Abstract
Background
A clinical challenge in esophageal squamous cell carcinoma (ESCC) remains the lack of applicable plasma biomarkers for screening and diagnosis. Circular RNAs (circRNAs) hold great potential as biomarkers for cancer. The study aims to explore a circRNA as a potential plasma biomarker for screening strategies and diagnostic approaches to ESCC.
Methods
Upregulated circRNAs were identified through RNA sequencing, with circCYP24A1 being identified as the target circRNA. Fluorescence in situ hybridization was employed to detect the expression of circCYP24A1 in ESCC tissue microarrays, aiming to assess the expression of circCYP24A1 in a large population and its correlation with clinical indicators. Subsequently, qRT-PCR analysis was performed on plasma samples from both ESCC patients and healthy controls to evaluate the expression levels of circCYP24A1, exploring its potential as a biomarker. Finally, the functions of circCYP24A1 were validated through CCK-8 assay, wound healing assay, trans-well assays and western blot assays.
Results
CircCYP24A1 demonstrated upregulation in both plasma and tissues, exhibiting correlations with lymph node metastasis, TNM staging, and prognosis in ESCC. The circCYP24A1 achieved a perfect area under the curve of 0.94 for the diagnosis of ESCC, and an area under the curve of 0.76 for the prediction of lymph node metastasis. Furthermore, functional loss assays revealed that circCYP24A1 effectively promotes the epithelial-mesenchymal transition and tumor metastasis in vitro.
Conclusions
CircCYP24A1 emerges as a potential plasma diagnostic biomarker and a predictive factor for LNM for ESCC.
Introduction
Esophageal cancer is the sixth leading cause of cancer-related deaths and esophageal squamous cell carcinoma (ESCC) is the most common histological type in Asia. 1 The prognosis of ESCC patients is closely associated with tumor staging at diagnosis. 2 It is reported that the 5-year survival rate after surgical resection of early ESCC can reach above 90%, but the overall 5-year survival rate of patients in the middle to late stages is only 6% to 15%. 3 However, ESCC patients usually have no obvious symptoms in the early stages, and the vast majority of patients are diagnosed only after experiencing progressive dysphagia or metastatic symptoms, by which time the tumor has often reached the middle to late stage, accompanied by regional lymph node metastasis (LNM) or even distant metastasis. 4 Therefore, screening for ESCC, especially among populations in high-incidence regions, is helpful for early detection, early diagnosis and early treatment of ESCC, thus reducing its mortality rate. Gastroscopy stands as the most effective screening method for ESCC, but its acceptance is low and its application in large scale population screening is limited due to its invasiveness and high cost. 5 Plasma tumor biomarker examination is handy and non-invasive, which is an ideal method for tumor screening. However, conventional blood biomarkers for ESCC such as squamous cell carcinoma antigen (SCC-Ag), cytokeratin 19 fragment, and carbohydrate antigen 72-4 lack satisfactory specificity and sensitivity. 6
LNM is the major cause of recurrence and poor prognosis in ESCC patients. 7 Treatment is no longer limited to the resection of the primary tumor lesion, but also requires lymph node dissection after developing LNM. Yet, it increases the complexity and risk of the surgery, while also requiring a more comprehensive treatment plan to deal with possible complications. At present, the evaluation of LNM in ESCC mainly relies on imaging examinations, such as endoscopic ultrasound, computed tomography (CT), fluorodeoxyglucose-positron emission tomography. 8 However, the existing technologies do not have satisfactory sensitivity and specificity in diagnosing LNM, thereby influencing the formulation of treatment plans and the prognosis of patients. Therefore, there is an urgent need to explore plasma biomarkers for diagnosis of ESCC and prediction of LNM, in order to improve screening and diagnostic strategies and facilitate treatment.
In recent years, circular RNA (circRNA) has gained widespread attention as a promising non-invasive biomarker for cancer. 9 CircRNAs are single-stranded circular molecules formed by pre-mRNA through reverse splicing which play significant roles in proliferation, apoptosis, migration, invasion, and drug resistance in various cancers.10‐13 The inherent characteristics of circRNAs endow them with immense potential as tumor biomarkers. First, circRNAs exhibit relatively high expression levels in cells and display tissue-specific and disease-specific patterns. 14 Besides, the circular structure of circRNA confers resistance to Ribonuclease R (RNase R) degradation, making it more stable in extracellular environments compared to linear RNA. 15 In addition, circRNAs can accumulate and be detected in extracellular vesicles, human peripheral blood and other bodily fluids. 16 At present, some circRNAs differentially expressed in tumors have been widely characterized as biomarkers in various studies. For example, cerebellar degeneration-related protein 1 antisense and hsa_circ_0015286 have been studied as biomarkers for diagnosis and prognosis in gastric cancer.17,18 and circIMMP2L has been reported as a biomarker for diagnosis, prognosis and LNM in ESCC. 3 But few circRNA biomarkers were utilized to predict ESCC in clinical practice until now. Consequently, circRNAs still deserve further investigation as ESCC biomarkers.
