Abstract
Head and neck malignancies are a significant global health concern, with head and neck squamous cell carcinoma (HNSCC) being the sixth most common cancer worldwide accounting for > 90% of cases. In recent years, there has been growing recognition of the potential role of alternative splicing (AS) in the etiology of cancer. Increasing evidence suggests that AS is associated with various aspects of cancer progression, including tumor occurrence, invasion, metastasis, and drug resistance. Additionally, AS is involved in shaping the tumor microenvironment, which plays a crucial role in tumor development and response to therapy. AS can influence the expression of factors involved in angiogenesis, immune response, and extracellular matrix remodeling, all of which contribute to the formation of a supportive microenvironment for tumor growth. Exploring the mechanism of AS events in HNSCC could provide insights into the development and progression of this cancer, as well as its interaction with the tumor microenvironment. Understanding how AS contributes to the molecular changes in HNSCC cells and influences the tumor microenvironment could lead to the identification of new therapeutic targets. Targeted chemotherapy and immunotherapy strategies tailored to the specific AS patterns in HNSCC could potentially improve treatment outcomes and reduce side effects. This review explores the concept, types, processes, and technological advancements of AS, focusing on its role in the initiation, progression, treatment, and prognosis of HNSCC.
Keywords
Introduction
Head and neck squamous cell carcinomas (HNSCCs) are pernicious tumors that originate from the mucosal epithelium in the oral cavity, pharynx, and larynx and are the most common malignancies of the head and neck region. HNCC is the sixth most common cancer worldwide, with a high recurrence rate and significant mortality, and has an estimated annual incidence of 600,000 new cases and 380,000 deaths.1,2 HNSCC is a malignant tumor originating from the mucosal tissue of the head and neck. It is the sixth most common cancer in the world, with a mortality rate of about 50% and a 5-year survival rate of only 40%–50%. The cause of the disease is closely related to unhealthy lifestyle habits, such as smoking and alcohol abuse, as well as human papillomavirus (HPV) infection.3–5Despite advancements in treatment modalities such as surgery, radiation therapy, chemotherapy, and immunotherapy, the 5-year survival rate has not improved over the past few decades. 6 This highlights the need for a deeper understanding of the molecular drivers of HNSCC, which could lead to the development of more effective targeted therapies.
RNA plays a crucial role in transmission of genetic information in cells and its stability is essential for ensuring that these genetic instructions are accurately transcribed and translated into functional proteins. Any disruptions in RNA stability can lead to errors in gene expression and protein production, which can have serious consequences for cellular function and organismal health. Understanding the mechanisms that regulate RNA stability is therefore fundamental to molecular biology and has important implications for various biological processes and diseases. 7 Most human precursor mRNAs are initially transcribed as a single molecule that includes both exons, which encode protein sequences, and introns, which do not code for proteins. RNA splicing, a complex biological process, removes the introns and join the exons to produce mature mRNA molecules that can be translated into proteins. This process is essential for generating protein diversity and ensuring proper functioning of the cell. Although intron sequences are not essential for protein translation, they are ubiquitously present in many vertebrate eukaryotic genes. Interestingly, the more evolutionarily advanced the species are, the longer and more complex its intron sequences tend to be.8,9 Pre-mRNA splicing can be regulated in a process known as alternative splicing (AS) to produce different types of mature mRNA transcripts from a single gene. 10 Splicing of pre-mRNA is necessary for over 95% of human gene expression. 11 AS increases protein diversity of proteins that can be produced from the limited number of genes in the human genome. It plays a critical role in cellular differentiation, development, and disease, and its dysregulation has been implicated in various human disorders, including cancer. 12 The study of AS variants of genes has made significant advancements in recent years. Researchers have developed standardized methods for bioinformatics analysis and prediction of AS events, which has greatly facilitated the study of AS in various biological processes and diseases. RNA sequencing (RNA-seq) is a high-throughput sequencing technology that enables comprehensive analysis of all RNA sequences and their abundances in a sample. It boasts high sensitivity, high throughput, and unbiased quantitative capabilities, making it particularly suitable for detecting and quantifying alternative splicing events. 13 As research on tumor occurrence and development mechanisms continues to evolve, the role of AS in cancer remains a prominent and dynamic area of study. 14 Splicing variations are a common feature in cancer, and these variations can be specific to different types of tumors. To fully understand the role of AS in cancer, it is essential to conduct precise and exhaustive analyses in selected patient samples and representative experimental models. The importance of AS in the research of HNSCC lies in its elucidation of the pathogenesis and its potential in the development of novel therapeutic strategies. Further research will continue to advance this field, bringing more precise and effective treatment methods.Therefore, this paper reviews the concepts, types, processes, and technological advancements of AS, focusing on its involvement in the onset, progression, treatment, and prognosis of HNSCC.
Forms and Mechanisms of mRNA Alternative Splicing
Forms of mRNA Alternative Splicing
The five classical modes of AS (Figure 1) include: alternative 5′ splice site, alternative 3′ splice site, mutually exclusive exons, exon skipping or cassette exon, and intron retention. 15 However, some researchers suggest expanding this classification to seven categories by including alternative promoters and polyadenylation (Poly A) sites. 16

Alternative splicing produces multiple combination of transcripts.
During gene expression, after DNA is transcribed into pre-mRNA, it undergoes a process of splicing events, which involve two types: constitutive splicing and alternative splicing. Constitutive splicing events provide stability and consistency in gene expression, while alternative splicing events provide diversity and adaptability in gene expression. The following are the five main modes of alternative RNA splicing:Exon Skipping: An exon can be skipped during splicing, resulting in its exclusion from the final mRNA;Alternative 5’ Splice Site: Different 5’ splice sites are used, leading to variations in the 5’ end of the mRNA;Alternative 3’ Splice Site: Different 3’ splice sites are used, resulting in variations in the 3’ end of the mRNA;Intron Retention: An intron is retained in the mature mRNA instead of being spliced out;Mutually Exclusive Exons: One of several exons is included in the mRNA while the others are excluded.These modes of alternative splicing greatly increase the diversity of proteins that a single gene can produce, allowing for complex regulation and adaptation to various physiological and environmental conditions. Different colored boxes represent different exons, and dashed lines represent different splicing events.
Mechanism of Alternative Splicing
The process of AS is closely associated with the activity of the spliceosome complexes, which recognize specific signals, called splice sites, at the boundaries between introns and exons in pre-mRNA. The spliceosome assembles on these signals and catalyzes the removal of introns and the joining of exons to generate mature mRNA. 17 The spliceosome undergoes four stages during the splicing process: construction, activation, catalysis, and disassembly. It consists of five small nuclear RNA molecules (snRNA) named U1, U2, U4, U5, and U6, along with numerous proteins, such as the NineTeen complex (NTC) comprising 19 proteins as well as NTC-related (NTR) proteins. Additionally, there are eight RNA-dependent ATPase/helicase enzymes that are highly conserved, along with various splicing factors and other proteins.18,19 These components work together to recognize splice sites, catalyze the splicing reaction, and ensure accurate splicing of pre-mRNA. The splicing process is highly regulated and involves the recognition of the highly conserved GU-AG sequence on the pre-mRNA; the 5’ splice site, beginning with GU, and the 3’ splice site, ending with AG. When catalyzing the splicing reaction, the spliceosome assembles step by step on the target pre-mRNA. 20 AS in eukaryotes requires the involvement of many factors, including cis- (sequences within the pre-mRNA that regulate splicing) and trans- (proteins or RNAs that bind to the pre-mRNA and regulate splicing) acting elements.
