Abstract
Background
Neural precursor cell expressed developmentally downregulated 9 (NEDD9) is considered an important factor in the progression of cancer, acting as a modulator of cellular migration, adhesion, and metastatic potential. Its significance as a prognostic factor, however, remains unclear, which necessitated a comprehensive review and meta-analysis.
Methods
Our study followed the PRISMA guidelines, analyzing studies from major databases including PubMed, Embase, and Cochrane. Our eligibility criteria included studies evaluating NEDD9 expression in relation to cancer prognosis and outcomes such as overall survival (OS), progression-free survival (PFS), disease-free Survival (DFS), recurrence-free survival (RFS), and cancer-specific survival (CSS). We used random-effects and fixed-effect models for meta-analysis, and we validated our findings by comparative analysis using data from external cohorts like The Cancer Genome Atlas (TCGA).
Results
The analysis of 27 studies with 3915 patients demonstrated a significant relationship between NEDD9 expression and poor OS as indicated by the pooled meta-analysis outcome across all included cancers (HR: 1.81, 95% CI: 1.38-2.37). A significant effect on PFS/DFS/RFS/CSS was also found (HR: 2.14, 95% CI: 1.42-3.23). Variations in survival across different types of cancer were indicated by subgroup analysis. NEDD9 expression was correlated with various immune cells across cancer types according to immune infiltration analysis. Protein-protein interaction (PPI) analysis indicated significant interactions involving NEDD9, suggesting mechanisms which influence tumor behavior and response to treatment.
Conclusion
Our results suggest that NEDD9 is a significant prognostic marker in several human cancers. As a result of its central role in cancer progression and prognosis, it presents a promising target for therapeutic interventions. Our study highlights the importance of further research into the biology of NEDD9 and its therapeutic potential.
Keywords
Introduction
Cancer is now one of the leading causes of death and a barrier to increasing life expectancy worldwide. 1 Metastasis, the spread of cancer cells to distant organs from their original site, represents the most fatal aspect of cancer. Most cancer-related deaths occur due to metastatic disease rather than the primary tumor. This process involves gaining the ability to penetrate through tissues, traveling via blood vessels, lymphatic system, or direct infiltration into nearby structures, establishing secondary growths in distant organs, and subsequently resuming proliferation to colonize these organs.2–4
NEDD9 is a gene that was first described by Kumar et al using subtractive cloning technology. 5 The protein encoded by this gene is alternatively called human enhancer of filamentation (HEF1) and Crk-associated substrate lymphocyte type (Cas-L). In 1996, an initial biological function of NEDD9 was defined, and it was subsequently identified to encode for a major Crk-associated substrate protein in lymphoid cells.6,7 The most notable structural features of NEDD9 protein include many SH2 binding sites (often termed the “substrate domain”) and a carboxyl-terminal domain. 8
While a portion of the NEDD9 proteins is found in the cytoplasm, they exhibit concentration at focal adhesions (FA) in response to both internal and external signals. 9 FAs connect cells to the extracellular matrix (ECM), transmitting forces and potentially sensing mechanical signals. 10 Focal Adhesion Kinase (FAK) and Src kinases regulate FA dynamics in response to these cues.11–14 Integrin binding activates FAK, which then phosphorylates NEDD9 at a tyrosine residue near its carboxy terminus. This phosphorylation creates a binding site for Src, leading to extensive tyrosine phosphorylation of NEDD9 in its substrate domain. This activation process, which has been studied in multiple cell types, enables NEDD9 to interact with downstream effectors, facilitating migration, invasion, and proliferation-related signaling.15,16
Increased expression of NEDD9 is associated with the progression of solid cancers in lung, breast, ovarian, glioblastoma carcinoma, and cervical cancers, promoting metastasis.17–22 This pro-metastatic activity of NEDD9 is attributed to its various biological functions, such as regulating the Src-FAK “migratory switch”.8,23,24 Evidence suggests that NEDD9 overexpression is linked to a poor prognosis in cancers such as gastric, colorectal, hepatocellular carcinoma, and lung cancers.25–29 While a previous meta-analysis underscored this association, 30 controversy persists regarding NEDD9's status as an independent predictive biomarker, wit some studies even indicating a tendency towards longer survival with higher NEDD9 levels. 31 Moreover, the limitation of many studies to small sample sizes and specific cancer types further complicates the interpretation of NEDD9's prognostic value. Therefore, we conducted a comprehensive meta-analysis and systematic review to clarify the true prognostic significance of NEDD9.
Methods
The meta-analysis was performed according to the PRISMA guideline and the statement for reporting systematic reviews and meta-analyses. 32 The review was registered with PROSPERO (CRD42024519825).
Study Search and Eligibility
We conducted a search across PubMed, EMBASE, Web of Science, Scopus, and Cochrane to identify studies examining NEDD9 expression and prognosis in human cancers until March 2024. The search terms included (“NEDD9” OR “neural precursor cell-expressed developmentally down-regulated 9” OR “HEF1” OR “human enhancer of filamentation 1” OR “CAS-L” OR “Crk-associated substrate lymphocyte” OR “E230025G09Rik”) AND (survival OR prognosis OR prognostic). Studies were limited to English language focusing on human studies involving different cancers. Only articles that were published in peer-reviewed international journals or presented at conferences were included in our analysis resulting in the identification of 505 relevant papers.
The inclusion criteria for this study involved an assessment of NEDD9 expression in relation to various prognostic outcomes, including OS, DFS, PFS, recurrence-free survival (RFS), or cancer-specific survival (CSS). The selection process involved a review of references from relevant studies, with potential candidates being further screened for eligibility. Studies without sufficient data to compute hazard ratios (HR) and their respective 95% confidence intervals (CI) required for meta-analysis were excluded. Univariate and multivariate HR estimates were pooled from different studies. For the study that provides both univariate and multivariate HRs, we have prioritized multivariate HR. This is because multivariate HR accounts for potential confounders and provides a more precise estimate of the true effect. Subgroup analysis was done based on different types of cancers and they were classified into categories.
Data Collection and Quality Assessment
Titles and abstracts were screened independently by two reviewers, to identify studies that potentially meet the inclusion criteria. The full text of these potentially eligible studies was retrieved and independently assessed by two review team members. Any disagreements were resolved through discussion between the two reviewers and a third reviewer. Data have been extracted using a standardized form.
We validated our findings using data from TCGA. For this validation work, we extracted relevant NEDD9 expression data from TCGA for different cancer types included in our study. As a result, we performed comparative analyses to assess the reliability of our findings with TCGA data, enhancing the generalizability of our results.
We employed the Quality in Prognosis Studies (QUIPS) tool, recommended for quality assessment of prognostic studies. 33 The QUIPS tool evaluates various domains including study participation, study attrition, prognostic factor measurement, outcome measurement, confounding, and statistical analysis and reporting. Each domain was assessed for risk of bias, with an overall score reflecting the methodological quality of the study, classified as high, moderate, low, and no information studies. Studies achieving a score indicative of low risk of bias, typically above a predefined threshold, were considered to have a high-quality methodology.
