Abstract
Background
Exosomal miRNAs have emerged as promising biomarkers for cancer. However, little is known about the role of exosomal miRNAs in the response prediction of esophageal squamous cell carcinoma (ESCC) patients treated with chemoradiotherapy (CRT).
Methods
In this prospective study, 40 ESCC patients treated by CRT were enrolled from January 2021 to June 2022. Exosomes were isolated from plasma through EXODUS platform. We used small RNA sequencing in 14 samples of ESCC patients (7 responders, 7 non-responders) and the selected exosomal miRNAs were further validated in the extended cohort of 40 ESCC patients by quantitative real-time polymerase chain reaction (qRT-PCR).
Results
In the discovery phase, we identified five significantly differentially expressed exosomal miRNAs from miRNA sequencing data between the responder and non-responder patients. In the extended groups of responders (n = 27) and non-responders (n = 13), only miR-23b-3p (
Conclusion
Exosomal miR-25-3p and miR-23b-3p serve as promising biomarkers for predicting the early effectiveness of CRT in locally advanced ESCC patients, whereas miR-25-3p is a novel prognostic marker for ESCC. However, further larger prospective studies are needed to confirm their utility for individualized treatment decision in ESCC.
Background
Esophageal cancer ranks the sixth cause of malignancy-related deaths worldwide. 1 Esophageal cancer is a common malignant tumor in China and esophageal squamous cell carcinoma (ESCC) is the major pathological type. 2 More than half of new cases of ESCC occur in China. ESCC is an aggressive cancer with early lymphatic and hematogenous metastasis. ESCC is usually diagnosed at an advanced stage, showing rapid progression and extremely poor prognosis. Concurrent chemoradiotherapy (CRT) is the generally acknowledged treatment measure for patients with locally advanced ESCC, but the 5-year overall survival (OS) rate is less than 40%.3,4 Local/locoregional failure remains the dominant cause for poor outcome. 5 Early prediction of therapeutic effect will be helpful for adjusting the treatment measures timely and improving therapeutic effect. At present, the effective biomarkers to screen ESCC patients insensitive to CRT remains absent in clinical practice. Therefore, novel biomarkers that can reliably predict individual responses to CRT are needed.
Liquid biopsy has emerged as an emerging method to screen for malignancies and predict treatment outcomes through detecting circulating biomarkers in biofluids such as blood, saliva, and urine. 6 Exosomes have become a central focus of liquid biopsy research. 7 Exosomes are nano-sized extracellular vesicles. They are rich in bioactive substances, including proteins, lipids, mRNAs, and miRNAs derived from their cells of origin. 8 Exosomes are excreted only by living cells, with better stability and higher concentration in the circulation than CTCs and ctDNA. The RNA cargo of exosomes is mainly microRNAs (miRNAs), and exosomal miRNAs may carry more cancer-specific molecular biomarkers than dissociative miRNAs. 9 The function of exosomal miRNAs is brought into sharp focus in recent years, due to their non-invasiveness, high accessibility, and stability. Exosomal miRNAs have been applied as useful early diagnostic markers. Liu et al 10 identified decreased plasma exosomal miR-139-3p levels in colorectal cancer (CRC) patients may become a new biomarker for the screening for early-stage tumors and metastatic lesions. High diagnostic value was observed with significantly reduced levels of exosomal miR-5684 and miR-125b-5p in non-small cell lung cancer (NSCLC) patients compared to healthy controls. 11 Exosomal miRNAs could be used as biomarkers for cancer staging and patient prognosis prediction. Significantly higher levels of miR-21 were found in the serum exosomes of breast cancer patients with bone metastases compared to those with earlier stage or with metastases in other organs. 12 Zou et al 13 demonstrated that downregulation of serum exosomal miR-150-5p was related to shorter survival in patients with CRC. Furthermore, exosomal miRNAs were essential tools to monitor therapy responses. Serum exosomal miR-574-3p was identified as a biomarker for predicting efficacy of radiotherapy in glioma. 14 Circulating exosomal miR-425-3p could reflex the therapeutic efficacy of platinum-based chemotherapy in patients with NSCLC. 15 Despite these promising findings, limited research has been conducted specifically on exosomal miRNAs as predictive biomarkers in ESCC patients treated with radical CRT, prompting our investigation. In our previous study, we have extracted high yield and purity plasma exosomes from 40 patients with locally advanced ESCC who received definitive CRT through exosome detection method via the ultrafast-isolation system (EXODUS) platform. 16 The objective of this study was to identify differentially expressed plasma exosomal miRNAs in ESCC patients showing different response to CRT using RNA sequencing. Furthermore, we aimed to investigate the potential of these differentially expressed exosomal miRNAs as biomarkers for predicting the efficacy of CRT. The flowchart of the study is shown in Supplementary Figure S1.
