Abstract
Keywords
Introduction
Triple-negative breast cancer (TNBC) stands out as a distinct subtype characterized by the absence of estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2), 1 which has been associated with heightened aggressiveness and an elevated risk of metastasis compared to other types. 2 Traditional hormonal therapies and HER2-targeted drugs, effective for other breast cancer types, are typically ineffective against TNBC. 3 The paucity of targeted therapies for TNBC leaves treatment options reliant on chemotherapy, radiation therapy, and surgery. 4 Consequently, there exists an urgent imperative to uncover more efficacious treatments for TNBC.
Numerous studies have highlighted the potential health advantages linked to experiencing moderate hunger, with IF emerging as a dietary strategy that emphasizes timing rather than specific food choices. 5 IF has garnered attention for its potential benefits in metabolic health, weight management, and its impact on chronic diseases.6–8 Recent investigations have explored the influence of IF on breast cancer, especially the aggressive TNBC subtype,9,10 revealing promising outcomes in restraining cancer growth and improving treatment effectiveness.11,12 However, this field remains dynamic, with ongoing research necessary to comprehensively understand the intricate relationship between IF and TNBC.
Recognizing that IF induces systemic shifts across various organs and tissues,13,14 and given the critical role of metabolites in reflecting metabolic states, our study firstly utilized a 5:2 diet model to demonstrate the functional mechanism of IF in high-fat diet-fed mice on TNBC progression. Metabolomic sequencing holds a crucial role in investigating a spectrum of metabolic diseases and associated research.15,16 Decanoylcarnitine, a derivative of carnitine, a naturally occurring compound facilitating the transport of fatty acids into the mitochondria for energy production, 17 comprises a 10-carbon fatty acid chain linked to the carnitine molecule. 18 This compound has gradually become a research hotspot, especially in the context of metabolic studies and its potential implications in various physiological processes.
In this study, we employed serum-targeted metabolomic sequencing to know the metabolic reprogramming in mice undergoing IF, notably a marked increase in decanoylcarnitine levels. We unveiled that decanoylcarnitine could effectively inhibit TNBC cell proliferation and migration. RNA-sequencing data indicated downregulation of extracellular matrix proteins, including Mmp9, suggesting Mmp9 as a potential target.
Materials and Methods
Reagents
DMEM, RPMI1640, FBS, 100 U/mL penicillin and 100 μg/mL streptomycin, and 0.25% trypsin were purchased from Thermo Scientific (Gibco, Grand Island, NY, USA). Diosmetin (HY-N0125), decanoylcarnitine (C10, HY-135035), curcumin (HY-N0005), acetic acid (HY-Y0319), 3-hydroxybutyric acid (HY-113378), taurohyodeoxycholic acid (HY-114360), hippuric acid (HY-W016562), nordeoxycholic acid (HY-N8268), desaminotyrosine (HY-W015346), myo-inositol (HY-B1411), 2-phenylpropionic acid (HY-W015608), hydrocinnamic acid (HY-Y1088), 3-indolepropionic acid (HY-W015229), and polybrene (HY-112735) were purchased from MCE (Shanghai, China). Ilomastat (Cat#S7157) was purchased from Selleck (Shanghai, China). Anti-Mmp9 (Cat#3852) and anti-Ki67 (Cat#9129) antibodies were purchased from Cell Signaling Technology (Boston, MA, USA). Anti-Gapdh antibody (Cat#AF1186) and CCK8 kit (Cat#C0038) were purchased from Beyotime (Shanghai, China). RNAiso Plus (Cat#9109) and PrimeScript™ RT Master Mix (Cat#RR036A) were purchased from Takara (Beijing, China). SYBR Green qPCR Master Mix (Cat#11202ES08) was purchased from Yeason (Shanghai, China). Puromycin (Cat#540411) was purchased from Sigma (St Louis, MO, USA).