Meta-Analysis
Data from the primary studies were extracted and analyzed using R version 4.3.3 (R Core Team (2023). R: A Language and Environment for Statistical Computing), R ‘meta’ package. 34 We used HR and corresponding 95% CI to assess the association between NEDD9 expression and OS as well as DFS, PFS, RFS, and CSS. A HR greater than 1 indicated worse prognosis among patients exhibiting NEDD9 overexpression. Both random-effects and fixed-effects models were applied, following ‘DerSimonian’ and Laird's method and Mantel-Haenszel's method, respectively. Assessment of statistical heterogeneity was performed using the I2 value, with thresholds of 25%, 50%, and 75% representing low, moderate, and high heterogeneity, respectively. Heterogeneity identified as relatively low or moderate (I2 < 50%), the fixed-effects model was employed for result pooling, whereas the random-effects model was utilized for I2 values ≥ 50%. Subgroup analyses based on cancer characteristics were performed to identify possible sources of heterogeneity. The reliability of the results was confirmed by sensitivity analysis and publication bias assessed by visual inspection of funnel plots, with asymmetry determined using regression testing Egger's linear regression. A significance level of P < 0.05 was considered statistically significant.
The Cancer Genome Atlas (TCGA) Analysis and Validation
We conducted several analyses utilizing TCGA datasets, using GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. 35 Firstly, we performed Kaplan-Meier (KM) survival analysis using TCGA data to validate the prognostic implications of NEDD9 expression identified in the meta-analysis. TIMER, a web server for comprehensive analysis of tumor-infiltrating immune cells,36–38 was employed to explore somatic copy number alterations (sCNAs) and differential gene expression profiles of NEDD9 across various cancer types.
TCGA analysis was conducted on 33 different cancer types including: Acute Myeloid Leukemia (LAML), Adrenocortical carcinoma (ACC), Bladder Urothelial Carcinoma (BLCA), Brain Lower Grade Glioma (LGG), Breast invasive carcinoma (BRCA), Cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC), Cholangiocarcinoma (CHOL), Chronic Myelogenous Leukemia (LCML), Colon adenocarcinoma (COAD), Esophageal carcinoma(ESCA), Glioblastoma multiforme (GBM), Head and Neck squamous cell carcinoma (HNSC), Kidney Chromophobe (KICH), Kidney renal clear cell carcinoma (KIRC), Kidney renal papillary cell carcinoma (KIRP), Liver hepatocellular carcinoma (LIHC), Lung adenocarcinoma (LAUD), Lung squamous cell carcinoma (LUSC), Lymphoid Neoplasm Diffuse Large B-cell Lymphoma (DLBC), Mesothelioma (MESO), Ovarian serous cystadenocarcinoma (OV), Pancreatic adenocarcinoma (PAAD), Pheochromocytoma and Paraganglioma (PCPG), Prostate adenocarcinoma (PRAD), Rectum adenocarcinoma (READ), Sarcoma (SARC), Skin Cutaneous Melanoma (SKCM), Stomach adenocarcinoma (STAD), Testicular Germ Cell Tumors (TGCT), Thymoma (THYM), Thyroid carcinoma (THCA), Uterine Carcinosarcoma (UCS), Uterine Corpus Endometrial Carcinoma (UCEC), and Uveal Melanoma (UVM). To investigate the impact of NEDD9 expression on patient survival, we performed survival analysis using GEPIA, and NEDD9 expression levels were stratified using median and quartiles cutoffs.
Protein-Protein Interaction (PPI) Network and Functional Enrichment Analysis
To shed light on possible connections and interactions involving NEDD9, we built a PPI network using the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) database. 39 Enrichment analysis results on the PPI were extracted from STRING and visualized using bar plots in Python 3.8. The plots incorporated key parameters such as strength in network and FDR-adjusted p-value. In addition, GSCA, an Integrated Platform for Gene Set Cancer Analysis at Genomic, Pharmacogenomic, and Immunogenomic Levels,40,41 was employed to explore the relationship between NEDD9 expression and pathway activity in cancer. This analysis provided insights into its role in progression and tumorigenesis.
Immune Infiltration Analysis
We used GSCA to run the ‘ImmuCellAI’ algorithm for immunogenomic analysis with a focus on 24 different types of immune cells to clarify the relationship between NEDD9 expression and the immune microenvironment in cancer. Significant immune infiltration was considered as those having an FDR adjusted p-value < 0.05. Furthermore, TIMER 2.0 was employed to evaluate immune infiltration, with a specific emphasis on cancer-associated fibroblasts (CAFs), and to clarify the correlation between immune infiltration and NEDD9 mRNA expression levels.
Protein-Level Expression Analysis
Data from the Human Protein Atlas (HPA) 42 were used to quantify the percentages of patients with medium or high expression of NEDD9 protein in different human cancers. Results with experiments that used the antibodies HPA038768, HPA072955, and CAB009720 were reported. Additionally, differential NEDD9 protein expression analyses were performed using Clinical Proteomic Tumor Analysis Consortium (CPTAC) data, and the analysis on such data was run on The University of Alabama at Birmingham Cancer data analysis Portal (UALCAN).43,44
Results
Search Results
Our search strategy yielded 504 studies from Medline (via PubMed), Web of Science, Scopus, Cochrane, and Embase. Following the removing of duplicate records, 292 studies underwent screening based on title and abstract, resulting in the exclusion of 265 studies. The remaining 27 full texts were independently reviewed by the same investigators. Of these, 27 were deemed eligible for inclusion. Meta-analysis was conducted on 24 of these studies. The search and selection process are illustrated in Figure 1a.

(a) PRISMA flowchart; (b) QUIPS quality assessment.
Summary of the Included Studies
Our study included a total of 3915 participants from twenty-seven studies.19,25–29,31,45–64 Of these participants, 1898 were males and 1275 were females. Three studies did not mention the gender of the patients.50,60,64 Most studies used immunohistochemistry (IHC) as a detection method. However, three studies used enzyme-linked immunosorbent assay (ELISA). Studies were from China (n = 13),19,27–29,45–49,51,53,56,61,64 Croatia (n = 4),31,57,62,63 Egypt (n = 3),52,53,58 Turkey (n = 3),26,50,54 Greece (n = 1), 59 Germany (n = 1) 64 and Japan (n = 1). 55 One study did not mention the country. 60 Among all studies, six evaluated gastric cancer,26,28,46,51,58,62 four lung cancer (Two reported as lung adenocarcinoma31,60 and the other two reported as non-small cell lung cancer by the primary studies),29,55 three colorectal cancer,25,57,59 two melanoma (Including one uveal melanoma 64 ), 50 two bladder cancer,48,53 two pancreatic cancer (PC)45,54 and one study of each remaining cancers: Acute Myeloid leukaemia, 56 Osteosarcoma, 61 Head and neck squamous cell carcinoma, 63 Renal cell carcinoma, 49 Ovarian cancer, 19 Hepatocellular carcinoma, 27 Transitional cell carcinoma 52 and Triple-negative breast cancer 47 (Table 1).