Materials and Methods
Patient Characteristics, Treatment and Follow-up
In this prospective study, 40 newly diagnosed locally advanced ESCC patients were enrolled at Zhejiang hospital and Hangzhou cancer hospital between January 2021 and June 2022. All patients received radical concurrent chemoradiotherapy. Eligibility criteria included: patients with histologically confirmed medically inoperable or clinically unresectable ESCC, and the tumor staged as T2 to T4, N0/1, M0/1a (only supraclavicular/celiac lymph node metastasis and not any other distant metastasis) according to the sixth edition of the American Joint Committee on Cancer (AJCC); 18–70 years old; Eastern Cooperative Oncology Group (ECOG) performance status of 0–2; adequate renal, liver, and bone marrow reserve; and adequate nutritional intake. Ineligibility criteria included history of other tumors, tracheoesophageal fistula, metastatic disease, prior systemic treatment or thoracic radiation, active infection, and severe cardiovascular disease.
Radiotherapy was administered with intensity modulated radiotherapy (IMRT) technique to a total dose of 60 Gy in 30 fractions (5 days per week at 2 Gy/d). Gross tumor volume was defined as the primary tumor and the involved regional lymph nodes. Clinical target volume included the gross tumor volume plus a 3-cm expansion superiorly and inferiorly along the length of the esophagus and a 1-cm expansion of supraclavicular and mediastinal lymph node regions. All patients received chemotherapy comprising intravenous paclitaxel (135 mg/m2, day 1) and cisplatin (25 mg/m2, days 1-3) or oxaliplatin (130 mg/m2, day 2) every 4 weeks for two cycles.
Clinical response was assessed according to Response Evaluation Criteria in Solid Tumors (RECIST 1.1) 2 months after the completion of chemoradiotherapy. Response was evaluated by esophagography, esophagoscopy with biopsy, and computed tomography (CT) imaging. PET–CT imaging was optional. Tumor response assessed by CT scan was based on its vertical length, maximal transverse thickness, and maximum thickness of the esophageal wall. Patients were regularly followed up for 2 years every 3 months, followed by 3 years every 6 months, and then annually. Additional assessments were carried out if disease progression was suspected. Patients were considered responders if they had either a complete response (CR) or partial response (PR) and non-responders if they had stable disease (SD) or progressive disease (PD). The study was approved by the Medical Ethics Committee of Zhejiang hospital (approval number 2021-161 K) and Hangzhou cancer hospital (approval number HZCH-2020). All participants signed informed consent to participate in the study. We have de-identified all patients’ details. Patient and tumor characteristics are listed in Table 1 and Supplementary Table S1.
Demographics and Clinicopathological Characteristics of ESCC Patients

ESCC, esophageal squamous cell carcinoma; ECOG, eastern cooperative group.