The cis-acting elements include the exon splicing enhancers (ESE), intron splicing enhancers (ISE), exon splicing silencers (ESS), and intron splicing silencers (ISS). RNA binding proteins, including serine/arginine-(SR) rich protein and heterologous nuclear ribosome proteins (hnRNP), are the trans-acting elements, that interact with the splicing enhancers and silencers. 21 ESE, ISE, ESS, and ISS are 6–10 base sequences within the pre-mRNA, that play roles in regulating AS. They exhibit both specificity and degeneracy within each class, as well as overlapping and redundant features across the classes. These sequences exhibit relative specificity and degeneracy within each class; moreover, they can have cross-talk and redundant characteristics, where multiple sequences or factors can influence the same splicing event. SR proteins have highly conserved RNA recognition motifs and RS domains at the C-terminus (repeated sequences rich in Ser and Arg). By recognizing and binding to specific sequences in pre-mRNA, they promote the assembly of the spliceosome. hnRNP proteins are involved in various RNA processing steps from transcription to translation, including pre-mRNA splicing, mRNA stability, transport, and translational regulation 22 (Table 1).The enhancers and inhibitors promote or inhibit the selection of splicing sites, respectively, by combining them with splicing factor proteins. ESE and ISE enhance splice site activity and facilitate AS of exons by interacting with SR proteins. Conversely, ESS and ISS can inhibit splice site activity and impede the AS of exons by binding to the hnRNPs family proteins (Figure 2). Splicing factors, splicing-related proteins, RNA molecules, splicing enhancers, and splicing silencers all play important roles in regulating AS, by identifying splicing sites and improving splicing precision. ESEs, recognized by SR proteins, facilitate the binding of U1 and U2 small nuclear ribonucleoproteins (snRNP) to splice sites. This process is crucial for accurate splicing and the generation of functional protein isoforms. ESS and ISS can inhibit the splicing process in mRNA by preventing the binding of U1 snRNP and U2 snRNP to the splicing site. This inhibition leads to the exclusion of exons from mature mRNA, which is known as exon skipping. The structure of exons and introns plays a crucial role in determining the selection of splicing sites and facilitating the recognition of exons by forming an exon-defining complex.23,24
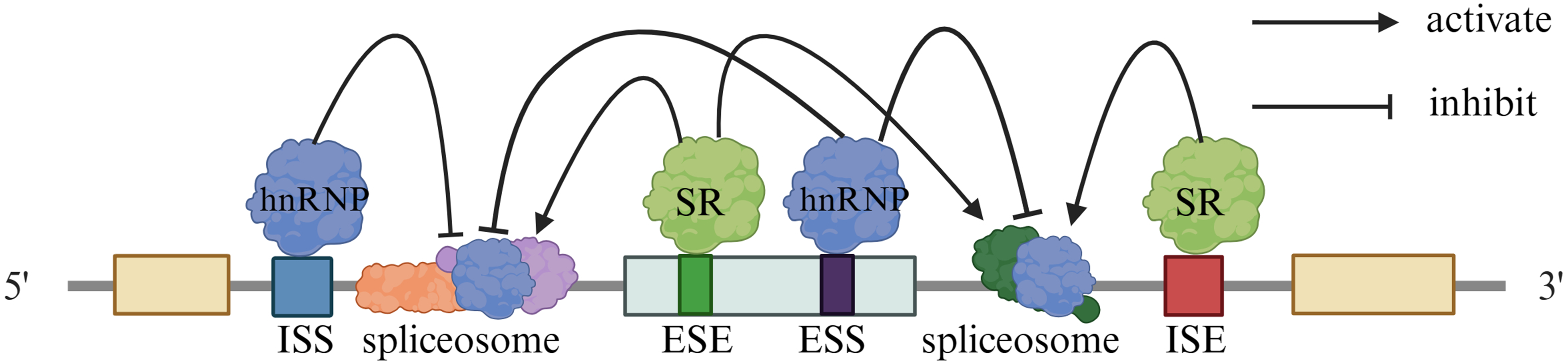
Alternative splicing regulatory sequences and factors.
Major Types, Functions, Structural Characteristics, and Roles of SR Proteins and hnRNP Proteins.

RNAs with non-coding or low coding potentials, known as non-coding RNAs (ncRNAs) play a crucial role in the post-transcriptional regulation of gene expression, particularly in AS events. ncRNAs can be detected with next-generation sequencing technologies. Recent studies have shown that long non-coding RNAs (lncRNAs) can interact with various splicing factors, altering the selective splicing patterns of target genes. 34 Most lncRNAs and splicing factors are localized in the nucleus, where lncRNAs modulate splicing activity through their interactions with splicing factors. LncRNAs can regulate AS through various mechanisms including: 1) interactions between with specific splicing factors; 2) formation of RNA-RNA duplexes with precursor mRNAs; and 3) influencing the chromatin structure; thereby fine-tuning the splicing process 34 (Figure 3). Understanding these mechanisms are crucial for comprehending the regulation of AS in iseases such as HNSCC.
