Abstract
Introduction
Colorectal cancer (CRC) is the world's third most frequent cancer, with a significant mortality rate due to late detection. There is a need to search for biomarkers that can detect colorectal cancer at an early stage. MicroRNAs (miRNAs) regulate several targets that function as oncogenes and/or tumor suppressor genes, so any change in microRNA expression level can predict abnormality.
Objective
The objective of the study was to evaluate the expression of miR-1290, and Suppressor of cancer cell invasion (SCAI) gene that may be used as biomarkers for early diagnosis of colorectal carcinoma.
Methodology
This study included 50 subjects consisting of newly diagnosed colorectal carcinoma patients (n = 25), and healthy controls (n = 25). After RNA isolation and reverse transcription, the expression level of miR-1290 and SCAI gene in the tissues and plasma samples of CRC patients were analyzed using real time PCR and compared with healthy individuals as normal controls. The 2−ΔΔCt formula was used to compute the fold-change, while using miR-16 and GAPDH as reference genes for normalization.
Results
We found that miR-1290 is upregulated, whereas SCAI gene is downregulated in both plasma and tissue samples of CRC patients. For miR-1290, the sensitivity was 96% and specificity was 100%, and for SCAI, 100% sensitivity and 88% specificity was calculated by ROC analysis.
Conclusion
The expression of miR-1290 and SCAI gene may be utilized as biomarkers for diagnosis of colorectal carcinoma.
Introduction
Colorectal cancer (CRC) that affects the rectum and colon, is the third most occurring type of cancer 1 and the fourth most common cancer causing death in patients. 2 The global cancer data showed that it is second cancer that is most often diagnosed in women and ranks third in men.3,4 It is contributing 9.7% of worldwide cancer burden and expected to rise with one million fatalities and over two million additional cases by the year 2030.
Commonly CRC remains asymptomatic until it has progressed to an advanced stage. It is unfortunate that up to 25% of patients are diagnosed at metastatic stage. 5 Survival rate and prognosis of colorectal cancer depends upon the stage at which it is diagnosed, at stage I, the 5 years survival rate is 90% - 92% compared to 10% or less than 10% in stage IV, when distant metastases are present. 6 Usually, when cancer is diagnosed with presenting symptoms, up to 70% are at an advanced stage like III-IV. For improved CRC survival rate, disease-free outcomes, and good quality of life, screening programs should be developed that can diagnose colorectal carcinoma at an early stage (I-II). In the UK, 16% decline in overall mortality rate was achieved with the use of screening colonoscopy and on fecal occult blood testing (FOBT). 7
CRC screening is more important as CRC grows slowly, and usually symptoms are not produced until reaching large size of many centimeters, which can block feces passage, and present additional symptoms like pain, cramping, and bleeding with stool or rarely “tarry” dark stool. Development of CRC is a multistep route which involves genetic, morphological, and histological changes which takes time to mount up. 8
Colonoscopy is an outstanding screening tool, still there are many limitations, such as bleeding, colon perforation, cost, sedation requirements and procedure invasiveness. 9 Also, improper bowel preparation and limited access are few more limitations of colonoscopy. 10 The FOBT is also considered imperfect due to low positive predictive value. 11 Blood biomarkers are alternative, and promising diagnostic approach for an early detection of colorectal carcinoma.12,13
Lack of early detection is the primary cause of cancer's high fatality rate. The biomarkers that can be detected in body fluids are easier to perform. 14 Biomarkers are molecular patterns, which can be helpful for early detection of colorectal carcinoma, and treatment as well. 15 As compared to the conventional tools such as colonoscopy, sigmoidoscopy, FOBT, double contrast barium enema, and computer tomographic colonoscopy, it is expected that molecular tests are more sensitive, specific, and patients report better tolerance. However, additional research is required to confirm authenticity. 16
MicroRNAs (miRNAs) are 18–25 bp ribonucleic acids, that inhibit the translation by binding 3’ ends of mRNA. By regulating translation, they affect cell growth, apoptosis, and differentiation. 17 MiRNAs are single stranded, noncoding small RNA which by interacting with messenger RNA at 3’ untranslated region suppresses the gene expression. 18 Each miRNA can potentially interrelate with diverse mRNAs, in response to them many miRNAs can be suppressed. Therefore, miRNAs are involved and affect angiogenesis, inflammatory response, and malignant transformation of cells. 17