Hence, we performed a study to find a potential plasma circRNA biomarker that could be used for the diagnosis of ECSS and prediction of LNM.
Materials and Methods
Specimen Collection
This study collected plasma samples from 70 patients with ESCC, 8 patients with esophagitis and 40 healthy individuals. CircCYP24A1 has been reported to be upregulated in prostate cancer (PCa) 19 and cutaneous squamous cell carcinoma (cSCC). 20 In China, the age-standardized incidence rate for PCa is 9.68%, while non-melanoma skin cancer is much lower. 21 Therefore, we also collected the plasma of 5 PCa patients in order to verify the expression of circCYP24A1 in the plasma of patients with other tumor types. All the volunteers were sourced from the Second Clinical Medical College of North Sichuan Medical College. All the samples were obtained after getting informed consents of patients and/or their legal guardian, and the utilization of clinical specimens has been granted approval by the Ethics Committee of North Sichuan Medical College in accordance with the Declaration of Helsinki on June 22, 2021 (Approval number: [2021]42) in Nanchong, China. The staging of ESCC follows the eighth edition of the UICC/AJCC TNM staging system for esophageal cancer, established in 2017. The “LNM positivity criterion” was established based on histopathological findings and imaging examination. Pathological biopsy of lymph nodes revealing ESCC cells, or ultrasound examination of supraclavicular lymph nodes and CT scanning of neck, chest and abdomen regions revealing abnormal lymph node size, morphology, echo characteristics, blood flow, density, enhancement mode and other manifestations such as enlarged lymph nodes, a morphological index >0.5, irregular morphology, blurred edges, disappearance of the portal structure, uneven echo, rich blood flow, irregular circular enhancement with central low-density shadow, etc were considered positive for LNM. N0 indicates the absence of regional lymph node involvement, while N1-N3 represent varying degrees of LNM.
The inclusion criteria for the healthy control group were adult individuals who had no evident abnormalities in chest X-ray or chest CT scans, gastroscopy, as well as tumor marker tests. The patient group of ESCC patients consisted of untreated primary ESCC patients diagnosed by histopathological examination, aged 18 to 80 years, with a clear LNM area and a complete medical history. The exclusion criteria for ESCC patients were patients with an unclear medical record diagnosis, patients with non-primary ESCC or primary non-squamous cell carcinoma, those with serious underlying disease, those who had received radical resection surgery or chemoradiotherapy, patients with diseases causing pathological changes of lymph nodes such as infection, autoimmune diseases, tuberculosis, etc, and those unable to complete the follow-up. The inclusion and exclusion criteria for Pca patients were consistent with those of ESCC. And patients with esophagitis were diagnosed through gastroscopy and brush biopsy.
Peripheral Blood Collection
Using EDTA anticoagulant vacuum collection tubes (purple tubes), approximately 2 mL of fasting venous blood was collected from patients and healthy adults in the morning, followed by centrifugation in a low-temperature high-speed centrifuge at 4 °C, 1900 ×g (3000 rpm) for 10 min. After centrifugation, the supernatant (plasma) was aspirated with a pipette and transferred to RNase-free tubes, which were then stored in a −80 °C freezer for future use.
Cell Culture and Transfection
ESCC cell lines TE-1 (CSTR TCHu89, RRID: CVCL_1759) and KYSE-150 (CSTR TCHu236, RRID: CVCL_1348) were obtained from the Cell Bank/Stem Cell Bank, Chinese Academy of Sciences (Shanghai, China). KYSE-410 (DSMZ ACC-381, RRID: CVCL_1352) was purchased from Fenghui Biological Co. Ltd (Changsha, China), which introduced KYSE-410 from German Collection of Microorganisms and Cell Cultures. Human esophageal epithelial cell line Het-1A (ATCC CRL-2692, RRID: CVCL_3702) and ESCC cell line KYSE-510 (JCRB JCRB1436, RRID: CVCL_1354) were obtained from Mengjin Biological Co. Ltd (Shanghai, China), which introduced KYSE-510 from the Japanese Collection of Research Bioresources Cell Bank and Het-1A from the American Type Culture Collection. All cell lines were authenticated by STR profiling and free of mycoplasma contamination. Cells were cultured in RPMI-1640 medium or DMEM (GIBCO, USA) supplemented with 10% fetal bovine serum (GIBCO, USA) at 37 °C in a 5% CO2 incubator. SiRNA targeting circCYP24A1 was designed and synthesized by GenePharma (Shanghai, China), the sequence of si-circCYP24A1 was as follows: sense AUAAUACGCCUCAGGGAAGTT, antisense CUUCCCUGAGGCGUAUUAUTT. After cells reached 60% confluency following plating, si-circCYP24A1 was transfected into the cells using Lipofectamine™ 2000 (Invitrogen, USA).