Baseline Characteristics of Studies Included in Qualitative and Quantitative Synthesis.
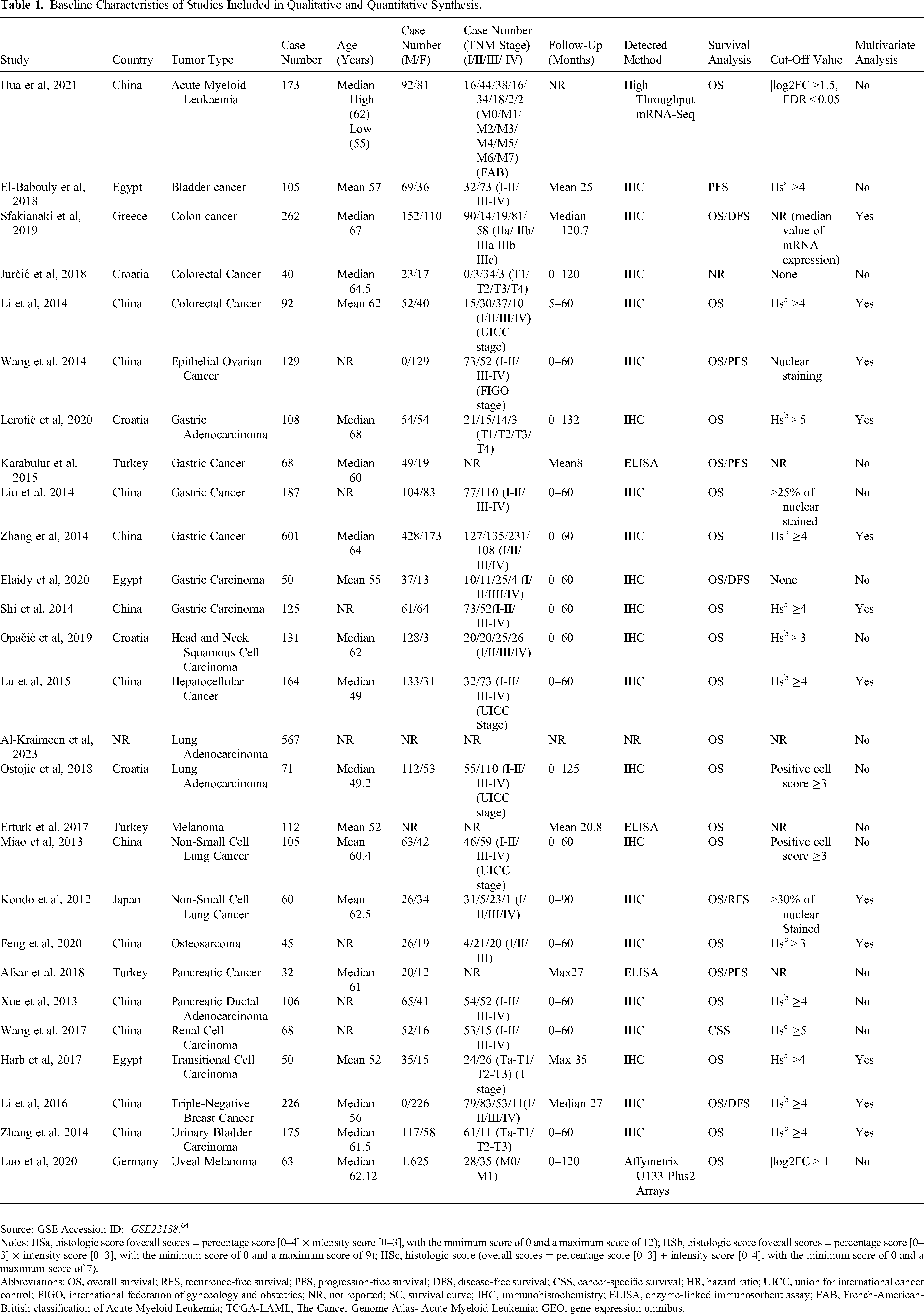
Source: GSE Accession ID: GSE22138. 64 Notes: HSa, histologic score (overall scores = percentage score [0–4] × intensity score [0–3], with the minimum score of 0 and a maximum score of 12); HSb, histologic score (overall scores = percentage score [0–3] × intensity score [0–3], with the minimum score of 0 and a maximum score of 9); HSc, histologic score (overall scores = percentage score [0–3] + intensity score [0–4], with the minimum score of 0 and a maximum score of 7). Abbreviations: OS, overall survival; RFS, recurrence-free survival; PFS, progression-free survival; DFS, disease-free survival; CSS, cancer-specific survival; HR, hazard ratio; UICC, union for international cancer control; FIGO, international federation of gynecology and obstetrics; NR, not reported; SC, survival curve; IHC, immunohistochemistry; ELISA, enzyme-linked immunosorbent assay; FAB, French-American-British classification of Acute Myeloid Leukemia; TCGA-LAML, The Cancer Genome Atlas- Acute Myeloid Leukemia; GEO, gene expression omnibus.
Quality Assessment
We performed a quality assessment for the 27 included studies. Following the QUIPS Quality Assessment Tool guidelines, 24 studies were considered of good quality and three were of moderate quality. Most studies are highly susceptible to bias from confounding factors. Also, eight studies do not have a justification about attrition. However, overall assessment revealed that most studies had good quality. The details of the quality assessment are summarized in Figure 1b.
Meta-Analysis Results
Overall survival (OS) and Progression-Free Survival (PFS). The pooled HR for OS demonstrates that there is a significant association between NEDD9 and poor prognosis (HR: 1.81, 95% CI (1.38-2.37), I2 = 82%) (Figure 2). The pooled HR for PFS demonstrated that a significant association with NEDD9 expression (HR: 2.14, 95% CI (1.42-3.23), I2 = 70%) (Figure 3a). Also, we conducted a subgroup analysis depending on the cancer location.