MicroRNA as a promising biomarker of cancer has been described in many cancers. 19 Abnormal expression of miRNA has been linked to many types of malignant and hematological tumors which also include colorectal carcinoma, emphasizing their use as a very potential biomarker for diagnostic, prognostic, and to classify many human malignancies as well. 20
MicroRNAs can be extracted from blood, plasma, stool, urine, tissue samples, saliva, serum, and various body fluids in frozen and paraffin-embedded tissues because of their stability against endogenous RNAs.21,22 Alterations in plasma microRNA expression in cancer patients are tumor-specific that can be used as promising diagnostic/prognostic non-invasive biomarkers. Plasma microRNAs are highly stable, this property highlighting their application as potential biomarker in plasma.23,24 MicroRNA released by plasma can be measured to check in many instances, track the initiation of medication resistance mechanisms and assess the treatment efficacy. 25
By regulating the numerous molecular signaling pathways, these small molecules can control the immune system and metabolic activities, and their dysregulation can result in a few illnesses, including cancer.26,27 Several miRNA expressions have been connected to different types of tumor biomarkers. Let-7a, miR-150, miR-142, miR-21, miR-223, miR-1246, miR-23a, miR-1229, and miR-378 are few examples of microRNAs, that all have been found to have high expression levels in CRC. Whereas, elevated levels of expression of miR-320, miR-596, miR-141, and miR-203 are mostly linked to tumor recurrence, malignant potential, and prognosis. Low levels of miR-484, miR-106a, and miR-130b expression are associated with poor prognosis, whereas high levels of these genes are associated with good predictive biomarkers. 28
Recent research has shown that miR1290 is important for determining the prognosis and an early diagnosis of patients with CRC. The expression of miR-1290 is elevated in colon cancer, which causes delayed cytokinesis and the development of multinucleated cells. 29 Colorectal cancer involves the carcinogenic functions of miR-1290. In CRC, cell proliferation has been inhibited by miR-1290 silencing. miR-1290 up-regulation increased cyclin D1 transcript, and protein levels while decreasing p27 expression. 30
A recently discovered, and extremely conserved protein known as Suppressor of cancer cell invasion (SCAI) uses a three-dimensional matrix to mediate metastasis. It has been shown that in human malignancies, SCAI serves as a target gene for several miRNAs. In many different types of cancers, SCAI is downregulated and functions as an anti-tumor gene. 31 SCAI was first described as a tumor suppressor preventing metastasis in several human cancer cells. It is still unclear how SCAI expresses itself and what function it serves during physiological and pathological processes. 32
In the present study, the relative expression of miR-1290 and SCAI gene was determined in colorectal carcinoma patients and matched healthy controls to explore their potential as biomarkers for early detection of colorectal cancer.
Methodology
Study Subjects
The present study included a total of 50 subjects involving newly diagnosed patients with colorectal carcinoma (n = 25), and healthy human controls (n = 25). Written consent and permission were taken from all the study subjects. History was taken from all participants enrolled in the study on the pre-designed proforma.
Ethical Approval
The study was approved by the Institute of Molecular Biology and Biotechnology Ethical Review Committee at The University of Lahore (Ref-IMBB/BBBC/22/) and ethical review board of Allama Iqbal Medical College/ Jinnah Hospital, Lahore (Ref no. 275/23/06/2022/S1 ERB).
Reporting of the Study
The reporting of this study conforms to STARD guidelines. 33
Blood Collection and Plasma Isolation
Blood samples of CRC patients were collected from Jinnah Hospital Lahore, Pakistan. Blood was collected in CBC vial, about 5 mL from each patient, and processed on the same day to isolate plasma and was kept at −20 °C until its next processing. The initial step involved separating the plasma by centrifugation at 3000 rpm for 10 min; the second step involved repeating the process at room temperature. Temperatures below −20 °C were used for storing plasma samples. 34
Biopsy Collection
The colonoscopy biopsy samples from CRC patients during routine colonoscopy examination were taken from Jinnah Hospital, Lahore, Pakistan. Samples were immediately placed in RNALater and were stored at −20 °C until further processing.