RNA Sequencing (RNA-seq)
A total of six matched ESCC tissues and normal tissues surrounding the tumor were collected for RNA-seq,22,23 the obtained fresh tissue had been cryogenically preserved in liquid nitrogen within 30 min after surgery for future use. And all the volunteers had received no prior treatment. Total RNA extraction and sequencing were performed by Aksomics Inc. (Shanghai, China) with standard procedures. The total RNA was checked by agarose electrophoresis and quantified by nanodrop. After removing rRNA and digesting linear RNA, the circRNA sequencing library was constructed by the KAPA Stranded RNA-Seq Library Prep Kit (Illumina, USA), and sequenced by Illumina Hiseq 4000 (Illumina, USA). The sequencing data were uploaded in the NCBI's Gene Expression Omnibus database (GSE263647). The junction sites of circRNAs were detected by comparing the sequencing data to the known transcriptome in the STAR, and then the expression quantification of circRNAs (the backsplice junction reads count) was calculated by CIRCexplorer2. The circRNAs with a CPM mean above 100 in each group were considered as expressed in the group.
Quantitative Real-Time Reverse Transcription-Polymerase Chain Reaction (qRT-PCR)
Plasma and cell total RNA were extracted using the MiRNeasy Serum/Plasma Kit (QIAGEN, Germany) and the FastPure Cell/Tissue Total RNA Isolation Kit (Vazyme, China), respectively. Reverse transcription of the RNA into complementary DNA (cDNA) was carried out using the Hiscript III first strand cDNA Synthesis Kit (Vazyme, China). PCR was conducted using the AceQ qPCR SYBR Green Master Mix (Vazyme, China).
Detection and Analysis of the Plasma circCYP24A1
Each 200 μL of plasma was extracted to obtain 10 μL of total RNA, all of which were reverse transcribed to obtain 20 μL of cDNA. The undiluted cDNA was amplified according to the cycle reaction parameters shown in Supplementary Table 2. In addition, total RNA from KYSE-510 cells was extracted, and 1μg of RNA was reverse transcribed, and the resulting cDNA was diluted 10 times and used for future use. The primer sequences were displayed in Supplementary Table 3. With GAPDH as the internal reference, total RNA of KYSE-510 cells was introduced into each batch of samples during the qRT-PCR process as the control group to facilitate data normalization. The relative changes in gene expression were determined using the 2−ΔΔCt method. We collected clinical data of 70 ESCC patients, divided them into low expression group (n = 35) and a high expression group (n = 35) based on the median expression level of plasma circCYP24A1, and analyzed the correlation between clinical data and plasma circCYP24A1. Patients were followed up once a month from the time of blood collection for 13 months to analyze the effect of plasma circCYP24A1 content on patients’ survival, and the follow-up rate was 98.6%. The receiver operating characteristic (ROC) curve analysis was used to calculate the area under the curve (AUC), sensitivity, and specificity to evaluate the ability to distinguish ESCC patients from health control groups and predict LNM.
RNase R Assay
RNase R (Biosearch, USA) is used to digest RNA. 1 μg of RNA was treated with 3 U of RNase R at 37 °C for 30 min. The changes of circCYP24A1 and CYP24A1 abundance before and after digestion were detected by qRT-PCR.
Tissue Microarrays (TMA) and Fluorescence in Situ Hybridization (FISH) Assay
The TMA (HEsoS180Su11) was obtained from Shanghai Outdo Biotech Co., Ltd (Shanghai, China) with the tissue samples obtained from the National Human Genetic Resources Sharing Service Platform. The chip contained data on 108 cases of ESCC with 72 cases having corresponding adjacent non-cancerous tissue. The human tissues obtained in TMA were approved by the Ethics Committee of Shanghai Outdo Biotech company in accordance with the Declaration of Helsinki on April 1, 2020 (Approval number: SHYJS-CP-2004003) in Shanghai, China. The data was anonymous, and thus the requirement for informed consent was waived. Probes mixes were designed for tissue FISH assay targeting circCYP24A1. The probe mixes sequences are displayed in Supplementary Table 3. The RNA FISH kit SA-Biotin System (paraffin sections) was purchased from GenePharma (Shanghai, China) and used according to the provided instructions. Panoramic multispectral scanning of slides was performed by the Tissue- FAXS Spectra (TissueGnostics, Austria) and we imported the data into Strata-Quest analysis software to calculate the positive rate of circCYP24A1. The results of FISH showed that one of the 108 cancer tissues lacked ESCC tissue and one of the 72 corresponding adjacent non-cancer tissues lacked normal esophageal epithelial tissue. So, there were a total of 107 ESCC tissues and 71 normal esophageal epithelial tissues, with 70 pairs of matched cancerous and adjacent normal epithelial tissues in our subsequent statistical analysis. Taking the median of the positive rate of circCYP24A1 as the cutoff value, ESCC patients were stratified by low (n = 54) and high (n = 53) expression of circCYP24A1, and correlation analysis was performed according to the collected clinical data and follow-up information. The 5-year survival rate is defined as the time from the date of surgery to the date of death or 60 months. The follow-up rate was 99.0%. The impact of different circCYP24A1 expression groups on the 5-year survival rate of ESCC patients was analyzed by survival analysis.