Animal Model
All animal experiments were conducted in strict accordance with the procedures approved by the ethics committee of Animal Experiments at Nanjing Medical University (Approval No. IACUC-2308039). 8-Week-old female Balb/C mice were procured from the Model Animal Research Center of Nanjing University. The mice were housed at a temperature of 22 °C under a 12-hour light/dark cycle, with free access to a high-fat diet (research diets, 12492) and water for 8 weeks. Subsequently, the mice were randomly assigned to experimental groups. To induce IF in mice, a 5:2 diet regimen was employed. This involved allowing the mice to eat freely for 5 days a week, interspersed with 2 days of fasting, with unrestricted access to water during fasting periods. For the normal diet (ND) group, mice (n = 6) had continuous free access to food. In the intermittent fasting (InD) group, mice (n = 6) were deprived of food for 24 hours every other 2 or 3 days, with ad libitum feeding on the intervening days until the end of the study. Two weeks into the diet regimen, 1 × 106 4T1 cells suspended in 100 µL of PBS were inoculated subcutaneously in the fourth mammary of all the mice. Tumor volume was measured regularly using a digital caliper, and the calculation was performed as length × width2 × 0.5. Mice were euthanized if they displayed any signs of sickness or when the tumor size reached 2000 mm3. Following euthanasia, mouse serum, tumor, lung, and liver tissues were collected for further research.
Cell Culture
The 4T1, MDA-MB-231, HEK293 T cell were purchased originally from American Type Culture Collection (Manassas, VA, USA). The 4T1 were cultured in RMPI1640, MDA-MB-231, and HEK293 T were cultured in high-glucose DMEM medium, all the medium was supplemented with 10% fetal bovine serum, 100 U/mL penicillin, and 100 mg/mL streptomycin, respectively. Cells are maintained in a humidified incubator at 37 °C with 5% CO2. The medium is refreshed every 2 to 3 days. Upon reaching 90% confluency, the old medium is aspirated, and trypsin is added for cell detachment and subsequent subculture.
Total RNA Isolation and Quantitative Real-Time PCR (RT-qPCR)
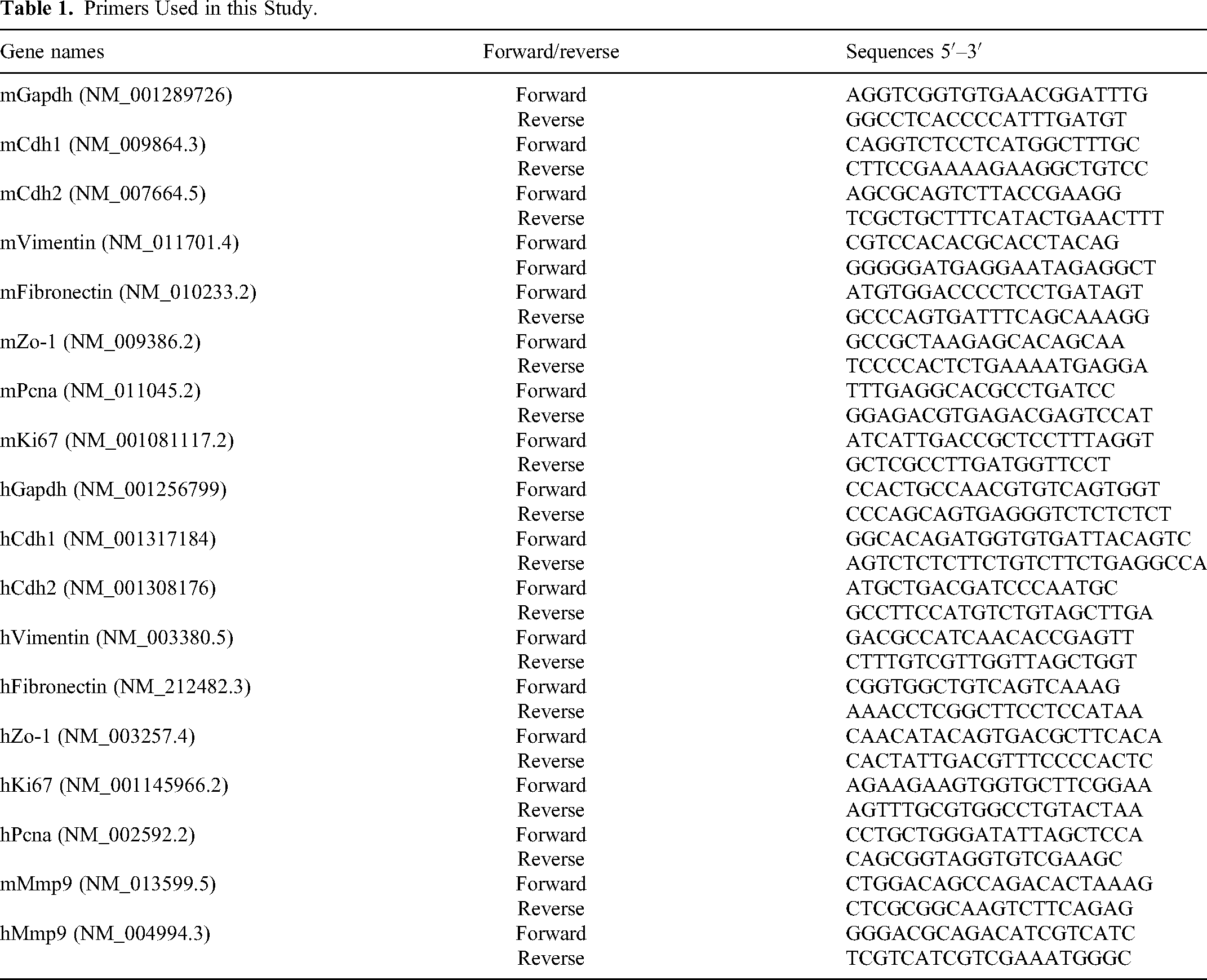
According to the instructions provided by RNAiso Plus (Takara, Beijing, China), total RNA were extracted and performed reverse transcription using PrimeScript™ RT Master Mix (Takara, Beijing, China). Subsequently, qPCR was performed utilizing SYBR Green qPCR Master Mix (Cat#11202ES08, Yeason, Shanghai, China). GAPDH was employed as the internal reference gene for mRNAs. The primer sequences utilized in this study are detailed in Table 1. The relative expression levels of each gene were determined using the 2−(ΔΔCt) method.