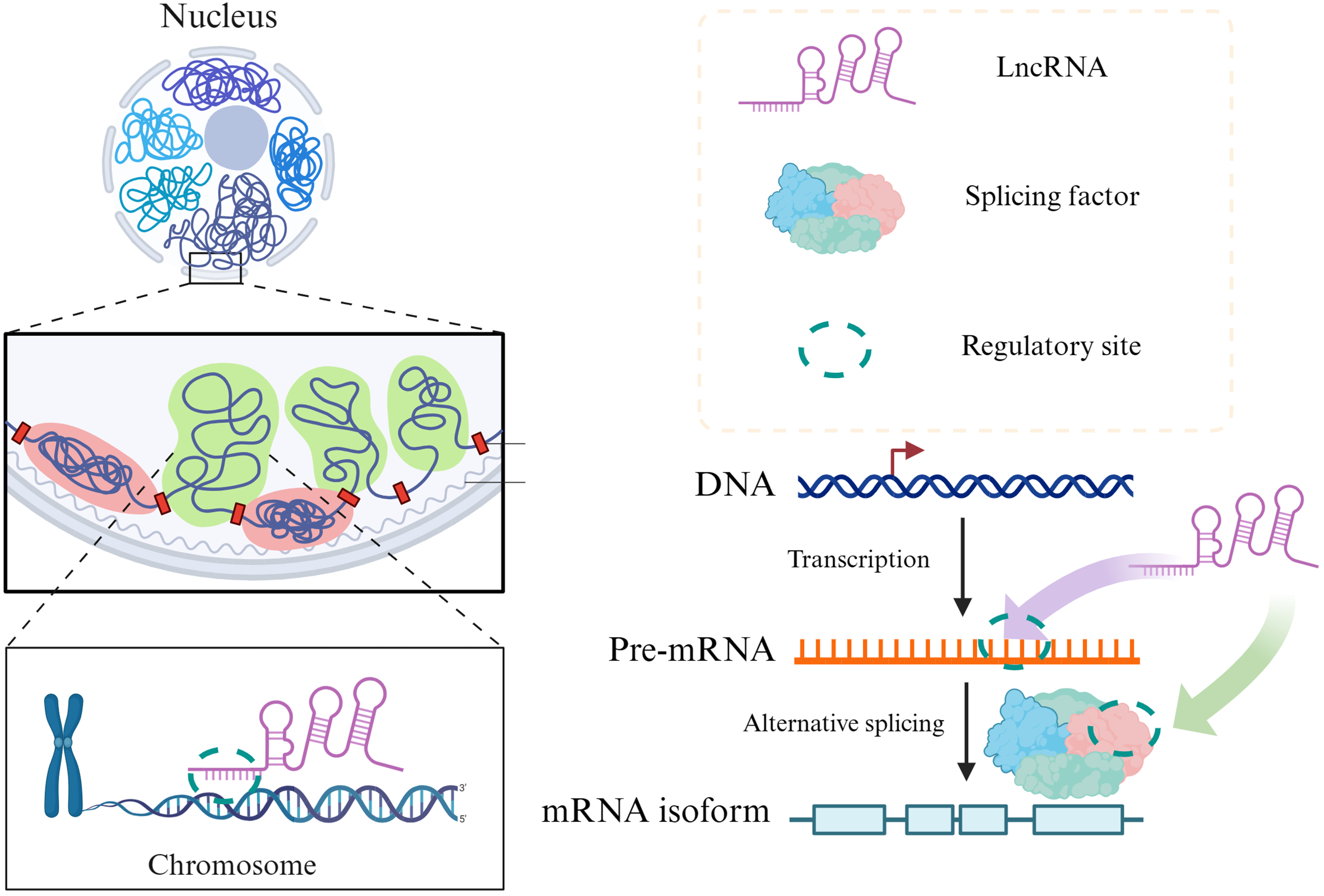
Alternative splicing regulatory sequences and factors.
The splicing regulatory mechanisms involving lncRNA mainly include the following three types: (1) lncRNA directly interacts with splicing factors, affecting the activity of these splicing factors or its binding to pre-mRNA to promote or inhibit the inclusion or deletion of specific exons, thereby changing the composition of mature mRNA. (2) lncRNA can complementary pair with specific precursor mRNA sequences to form a double-stranded RNA structure, blocking splicing sites or splicing regulatory elements (such as enhancers or silencers), affecting splicing efficiency and pattern. (3) lncRNA regulates the expression of certain gene regions by interacting with chromatin modification complexes, changing the recruitment of splicing factors or RNA polymerase, thereby affecting the splicing process.
Role of Alternative Splicing in HNSCC
Although the number of genes and regulatory mechanisms identified so far is impressive, they are still insufficient to provide effective HNSCC treatment strategies. 35 Splicing dysregulation is increasingly being identified as a crucial factor in the progression of HNSCC. This dysregulation may originate from variations in the expression levels of certain components of the splicing mechanism. Moreover, alterations in the proportions of splicing variants, or even the emergence of new variants, may contribute to the complex landscape of HNSCC development.36–38 The progress of systematic analysis in AS studies has been somewhat limited by the complexities of sequencing technologies and analytical methods. RNA-seq has emerged as a reliable technique for studying AS in tumorigenesis, providing comprehensive analysis of gene expression and splicing patterns, and offering valuable insights into cancer development and progression.39–41 A whole-genome investigation of 464 patients with HNSCC using RNA-Seq data from The Cancer Genome Atlas (TCGA) project revealed 32,309 AS events spanning 9844 genes, 42.4% of which were exon skipping events. 42 This underscores the applicability of RNA-Seq in researching AS variations in HNSCC and its ubiquity in the disease. Ye et al, through the analysis of expression profiles and clinical data from 486 HNSCC patients, identified 2726 AS events associated with survival. These events involve a variety of genes and pathways, underscoring the pivotal role of AS in the development and progression of HNSCC. 43 Similarly, Ding et al identified 3433 AS events significantly linked to overall survival through a systematic analysis of AS events in HNSCC. Their findings indicate that AS markers can not only accurately predict the prognosis of HNSCC but also offer new therapeutic targets. By constructing an SF-AS regulatory network, they unveiled the correlation between AS events and splicing factor expression, providing new insights into AS-targeted therapies. 44 In the following paragraphs, we have described the role of AS in HNSCC, emphasizing its relationship with tumor occurrence, development, treatment, and prognosis (Table 2).
Aberrant Splicing Events in HNSCC.

Splice events were identified through genome-wide analyses.
Splice events were identified by a candidate gene method.
Proliferation, Migration, and Invasion
AKT Serine/Threonine Kinase 3 (AKT3)
The AKT signaling pathway plays a crucial role in the context of HNSCC. Activation of the AKT pathway is commonly observed in HNSCC and is associated with various aspects of tumorigenesis, including cell proliferation, survival, migration, and invasion. In oropharyngeal squamous cell carcinoma (OPSCC), a novel functionally relevant splice variant of AKT3 has been identified. The expression of this splice variant is independent of the patient's HPV status and smoking history. The discovery of this variant may have significant implications for the biological understanding and therapeutic strategies of OPSCC. 39 This variant encompasses a distinct exon 1, which actively stimulates the proliferation of OPSCC. Suppression of the AKT3 isomer curbs the growth of various head and neck cancer cell lines. Moreover, this inhibition potentially occurs through the independent mediation of phosphorylated (p)AKT1 and the pAKT1 targets. 39
Serine/Arginine Protein Kinase 2 (SRPK2)
SRPK2, an arginine-serine rich protein-specific kinase, has the ability to phosphorylate the serine residues within the serine arginine-rich domain of the SR family splicing factor. Additionally, SRPK2 engages in phosphorylation of SRSF2 at distinct nuclear locations, thereby facilitating pre-mRNA splicing processes. 65 SRSF6, a frequent splicing factor and target of SRPK2, holds significant importance in the selection of alternative and constitutive splicing sites. 66 The hyperphosphorylation (≥ 1.5-fold compared to normal cells) and overexpression (2-fold increase compared to normal cells) of SRPK2 in the HNSCC cell lines has been confirmed by in vitro experiments. Inhibition of SRPK2 activity can decrease the growth, invasion, and migration activities of HNSCC cells. 37
Dedicator of Cytokinesis 5 (DOCK5)
DOCK5, a guanine nucleotide exchange factor, plays a critical role in regulating epithelial invasion and metastasis. An isomer of DOCK5 is found to be significantly elevated in patients with HPV-negative HNSCC. Individuals with increased expression levels have lower overall survival rates. Furthermore, it stimulates the activation of the p38 and MAPK/ERK signaling pathways in HPV-negative HNSCC, thereby facilitating the proliferation, migration, and invasion of tumor cells. 40
Laminin α3 (LAMA3)
LAMA3 or Laminin α, forms a vital component of Laminin-332 also known as Laminin-5, an extracellular protein that has demonstrated significant involvement in the invasion and metastasis of different types of cancer cells.67–69 Patients with HNSCC display a notable elevation in the expression of the splicing isomer LAMA3-A of LAMA5, whereas no correlation is observed between hypoxia-induced tumors and the full-length transcript of this gene, namely LAMA3-B. The invasive and metastatic characteristics of HNSCC are stimulated by the splicing isomer LAMA3-A. Moreover, the overexpression of LAMA3-A serves as an unfavorable prognostic indicator for patients with HNSCC following primary surgery and holds promise as a potential biomarker for disease prognosis. 47
Lysyl Oxidase-Like 2 (LOXL2)
LOXL2, also known as lysyl oxidase-like 2, is a crucial element in controlling the remodeling process of the extracellular matrix. This molecule plays a significant role in facilitating the invasion and metastasis of tumors. One of LOXL2's primary functions is to propel the local invasion of non-hypoxic HNSCC cells, while concurrently encouraging the creation of ecological niches preceding the transfer. 70 Liu et al 48 discovered a distinct splicing isomer called LOXL2 Var in patients with HPV-negative HNSCC. Interestingly, they observed the insertion of a novel 120 bp exon between exons 1 and 2, resulting in the formation of a unique isomer with distinct characteristics compared to the 5′ untranslated regions of LOXL2. Notably, LOXL2 Var exhibited high expression levels specifically in HPV-negative patients with HNSCC. Moreover, functional analyses revealed that LOXL2 Var promoted tumor cell proliferation, migration, and invasion by activating the FAK/AKT signaling pathway through phosphorylation.