Plasma Exosomes Isolation and Characterization
Peripheral blood specimens (5 mL) were collected from each patient prior to the start of treatment. The plasma was isolated, aliquoted, and stored at −80 °C until thawed for the following exosome isolation. Additionally, plasma exosomes were isolated using the EXODUS platform according to the manufacturer's instructions. The purified exosomes were fixed with 4% paraformaldehyde (PFA) and loaded on carbon-coated copper grids. Grids were washed with PBS and transferred to a 50 μL drop of 1% glutaraldehyde for 5 min. Afterward, the grids were washed thoroughly with PBS and then with 100 μL of distilled water. The grids were stained with uranyl acetate (2%) for 30 s and washed again in PBS. After air drying, the samples were observed by a transmission electron microscope (Talos F200S, Thermo). The size and concentration of plasma exosomes were measured through Nanosight NS300 (Malvern Instruments) following the manufacturer's instructions. Exosome markers, including CD63 (Abcam, Cat. ab134045), FLOT1(BD Transduction, Cat. 610820) and TSG101(HuaBio, Cat. ET1701-59), were detected by Jess automated Western blotting analysis as previously described. 17
RNA Extraction and Small RNA Sequencing
We isolated exosomal RNAs by a miRNeasy Serum/Plasma Kit (Qiagen) according to the manufacturer's instructions. Exosomal RNA was isolated from a total of 14 samples, comprising 7 non-responders and 7 responders. Total RNA was quantitated using Nanodrop 2000 (Thermo Fisher Scientific Inc., USA). RNA quality was assessed on an Agilent 2100 Bioanalyzer (Agilent Technology, USA). Small RNA library construction was performed using NEBNext Small RNA Library Prep Set for Illumina kit (Cat. No. NEB#E7330S, NEB, USA). The size distribution and quantity of sequencing libraries were determined using the DNA 1000 Kit (Agilent Technologies) on Agilent 2100 Bioanalyzer. The libraries were finally sequenced using the Illumina Novaseq 6000 platform. Volcano plot and clustered heat map were used to visualize the differentially expressed miRNAs.
qRT-PCR Validation
To verify the small RNA sequencing results, a qRT-PCR assay was carried out. The validation cohort was extended to 40 patients of which 27 were CRT responders and 13 were CRT non-responders. We further assessed the expression levels of the top five miRNAs. RNA isolated from exosomes was reverse transcribed to cDNA using the miRCURY LNA RT kit (Qiagen, Cat. 339340). qRT-PCR was performed in a reaction mixture comprising cDNA, primers, and SYBR GreenER qPCR SuperMix on LightCycler 480 System II (Roche, Swiss) according to the manufacturer's instructions. Each sample was run in triplicate. Melting curve analysis was conducted to check the amplicon specificity. Primers were designed and synthesized based on the mRNA sequences obtained from the NCBI database (Supplementary Table S2). The miRNA expression level was standardized to 5S RNA and calculated using the 2−ΔΔCT method.
miRNA Target Prediction, Gene Ontology and Pathway Enrichment Analysis
The target genes of the significantly differentially expressed miRNAs were predicted by 2 databases (miRanda and Target-scan). Only the miRNAs that could predict the targets within the 2 databases were further analyzed. Gene ontology (GO) and kyoto encyclopedia of genes and genomes (KEGG) pathway enrichment analysis were performed using target genes.
Statistical Analysis
All data were analyzed using the SPSS 26.0 statistical software and Graphpad Prism 9.0. All data were expressed as mean ± standard deviation or median according to data distribution. The Mann-Whitney U test was used for comparisons between the two groups. The area under the curve (AUC) of the receiver operating characteristic curve (ROC) was used to analyze the predictive value of plasma exosomal miRNAs. PFS was defined as the interval from ESCC diagnosis to the time of tumor progression (locoregional recurrence and/or distant metastasis)or death from any cause. To clarify the link between PFS and miRNAs, the allocation of patients in the high/low expression group was made using the median expression value of each miRNA. PFS was compared using the Kaplan-Meier method with a log-rank test. Univariate and multivariate Cox regression analysis were performed for prognostic factors of PFS.