Primers Used in this Study.
Metabolomics Analysis
Serum samples were meticulously collected from two groups of six mice each. For each analysis, A 20 μL of serum and 20 μL of standard were combined, and 120 μL of the sample release agent was added. The mixture was shaken at 1200 rpm for 30 minutes and then centrifuged at 18,000 g at 4 °C for 30 minutes. Following centrifugation, 30 μL of the supernatant was transferred to a 96-well plate. To this, we added 20 μL of derivatization reagent and 20 μL of EDC working solution. The 96-well plate was covered with aluminum film and placed in a constant temperature shaker, reacting at 40 °C and 1200 rpm for 60 minutes. Following the shaking process, centrifugation was carried out at 4000 g at 4 °C for 5 minutes. Subsequently, 30 μL was taken and added to a new 96-well plate. To each well, 90 μL of sample diluent was added, and the mixture was mixed at 600 rpm for 10 minutes. Centrifugation was then performed at 4000 g at 4 °C for 30 minutes. The plate was sealed with film in preparation for LC–MS/MS acquisition. Sample extracts were analyzed using Waters UPLC I-Class Plus (Waters, USA) equipped with QTRAP 6500 Plus (SCIEX, USA), and data analysis was conducted by BGI Genomics.
Mmp9 Overexpression
Mouse Mmp9 (NM_013599) overexpressed lentivirus plasmds was generated by ligating the full-length open-reading frame into the pCDHPuro-IRES-GFP vector (Addgene). High-titer lentivirus was packaged in HEK293 T cells using Lipofectamine transfection. Viral supernatants were collected twice at 24-hour intervals after transfection, filtered and applied to 4T1 breast cancer cells in the presence of 1 μg/mL polybrene for 16 hours. Cells were selected with puromycin for about 1 week. The expression level was assessed by immunoblotting.
Cell Migration Assay
Cells from their different groups were harvested, trypsinized for digestion, centrifuged, and resuspended in a culture medium. A cell count was performed, and cell density was adjusted to 1 × 106 cells/mL, and 200 µL of the cell suspension was seeded into the upper chamber of a Transwell apparatus. The lower chamber was filled with 500 μL of DMEM culture medium containing 10% serum, and the setup was incubated for 24 hours. Following incubation, the Transwell chamber was removed, and cells were fixed with 4% paraformaldehyde for 15 minutes. Hematoxylin and eosin (HE) staining was then performed according to the manufacturer's instructions (Beyotime). Ten random fields of view were selected for cell counting under an inverted microscope.