Actinomycin D Assay
To assess RNA stability, 2 μg/mL Actinomycin D was added to KYSE510 and KYSE150 cells when their fusion degree reached approximately 50%. The cells were treated with Actinomycin D for 4, 8, 12 and 24 h, respectively. A control group without intervention was set at each time point. After reaching the corresponding time points, the cells were collected and total RNA was extracted. Reverse transcription was performed to synthesize cDNA, followed by qRT-PCR to detect the molecular levels of circCYP24A1 and CYP24A2.
Cell Proliferation Assay (CCK-8)
Cell viability was assessed via the CCK-8 assay (Kyegen, China) following kit directions. In brief, 100 μL cells/well (approximately 3 × 103 cells) were seeded in 96-well plates and incubated at 37 °C in a 5% CO2 cell culture incubator. At different time points (0, 24, 48, 72, 96 h), after preparing the culture medium and CCK-8 reagent in a ratio of 10:1, 100 μL of the mixture was added to each well and incubated for 1 h in the dark at 37°C. Absorbance was measured at 450 nm via microplate reader.
Wound Healing Assay
Cells treated with siRNA were seeded in a 6-well plate. After 24 h of incubation, two perpendicular baseline lines were drawn in the cells using a 10 μL tip. Photographs were taken at 0 and 24 h after scratching. Scratch healing rate (%) = [(0 h scratch width-24 h scratch width)/ 0 h scratch width] × 100%.
Trans-Well Assay
100 μL of serum-free culture medium containing 3 × 104 cells was added to the upper chamber of the trans-well. In the lower chamber, 600 μL of 1640 culture medium containing 20% fetal bovine serum was added. For the invasion assay, Matrigel (BD biosciences, USA) was diluted 1:4 with serum-free culture medium, and 40 μL of the mixture was evenly spread on the upper chamber. After being kept at a temperature of 37 °C for 1 h, the trans-well chamber containing the mixture added 100 μL of serum-free medium containing 5 × 104 cells. After 24-48 h of incubation (48 h for KYSE-510, 24 h for KYSE-150), the cells were fixed with 20% methanol for 30 min, followed by staining with 0.1% crystal violet solution. The cells in the upper chamber were gently wiped off, photographed, and counted.
Western Blot Analysis
ESCC cells were lysed using RIPA protein lysis buffer, followed by a 30-min incubation on ice, centrifugation and collection of the supernatant. Proteins quantitated by BCA were subjected to 10% SDS-PAGE analysis. The proteins were transferred onto a PVDF membrane (Millipore, USA). The membrane was blocked with 5% skim milk for 1 h and incubated overnight at 4 °C with antibodies against N-cadherin (1:500, 22018-1-AP, Proteintech, China), E-cadherin (1:10000, #ET1607-75, HUABIO, China), snail (1:1000, #13099-1-AP, HUABIO, China), vimentin (1:500, #ET1610-39, HUABIO, China), and GAPDH (1:50000, ET1601-4, HUABIO, China). After incubation, the membrane was treated with HRP-conjugated secondary antibodies at room temperature for 1 h, washed with PBST and exposed in a darkroom after adding the ECL chromogenic solution.
Statistical Analysis
The experimental data were analyzed using GraphPad Prism 9.0 statistical software. Normally distributed measurement data were expressed as (
Results
Identification and Characterization of the circCYP24A1
We had obtained the expression profile of circRNAs in ESCC tissues by conducting RNA-seq analysis on six pairs of ESCC tissues and adjacent normal tissues. A total of 2228 circRNAs were identified, and 131 down-regulated circRNAs and 54 up-regulated circRNAs were selected by the criteria of |Log2FC|≥1 and P < .01 (Figure 1A-B). We focused on the up-regulated circRNAs in ESCC, as they exhibited higher expression levels which makes them more detectable in tissues and blood. CircCYP24A1 emerged as the most significantly up-regulated among the differentially expressed circRNAs (Figure 1B).

Identifcation of the circCYP24A1 up-regulated in ESCC and its characterization. A Clustered heatmap for differentially expressed circRNAs. Cut of is |Log2FC|≥1 and P < .01. B Volcano plot compared the expression fold changes of circRNAs for six paired samples of ESCC by RNA-Seq. Right upper panel represented the up-regulated circRNAs, the dot pointed by the arrow indicate circCYP24A1. C The expression of circCYP24A1 in Het-1A and ESCC cells was detected by qRT-PCR. D The schematic representation of circCYP24A1 formation and Sanger sequencing results of the circCYP24A1 backsplice junction site. E The gel data indicating the detection of circCYP24A1 using divergent and convergent primers from complementary DNA or genomic DNA . F The resistance of circCYP24A1 and CYP24A1 to RNase R cleavage was evaluated by qRT-PCR. G The relative RNA levels of circCYP24A1 and CYP24A1 treated with Actinomycin D (5 μg/mL) was detected by qRT-PCR. Data are presented as the mean ± SD (n = 3). *P < .05, **P < .01, ***P < .001, ****P < .0001. P values were determined using the unpaired Student's t-test.