RNA Isolation for Colorectal Cancer Patients and Healthy Controls
Total RNA was isolated from colon biopsy and plasma samples according to the manufacturer's instructions using the mirVana™ miRNA Isolation Kit (catalog no. AM1560) by Life Technologies. For RNA extraction, 5–10 mg of frozen tissue samples were utilized, which were treated on ice to avoid thawing. The extracted RNA was then dissolved in 35 μL of RNase-free water and kept at −80 °C for future use. The concentration and purity of the isolated RNA was measured by using a NanoDrop spectrophotometer (Thermofisher Scientific). Electrophoresis to make sure the RNA was pure and undamaged was used.29,34–36
Reverse Transcription
The Thermo Scientific™ RevertAid™ First Strand cDNA Synthesis Kit (Cat no. K1622) for whole RNA reverse transcription was utilized to reverse transcribe the extracted total RNA, which served as the microRNA template. All the procedures were performed according to the manufacturer's instructions. The steps for the reaction were as follows: first, 5 min at 65 °C, then chilled on ice. Then, incubation at 25 °C for 5 min, 42 °C for 60 min, and at the end, 70 °C for 5 min to stop the reaction. The reverse transcription mixture included 1 µL of random hexamer primers, 1 µL of Ribo-lock RNase inhibitor 200U/µL, 2 µL of 10 mM dNTP Mix, 1 µL of RevertAid M-MuLV RT (200 U/µL) reverse transcriptase, and 5 µL of the 5X reaction buffer. The final volume was 20 µL, in which 1 µg of total RNA and nuclease-free water was also included. The cDNA that was synthesized was kept at −20 °C until needed. 37
Quantitative Real Time PCR
Thermo Scientific Maxima SYBR Green/ROX qPCR Master Mix (2X) (cat no. K0221) kit was used to amplify the cDNA using specific primers. StepOne Plus system (Thermo Fisher Scientific) was used to perform quantitative real time PCR (qRT-PCR). Using SYBR Green master mix qRT-PCR was run to assess the changes in the expression of the miRNA and its targets. For every group, there were a minimum of two replicates used for the qPCR tests. The quantitative data was normalized using GAPDH against SCAI, while hsa-miR-16 was used to normalize the hsa-miR-1290 expression. Following conditions were followed for all experiments, initial denaturation was at 95 °C for 10 min, and in cycles denaturation at 95 °C for 15 s, annealing at 54 °C for 60 s, and extension at 60 °C for 20 s. The 2−ΔΔCt formula was used to calculate normalized expression and fold change. 38 Following primer sequences were used: miR-1290 mature sequence 5’UGGAUUUUUGGAUCAGGGA 3’, SCAI sequence was 5’AAATATGTGTGAGGTGGCTG 3’.
Statistical Analysis
The data analysis was carried out using various statistical software, such as GraphPad Prism (version 8.0.2) for graphical representation, MedCalc (version 22.021) for the production of ROC curves, and Cytoscape (version 3.10.1) for the determination of miR-1290 targets. We checked the data on miRDB online website for miRNA target prediction while selecting the gene.
Results
Characteristics of Study Subjects
Overall, 50 subjects were included in the study comprising patients (n = 25), normal controls (n = 25). Patients with colorectal cancer who participated had an average age of 49.41 years and a range of 25–75 years. There were 25 CRC patients in our study, out of which 13 (52%) were males, and 12 (48%) females. From the diagnosed patients, most patients were at advanced stage ie, 6 (24%) patients at pathological stage II, 9 (36%) stage III, and 10 (40%) patients were at stage IV. The demographics of the patients and clinical details are detailed in Table 1. There were 25 healthy controls in the study, 8 were female, and 17 were male, making up 68% of the total. The average age of the healthy control subjects was 40.44, and their ages varied from 21 to 67. Hematological and other blood characteristics of healthy controls were also displayed in Table 2.
Demographic Representation and Medical History of the Study Subjects

Values are expressed as n = number or mean ± standard deviation (minimum-maximum).