Immunoblotting Analysis
Cell lysates were prepared using RIPA lysis buffer, and the protein concentration in the supernatants was quantified. We loaded 30 μg of protein per sample onto SDS–PAGE gels for electrophoresis, followed by transfer onto membranes. To block nonspecific binding, the membranes were incubated with 5% skimmed milk at room temperature for 2 hours. We then probed the membranes with a primary antibody against Mmp9 at a dilution of 1:1000 and incubated them overnight at 4 °C. This was followed by incubation with the appropriate secondary antibody at a dilution of 1:5000 for 1 hour at room temperature. Protein bands were visualized using a ProteinSimple FluorChem E gel imaging system (ProteinSimple, CA, USA), allowing for the identification and analysis of target proteins.
RNA Extraction, Library Preparation and Sequencing
Total RNA was extracted from 4T1 or MDA-MB-231 cells treated with various metabolites or Ilomastat using TRIzol® Reagent (Takara, Beijing, China), following the manufacturer's protocol. RNA quality was assessed using a Nanodrop ND-2000 system (Thermo Scientific, USA) for A260/A280 absorbance ratio and an Agilent Bioanalyzer 4150 (Agilent Technologies, CA, USA) for RNA integrity number (RIN). Only high-quality RNA samples were used for library construction. The ABclonal mRNA-seq Lib Prep Kit (ABclonal, China) was utilized to prepare paired-end libraries, according to the manufacturer's instructions. mRNA was isolated from 1 μg of total RNA, fragmented, and used for first-strand cDNA synthesis with random hexamer primers and Reverse Transcriptase (RNase H). Second-strand cDNA was synthesized using DNA polymerase I and RNaseH. The resulting double-stranded cDNA fragments were adapter-ligated and PCR amplified. PCR products were purified using the AMPure XP system. Library quality was assessed on an Agilent Bioanalyzer 4150. Libraries were then sequenced on an Illumina Novaseq 6000, generating 150 bp paired-end reads.
Differential Expression Analysis
Differential expression analysis was performed using DESeq2 (http://bioconductor.org/packages/release/bioc/html/DESeq2.html). Genes exhibiting an absolute log2 fold change (|log2FC|) greater than 1 and an adjusted p-value (Padj) of less than 0.05 were considered significantly differentially expressed. This rigorous process allowed for the accurate identification of key genes impacted by the treatment in the study, providing vital insights into the molecular mechanisms underpinning the observed cellular responses.
Enrichment Analysis
For our analysis, we utilized the clusterProfiler R software package, which is designed for comprehensive gene ontology (GO) function enrichment and Kyoto encyclopedia of genes and genomes (KEGG) pathway enrichment analysis. We set a significance threshold at P < 0.05 to identify significantly enriched GO terms or KEGG pathways. This approach allowed us to effectively categorize genes based on their biological functions and pathway involvement, providing a deeper understanding of the underlying molecular processes.
Statistical Analysis
Experimental data were presented as mean ± standard deviation (x ± s) and analyzed using GraphPad Prism 8.0 software (GraphPad, San Diego, CA, USA). For comparisons between two groups, Student's t-test was applied, while one-way ANOVA was used for multiple group comparisons. A P-value of less than 0.05 was considered statistically significant. This statistical approach ensured the robustness and reliability of our findings, allowing for meaningful interpretation of the data.
Results
IF Improves Obesity-Related TNBC Progression
To assess the impact of IF on obesity-associated TNBC, firstly, we established a high-fat diet (HFD)-induced mouse model and implemented an IF regimen (5:2 diets). Two weeks later, 4T1 cells were inoculated into the fourth intramammary gland, and we evaluated tumor growth and metastasis to liver and lung tissues to assess breast cancer progression (Figure 1A). Notably, a marked reduction in body weight was observed after 2 weeks of IF and a further significant decrease in body weight was observed 2 weeks after tumor cell inoculation (Figure 1B). Importantly, we found IF led to the inhibition of tumor growth and reduced metastasis to the lung and liver (Figure 1C–F). Taken together, these findings suggest that IF inhibits the TNBC progression in obese mice.