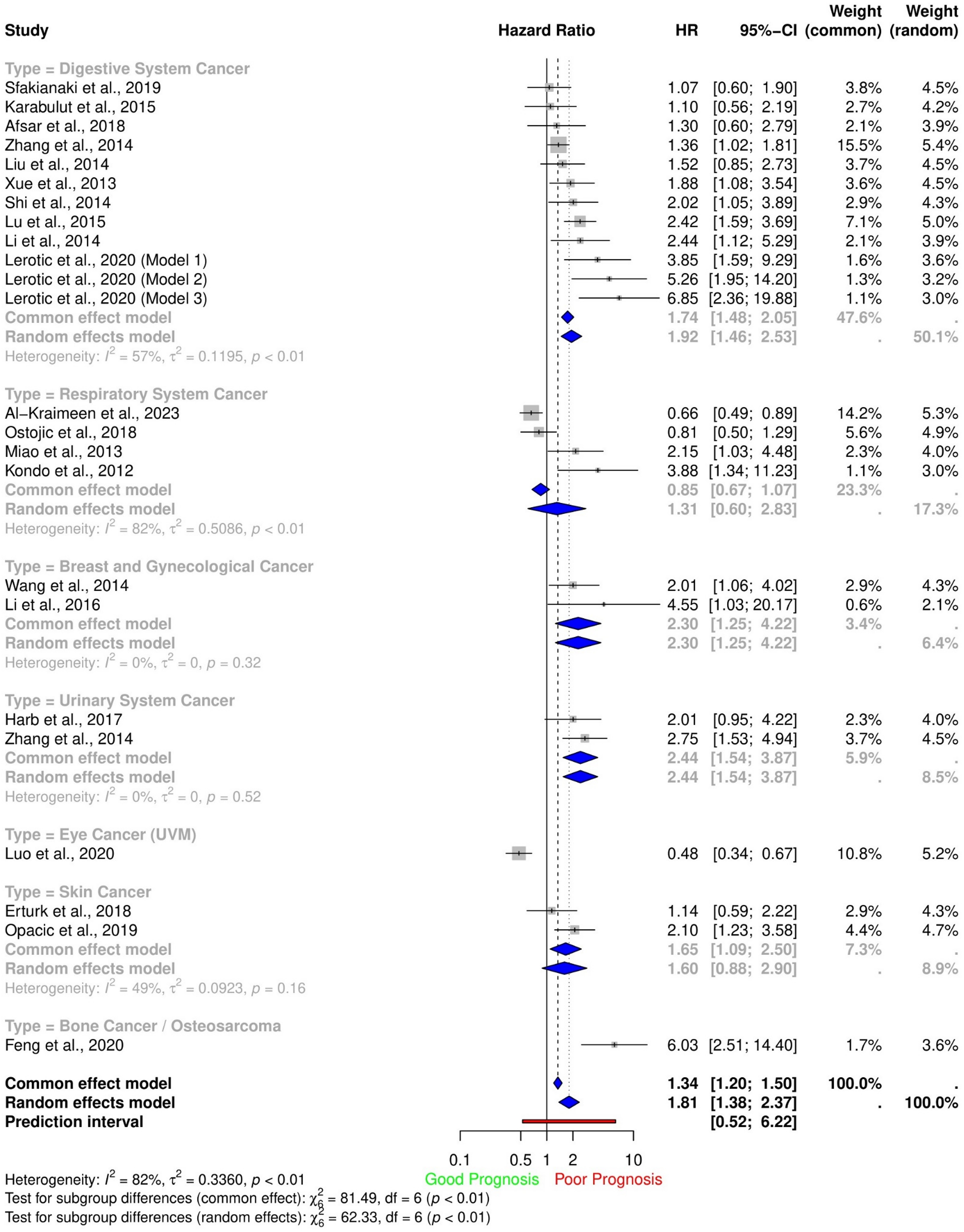
Meta-analysis of hazard ratios for overall survival.

Forest plots for meta-analysis on (a) PFS/RFS/CSS, (b) OS for lung cancers, and (c) OS for gastrointestinal cancers.
Digestive system cancer. The pooled HR for OS in the digestive system cancer group was (HR: 1.92, 95% CI (1.46-2.53), I2 = 57%) (Figure 2). Also, we conducted a subgroup analysis depending on cancer types (Figure 3c). The pooled HR for OS in gastric cancer was (HR: 1.88, 95% CI (1.28-2.76), I2 = 62%), PC (HR: 1.64, 95% CI (1.02-2.62), I2 = 0%), colon cancer (HR: 1.54, 95% CI (0.69-3.45), I2 = 64%) and hepatocellular carcinoma (HR: 2.42, 95% CI (1.59-3.45)).
The pooled HR for PFS in the digestive system cancer group was (HR: 1.35, 95% CI (0.90-2.02), I2 = 46%) (Figure 3a).
Respiratory system. The pooled HR for OS in the respiratory system tumors consisting of non-small cell lung cancers was insignificant (HR: 1.31, 95% CI (0.60-2.83), I2 = 82%) (Figure 2). Also, we conducted a subgroup analysis depending on the cancer type (Figure 3b). For the identified lung adenocarcinoma (LUAD) subgroup, a significant association was found (pooled HR: 0.70, 95% CI (0.54-0.90), I2 = 0%).
One study by Kondo et al, reported the HR for PFS in lung cancer (HR: 3.08, 95% CI (1.37-6.93). 16
Breast and gynaecological cancer. The pooled HR for OS in the Breast and gynaecological cancer group was (HR: 2.30, 95% CI (1.25-4.22), I2 = 0%) (Figure 2). Also, the pooled HR for PFS was (HR: 2.70, 95% CI (1.62-4.49), I2 = 0%) (Figure 3a).
Urinary system cancer. The pooled HR for OS in the Urinary system cancer group was (HR: 2.44, 95% CI (1.54-3.87), I2 = 0%) (Figure 2). Also, the pooled HR for PFS was (HR: 3.64, 95% CI (1.36-9.75), I2= 34%) (Figure 3a).
Skin cancer. The pooled HR for OS in skin cancer was (HR: 1.60, 95% CI (0.88-2.90), I2 = 49%) (Figure 2).
Bone cancer. One study reports the HR for OS in osteosarcoma (HR: 6.03, 95% CI (2.51-14.40) 61 (Figure 2).
Eye cancer. One study reports the HR for OS in uveal melanoma (HR: 0.48, 95% CI (0.34-0.67) 64 (Figure 2).
Publication bias. In evaluating publication bias for the prognostic significance of NEDD9 in human cancers, Egger's test showed different findings across different survival metrics. For OS, a significant intercept of 4.09 (95% CI: 2.32 to 5.85, P < 0.001) suggested the presence of publication bias, as funnel plot (Figure 4a). On the other hand, for the combined metric of PFS, DFS, RFS, and CSS, the test indicated no significant publication bias (intercept: 2.57, 95% CI: −0.23 to 5.37, P = 0.121) (Figure 4b).

Funnel plots for (a) OS, and (b) PFS/RFS/CSS.