Hematological Parameters and Liver Function Test Details of Study Subjects

The values are n = number and/or mean ± standard deviation (minimum–maximum)
Circulating miR-1290 is Upregulated in Colorectal Cancer
By comparing the plasma miR-1290 levels of healthy individuals with those of cancer patients, we were able to determine the significance of miR-1290 in colorectal carcinoma. By employing RT-qPCR for relative quantification, the fold expression was ascertained. The difference in fold was calculated using the 2−ΔΔCt formula. The difference of miR-1290 expression in patients and healthy controls was statistically significant with a p value of <0.05 as demonstrated by scatter plots. Figure 2(A) shows that, in comparison to plasma from healthy individuals, cancer patients had a significant increase in the expression of miR-1290.
MiR-1290 is Upregulated in the Tissue Biopsy of Colorectal Cancer
After collecting samples from patients with colorectal cancer during colonoscopies, miR-1290 expression level was checked in the pinch biopsy. RT-qPCR was used for relative quantification of miR-1290 in CRC patients and controls and normalized fold change in miR-1290 expression was determined by 2−ΔΔCt formula. A graph was generated to show the relative expression of miR-1290 in colorectal cancer patients compared to healthy control subjects. As compared to healthy individuals, cancer patients exhibited a significant increase in the expression of miR-1290 Figure 2(B).
Circulating SCAI is Downregulated in Colorectal Cancer
Compared to normal plasma, CRC plasma has a decreased expression of SCAI. A patient's SCAI expression level was established using RT-qPCR. The fold change was calculated using the 2−ΔΔCt formula. Figure 2(C) showed that compared to healthy plasma, the SCAI expression level was much lower in colorectal cancer patients’ plasma.
Target Prediction Analysis of miR-1290 by Cytoscape Software
Several tumor suppressor genes have been implicated in CRC aetiology, miR-1290 has showed a great potential to inhibit their expression. As demonstrated in Figure 1, Cytoscape was used to display the targets of miR-1290. The gene network analysis showed that the predicted target genes were involved in cell development, cell cycle, DNA replication and repair, cell death, and other related processes. In order to anticipate new miR-1290 targets, we also built a hypothetical pathway comprising these targets using Cytoscape software (Figure 1). Red round ball showed the target genes of miR-1290 which played role in CRC, green round ball showed all the genes which are targeted by miR-1290, while purple ball showed that SCAI expression is downregulated in CRC. There were a number of genes regulated by miR-1290, of which most common were BCL7A, THBS, DKK3, SMEK1, SCAI, TOMM20, HM13, ADD2, PYGO1, CREB1, CDC73.

SCAI is Downregulated in Tissue Biopsy of Colorectal Cancer
Tissues affected by colorectal cancer express less SCAI than healthy tissues. RT-qPCR was used to ascertain the degree of SCAI expression in patient tissues. To calculate the fold change, the 2−ΔΔCt formula was employed. The results showed that compared to healthy tissue, cancer tissue had a significant decreased expression level of SCAI Figure 2(D).

Comparison of Circulating and Tissue Expression
Comparison of both miR-1290 and SCAI in plasma samples and as well as matched tissue biopsy samples were shown in Figure 2. MiR-1290 was upregulated in both plasma and tissue samples while SCAI was downregulated in plasma and tissue samples as shown in Figure 2.
Comparison of Expression at Different Stages of Colorectal Cancer
When we compared the results of miR-1290 and SCAI in plasma samples and as well as matched tissue biopsy samples, we noticed that there is a gradual increase in expression of miR-1290 at all stages of colorectal cancer, and decrease in expression of SCAI in all stages of colorectal cancer, both in plasma and tissue, this differential expression is shown in Figure 3. MiR-1290 is upregulated in the plasma samples of CRC patients at all stages (II-IV) compared to the healthy control (p < 0.05) as shown in Figure 3(A). Figure 3(B) showed the results of tissue samples, miR-1290 is upregulated in the tissue samples of CRC patients compared to the healthy control tissue samples (p < 0.05). On the contrary, SCAI expression in plasma samples, and matched tissue biopsy samples is markedly reduced in colorectal cancer patients compared to the healthy control (p < 0.05) as shown in Figure 3(C) and 3(D).