Impact of IF on obesity-related TNBC progression. (A) Schematic of the animal experiment procedure. Mice were fed a HFD for 8 weeks, then randomly divided into normal diet (ND, n = 6) and IF diet (InD, n = 6) groups. Two weeks later, all mice were inoculated with 1 × 106 4T1 cells in the fourth mammary pad for tumor progression analysis. (B) Graph showing body weight changes throughout the mouse modeling process. (C) Tumor volume comparison in different mice. (D) Weights of tissues from different mice. (E) Number of tumor metastases of different mice. (F) Hematoxylin and eosin (HE) staining and immunohistochemistry (IHC) of Ki67 in various tissues. Data are mean ± SEM. *P < 0.05; **P < 0.01. Scale bars, 200 μm.
Targeted Metabolomics Reveals the Serum Metabolic Profiles Involved in the Process of IF Improving TNBC in Mice
Next, we set out to unravel the mechanism of IF inhibits cancer cell proliferation and migration. As IF can induce systemic metabolic changes in the body, and the metabolites in the blood can directly reflect the metabolic status of the organism. We analyzed serum metabolites of IF mice using HM700 targeted metabolomics, and identified 349 metabolites (Figure 2A). Principal component analysis (PCA) demonstrated clear clustering and good reproducibility between control and IF groups (Figure 2B). The top three major categories among the 349 metabolites were amino acids and peptides, fatty acids, and bile acids (Figure 2C). Categorization based on KEGG metabolic pathways revealed amino acids metabolism, lipid metabolism, and carbohydrate metabolism as the top three major categories (Figure 2D and E). This comprehensive metabolite analysis provides insights into the blood metabolome of a TNBC mouse model, indicating the beneficial effects of IF may be achieved through alterations in the metabolite profile.

Serum-targeted metabolomics in TNBC mice undergoing IF. (A) Workflow of serum-targeted metabolomics. (B) PCA plot summarizing biological sample data. (C) Bar chart classifying metabolites. Vertical axis: metabolite categories; Horizontal axis: metabolite count. (D) Bar chart of metabolic pathway classification. Vertical axis: pathway categories; horizontal axis: metabolite count. (E) Ring chart categorizing metabolites. Different colors represent various metabolite categories, with percentages indicating their proportion.
In-Depth Screening of Differential Metabolites
To explore the crucial metabolite involved in the IF-mediated inhibition of TNBC, we conducted a comprehensive analysis of all metabolites. Among the 349 detected serum metabolites, 69 were identified as differential metabolites (27 upregulated and 42 downregulated) after IF (Figure 3A). The top 10 upregulated metabolites included diosmetin, decanoylcarnitine (C10), curcumin, acetic acid, 3-hydroxybutyric acid, taurohyodeoxycholic acid, 3-nitrotyrosine,

Detailed analysis of differential metabolites. (A) Volcano plot showing differential metabolites. (B) Top 10 differential metabolites. (C) Heatmap illustrating correlations among differential metabolites. (D) Network diagram of metabolite correlations. (E, F) Concentrations of top upregulated metabolites (E) and downregulated metabolite (F), analyzed by HM700 metabolisome. Data were presented as mean ± SEM. *P < 0.05; **P < 0.01.
Decanoylcarnitine (C10) Inhibits the Migration and Proliferation of TNBC Cells
The highly invasive and metastatic traits of TNBC significantly contribute to its high mortality rate. Our research thus focused on the effects of differential metabolites on cell proliferation and epithelial-to-mesenchymal transition (EMT), a key factor for cell migration, of 4T1 cells. Assessment of the top three upregulated and downregulated metabolites revealed that decanoylcarnitine could effectively inhibit cell proliferation-associated genes Ki67 and Pcna (Figure 4A and B). It also suppressed mesenchymal-specific genes Cdh2 and Vimentin (Figure 4C and D), while promoting epithelial-specific genes Cdh1 and Zo-1 (Figure 4E and F). Migration assays further confirmed Decanoylcarnitine's significant role in curtailing cell migration (Figure 4G). These results strongly suggest that decanoylcarnitine is a key modulator of cell proliferation and migration in the IF model of TNBC.