TCGA Analysis and Validation
sCNAs were analyzed on 33 TCGA cohorts, and NEDD9 has shown significant deletions (deep or arm-level) and gains (arm-level or high amplifications) in some cancers. In addition, further mutational analysis revealed that NEDD9 (CAS-L) protein mutations are most abundant in stomach, colorectal, endometrial, bladder, head/neck, prostate, squamous lung, and clear cell kidney cancers (Figure 5a). NEDD9 expression levels were also analyzed between cancer and normal samples, and NEDD9 was revealed to be differentially expressed in several cancers, including BLCA, BRCA, CJOL, COAD, GBM, HNSC, KICH, KIRC, KIRP, LIHC, LUSC, PRAD, READ, SKCM, STAD, and THCA (Figure 5b).
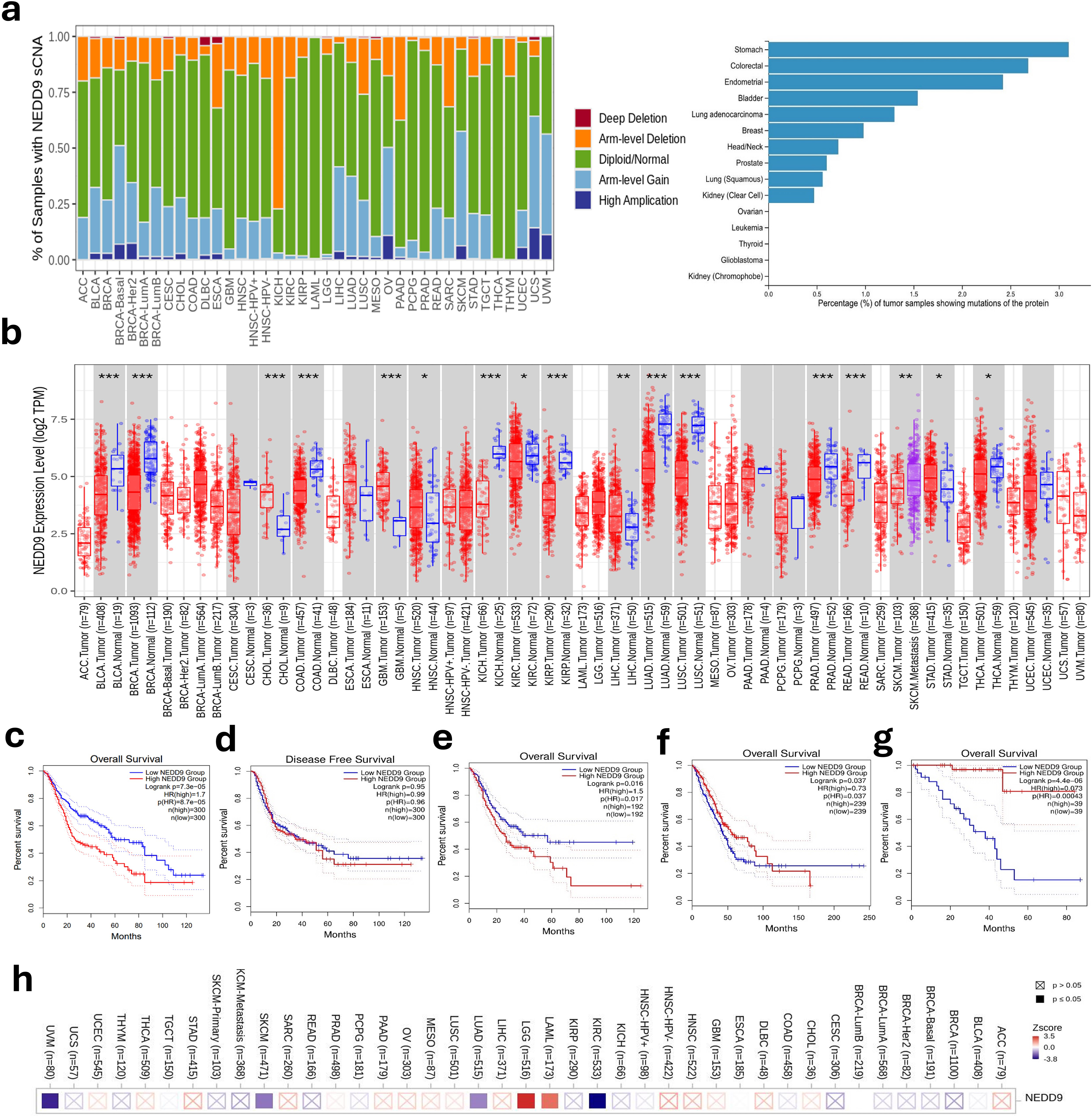
TCGA analysis and validation across different human cancers. (a) Proportions of somatic copy number alterations (sCNAs) for NEDD9, (b) NEDD9 expression profiles in tumor/normal samples, (c) Kaplan meier (KM) curve for selected gastrointestinal (GI) cancers (Outcome: OS), (d) KM curve for selected GI cancers (Outcome: DFS), (e) KM curve for STAD (Outcome: OS), (f) KM curve for LUAD (Outcome: OS), (g) KM curve for UVM (OS), and (h) heatmap for NEDD9 expression and death risk-related scores adjusted to cancer stage.
Validation from selected TCGA cohorts has shown that the HR in gastrointestinal cancers (STAD, LIHC, PAAD, COAD) combined was (HR: 1.7, P < 0.001, quartiles cutoffs) and (HR: 0.99, P = 0.96, quartiles cutoff) for OS and DFS, respectively (Figure 5c and Figure 5d). For TCGA-STAD gastric adenocarcinoma cohort individually, the HR was (HR: 1.5, P = 0.017, median cutoff) (Figure 5e). In addition, the HR for TCGA-LUAD cohort was (HR: 0.73, P = 0.037, median cutoff) (Figure 5f). And for TCGA-UVM, the HR was (HR: 0.073, P < 0.001, median cutoff) (Figure 5g). The death risk-related scores associated with NEDD9 expression adjusted to cancer stage are illustrated in Figure 5h. With this adjusted model, NEDD9 expression was significantly associated with poor prognosis in cancers like LAML and LGG, and it was significantly associated with good prognosis in UVM and LUAD.