MKI67(Ki67)
Ki67, a significant nuclear protein, is encoded by the MKI67 gene in the human genome and serves as a widely recognized marker for proliferation in numerous cancer types. 71 The process of alternative exon 7 splicing in Ki67 generates two distinct isotypes: the full-length isotype, Ki67-F, which includes exon 7, and the shorter isotype, Ki67-Δ7, which lacks exon 7. The presence of exon 7 in this subtype is found to be highly expressed in cancerous tissues and is strongly associated with patients’ survival rates. SRSF3 actively promotes the inclusion of exon 7 of Ki67 by enhancing the splicing of two adjacent exons within exon 7. Consequently, the depletion of this isomer found in exon 7 exhibits a substantial reduction in cell proliferation and tumorigenesis of cancerous cells. 49
Cyclin-Dependent Kinase 2 (CDK2)
CDK2 is important in monitoring various aspects of the cell division cycle. 72 In comparison to typical oral mucosal tissue or primary epithelial cells grown in culture, the expression level of hnRNP A1 is higher in HNSCC tissue or cultured cells, potentially influencing cell proliferation through the regulation of specific splicing events. For instance, the addition or omission of exon 5 of CDK2 might yield subtype 1 or 2 respectively, with increased CDK2 subtype 1 expression supporting cell growth. The findings suggest that hnRNP A1 could govern cell cycle progression by controlling the expression and AS of the CDK genes associated with the G2/M phase of the cell cycle. 50
Signal Transducers and Activators of Transcription 3 (STAT3)
STAT3 is a common oncogene continuously activated in HNSCC through abnormal signals from various growth factor receptors. 73 Selective splicing of exon 23 produces two isoforms of STAT3, namely STAT3α and STAT3β. STAT3α, known as the longer isoform, encodes a protein that comprises the full-length oncogenic STAT3α. On the other hand, STAT3β, is shorter in length and encodes a truncated protein known as the tumor suppressor STAT3β. 74 Wang et al observed a remarkable decrease in the STAT3α/STAT3β isoforms ratio and increase in expression of STAT3α protein upon the excessive expression of splicing factor Poly C Binding Protein 1 (PCBP1) in HNSCC. This demonstrates the crucial role of PCBP1 in the modulation of cellular proliferation and the metastatic processes. 51
Vesicle Transport Factor (USO1)
USO1, a pivotal protein, actively binds vesicles within the endoplasmic reticulum (ER)-Golgi complex, facilitating vital interphase vesicle transport and trafficking.75,76 USO1 possesses two transcript variations: Variant 1 encompasses 26 exons, while Variant 2 comprises 24 exons with the exclusion of exons 5 and 15 found in Variant 1. The enhanced presence of splicing factor Lsm12 in HNSCC results in the augmentation of USO1. Consequently, the overexpression of splicing factor Lsm12 in HNSCC intensifies the incorporation of exon 15, thereby stimulating cellular proliferation and migration. 52
LncRNA H19
Human cancer's AS regulation also involves ncRNAs . The lncRNA H19 is implicated in numerous cancer-related biological processes such as cell proliferation, differentiation, apoptosis, and metastasis. H19 functions as an oncogenic lncRNA in various cancers, enhancing tumor growth and progression. 77 Research indicates that lncRNA H19 facilitates the proliferation, invasion, and migration of tongue squamous cell carcinoma. 78 The H19 allele presents various splicing variants, namely the complete H19 transcript H19-L and the transcript H19-S which lacks exon 4. H19-S modulates the levels of zinc finger E-box-binding homeobox 1 (ZEB1) protein through its binding to ZEB1 mRNA, thus triggering epithelial-to-mesenchymal transition (EMT) and promote the development of oral cancer. 53
Oral Cancer Overexpressed 1 (ORAOV1)
Oral Cancer Overexpressed 1 (ORAOV1), which is alternatively referred to as Tumor Amplified and Overexpressed Sequence 1 (TAOS1), is now recognized as a potential oncogene implicated in oral squamous cell carcinoma (OSCC). It is believed to play a crucial part in cell proliferation, tumor expansion, and tumor angiogenesis, with its amplification and overexpression being observed on chromosome 11q13.79,80 Investigation of the function of ORAOV1 in OSCC led to discovery of two splicing variants of ORAOV1, ORAOV1-A and ORAOV1-B. ORAOV1-A encompasses all exons excluding exon 3, resulting in expression of a truncated 70 amino acid protein that is associated with the level of tumor differentiation. On the other hand, ORAOV1-B is an lncRNA generated through the fusion of exon 2 and exon 3 from the ORAOV1 pre-mRNA. This particular variant exerts a substantial influence on OSCC invasion and metastasis by interacting with heat shock protein 90 (Hsp90) and triggering the nuclear factor kappa (NF-κB)-tumor necrosis factor-alpha (TNFα) signaling pathway.54,81
Autophagy
Transforming Acidic Coiled-Coil Containing protein1 (TACC1)
TACC1 is a constituent of the TACC lineage. It possesses a characteristic enlarged spiral domain of the TACC gene family, commonly referred to as the TACC domain. TACC1 demonstrates close associations with transcription, translation, and the dynamics of the centrosome by interacting with diverse components of elaborate complexes involved in these processes, contributing to their regulation and coordination.82,83 The occurrence and development of different malignant tumors are associated with the dysregulation of TACC1. 84 The nucleus primarily regulates cell proliferation through extracellular signal-regulated kinase (ERK) activity, while c-Fos, a transcription factor downstream of ERK, plays a crucial role in tumor development. Xu et al observed downregulation of an AS variant of TACC1variant25 (TACC1v25) in both HNSCC tissues and cell lines. They further demonstrated that the modulation of ERK phosphorylation and the AKT/mTOR signaling pathway controlled autophagy, thereby exerting anti-proliferative effects. 55
Apoptosis
Myeloid Cell Leukemia-1 (McL-1)