The reporting of this study conforms to STROBE guidelines. 18
Results
Identification and Characterization of the Plasma Exosomes
The exosomes isolated from plasma of ESCC patients displayed as cup-shaped bilayered membrane-bound morphology under TEM in both groups (Figure 1A, B). The average diameter of exosomes was 103.6 ± 18.4 nm in responders and 105.2 ± 19.7 nm in non-responders (Figure 1C, D). Western blot analysis confirmed the presence of exosome markers CD63, FLOT1, and TSG101 in the isolated exosomes (Figure 1E, F).
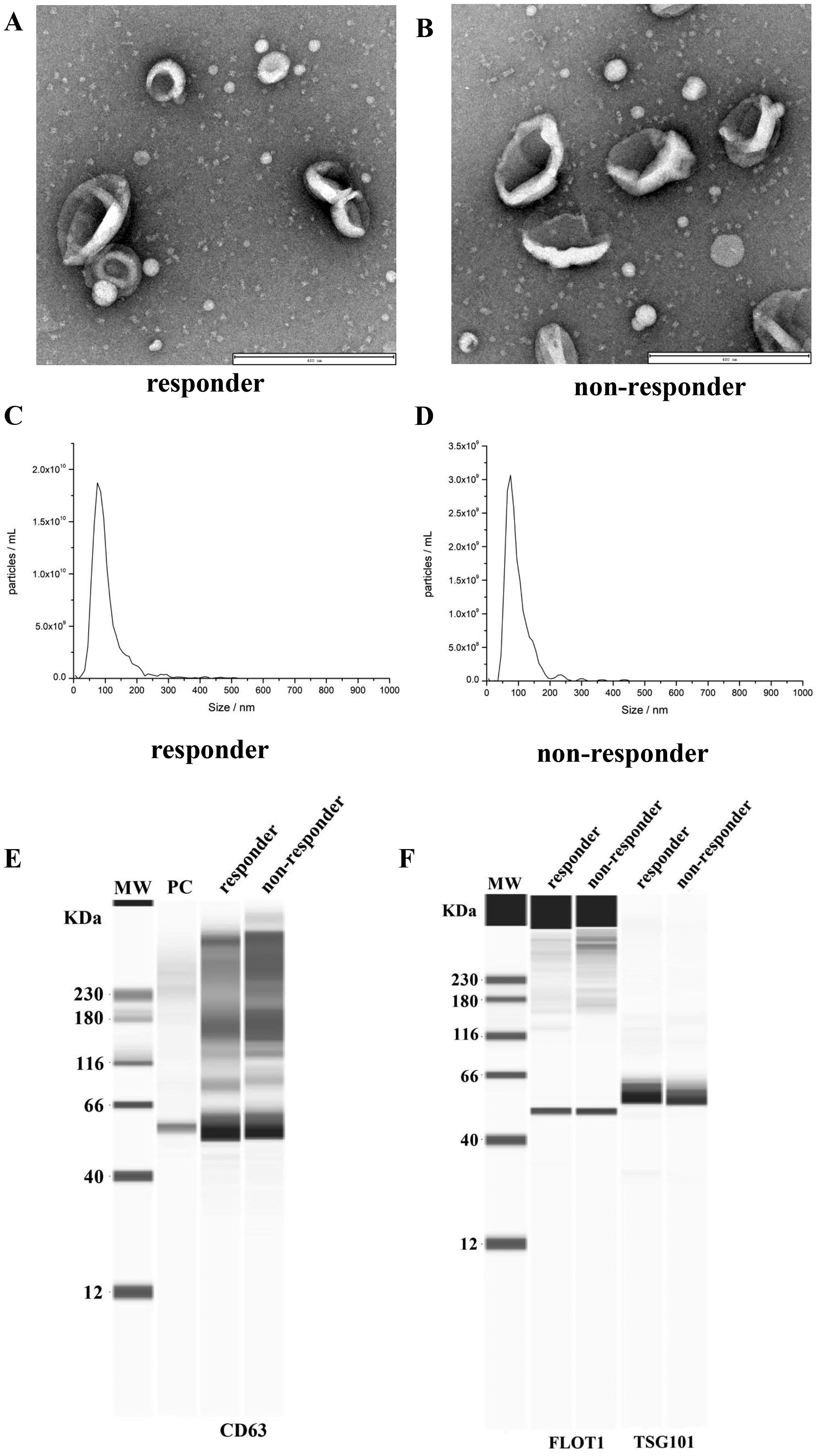
Plasma-derived exosomes identification in responder and non-responder ESCC patients treated with CRT. The morphology of exosomes observed by TME in responder patients (A) and non-responder patients (B). Particle diameter analysis of exosomes in responder patients (C) and non-responder patients (D). (E, F) Identification of CD63, FLOT1, and TSG101 marker proteins on exosomes. ESCC, esophageal squamous cell carcinoma; CRT, chemotherapy; TME, transmission electron microscopy; PC, positive control.