Protein-Protein Interaction (PPI) and Functional Enrichment Analysis
We constructed a STRING PPI network with 21 nodes and 87 edges. The network revealed interactions between NEDD9 and MICAL1, AURKA, BCAR3, DOCK3, SRC, CRKL, SMAD3, PTK2, CRK, HDAC6, and several other proteins (Figure 6a). Functional enrichment analysis on NEDD9-related proteins in the PPI network had GO terms related to signaling by receptor tyrosine kinases and the VEGFA-VEGFR2 pathway, which are critical to tumor angiogenesis and proliferation, as well as the hepatocyte growth factor receptor signaling, EGF/EGFR signaling, and signaling to ERKs. Furthermore, NEDD9 and its related proteins were demonstrated to play a role in pathways like MET enhancing cell motility, signaling by PDGF, focal adhesion, and regulation of actin cytoskeleton (Figure 6b). Tissue-specific terms, with enrichment in zones like cervical carcinoma cells and bone marrow cancer cells, emphasizing the impact of NEDD9 and its related genes in different organ systems (Figure 6c). Enrichment analysis on NEDD9-related PPI network has also shown significant associations with processes such as SH3 domain binding and protein kinase binding underscore the potential mechanisms by which NEDD9-related proteins might interact with components of growth factor signaling, in addition to aberrant PI3 K, PTK6 regulating Rho GTPase, Ras GTPase, and MAP Kinase. Some KEGG terms have shown relations to renal cell carcinoma and pathways in cancer (Figure 6d). Finally, enriched terms related to immune-related pathways were observed, such as positive regulation of immune system process, positive regulation of substrate adhesion-dependent cell spreading, helper T cell diapedesis, T cell receptor signaling, B cell receptor signaling, and cytokine signaling in immune system (Figure 6e). The percentage of cancers in which specific gene's mRNA expression has potential effect on pathway activity was calculated among the 33 TCGA cancer types using the GSCA tool. NEDD9 expression was significantly associated with the activation of epithelial-mesenchymal transitions EMT (22% of selected cancers), RAS/MAPK pathway (22%), and RTK (33%). In contrast, NEDD9 expression was significantly associated with the inhibition of apoptosis in 28% of cancers. Other relations between NEDD9 expression and pathway activity are shown by the heatmap in (Figure 6f). All enrichment terms mentioned were statistically significant (FDR P < 0.05).

(a) Protein-protein interaction (PPI) network for NEDD9 using STRING, (b) general functional enrichment analysis, (c) tissue-related enrichment, (d) cancer-related enrichment, (e) immune-related enrichment, and (f) heatmap for cancer-related pathway activation/inhibition and NEDD9 expression.
Immune Infiltration Analysis
For ImmuCellAI immune-infiltration analysis, the heatmap illustrated the Spearman's correlations between NEDD9 mRNA expression and immune cell infiltration across 33 cancer types. For instance, BLCA showed positive correlations for CD4_T cells (rho = 0.210), central memory (rho = 0.222), and Th17 (rho = 0.184). Negative correlations were observed with effector memory (rho = -0.195), monocytes (rho = -0.186), and neutrophils (rho = -0.163). For BRCA, a significant positive correlation with CD4+ T cells (rho = 0.380) was revealed but a noteworthy negative correlation with several immune cells such as macrophages (rho = -0.264), monocytes (rho = -0.242), and effector memory (rho = -0.337). For digestive system cancers like COAD, a predominantly positive correlation with CD4+ T Cells (rho = 0.337), Th2 (rho = 0.439), and Th17 cells (rho = 0.302) was noted. However, it has shown a negative correlation between NEDD9 expression and monocyte (rho = -0.328) and exhausted cells (rho = -0.267) infiltrations. In addition, KIRC demonstrated a diverse effect, with positive correlations in gamma delta (rho = 0.149) and Th17 cells (rho = 0.155), but negative correlations with effector memory (rho = -0.126) and macrophages (rho = -0.140). Furthermore, analysis on LIHC indicated a positive correlation between NEDD9 expression and central memory cells (rho = 0.197) infiltration but a negative effect on natural killer (NK) cells (rho = -0.135). LUAD and LUSC both display positive correlations with CD4_T cells (rho = 0.219 and 0.324, respectively) and negative interactions with effector memory (rho = -0.350 and −0.410, respectively). OV and PAAD are both characterized by a positive correlation with CD4_T cells (rho = 0.184 and 0.184, respectively) but show negative correlations with effector memory cells (rho = -0.311 and −0.242, respectively). READ and STAD both indicate positive correlations with Th17 cells (rho = 0.344 and 0.162, respectively) and negative interactions with several other immune cell types such as nTregs (rho = -0.346 and −0.179, respectively). UVM displays predominantly negative correlations, notably with CD8_T cells (rho = -0.410) and effector memory (rho = -0.427), suggesting a suppressive influence of NEDD9 on these cells (Figure 7a). In addition, analysis using TIMER revealed a positive correlation between CAFs and NEDD9 expression in several cancers including but not limited to: STAD, CHOL, BRCA, BLCA, PAAD, and HNSC. However, CAFs infiltration was shown to be insignificantly correlated with NEDD9 expression levels in UVM (Figure 7b). Finally, myeloid derived suppressor cells (MDSCs) infiltration was shown to have significant positive and negative correlations with NEDD9 expression in LIHC and LUAD, respectively, and a negative correlation in BRCA (Figure 7c).

(a) ImmuCellAI immune-infiltration heatmap (* indicates P < 0.05, # indicates FDR < 0.05), (b) TIMER cancer-associated fibroblasts (CAFs), and (c) MDSC infiltration heatmap. Abbreviations: DC, dendritic cells; MAIT, mucosal-associated invariant T cells; NK, natural killer cells; NKT, natural killer T cells; Th, T helper cells; MDSC, myeloid derived suppressor cells.
Protein-Level Expression Analysis
Figure 8a shows the percentages of patients with medium or high expression of NEDD9 protein in human cancers, where the three bar plots demonstrate the results for three different antibodies. For HPA038768, non-Hodgkin lymphomas (63.6%, 7 of 11) and prostate cancers (100%, 10 of 10) exhibited moderate to strong cytoplasmic staining. A small number of breast, cervical, and endometrial cancers showed moderate cytoplasmic staining, while hepatocellular carcinomas rarely demonstrated strong positivity. A rare instance of ovarian cancer showed strong cytoplasmic and membranous staining. Immune cells were clearly stained, whereas the remaining cancer tissues displayed weak staining or no staining at all. Nearly all cancers exhibited moderate to strong cytoplasmic and membranous positivity, while gliomas, lymphomas, and melanomas showed no staining with HPA072955. Cancers exhibited weak to moderate nuclear and cytoplasmic staining, with most glioma cases showing medium positivity (8 of 11) and the rest showing high positivity (3 of 11) with CAB009720. Differential protein expression analysis revealed that NEDD9 is significantly upregulated in breast cancer, pancreatic adenocarcinoma, head and neck squamous cell carcinoma, and glioblastoma multiforme (P < 0.05) (Figure 8b).

(a) Human Protein Atlas (HPA) analysis of the percentages of patients with medium/high NEDD9 protein expression in different human cancers and with different antibodies. (b) Differential protein expression analysis of NEDD9 in selected human cancers from CPTAC database. Abbreviations: NS, non-significant; * P < 0.05.
Discussion
In general, the relationship between NEDD9 expression and prognosis is very complex. Our findings support the fact that the relationship is also cancer dependent. This meta-analysis highlighted a significant association of NEDD9 with prognosis in several cancer types. OS data indicated a strong link to poor, and sometimes, good outcomes. Additional bioinformatics analyses underscored NEDD9's role in cancer progression and how it impacts gene expression and interacts with key cellular pathways. We also found that NEDD9 expression is significantly associated with immune cell presence in tumors, suggesting its fundamental role in tumor-immune interactions. These findings align with those of previous studies; however, more data with larger size and longer follow-up might be needed.