Clinical Correlation of miR-1290 with Blood Parameters
An exceptional characteristic of a diagnostic marker is its ability to remain unaffected by clinical factors and haematological variables, thereby demonstrating its independence from clinical relevance. This was validated by a correlation study between haematological markers and miR-1290 levels in plasma. There was no statistically significant correlation between haemoglobin, total bilirubin, ALP, ALT, AST, MCV, platelet count, or total bilirubin (Figure 4A–H) that demonstrated that plasma miR-1290 is not affected by any of the markers and may be utilized as an independent predictive biomarker for CRC.
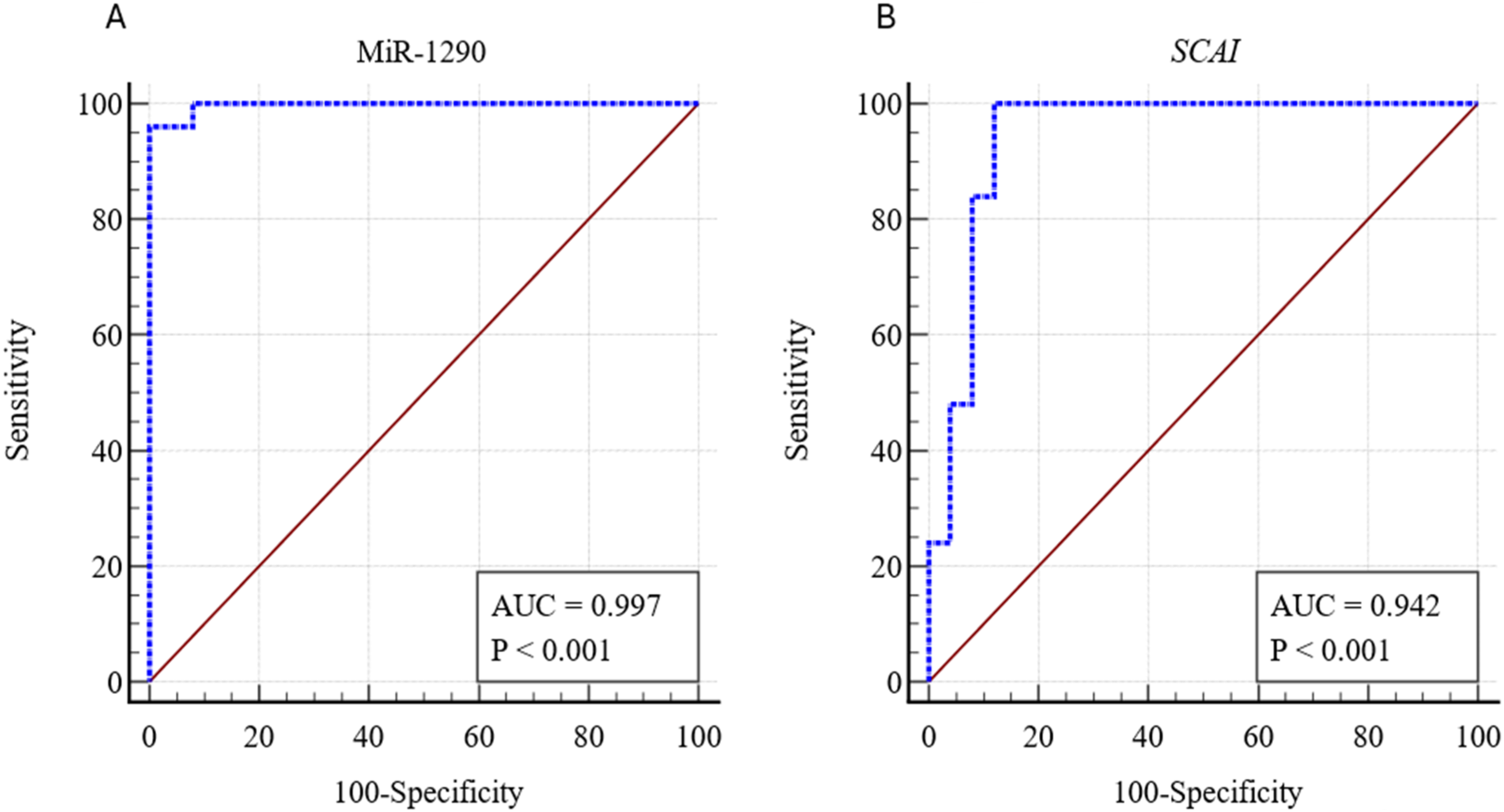
MiR-1290 and SCAI as Diagnostic Marker for CRC
To assess plasma's diagnostic potential of miR-1290 and SCAI in CRC patients, analysis of receiver operating curve (ROC) was performed. It was found that miR-1290 showed an area under the curves (AUC = 0.997, 95% CI 0.923 to 1.00), p < 0.001, representing 96% sensitivity and 100% specificity as shown in Figure 5(A). In the case of SCAI the area under the curve (AUC) was 0.942 (95% CI 0.838 to 0.988), and exhibited 100% sensitivity, 88% specificity (p < 0.001) for CRC as shown in Figure 5(B).