Due to the limitations of high-throughput sequencing, the identification of circRNAs often comes with a higher rate of false positives. 24 Thus, to demonstrate the authenticity of circCYP24A1 as a up-regulated circRNA in ESCC, a series of validations were conducted on its characteristics. Firstly, the sequencing results were further validated by qRT-PCR, which confirmed that circCYP24A1 was highly expressed in ESCC lines compared to Het-1A (Figure 1C). According to circBase annotation, CircCYP24A1 is located at chr20: 52773707-52788209 and is derived from the exons 3-11 of cytochrome P450 family 24 subfamily A member 1 (CYP24A1), with a length of 1106 bp. Divergent primers were used to amplify circCYP24A1, Sanger sequencing confirmed the splicing junctions which consistent with the annotation in circBase (Figure 1D). Next, convergent and divergent primers were used to amplify cDNA and genomic DNA from KYSE-510, Agarose gel electrophoresis revealed that both primer pairs yielded products when using cDNA as the template, while only the convergent primer pair showed amplification when using genomic DNA as the template, indicating that the detected circCYP24A1 was generated through reverse splicing rather than genomic recombination (Figure 1E). Further, the stability of circCYP24A1 was assessed through RNase R digestion and actinomycin D treatment. The results demonstrated that circCYP24A1 exhibited resistance to RNase R while CYP24A1 was degraded by RNase R (Figure 1F). Finally, after treatment with actinomycin D, circCYP24A1 displayed greater stability compared to its parental host gene CYP24A1 (Figure. 1G). Collectively, these results indicated that circCYP24A1 is a real circRNA.
Elevated Expression of circCYP24A1 is Related to LNM, Advanced Clinical Stage, and Poor Outcomes of ESCC
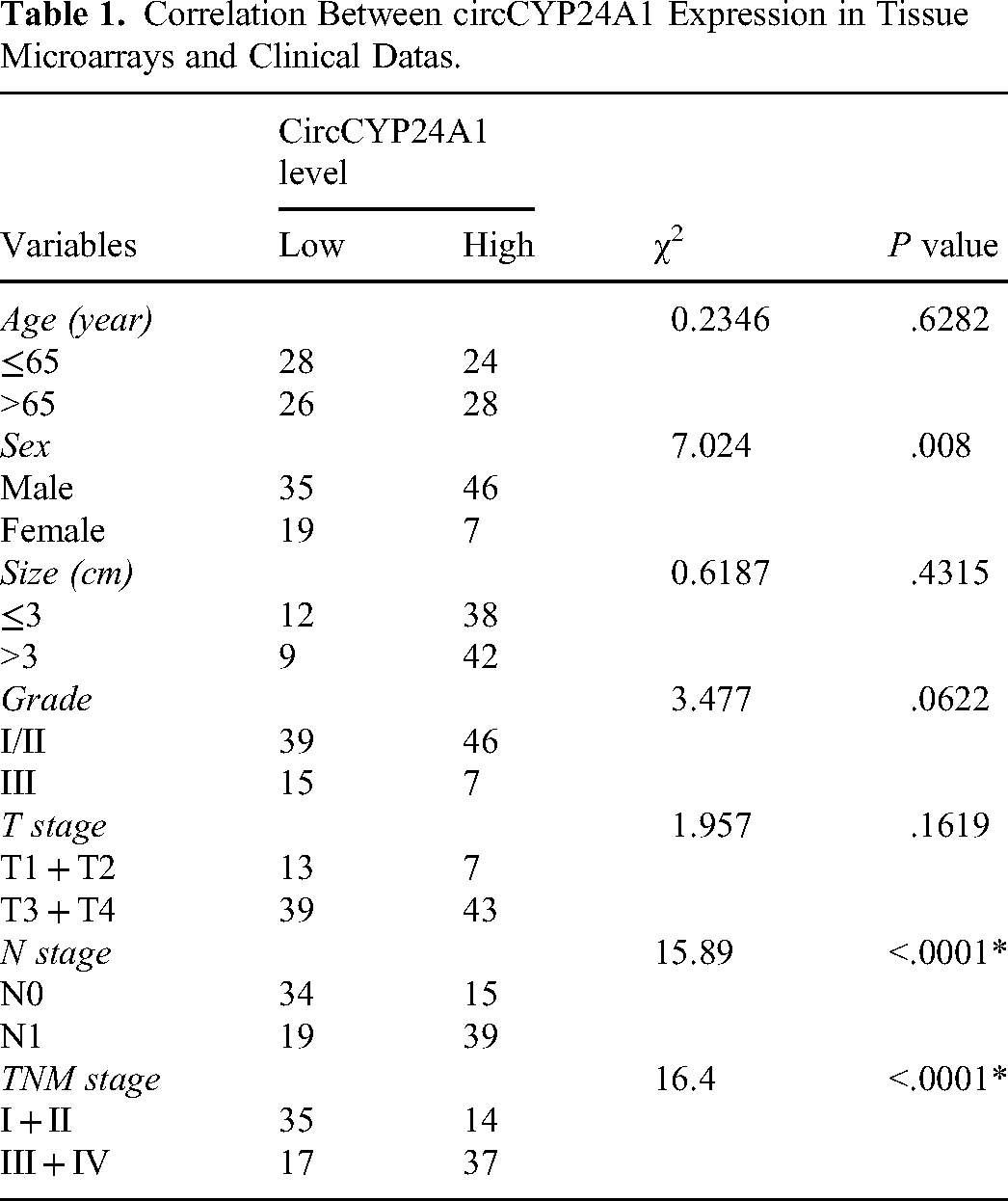
To assess the expression level of circCYP24A1 in a larger population, we designed probes specifically targeting circCYP24A1 and utilized FISH to detect the expression of circCYP24A1 in tissues based on TMA. The results demonstrated a significant up-regulation of circCYP24A1 expression in ESCC tissues compared to adjacent normal tissues (Figure 2A-B). The median of positivity rate of circCYP24A1 was 2.24% as a cut-off value, the circCYP24A1 positivity rate ≤ 2.24% was categorized as the “Low level circCYP24A1” group, while the positivity rate >2.24% was classified as “High level circCYP24A1” group. A high level of circCYP24A1 was associated with N and TNM stage, but not with age, tumor size, and distant metastasis in ESCC tissues (Table 1). Kaplan-Meier analysis indicated that a high level circCYP24A1 had a shorter 5 – year survival rate (Figure 2C). The aforementioned findings indicated that circCYP24A1 exhibited elevated expression in ESCC tissues, which was significantly associated with LNM, advanced clinical stage, and poor prognosis.