The molecular basis for NEDD9 in carcinogenesis arises from its role as a multifunctional coordinator of invasion, apoptosis, and cell cycle, as well as a crucial signaling intermediary in mesenchymal cell invasion. 8 This is consistent with what was indicated by our PPI and enrichment analysis, where NEDD9 and its related interacting proteins had relations to EMT, growth factor signaling, cell adhesion, and cytoskeleton. Acting as an adhesion docking protein, NEDD9 forms a part of a signaling hub at integrin-mediated adhesion sites. 8 Phosphorylation of serine, threonine, and tyrosine residues within the NEDD9 sequence modulates its interaction with downstream signaling partners, thereby regulating it signaling function. 65 NEDD9 phosphorylation is induced by factors related to cancer progression, including FAK, TGF-β, platelet-derived growth factor, Aurora A, and BCR-ABL.16,22,66,67 NEDD9 interacts with the guanine nucleotide exchange factor DOCK3 to facilitate Rac activation and promote elongated, mesenchymal-type tumor cell invasion. 68 Signaling through increased NEDD9 levels necessitates integrin β3 signaling, leading to heightened phosphorylation of integrin β3 and subsequently increased Src and FAK activity, thereby driving elongated, mesenchymal-type invasion in environments containing vitronectin. 68
The annual rise in cancer patients underscores the imperative for uncovering novel therapeutic targets.69–71 Discovering novel biomarkers is paramount for early detection, treatment, and improved patient outcomes. These biomarkers serve as crucial indicators of disease presence, progression, and response to therapy.72,73 Recently, biomarkers were found to have a prognostic role in different gastrointestinal cancers such as HOX transcript antisense intergenic RNA (HOTAIR), astrocyte elevated gene-1 (AEG-1), Human leukocyte antigen class I (HLA- I), and sequence-binding proteins 1 (SATB1).74–77 Our results revealed that NEDD9 plays a vital role in digestive system cancers (pooled HR: 1.92, 95% CI (1.46-2.53)). NEDD9 overexpression significantly predicts poor prognosis in gastric cancer, PC, and hepatocellular carcinoma. Zhang et al, 2014 51 study showed that the expression of NEDD9 was upregulated in gastric cancers compared to adjacent non-malignant gastric mucosae. Furthermore, it was correlated significantly with age, tumor size, tumor location, depth of invasion, TNM stage, lymph node involvement, and distant metastasis. Studies have also shown that NEDD9 facilitates hypoxia-induced gastric cancer cell migration via MICAL1 Related Rac1 Activation. 78
In colorectal cancer, NEDD9 emerges as a target gene of the TCF–β-catenin pathway, where heightened expression aligns with augmented migration and cancer advancement. 79 NEDD9 serves as a facilitator of epithelial-mesenchymal transition and fosters invasion in aggressive breast cancer contexts. 17 Recent research offers in vivo validation of NEDD9's role ins fueling oncogenic signaling pathways such as AKT, ERK, and SRC during breast cancer progression. 79 Our findings suggest that NEDD9 overexpression is associated with a poor prognosis in colorectal cancer, although not significantly. Limited information exists regarding the role of NEDD9 in PC, but existing data indicate elevated NEDD9 levels in PC patients, potentially serving a prognostic function.45,80–83 High NEDD9 expression levels were significantly associated with clinical staging, lymph node metastasis, and histological differentiation in PC patients. 45 However, in another study by Radulović et al, NEDD9 expression did not show statistically significant correlations with tumor stage and grade, gender, or patient survival. 82 Our findings suggest that NEDD9 overexpression is significantly associated with poor prognosis in PC (pooled HR: 1.64, 95% CI (1.02-1.62)).
In hepatocellular carcinoma, AFP is widely utilized in clinics worldwide for screening and serves as a crucial predictor of survival and tumor recurrence among serum AFP-positive HCC patients. 84 Lu et al, 2015 27 made a comparison between AFP and NEDD9. Among the patient subgroup where AFP levels did not predict prognosis or cancer recurrence, NEDD9 emerged as a robust indicator of both overall (HR: 2.42, 95% CI (1.59-3.69)). However, the generalizability of the results is difficult because there is limited information about the association between NEDD9 and prognosis.
Feng et al, 2012 85 showed that there is a significant correlation between NEDD9 expression and smoking. Recently, Miao et al, 2013 29 revealed an inverse correlation between NEDD9 expression and overall survival in patients with respectable NSCLC. Intriguingly, this correlation was observed independent of the TNM stage. Our pooled estimates showed that NEDD9 insignificantly predicts poor OS in NSCLC, overall, with a pooled HR effect of 1.31 (95% CI (0.60-2.83), I2 = 82%), but predicts good prognosis in LUAD subgroup (pooled HR = 0.70, 95% CI (0.54-0.90)). Our survival analysis consistently indicated favorable effect of NEDD9 overexpression on overall survival in TCGA LUAD cohort. However, we found insignificant prognostic outcomes with NEDD9 in LUSC cohort.
NEDD9 also strongly influences TGFβ signaling in breast cancer86–88; Fashena et al, 2002 89 demonstrated that NEDD9 induced cell motility and spreading in MCF7 cell lines. Additionally, Izumchenko et al, 2009 20 reported that the NEDD9-null genetic background significantly reduced mammary tumor cell initiation in the MMTV-polyoma virus middle T genetic model. Kong et al, 2011 17 identified NEDD9 as highly expressed in human breast cancers and facilitator of migration and invasion in TNBC. Furthermore, NEDD9 overexpression induced EMT and promoted interactions in-vivo with the E-cadherin promoter. These findings suggest that NEDD9 is associated with tumor progression and metastasis, and it also acts as a regulator in the EMT. 17 Our findings showed that NEDD9 was significantly associated with poor OS in patients with breast and gynaecological cancer (pooled HR: 2.30, 95% CI (1.25-4.22)). Also, NEDD9 expression significantly affected PFS (pooled HR: 2.70, 95% CI (1.62-4.49)).