Discussion
Colorectal cancer (CRC) ranks high among cancer-related deaths. In the last several decades, there has been a notable rise in both the frequency and mortality rate of CRC. 29 It is also becoming a main issue in Pakistani public health due to the sharp rise in cancer incidence rates. The overall burden of colorectal cancer in Pakistan, according to recent studies, was 4.27%, with 1.66% of women and 2.61% of males affected. 39 A lower death rate can be achieved with the early diagnosis of colorectal cancer and precancerous lesions. 40 An integral part of cancer progression is the involvement of molecular biomarkers. 41 Early detection, tracking of illness development, and evaluation of therapy efficacy can all be accomplished with the use of biomarkers. The clinical use of microRNAs (miRNAs) as novel biomarkers for diagnosis or prognosis has been suggested due to their great specificity and sensitivity.41,42
The available methods for CRC detection are, stool-based, imaging, endoscopic testing,43,44 and markers such as carcinoembryonic antigen (CEA) and carbohydrate antigen (CA19-9). Their utility remains debatable due to certain limitations, as (CEA) and, (CA19-9) are non-specific. 45 While colonoscopy has its own limitations including bleeding, colon perforation, cost, sedation requirements, procedure invasiveness, improper bowel preparation and limited access. 9 The biomarkers that can be detected in body fluids are easier to perform.15,45 MicroRNAs (miRNAs) in the blood can be utilized as biomarkers and diagnostics in CRC as in CRC often the miRNAs are deregulated. 46 The advantages of miRNA biomarkers are that these are noninvasive, with a high degree of specificity, and sensitivity, miRNA biomarkers that are specific to CRC or combinations of miRNA biomarkers that are detectable in serum or plasma. 47
This study showed that miR-1290 is dysregulated in the bloodstream, and it may be useful to detect colorectal cancer at an early stage. The findings suggested that elevated levels of miR-1290 are strongly connected with the advancement of the illness and that circulating miR-1290 is a possible biomarker for CRC tumors. Valuable early screening and monitoring can provide two benefits ie, reducing the likelihood of overdiagnosis-related overtreatment and improving patient prognosis. 38 Circulating miRNAs exhibit greater stability than cellular RNA species under extreme conditions, such as boiling, numerous cycles of freezing and thawing, treatment with RNase, and extremes of pH. For this reason, evaluating circulating miRNAs is a crucial component of liquid biopsy. 48 Few studies have demonstrated changes in miR-1290 expression in colorectal cancer (CRC) tissue and serum, indicating that miR-1290 may be a useful diagnostic tool for CRC. This suggests that miR-1290 overexpression could potentially contribute to the development of CRC.48,49 This study's results have demonstrated that miR-1290 promotes tumor progression because it was found to be considerably increased in colon cancer tissue compared to normal tissue of healthy control. MiR-1290 elevated in blood plasma, and tissue biopsy of the same patients, suggesting that detection of this microRNA in the plasma can be helpful for the early diagnosis of CRC. In addition, we found no significant link between haemoglobin, total bilirubin, platelet count, ALP, ALT, AST, MCV, or any of the haematological markers that we tested for miR-1290's clinical relevance. It can be inferred from these results that the clinical features did not impact miR-1290 levels.