The Bcl-2 family includes vital regulators of cell apoptosis, comprising both pro-apoptotic and anti-apoptotic members, which play critical roles in overseeing cell survival. 85 Mcl-1, an essential member of the Bcl-2 gene family, has a crucial role in development, differentiation, and proliferation, serving as a significant anti-apoptotic factor. 86 Mcl-1 possesses three distinct splicing variants having diverse functionalities: Mcl-1L, Mcl-1S, and Mcl-1ES, each serving different roles with respect to apoptosis. Among these variants, Mcl-1L transcript exhibits a noteworthy elevation in both oral tumors and neighboring normal cells. This overexpression contributes to the development of radiation resistance and anti-apoptotic capabilities within cancer cells. Consequently, Mcl-1L transcript emerges as a potential prognostic indicator for disease-free survival among patients afflicted with oral cancer.56,87
Survivin
Survivin, belonging to the inhibitor of apoptosis (IAP) protein family, which plays a key role in regulating cell survival and apoptosis. The gene encoding Survivin comprises four exons and the protein itself is involved in inhibiting apoptosis (programmed cell death) and promoting cell division, making it an important factor in cancer biology and other cellular processes. Survivin undergoes selective splicing, leading to the generation of different splice variants from its precursor mRNA, including survivin wt (wild-type), survivin ΔEx3 (lacking exon 3), survivin 2B, and survivin 3B. These variants exhibit anti-apoptotic properties, except for survivin 2α, survivin-3α, and survivin 2B, which demonstrate proapoptotic functions. 88 In oral cancer, the anti-apoptotic survivin protein variant was found to be overexpressed, whereas the variant that inhibits apoptosis showed reduced expression. 89 Obatoclaxis (GX15-070), a Bcl-2 inhibitor small molecule, can counteract Mcl-1 and overcome the resistance to apoptosis mediated by Mcl-1.58,59,90 The combination of Obatoclax and cisplatin exhibits a synergistic impact on the viability and growth of oral cancer cells, indicating the significance of eliminating the anti-apoptotic Mcl-1L protein for enhancing the chemosensitivity of these cells.
Angiogenesis
Vascular Endothelial Growth Factor A (VEGF-A)
Tumor growth and development depend on an adequate blood supply, which provides essential oxygen and nutrients and supports metabolic processes. This requirement is typically satisfied through the process of angiogenesis. 91 However, when the balance between pro-angiogenic and anti-angiogenic regulatory factors is disrupted, it results in abnormal angiogenesis, potentially triggering tumor onset. 92 The pre-mRNA of VEGF-A undergoes AS of exon 8, resulting in two variants : VEGF-Axxx and VEGF-Axxxb (xxx demonstrates the amino acid number of mature proteins).VEGF- Axxxb exhibits strong anti-angiogenic activity in vitro and in vivo. 93 VEGF-Ax isomers are usually expressed in all tissues, and VEGF-A165a and VEGF-A165b are the main types of expression isomers. VEGF-Axxx and VEGF-A165b are overexpressed in HNSCC, while VEGF-A isoforms exhibit differential expression in various anatomical sites of head and neck tumors. VEGF-Axxx is overexpressed in pharyngeal tumors, while VEGF-A165b isomer is upregulated in oral tumors. Additionally, the VEGF-A165b isoform shows a direct association with the expression of splicing factors SRSF1, SRSF6, and SRSF5. 60 Novel anti-tumor drugs targeting angiogenesis can not only reduce the vascular network essential for tumor growth and metastasis but also reshape the tumor's immune microenvironment. 94
Immune escape
C-X-C Motif Chemokine Receptor 3 (CXCR3)
CXCR3, a receptor of chemokines, has been demonstrated to stimulate T cells that combat tumors and also other immune cells, such as natural killer (NK) cells, within the tumor microenvironment. This process has the potential to convert tumors that lack immune activity (“cold” tumors) into tumors with enhanced immune response (“hot” tumors), consequently boosting their ability to fight against tumors. 95 There are three isoforms of CXCR3, namely CXCR3-A, CXCR3-B, and CXCR3-Alt. The CXCR3-A isoform is generated through the process of splicing exon 1 with exon 3 of the CXCR3 gene, whereas the CXCR3-B isoform is produced by splicing exon 2 with exon 3. In the tumor microenvironment, CXCR3-A facilitates tumor growth and metastasis, whereas CXCR3-B exhibits anti-proliferative properties and possesses a N terminus that is longer in length in comparison. 96 Immunosuppression of the chemokines CXCL10 and CXCL9 and the two splice isoforms (CXCR3-A, CXCR3-B) of their receptor CXCR3 are upregulated in patients with HNSCC and interferon-α 2b (IFN-α 2b) restores the impaired chemotactic activity of peripheral blood mononuclear cells in patients with HNSCC by regulating the interaction of CXC receptor ligands. The upregulation of CXCR3 was observed in the peripheral blood mononuclear cells of patients with HNSCC, leading to a decrease in their chemotactic function. CXCR3 was downregulated after IFN-α 2b therapy, while CXCR3-A and CXCR3-B were upregulated, and their chemotactic activity was restored, thereby triggering overexpression of CXCR3-A isoforms and driving cell migration, and interaction with CXCL10/CXCL9. CXCL10 affects the immune response to tumors by increasing the number of T lymphocytes/NK cells and boosting the migration of NK/T lymphocytes within the tumor region. 61
Programmed Cell Death Ligand 1(PD-L1)
Immune checkpoints have attracted much attention as one of the important characteristics of the immune microenvironment. The programmed death protein 1 (PD-1)/PD-L1 signaling pathway is one of the key mechanisms of cancer immune escape, which is mainly achieved by inhibiting the activity of T cells in the immune microenvironment. 97 The growth of tumors and the survival rate of solid tumors can be effectively reduced by inhibiting the PD-1/PD-L1 pathway. In the case of HNSCC, a novel mechanism of immune evasion has been found, where the oncogenic virus HPV integrates into CD274 (PD-L1). This integration occurs within the gene locus between exons 4 and 5, leading to an overexpression of the 5′ exon (exons 1-4) of PD-L1. Consequently, a truncated form of PD-L1 is produced, which acts as an isomer inhibiting the function of T lymphocytes via the PD-L1 receptor. Moreover, the presence of this PD-L1 isomer might potentially impact the efficacy of PD-1/PD-L1 blocking antibodies. 62 Furthermore, exon 3 of PD-L1 encodes the structural domain of the immunoglobulin variable (IgV), resembling the IgV structural domain found in antibodies and the T cell receptor. This structural domain serves as a connection between PD-L1 and PD-1. 98 There is limited observation of exon-3 jumping in HNSCC. Yan et al identified a PD-L1 splice variant that did not include exon 3. This variant consisted of the two crucial ESEs. To suppress the function of both ESEs, two highly effective antisense oligonucleotides (ASO) were employed. The blockade resulted in exon 3 jumping, leading to a decrease in PD-L1 production. Consequently, the tumor's ability to escape the immune system was suppressed.62,99