Differential Expression of Exosomal miRNAs Between Responders and non-Responders
In the discovery stage, plasma exosomal miRNA sequencing data revealed that 128 exosomal miRNAs were differentially expressed between 7 responder patients and 7 non-responder patients. Five significantly differentially expressed miRNAs were identified and screened by using the criteria of |log2FoldChange (FC)|>2.0 and false discovery rate (FDR)-adjusted

miRNA sequencing of plasma exosomes in responder and non-responder ESCC patients treated with CRT. (A) Heatmap illustrating the top differentially expressed miRNA between responder and non-responder groups. The columns represent samples, and the rows represent miRNAs. (B) Volcano plot of differentially expressed miRNA between responder and non-responder groups. The red dots are significantly up-regulated miRNAs. The blue dots are significantly down-regulated miRNAs. ESCC, esophageal squamous cell carcinoma; CRT, chemotherapy.
Validating Candidate Exosomal miRNAs and Exploring Their Predictive Values for CRT Early Efficacy
The expression levels of these 5 miRNAs were further quantified using qRT-PCR assays and statistically evaluated between the extended groups of responders (n = 27) and non-responders (n = 13). By using a non-parametric Mann-Whitney U test, only miR-23b-3p (
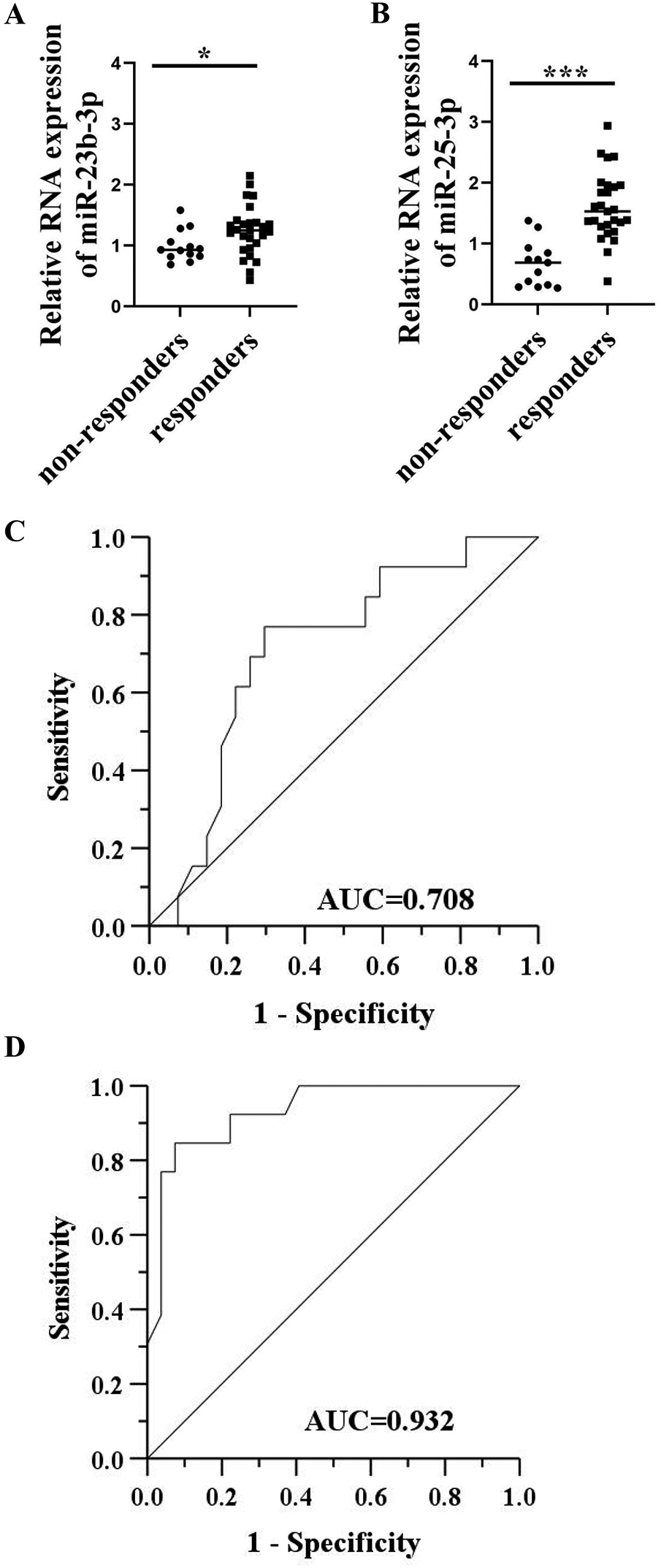