In urinary bladder cancer, Harb et al, 2017 52 found that NEDD9 expression is related to aggressive behavior. It has been found that NEDD9 regulates MMP-2 and MMP-9 expression in transitional cell carcinoma, thereby enhancing invasion and metastasis. 90
In patients with melanoma, NEDD9 was not found to be correlated with any of the following clinicopathological parameters: lymph node involvement, site of lesion, stage, sex, histology, Clark invasion level, Breslow thickness, response to therapy and presence of ulceration or regression. 50 However, contradictory effects have been observed; for example, while overexpression was highly protective in uveal melanoma, it was harmful in HNSCC and melanoma. In addition, our bioinformatics analysis has indicated that NEDD9 is involved in the activation and sometimes inhibition of the same cancer-related pathways in different cancers. Such pathways included apoptosis, hormone (AR/ER) signaling, EMT, and some kinase pathways like RTK and PI3 K/Akt signaling. This could indicate a biological role through different signaling pathways, but more research is needed to elucidate the precise molecular mechanisms accounting for the variations and paradoxical relations across different cancers. Limited information exists regarding the role of NEDD9 in bone cancer (Osteosarcoma), but previous data indicated elevated NEDD9 levels in osteosarcoma patients, potentially serving a prognostic function. 61 In addition, for LAML, a previous study showed that NEDD9 overexpression predicts poor prognosis and has significant links with FAB classifications. 56
High NEDD9 expression in tumors correlates with altered immune infiltration, impacting cancer progression. 20 NEDD9 influences immune checkpoints like PD-1 and CTLA-4, promoting immune evasion. 91 It also regulates cytokine secretion, affecting immune cell recruitment and activation within the tumor microenvironment.92,93 Also, our results showed that immune infiltration was variably affected by NEDD9 expression in different cancers, indicating a potential immunomodulatory role that requires further investigation. Notably, NEDD9 expression showed a robust positive correlation with CD4+ T cells in cancers such as Breast, Colon Adenocarcinoma, and Lung Adenocarcinoma, and consistent negative correlation with effector memory T cells (TEM) across Bladder, Breast, and Lung Squamous Cell Carcinoma. These findings align with the patterns observed in broader prognostic studies, where higher NEDD9 expression often correlates with poorer outcomes in terms of OS and PFS, given that in colon and lung carcinomas, CD4+ T cell infiltration correlates with a favorable prognosis, whereas in breast and renal cancers, including bladder cancer, CD4+ T cell infiltration correlates with poor survival.94,95 TEM cells have been associated with better prognosis in cancer patients. 96 The detailed correlation patterns across cancers, revealing both positive and negative associations with various immune cells, may reflect NEDD9's role in modulating the tumor microenvironment, potentially impacting immune surveillance and tumor progression. For instance, the positive correlation with CD4+ T cells might indicate a role in enhancing immune response, which theoretically could correlate with better outcomes; however, the negative correlation with effector memory cells suggests a potential suppression of long-term immunity, and this might contribute to the poor prognostic outcomes observed in the meta-analyses. Furthermore, CAFs, previously shown to have roles in cancer progression and metastasis, 97 consistently and robustly had increased infiltration with higher NEDD9 expression in several cancers, as indicated by our analysis, supporting the poor prognostic outcomes associated with NEDD9 overexpression. However, CAFs infiltration was not significantly associated with NEDD9 expression in UVM, aligning with the favorable prognosis of NEDD9 in this cancer. Furthermore, MAIT cell infiltration correlates with NEDD9 expression, especially the negative correlations in UVM and BLCA, and the positive correlations in COAD, STAD, LUAD, LUSC, which are consistent with the previous findings indicating that MAIT cells can have multifaceted dual roles in cancer, in addition to their role in modulating immune molecules like PD-1, CTLA-4, and TM-3, 98 thus suggesting a route through which NEDD9 can have effect on response to immunotherapies. Finally, the negative correlation between NEDD9 expression and MDSC infiltration in cancers like LUAD, and the positive correlation in LIHC, are consistent with what is known about MDSCs having tumor-promoting activity via immune suppression and indirect CAF activation.99,100 The prognostic outcomes associated with NEDD9 in such cancers show some paradoxical relations in cancers like BRCA, where high NEDD9 expression predicts poor prognosis but is associated with lower MDSC infiltration; this supports the idea that the immune-modulating role of NEDD9 in cancers is very complex and not straightforward, with several immune cells infiltrating and having similar or opposing effects on tumor progression and prognosis.
To the best of our knowledge, our study is the first meta-analysis to comprehensively evaluate the prognostic role of NEDD9 in human cancers along with TCGA and bioinformatics validation. However, there are some limitations to our study. First, we did not include any studies from the Middle East, Africa, or Australia due to limited publications about NEDD9 there. Second, not all studies adjusted for stage in their multivariate models, which may affect the results, and some studies have only reported univariate statistics, thus presenting a potential bias especially if late-stage patients were included. Between-study heterogeneity in terms of detection methods used for NEDD9 expression profiling is significant limitation also. Third, despite utilizing subgroup analysis, the high heterogeneity of the studies and potential publication bias remain concerns, especially for overall survival estimates. To address these issues, future research should focus on multicenter trials with standardized outcome measures and include more diverse patients.
Conclusions and Future Perspectives
Our study examined the prognostic profile of NEDD9 in various human cancers, supported by meta-analysis and bioinformatics validation. We suggest NEDD9 as a significant prognostic biomarker, especially in gastric, breast and gynecological, and urinary system cancers, where its overexpression predicts poor prognosis. Our meta-analysis also suggested better overall survival with NEDD9 expression in cancers like lung adenocarcinoma. Despite some heterogeneity across cancer types, NEDD9 dysregulation is generally associated with survival outcomes. Our results underscore NEDD9's roles in tumorigenesis, from promoting metastasis to affecting immune infiltration, highlighting it as a potential therapeutic target. Further research is needed to explore NEDD9's complex interactions and mechanisms, particularly in cancers with opposing results. Future studies should include diverse populations and standardized methods to validate NEDD9's prognostic significance. Additionally, developing targeted therapies, such as NEDD9 inhibitors or monoclonal antibodies, could be beneficial once its role in cancer prognosis is sufficiently understood.
Footnotes
Abbreviations
Acknowledgements
None.
Conflict of Interest Statement
Anwaar Saeed reports a consulting or advisory board role with AstraZeneca, Bristol-Myers Squibb, Merck, Exelixis, Pfizer, Xilio therapeutics, Taiho, Amgen, Autem therapeutics, KAHR medical, and Daiichi Sankyo; institutional research funding from AstraZeneca, Bristol-Myers Squibb, Merck, Clovis, Exelixis, Actuate therapeutics, Incyte Corporation, Daiichi Sankyo, Five prime therapeutics, Amgen, Innovent biologics, Dragonfly therapeutics, Oxford Biotherapeutics, Arcus therapeutics, and KAHR medical; and participation as a data safety monitoring board chair for Arcus therapeutics. The remaining authors have no relevant financial interests to disclose.
Credit Authorship Statement
RO, JY, FQ: Data extraction, analysis, screening, and scientific writing. MT, MA: Screening, and scientific writing. RO, JY, OY, LK, AS: Study concept, design, and drafting of the manuscript. All authors contributed to the article and approved the submitted version.
Data Availability Statement
The data generated in this study are available upon request from the corresponding author.
Ethical Considerations
No ethical approval was required for this meta-analysis and secondary research based on data from public databases.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