Exosomes
Anilin Actin-Binding Protein (ANLN)
ANLN is situated on chromosome 7p14.2 and codes for a 1125 amino acid protein that binds to actin, contributing significantly to the dynamics of the actin cytoskeleton. 100 Scientific investigation demonstrates that ANLN exhibits elevated expression levels across different types of cancers, including breast carcinoma, ovarian neoplasms, hepatic malignancies, among others.101–103 Dysregulation of ANLN can promote the occurrence and development of tumors. 104 In HNSCC tissues and cell lines, the presence of two primary splicing isoforms of ANLN was identified: ANLN-201 and ANLN-210 transcripts, both exhibiting high expression levels. Disrupting ANLN function effectively inhibits the proliferation, migration, and invasion of OSCC cells such as the SCC-9 cells. On a mechanistic level, ANLN-201 interacts with C-myc, ensuring the stability of its corresponding protein, thereby contributing to the carcinogenic process in HNSCC. ANLN-210 interacts with the RNA binding protein (hnRNPC) and subsequently transfers to macrophages via exosomes. Following secretion of ANLN-210 by macrophages, the exosomes induce polarization of macrophages via the PTEN/PI3 K/AKT signaling pathway, thus stimulating the proliferation of HNSCC.63,105
HPV Infection
HPV infection is known as an independent risk factor for HNSCC. HPV-negative and HPV-positive patients with HNSCC have different molecular characteristics, treatment plans, prognoses, and sensitivity to chemotherapy drugs. HPV-positive patients respond better to treatment (radiotherapy, chemotherapy, surgery) than HPV-negative patients. The regulation of host splicing regulatory factors is crucial for the expression of HPV protein. The hnRNPA1 and hnRNPA2 are the primary factors responsible for regulating HPV expression in HNSCC. They exert their control by modulating the expression level of the E6 protein 106 ; the production of E7 mRNA is facilitated by the 5′ splice site SD226 and the 3′ splice site SA409, whereas E6 mRNA is generated by unspliced viral mRNA. The utilization of SA409 splice sites is suppressed by hnRNPA1, resulting in decreased levels of E7 and increased expression of E6. Conversely, hnRNPA2 exerts an contrasting influence by inhibiting E6 expression.64,107
AS-Related Treatment Strategies
An essential characteristic of head and neck squamous cell carcinoma is the emergence of irregularities in AS. Hence, developing therapeutic approaches for AS has emerged as a major area of interest in the ongoing research of HNSCC. The treatment of AS levels provides remarkable benefits. The spliceosome being a complex machinery responsible for mRNA splicing, its suppression can lead to widespread changes in splicing patterns. Dysregulation of splicing is increasingly recognized as a key pathological event in cancer, as aberrant splicing can lead to the production of cancer-promoting proteins or the loss of tumor-suppressing proteins. Therefore, understanding the role of the spliceosome and splicing dysregulation in cancer hold significant implications for cancer research. Targeting abnormal AS may provide valuable insights for developing new cancer treatment strategies, including antisense oligonucleotides, small biological molecules, Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas gene editing technology, and immunotherapy that can regulate splicing. RNA interference (RNAi) and antisense oligonucleotide approaches are both well-established methods for modulating gene expression. RNAi uses short RNA molecules to target and degrade specific mRNA molecules, while antisense oligonucleotides, are short, single-stranded DNA or RNA molecules complementary to specific mRNA sequences, which can bind to their target mRNA and inhibit translation or promote degradation; both techniques can lead to downregulation of gene expression. In recent years, the newly emerging strategy of steric-blocking oligonucleotides has been to explore new ideas for disease treatment at the level of pre-mRNA splicing.108,109 Figure 4 illustrates some of the treatment strategies related to AS, which are.

Alternative splicing-related therapeutic strategies.
Alternative splicing has the potential to function as a diagnostic or therapeutic objective in clinical practice, and to effectively translate this potential into clinical applications, fundamental research is essential. Bioinformatics technology allows rapid comparisons between normal and pathological conditions, providing insights into AS's role in disease processes and potential biomarkers or therapeutic targets. By leveraging omics data and bioinformatics tools, researchers can enhance our understanding of AS and its implications for disease, ultimately paving the way for the development of novel diagnostic and therapeutic strategies, through in vivo, in vitro, and clinical trial experiments.
Regulators
Compounds or small molecules classified as regulators play a crucial role in the modulation of AS occurrences within tumor cells. These regulators can impact the occurrence and function of AS events by targeting and regulating AS factors or other regulatory mechanisms. By influencing AS, they can alter protein isoform expression, affecting tumorigenesis processes like proliferation, apoptosis, and metastasis, offering a promising approach for cancer therapeutics. In recent times, numerous regulators of AS have been discovered to exhibit inhibitory impacts on the proliferation and metastasis of tumor cells in HNSCC.