Validation of plasma exosmal miR-23b-3p and miR-25-3p as potent predictive markers in extended cohort of responder and non-responder ESCC patients treated with CRT. The relative expression of miR-23b-3p (A) and miR-25-3p (B) in the responder patients (n = 27) and non-responder patients (n = 13) by qRT-PCR. *p < 0.05; ***p < 0.001. ROC curves exploring the predictive values of miR-23b-3p (C) and miR-25-3p (D) in the early efficacy of CRT. ESCC, esophageal squamous cell carcinoma; CRT, chemotherapy; qRT-PCR, quantitative real-time polymerase chain reaction; ROC, receiver-operating characteristic; AUC, area under the curve.
Exosomal miR-25-3p is an Independent Predictor of PFS in ESCC
We further explored the relevancy of these exosomal miRNAs expression with PFS. The patients were divided into high and low groups based on the levels of miR-23b-3p and miR-25-3p. The median follow-up was 16.2 months. In the univariate survival analysis, miR-25-3p (HR = 0.322, 95% CI = 0.105-0.984,

Kaplan-Meier survival curve for PFS in 40 ESCC patients treated with CRT. (A) Patients were divided into low and high miR-25-3p expression groups according to the median miR-25-3p concentration. (P = 0.035, log-rank test). (B) Patients were divided into low and high miR-23b-3p expression groups according to the median miR-23b-3p concentration. (P = 0.730, log-rank test).
Multivariable Cox Regression Analysis of Clinical Characteristics and miR-25-3p and miR-23b-3p Level Effects on Progression-Free Survival (PFS) of ESCC Patients

ESCC, esophageal squamous cell carcinoma; ECOG, eastern cooperative group; HR, hazard ratio; CI, confidence interval.