Although the technology for cell-free miRNA identification is developing, there are still a lot of issues that need to be resolved before it can be used in clinical practice. Since circulating miRNA concentrations are relatively low, as well as miRNA integrity and detection stability, a high extraction efficiency should be ensured. Confirmation of procedures prior to analysis influencing miRNA quantification is highly significant, especially when a single miRNA is selected as a clinical biomarker. Most of the indicators’ clinical utility is dependent on resolving the mentioned issues, even though strongly up- or down-regulated miRNAs offer advantages as biomarkers. Methodological verification hasn't been done in many studies. 50 Given that the molecular mechanism and miRNA expression differ, a detection strategy that works for one study may not work for another. This means that the detection process (accuracy, repeatability, and reliability) for a particular miRNA needs to be checked and enhanced. 51
Interestingly, we found that SCAI could be a miR-1290 target. Consistent with an earlier study that also discovered a reduction in SCAI in cancers of the human body, the current research showed that SCAI was downregulated in tumor tissues as compared to normal tissues suggesting a tumor suppressive role. 52 The main cause of cancer death is metastasis, a complicated process brought on by mutations in tumor suppressor, and proto-oncogene genes that are linked to angiogenesis, invasion, proliferation, and migration of cancer cells. 53 The possible pathway through which upregulation of miR-1290 and SCAI can regulate the cell programming is shown in the Figure 6.

The possible pathway through which upregulation of miR-1290 and SCAI can regulate the cell programming. Upregulation of miR-1290 blocks the DDK and blockage of DDK3 blocks wnt signaling pathway, and beta catenin gets activated and causes increased proliferation, migration, and angiogenesis.
In a study conducted by Li et al, it was found that miR-1290 stimulated the growth, and spread of esophageal squamous cell carcinoma cells via blocking SCAI. Based on our findings, miR-1290 likely triggered an increase in CRC cell migration, and invasion by inhibiting SCAI. The results of different studies showed that miR-1290 inhibited the SCAI, and promoted cell proliferation and metastasis in cancer. 54 The function of miR-1290 in colorectal cancer (CRC) remains unclear, despite its demonstration as an onco-miRNA in multiple cancer types. In an in-vitro study it has been demonstrated the miR-1290's potential to promote cancer and blood vessel formation in colorectal cancer cells and human umbilical vein endothelial cells (HUVECs). Increased proliferation, and migration of HUVECs are caused by miR-1290 overexpression while downregulation of SCAI. 55 Here, our findings suggest that miR-1290 expression is significantly increased in the plasma and tissue of colorectal cancer patients, while expression of SCAI is downregulated in both plasma and tissue. So, the expression level of miR-1290 and SCAI may be a possible biomarker for the detection of CRC.
Conclusion
We have validated novel molecular markers comprising miR-1290 and SCAI in plasma and tissue for CRC detection. Expression of miR-1290 was upregulated in tissue, and plasma samples of CRC patient's while SCAI expression in the plasma and tissue was significantly downregulated. Therefore, expression levels of miR-1290 and its target gene SCAI may be utilized as potential biomarkers for detection of colorectal cancer.
Further research is needed before miRNA profiling may be used in clinical practice. Finding specific miRNA signatures to differentiate between subtypes will be a major challenge in the coming years.
Footnotes
List of Abbreviations
Acknowledgements
None declared.
Author Contributors
Conceptualization and designing and Conduction of experiment: SS, RN Instrumentation and analysis: RN, SS. Data collection, visualization and interpretation: RN, SS, WS, GA. Preparation of initial draft: RN, SS. Proof reading and approval of the final version: RN, SS, WS, GA. All authors have critically reviewed and approved the final draft and are responsible for the content and similarity index of the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
For this study involving human subjects, a formal ethical approval has been taken from ethical review committee of Institute of Molecular Biology and Biotechnology at University of Lahore, Lahore, Pakistan (Ref. No. IMBB/BBBC/22) and Ethical review Board of Allama Iqbal Medical College/Jinnah Hospital Lahore (Ref. No. 275/23/06/2022/S1 ERB).
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Informed Consent
Written informed consent has been taken from all the subjects to participate in the study.