Spliceostatin A (SSA) is a small-molecule compound extracted from the fermentation broth of Pseudomonas bacteria. It inhibits the splicing process by binding with the SF3B subunit of the U2 small nuclear ribonucleoprotein in the spliceosome. 110 Studies indicate that SSA not only has strong cytotoxicity against numerous human solid tumor cell lines, but it also significantly increases the life span of mice harboring tumors. Correct pre-mRNA splicing occurs within the nucleus to remove introns and join the exons, forming mature mRNA. Binding of SSA to SF3B inhibits this splicing process, leading to export of pre-mRNA containing introns outside the nucleus, thereby blocking further splicing.111,112SF3B1 is a vital component of the SF3B spliceosome complex. It plays a crucial role in splicing HPV16 E6 to generate non-carcinogenic HPV16 E6*, which may inhibit HPV16-induced tumorigenesis. SF3B1 increases HPV16 E6 mRNA splicing, favoring the E6*I isoform. This suggests that SF3B1 has the potential to impede HPV16-induced HNSCC. PLAD-B (Pladienolide-B), an inhibitor of SF3B1, can induce aberrant splicing patterns, including changes in myeloid cell leukemia-1(MCL1) splicing. In the case of MCL1, a key anti-apoptotic gene, altered splicing promotes the production of MCL1-S, a proapoptotic variant, while reducing the levels of the anti-apoptotic variant, MCL1-L. Moreover, paclitaxel (PTX), which is widely employed in chemotherapy for various types of cancers, including breast cancer and OSCC. However, its efficacy is limited in many patients with OSCC. To address this issue, the ASO SR-3 is utilized to downregulate the expression of SRSF3. This intriguing approach reveals that PTX treatment can suppress the expression of SRSF3 exon 4, thereby reducing the levels of SRSF3 protein. Interestingly, overexpression of SRSE3 reinstates the growth inhibition effect of PTX on OSCC. These findings emphasize that SR-3 effectively reduces the expression of SRSF3 protein, thus enhancing the sensitivity of cancer cells to PTX therapy. Importantly, the expression of SRSF3 exon 4 appears to be closely associated with poor prognosis in OSCC patients. Although nonspecific inhibition of splicing may cause cellular damage, the approach of broadly inhibiting splicing with small molecule compounds may aid in correcting abnormal splicing in cancer. 113 Therefore, ASO targeting of splicing factors may offer a promising new avenue for the treatment of oral cancer. 114
Targeting AS Events
Another treatment strategy related to AS is targeting the expression of molecules involved in AS events. In addition to regulating splicing through inhibition of the core spliceosome, splicing regulatory proteins also promote tumor development through their overexpression and mutations that alter function. These findings underscore the potential of these proteins as novel targets for cancer therapy.66,115,116This method uses RNAi technology or other gene therapy tools to target and inhibit or enhance certain AS processes, altering tumor cell growth and proliferation. The expression of the splicing factor SRSF5 is notably high in both HNSCC tissues and cells. Overexpression of SRSF5 induces transformation of rodent fibroblasts to form tumors in nude mice, while downregulation of SRSF5 in HNSCC impedes cell growth, cell cycle progression, and tumor growth. Additionally, it has been found that SRSF3 (a confirmed oncogene) disrupts the self-regulation of SRSF5 and stimulates its overexpression in cancer cells. This evidence suggests that SRSF5 is a novel target of SRSF3 and is upregulated in HNSCC cells due to the influence of SRSF3. In summary, as an emerging oncogenic splicing factor, SRSF5 plays a crucial role in facilitating HNSCC cell proliferation and promoting tumor formation. 117
Steric-Blocking Oligonucleotides (SBOs)
SBOs can alter the pre-mRNA splicing process while preserving the target mRNA and preventing erroneous exon splicing due to spatial structure. In addition, they promote the generation of accurate subtypes and, in the best-case scenario, can even restore the expression of normal proteins. 118 For example, some exons are difficult to include in mRNA, primarily because they contain ESS sequences or adjacent ISS sequences that recruit splicing repressors, thereby preventing exon inclusion. It is anticipated that SBO binding to these sites will block the recruitment of splicing repressors, thereby promoting exon inclusion. 119 Several instances have demonstrated the usefulness of this strategy. As an illustration, the fusion of particular ligands with receptor for advanced glycation end products (RAGE), can trigger related pathways for inflammatory signaling. Consequently, this can lead to the onset of degenerative ailments or the induction of liver malignancies. 120 The specific ligands can be neutralized by the soluble RAGE subtype (RAGEv1) found in peripheral blood, leading to the inhibition of pro-inflammatory signaling pathways. Modifying the splicing pattern of the RAGE gene to enhance the expression of soluble RAGEv1 presents a promising strategy for the treatment of liver cancers. Additionally, SBOs, when compared to the initial two, exhibit higher resilience towards chemical modifications, thereby possessing drug-like characteristics and offering favorable prospects for clinical application. 108
Conclusions and Future Perspectives
As discussed above, several factors such as heredity, epigenetics, and splicing factors network differential expression tightly influence AS events in HNSCC, resulting in splicing factors or isotypes with distinct differential expression in tumor tissue. Effector molecules frequently assume critical functions within biological mechanisms, including, but not limited to tumor growth, invasion, apoptosis resistance, immune evasion, while also offering potential applications in early detection, prognostic tracking, and targeted therapeutic strategies. This suggests new strategies for cancer monitoring and treatment (Figure 5).
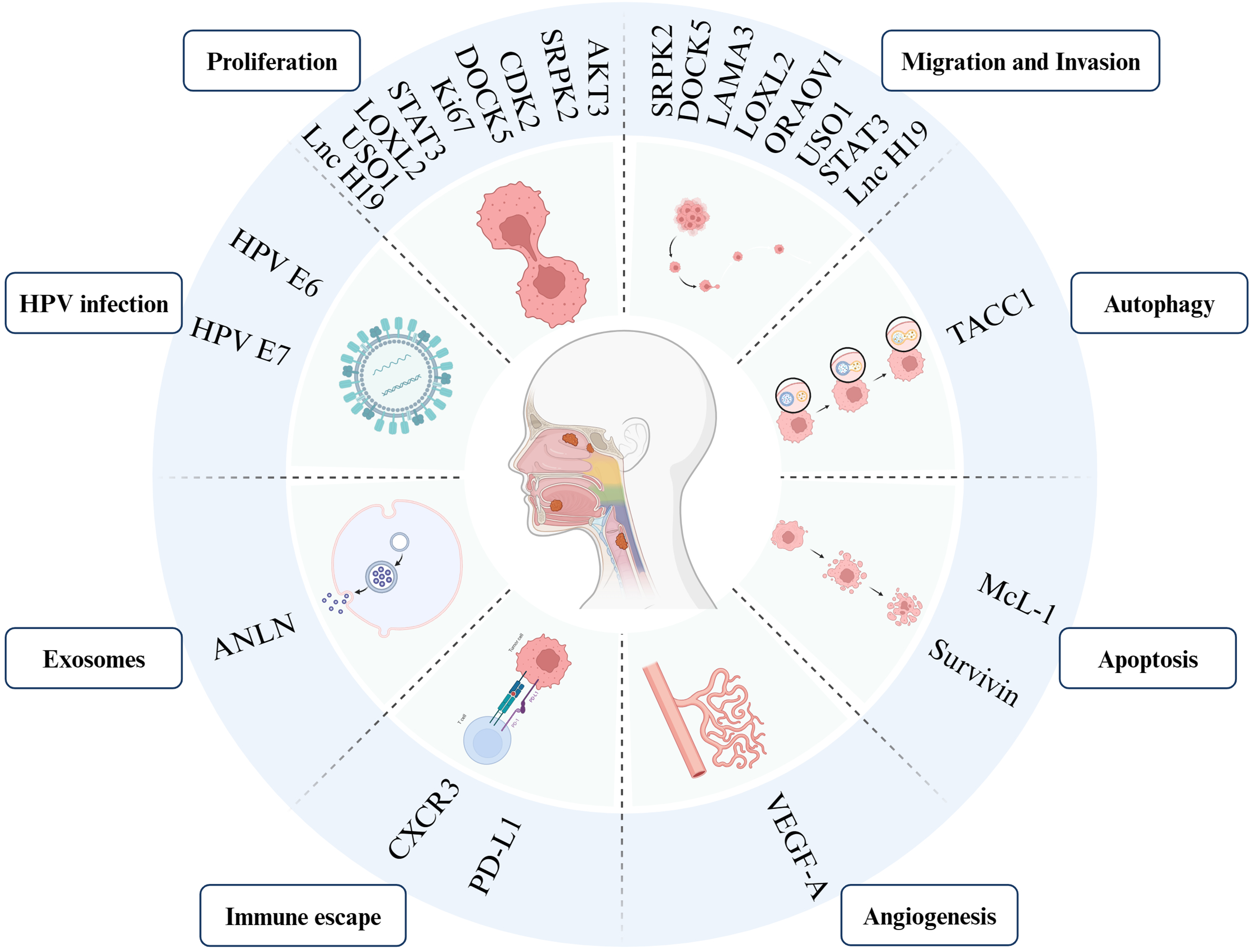
Involved genes affect the biological function of alternative splicing in head and neck squamous cell carcinoma.
