validate our knowledge and provide more insight into the future use of ctDNA for patients with early-stage colon cancer.
Footnotes
Author Contributions
RY and IHS developed the article concept and outline, TS created the figure, and all authors contributed to the writing and development of the manuscript.
Conflict of Interest
I.H.S. received Advisory Board fees in Seattle Genetics, GSK, Guardant Health, and Lumanity; Research Grants from BAYER and GSK. Speaker: Pfizer and Amgen. A.D.S received Honorarium from Foundation One; A.S. received research grants (to institution) from Merck, AstraZeneca, Bristol Myers Squibb, Exelixis, Clovis, Biontech, Astellas, Dragonfly therapeutics, Innovent biologics, KAHR medical, Amgen, Celgene, Actuate therapeutics, Incyte corporation, Seagen, Daiichi Sankyo, and Advisory board fees from Bristol Myers Squibb, AstraZeneca, Exelixis, Pfizer, and Daiichi Sankyo. Ad Board for Guardant Health. All COI detailes are provided the manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