CircCYP24A1 is up-regulated in ESCC tissues and associated with poor prognosis. A The representative fluorescence images of circCYP24A1 (red) in ESCC and adjacent normal tissues by FISH analysis based on TMA. Scale bar, 100 μm. B The relative expression of circCYP24A1 in paired and unpaired ESCC tissues and adjacent normal tissue in TAM. C Kaplan-Meier curves show the association between circCYP24A1 expression and 5-year overall survival in 106 ESCC patients. Data are presented as the mean ± SD. *P < .05, **P < .01, ***P < .001, ****P < .0001. P values were determined with the two-tailed paired and unpaired Student's t-test (B), log-rank test (C).
Correlation Between circCYP24A1 Expression in Tissue Microarrays and Clinical Datas.
Plasma circCYP24A1 is a Promising Potential Biomarker for the Diagnosis and Prediction of LNM in ESCC
According to the exoRBase database (http://www.exorbase.org/), we learned that circCYP24A1 is an exosomal circRNA that can be detected in human body fluids. We assessed the expression levels of circCYP24A1 in the plasma of ESCC patients, PCa patients, esophagitis patients and healthy individuals. Compared with the healthy controls, the expression of circCYP24A1 in plasma of ESCC patients was significantly upregulated (Figure 3A), while there was no significant difference in circCYP24A1 levels in the plasma of PCa patients and esophagitis patients (Figure 3B). The expression of circCYP24A1 in plasma was normalized, the median expression circCYP24A1 in the plasma of 70 ESCC patients was 1.565 as the cut-off value, samples with circCYP24A1 expression ≤ 1.565 were classified as the “low expression” group, while those >1.565 were classified into the “high expression” group. Notably, it was found that the high expression level of circCYP24A1 was associated with LNM and advanced clinical stage of the ESCC patients, while being independent of age, gender, smoking history and tumor size (Table 2). Further analysis revealed that patients in advanced stages (III/ IV) had significantly higher expression level of circCYP24A1 than patients in earlier stages (I/ II) (Figure 3C), and the level of circCYP24A1 in the LNM patients (N1-3) was significantly higher than patients non-LNM (N0) (Figure 3D). When ESCC patients were stratified according to the median expression of circCYP24A1 during the follow-up period of 13 months, Kaplan-Meier analysis revealed that patients with high circCYP24A1 levels had significantly lower overall survival compared to those with low levels (Figure 3E).