Human genetic variations and mutations significantly impact selective splicing, which is becoming an important area of focus in genetics and medical research. Most human disease mutations disrupt precursor mRNA splicing site signals or splicing regulatory elements, thereby producing abnormal mRNA and proteins. 121 It is estimated that these cis-acting splicing mutations account for 60% of human disease mutations. 122 Meanwhile, mutations that affect trans-acting splicing regulatory factors can impair the splicing of multiple downstream target genes, leading to various diseases. 123 Decades of genetic and medical research have firmly established the dominant role of aberrant splicing in Mendelian diseases 123 . However, our understanding remains limited regarding the naturally occurring variable splicing differences among individuals and their impacts on population phenotypes and disease susceptibility. Future research might employ isoform-specific gain- or loss-of-function experiments to reveal how genetic variations in selective splicing influence gene function and thereby affect the phenotype of cells and the entire organism.
Alternative splicing events are typically regulated by cell- or tissue-specific mechanisms. The molecular mechanisms involved in these splicing processes, including changes in the expression of various splicing factors and regulatory proteins, have been extensively detailed in studies of various diseases, particularly in tumor pathology. These molecular changes play a crucial role in regulating gene expression during disease progression. 124 AS therapeutic strategies, as an emerging approach in molecular medicine, show promising clinical applications. By regulating gene splicing patterns, it is possible to specifically alter the expression of mRNA and proteins, correcting disease-associated gene mutations. In cancer treatment, AS strategies inhibit tumor growth and metastasis by modulating tumor cell-specific splicing events. Despite the enormous potential, the clinical application of AS therapeutic strategies faces numerous challenges. For instance, the mechanisms of AS regulation are complex, and there is significant inter-individual variability in splicing patterns, affecting therapeutic efficacy. Additionally, the safety and specificity of AS strategies need further validation, and delivery systems must be optimized to ensure efficient and specific targeting of molecules to the target cells.In HNSCC, systematic analyses of cancer-associated AS events have identified 18 AS events as independent prognostic factors for overall survival and disease-free survival, underscoring the significance of AS in predicting clinical outcomes for patients with HNSCC. 42 With the advent of new strategies, including chemotherapeutic agents, immunotherapies, and molecularly targeted therapies, substantial advances have been achieved in cancer treatment. The interplay between AS and immunotherapy represents a burgeoning area of research, underscoring the potential of AS events in the development of novel targets for cancer immunotherapy. 125 AS can produce unique tumor-specific antigens that are absent in normal cells, thereby enhancing the specificity and efficacy of treatments. 126 Nevertheless, drug resistance in tumors has emerged as a significant barrier in cancer treatment, impeding the efficacy and advancement of therapies. The phenomenon of AS holds the potential to exert a substantial influence on both the expression levels and functionality of drug targets associated with cancer, ultimately bolstering the resistance of tumor cells against chemotherapeutic agents. 113 Cancer drug resistance has been linked to the presence of various isoforms generated by AS in VEGF, FGFR, CD44, androgen receptor, BRCA1, c-FLIP, HER2, and ERα/β. 127 In OSCC, the activation of Dendritic Cells (DC)-mediated tumor immune response by the exon skipping event involving CD44 leads to enhanced cisplatin resistance. 128 Adenoid cystic carcinoma (ACC) is the second most prevalent cancer among HNSCC and exhibits limited responsiveness to existing therapeutic treatments. 129 RNA sequencing revealed that expression of a shortened FGFR1, FGFR1v, makes ACC cells resistant to FGFR1 inhibitors. 130 Hence, a fresh strategy to combat drug resistance in diverse tumors can be found through exploration of the involvement of AS in tumor resistance. It is noteworthy that STF-080310, an innovative inhibitor of Inositol-Requiring Enzyme 1 alpha (IRE1α)/ X-box Binding Protein 1(XBP1), has the ability to enhance the sensitivity of tamoxifen in drug-resistant MCF7 cancer cells by selectively disrupting the splicing of the transcription factor XBP1, which undergoes AS, subsequently reducing the expression of XBP1s. 131 The splicing of serine/threonine kinase can be inhibited by splicing modulators like SSA or its analog meayamycin B, which can target SF3B1. In turn, this sensitizes drug-resistant melanoma to Vemurafenib. 132 Considering the extensive occurrence of drug resistance caused by pre-mRNA splicing, focusing on pre-mRNA splicing could potentially serve as a viable strategy to surmount drug resistance in individuals diagnosed with HNSCC.
With the advancement of technology and extensive research on AS mechanisms, the future directions for therapeutic developmental strategies may include the following:
① Accurate analysis of the functions of AS isomers: There are still many unclear functions of AS isomers, so precise analysis of these isomers is needed. Using high-throughput sequencing technology, proteomics, bioinformatics, and various other methodologies, it may be conceivable to investigate the role and regulatory mechanisms of AS isomers in diverse biological processes in the future.
② Developing new therapeutic drugs targeting AS: In recent years, new therapeutic drugs targeting AS mechanisms have been developed, though, currently no drug can be widely applied in clinical practice. Although there are limited drugs and clinical trials targeting AS-related mechanisms, organ-on-a-chip technology offers new opportunities by simulating the physiological environment and functions of human organs for disease research and drug target screening at the animal and cellular levels. 133 Moreover, mRNA vaccines, as tools for disease therapy, are distinguished by their safety, efficacy, and the convenience of industrial production. AS, as a significant source of novel antigens in immunotherapy, has spurred research into combining immunotherapy with targeting AS mechanisms, opening up potential new avenues. In the future, we may continue to explore the mechanism of AS and develop more effective therapeutic drugs targeting it. ③ Building an AS database: By directly comparing tumor samples with paired normal tissues, deeper insights into the core events of tumor biology can be achieved. However, adjacent normal tissues may not always reflect the origin of tumor cells, and mere changes in expression are not always necessary for splicing variants to function effectively. 134 Single-cell (sc) RNA seq has been a popular technology in biology. With its exceptional resolution, this technology enables precise examination of cellular composition within samples. Consequently, it unveils the genetic blueprint and expression status of individual cells on a mass scale, highlighting the significant diversity that exists among them. 135 This technology has opened up a new fields beyond the study of cell state description. Therefore, a comprehensive database is critical for understanding AS mechanisms and creating new therapeutic medicines.
Our comprehension regarding the regulatory and functional elements of AS in oncology is merely the surface level insight into this complex phenomenon. Many excellent studies have emerged during the last decade of rapid development in AS knowledge-base. Significant advancements have been achieved in the field of AS and the regulation of transcription, as well as translational and post-translational modifications. Additionally, remarkable strides have been made in the realm of genetic disease treatment.108,136,137 These have broken away from the old way of analyzing single AS events and continue to unravel fascinating new properties of AS at a systemic level. It is foreseeable that, with the deciphering of one “splicing code” after another, the study of AS at the histological level will gradually deepen and refine, leading humankind to take a solid step toward uncovering the deepest secrets of the genome.
Footnotes
Abbreviations
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Heilongjiang Province, (grant number LH2021H033).