Pathway Enrichment Analysis of Exosomal miR-25-3p
To elucidate the molecular mechanism of the exosomal miR-25-3p, we performed GO clustering and KEGG signaling pathway enrichment analysis of the identified target genes. Figure 5A presented the top 10 prominently enriched biological processes (BPs), cell components (CCs), and molecular functions (MFs). The KEGG pathway analysis in Figure 5B revealed that the target genes mainly involved in pathways related to proteoglycans in cancer, forkhead box O (FoXO) signaling pathway, signaling pathways regulating pluripotency of stem cells, mammalian target of rapamycin (mTOR) signaling pathway, and ferroptosis.

Go (A) and KEGG pathway (B) analysis of target mRNAs of the exosomal miR-25-3p. GO: gene ontology; KEGG: Kyoto encyclopedia of genes and genomes.
Discussion
Currently, the limitation of contrast CT scan and endoscopy/biopsy has been progressively realized in the area of guiding personalized treatment managements for ESCC. Real-time dynamic efficacy assessment is important for individualized treatment adaptation. Repeated tumor biopsy would be a reliable approach. However, it is not clinically feasible for dynamic assessments due to its invasiveness. Exosome-based liquid biopsy is superior to traditional biopsy for its less invasion, convenience, and real-time monitoring of tumor progression and treatment effect.19,20 Due to high abundance and good stability, tumor-derived exosomal miRNAs seem to be appealing as potential biomarkers. However, exosomes have not been extensively applied in clinical practice due to lack of appropriate isolation methods. 21 Developing reliable isolation and purification methods to separate exosomes from tumor will promote clinical translation of exosome-based liquid biopsy for cancer precision medicine. Chen et al 22 reported an efficient exosome isolation system- EXODUS which was highly efficient and stable to purify exosomes from body fluids.
At present, there have been several studies about exosomal miRNAs in esophageal cancer. Tanaka et al 23 revealed that miRNA-21 expression from plasma exosomal obviously declined in ESCC patients and strongly linked to a worse clinical profile. High plasma exosomal miR-182 level was also closely related to the tumor burden of ESCC patients. 24 Recently, Liu et al 25 found that exosomal miRNA could predict the status of lymph node metastasis. However, the majority of studies concern exploration of exosomal biomarkers for tumor early detection rather than exosomal markers for the assessment of therapeutic response in ESCC.
Here, we assessed the profile of plasma exosomal miRNAs from ESCC patients with different response to CRT by RNA sequencing. This approach identified 5 significantly differentially expressed exosomal miRNAs. In the further extended 40 plasma exosome samples cohort, we confirmed the low level of miR-23b-3p and miR-25-3p in plasma exosomes correlated with unfavorable response to CRT treatment. Based on the results, ESCC patients with high level of miR-23b-3p and miR-25-3p might be better candidates for CRT treatment. Low level of miR-25-3p was finally revealed to serve as an independent predictive biomarker associated with poor PFS. These results suggest that exosomal miRNAs have the potential to predicting the chemoradiotherapy sensitivity of locally advanced ESCC patients. The real-time dynamic assessment of exosomal miRNAs will help for tailoring therapeutic decisions. Patients predicted to have poor response can intensify or switch treatment strategy, whereas patients with good response can use the standard treatment more reliably.
Recent research revealed exosomes might mediate intercellular communication by delivering miRNAs to influence the expression of target genes. It was shown that exosome-derived miRNAs were involved in the development of cancer treatment resistance. Qi et al 26 revealed that cancer-associated fibroblasts play important role in contributing chemoresistance in pancreatic ductal adenocarcinoma through secretion of exosomal miR-3173-5p inhibiting ferroptosis. Exosomal miRNA-205 could promote tamoxifen resistance and tumorigenesis in breast cancer through targeting E2F1. 27 In addition, exosomes may serve as efficient miRNA delivery systems. 28 miRNAs can be packaged in exosomes and delivered as an effective cancer treatment molecule for increasing cancer cell susceptibility to chemoradiotherapy.