CircCYP24A1 showed elevated expression in the plasma of ESCC patients, exhibiting a positive correlation with LNM, advanced stage, and poorer survival rates. A The relative expression levels of circCYP24A1 in the plasma of ESCC patients (n = 70) and healthy controls (n = 40). B The relative expression levels of circCYP24A1 in the plasma of PCa patients (n = 5), esophagitis patients (n = 8) and healthy controls (n = 40). C The relative expression levels of circCYP24A1 in the plasma of patients with different stages of ESCC and healthy controls. Early stage (I/II, n = 49) and advanced stage (III/IV, n = 20). D The relative expression levels of circCYP24A1 in the plasma of patients with non-LNM (n = 46) and LNM (n = 23) compared to the healthy control group. E The correlation between the expression of circCYP24A1 in the plasma of 70 ESCC patients and the overall survival rate was demonstrated by the Kaplan-Meier curve. F The ROC curve analysis of circCYP24A1 differentiating ESCC patients and healthy controls. G The ROC curve analysis of circCYP24A1 differentiating ESCC patients with LNM and without LNM. **P < .01, ***P < .001, ****P < .0001. Data are presented as the mean ± SD. P values were determined with the unpaired Student's t-test(A-D), log-rank test (E).
The Correlation Between circCYP24A1 Expression in Plasma and Clinical Data.

We evaluated the ability of circCYP24A1 to diagnose ESCC and predict LNM through ROC analysis. As shown in Figure 3F, the ROC curve analysis of circCYP24A1 in distinguishing ESCC patients and healthy controls showed an AUC of 0.94 (95% CI 0.90-0.98), the sensitivity and specificity were 0.84 and 0.93, respectively. And Figure 3G showed that circCYP24A1 could be used as a significant parameter to discriminate between non-LNM and LNM ESCC patients with an AUC of 0.76 (95% CI: 0.65∼0.87), the sensitivity and specificity both were 0.70.
In conclusion, elevated circCYP24A1 in the plasma of ESCC patients was significantly related to LNM and advanced clinical stages as well as shorter overall survival, and circCYP24A1 was a potential biomarker for diagnosis and predicting LNM.
CircCYP24A1 Promotes ESCC Cells Migration, Invasion and EMT
In order to investigate the biological functions of circCYP24A1 in ESCC, we successfully knocked down circCYP24A1 in KYSE-510 and KYSE-150 cells (Figure 4A). Regrettably, we found that the knockdown of circCYP24A1 did not affect the proliferative capacity of ESCC cells (Supplementary Figure 1). To determine whether the expression level of circCYP24A1 influences cell motility, we conducted the wound healing assay, in which the control group showed significantly smaller gap sizes than the knockdown group, with the healing rate being the opposite (Figure 4B). The transwell assay also indicated that the depletion of circCYP24A1 reduced the migratory activity of cells and decreased the invasive activity when a matrigel was added (Figure 4C). Furthermore, we assessed whether circCYP24A1 could enhance the epithelial-mesenchymal transition (EMT) in ESCC cells. Western blot analysis revealed that the knockdown of circCYP24A1 in ESCC cells resulted in an increase in the expression of the epithelial marker E-cadherin, while N-cadherin, vimentin and snail significantly decreased (Figure 4D). Taken together, circCYP24A1 promotes ESCC cells’ migration, invasion, and EMT.

CircCYP24A1 promotes migration, invasion and EMT of ESCC cell. A QRT-PCR analysis of circCYP24A1 after knocking down circCYP24A1 in KYSE510 and KYSE150. B Scratch healing assay was performed on ESCC cells from different treatment groups. C Transwell assay was used to determine cell migration and invasion abilities. D Levels of EMT marker proteins in ESCC cells from different treatment groups were assessed. Data are presented as the mean ± SD (n = 3). **P < .01, ***P < .001, ****P < .0001.P values were determined with the unpaired Student's t-test (A, C and D), two-tailed paired Student's t-test (B).
Discussion
Esophageal cancer has consistently posed a significant threat to human health. 25 The two major histological types of esophageal cancer are ESCC and esophageal adenocarcinoma, among which ESCC is the most common histological type in China. 26 We need to intensify screening for ESCC, strive for earlier detection and treatment to improve prognosis. In recent years, due to their stability, abundance, cell and tissue specific expression, as well as detectability in liquid biopsy samples, circRNAs have strong potential as diagnostic,and prognostic biomarkers.27,28 CircCYP24A1 has been reported to play a significant role in various types of tumors. For example, circCYP24A1 is up-regulated in PCa, cSCC and ESCC, promoting the proliferation and migration of cSCC and ESCC cells, and enhancing Docetaxel resistance in PCa cells.19,20,29 However, circCYP24A1 is down-regulated in renal cell carcinoma and inhibits its malignant phenotype. 30 This may be due to the tissue and disease specificity of circCYP24A1. But, these studies investigated the underlying mechanism, while the diagnostic performance based on circCYP24A1 remains largely undetermined in ESCC.
In this study, we have confirmed circCYP24A1 is up-regulated in the plasma and tissues of ESCC patients. Especially, in the plasma, the expression levels of circCYP24A1 in ESCC patients were significantly higher than those of healthy individuals, and in advanced patients were higher than those in early patients. Based on our ROC curves, the AUC for plasma circCYP24A1 in diagnosing ESCC was 0.94, sensitivity and specificity were 0.84 and 0.93, respectively. Compared to the classic ESCC biomarker SCC-Ag (AUC: 0.72, sensitivity: 0.91, specificity: 0.69), 31 circCYP24A1 exhibits higher specificity and showed strong potential to diagnose ESCC. Furthermore, in the plasma of patients with other up-regulated tumors such as PCa or benign esophageal conditions like esophagitis, there was no significant difference in the expression of circCYP24A1 compared to healthy individuals, indicating disease specificity of circCYP24A1. Our study establishes circCYP24A1 as a major prospective biomarker of ESCC diagnostic and uncovers a non-invasive detection method.
Metastasis is the leading cause of cancer-related mortality, and the majority of cancer patients succumb to metastatic disease rather than the primary tumor. 32 EMT is a crucial process in inducing cancer cell metastasis, 33 and biological studies on circCYP24A1 indicated that circCYP24A1 may promote tumor dissemination by regulating EMT. So, circCYP24A1 may serve as a potential target for the treatment of ESCC, especially metastatic ESCC. The elevation of circCYP24A1 may be associated with ESCC progression; the high expression of circCYP24A1 was closely related to the clinical advanced stage, LNM and poor prognosis in the plasma and tissue cohorts. Lymph nodes are important hubs for metastatic cell arrest and growth, immune modulation, and secondary dissemination to distant sites through a series of mechanisms. 34 As reported by Reticker-Flynn, 35 LNM of tumors plays a critical role in metastatic progression by inducing tumor-specific immune tolerance that renders distant tissues amenable to metastatic colonization. In our plasma and tissue cohorts, there were fewer ESCC patients with distant metastasis, and circCYP24A1 did not show significant correlation with distant metastasis. This result may be biased due to the limited sample. But, we found the levels of circCYP24A1 in the plasma of ESCC patients with LNM were significantly higher than those of patients without LNM. Furthermore, circCYP24A1 had an AUC of 0.76 as an LNM prediction model, sensitivity and specificity both were 0.70. This indicates that circCYP24A1 has superior predictive performance for LNM and provides valuable indication for ESCC patients with LNM.
Although we have done considerable work, there are still some limitations in our study. First, the verified samples, particularly the plasma samples, had a relatively small size and a single source, which may have a selection bias, and this may limit the generalizability of our findings. Second, circCYP24A1 has been reported in other cancers. It is necessary to assess the expression levels of circCYP24A1 in various other malignancies, aiming to determine its specificity in ESCC and its potential role in other tumors. Therefore, further investigation is required for circCYP24A1 to be utilized as an effective biomarker for ESCC in clinical practice. In our future endeavors, we plan to collect more plasma samples from patients with ESCC or other diverse tumor patients in different regions and multiple hospitals to validate and expand on our findings in a larger and more diverse population.
Conclusion
In conclusion, the study revealed that plasma circCYP24A1 is relatively high among ESCC patients and has predictive performance for ESCC in diagnosis and LNM. Hence, circCYP24A1 is a potential non-invasive biomarker for diagnosis and predictor of LNM in ESCC.
Supplemental Material
sj-docx-1-tct-10.1177_15330338241295313 - Supplemental material for Plasma CircCYP24A1 as a Novel Biomarker of Esophageal Squamous Cell Carcinoma
Supplemental material, sj-docx-1-tct-10.1177_15330338241295313 for Plasma CircCYP24A1 as a Novel Biomarker of Esophageal Squamous Cell Carcinoma by Ruolan Zhang, M.M., Jianlin Liu, M.M., Hang Yang, M.M., Jinsong Tan, M.M., Rong Xiong, M.M., Yun Liu, MM, Gang Feng, PhD, Guiqin Song, PhD, and Kang Liu, PhD in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Acknowledgements
Not applicable.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by National Natural Science Foundation of China [grant number 82203851], Sichuan Science and Technology Program [grant number 2023NSFSC0731, 2023YFSY0045], Nanchong Science and Technology Program [grant number 22SXQT0334, 22SXQT0336, 20SXQT0328, 20SXPTJS0003] and Doctoral Fund of North Sichuan Medical College [grant number CBY22-QDA01].
Ethical Approval
Ethical approval for the utilization of human peripheral blood was obtained from the Ethics Committee of North Sichuan Medical College in accordance with the Declaration of Helsinki. (APPROVAL NUMBER: [2021]42). Ethical approval for the human tissues was approved by the Ethics Committee of Shanghai Outdo Biotech company in accordance with the Declaration of Helsinki. (APPROVAL NUMBER: SHYJS-CP-2004003).
Informed Consent
Written informed consent was obtained from the patients for their anonymized information to be published in this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