Several studies have indicated that miR-23b-3p is highly expressed in cancer. miR-23b-3p could induce the proliferation and metastasis of ESCC. 29 Serum exosomal miR-23b-3p may serve as biomarker with a high diagnostic efficacy for NSCLC. 30 miR-25-3p (the miR-106b-25 cluster) has been found to be aberrantly expressed in cancers, Alzheimer's disease, and ovarian endometriosis, etc31–33 The expression of miR-25-3p was low in tongue squamous cell carcinoma, 34 prostate cancer, 35 and cervical cancer, 36 while overexpression of miR-25-3p inhibited the proliferation and metastasis of cancer cells. The discoveries indicate that miR-25-3p may participate in inhibiting cancer cells. However, the role of miR-23b-3p and miR-25-3p as biomarkers for prediction response to concurrent chemoradiotherapy in ESCC has not been reported.
To clarify potential molecular mechanisms of miR-25-3p, we performed GO functional and KEGG pathway enrichment analysis of the miR-25-3p target genes. The target genes in clusters involved in positive regulation of cellular catabolic process, regulation of autophagy, proteoglycans in cancer, FoXO signaling pathway, mTOR signaling pathway, and ferroptosis. Ferroptosis is characterized by lipid reactive oxygen species accumulation and iron dependency. The crosstalk between metabolic reprogramming and ferroptosis contribute to promote tumor progression and chemoresistance.37,38 Autophagy was reported to promote chemoresistance in cancer cells by protecting against apoptosis and driving senescence. 39 Low FoXO transcriptional activity has been shown to drive chemoradiation resistance in esophageal adenocarcinoma. 40
Consistent with our results, miR-25-3p, has been previously shown to be taken part in the drug resistance. miR-25-3p/MEK4/JNK1 regulatory axes played critical roles in the development of adriamycin resistance of breast cancer. 41 miR-25-3p also played an important part in regulating EMT and chemoresistance in cervical cancer. 42 Studies aimed at demonstrating the accurate mechanisms underlying the miR-25-3p both in vitro and in vivo are needed.
There are several limitations of this research. One is the small number of patients and this is a single-center-based study. A relatively short follow-up period is another limitation. Finally, the optimal cut-off value of this laboratory marker for translation into clinical practice is needed to be defined in future studies. In the future, we still need more prospective studies containing multicenter larger patient cohorts and long follow-up time to further validate the value of miR-25-3p in ESCC.
Conclusion
Our findings demonstrate that exosomal miR-25-3p and miR-23b-3p may predict therapeutic response to CRT, highlighting their potential as biomarkers for early effectiveness in locally advanced ESCC patients. Validation of the results in a larger independent cohort is needed. Functional investigation will help to describe precise mechanisms of miR-25-3p in regulation of response to CRT in ESCC patients and develop more effective therapeutic strategies.
Supplemental Material
sj-docx-1-tct-10.1177_15330338241289520 - Supplemental material for Plasma-derived Exosomal miR-25-3p and miR-23b-3p as Predictors of Response to Chemoradiotherapy in Esophageal Squamous Cell Carcinoma
Supplemental material, sj-docx-1-tct-10.1177_15330338241289520 for Plasma-derived Exosomal miR-25-3p and miR-23b-3p as Predictors of Response to Chemoradiotherapy in Esophageal Squamous Cell Carcinoma by Zhao Jing, Zhen Guo and Chuanfeng Zhang in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by the Zhejiang Province Public Welfare Technology Application Research Project (grant numbers LGC21H160001) and the General Research Program of Zhejiang Provincial Department of Health (grant numbers 2022KY472). The funders were not involved in the design or interpretation of the results.
Ethical Approval and Informed Consent Statements
The study protocol was approved by the Medical Ethics Committee of Zhejiang hospital (2021-161 K) and Hangzhou cancer hospital (HZCH-2020). The study was conducted according to the principles of the Helsinki Declaration. Written informed consent was obtained from all participants.
Data Availability Statement
The data that support the findings of this study are available on request from the corresponding author. The data are not publicly available due to privacy or ethical restrictions.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