Decanoylcarnitine's role in inhibiting TNBC cell migration and proliferation. (A–F) qPCR analysis of proliferative genes (Ki67, Pcna), mesenchymal genes (Cdh2, Vimentin), and epithelial genes (Cdh1, Zo-1) in 4T1 cells treated with top differential metabolites. (G) Transwell assay of cells treated with top differential metabolites. The concentration of each metabolite is 10 μM, and the vehicle used is an equal volume of DMSO. Data were presented as mean ± SEM. *P < 0.05; **P < 0.01. Scale bars, 50 μm.
RNA-seq Analysis of Decanoylcarnitine-Treated TNBC Cells
For a deeper understanding of Decanoylcarnitine's role in TNBC, we treated 4T1 cells with this compound and conducted RNA-sequencing (RNA-seq) analysis (Figure 5A). This revealed 539 differentially expressed genes, with 174 upregulated and 365 downregulated (Figure 5B and C). GO analysis of the downregulated genes indicated significant involvement in cell adhesion and extracellular matrix binding (Figure 5D), with key transcription factors like zf-C2H2, TF-bZIP, and bHLH identified (Figure 5E). KEGG pathway analysis underscored the importance of ECM-receptor interaction in decanoylcarnitine-treated cells (Figure 5F). This evidence suggests that decanoylcarnitine inhibits breast cancer cell migration by affecting the cell-extracellular matrix process.

RNA-seq analysis of decanoylcarnitine-treated TNBC cells. (A) Schematic of RNA-seq in 10 μM decanoylcarnitine-treated 4T1 cells. (B) Differentially expressed genes in RNA-seq data. (C) Heatmap of 539 differentially expressed genes (DEGs). (D) Gene ontology (GO) enrichment for downregulated genes. (E) Transcription factor annotation for DEGs. (F) KEGG analysis of downregulated DEGs.
Mmp9 is a Potential Target of Decanoylcarnitine in TNBC Cells Progression
To validate the RNA-seq data, we assessed the differential expression profiles, revealing a substantial downregulation in the expression levels of various genes associated with the extracellular matrix. Notably, Mmp9 stood out as the gene with the most significant difference (Figure 6A). It is widely acknowledged that the functional roles of proteins necessitate regulation and mediation by other proteins, typically facilitated through binding or interactions. The interactome data unveiled that Mmp9 engaged with various extracellular matrix proteins (Figure 6B). An in-depth examination of Mmp9 expression in breast cancer cells revealed significant suppression with the administration of decanoylcarnitine (Figure 6C and D). Additionally, an analysis of expression levels in primary tumor, lung, and liver tissues from IF breast cancer mice demonstrated significant attenuation of Mmp9 expression (Figure 6E). This evidence indicates Mmp9 is a potential target of decanoylcarnitine in TNBC cancer cells progression.

Mmp9 is involved in inhibition of TNBC cell migration by decanoylcarnitine. (A) Volcano plot showing gene expression distribution. (B) Interaction network of the top DEGs. (C, D) qPCR and Immunoblotting analysis of Mmp9 expression in 10 μM Decanoylcarnitine-treated cells. (E) IHC analysis of Mmp9 in tissues from IF mice. Data are presented as mean ± SEM. *P < 0.05; **P < 0.01. Scale bars, 200 μm.
Mmp9-Mediated Decanoylcarnitine-Inhibited TNBC Cells Progression
To further investigate the role of Mmp9 in the process of TNBC progression. We adopted Ilomastat, a Mmp9 inhibitor, to know the role of Mmp9 in TNBC progression. The results show that Ilomastat could significant inhibits cell proliferation and cell migration (Figure 7A–D). To further confirm the role of Mmp9 in the process of decanoylcarnitine-inhibited TNBC cells progression. We overexpressed Mmp9 in decanoylcarnitine-treated TNBC cells, and found Mmp9 could rescue the inhibitory effect of decanoylcarnitine in TNBC cells (Figure 7E–G). These collective findings indicate that IF upregulates decanoylcarnitine in the circulatory system, subsequently suppressing the expression of Mmp9 (Figure 7H).
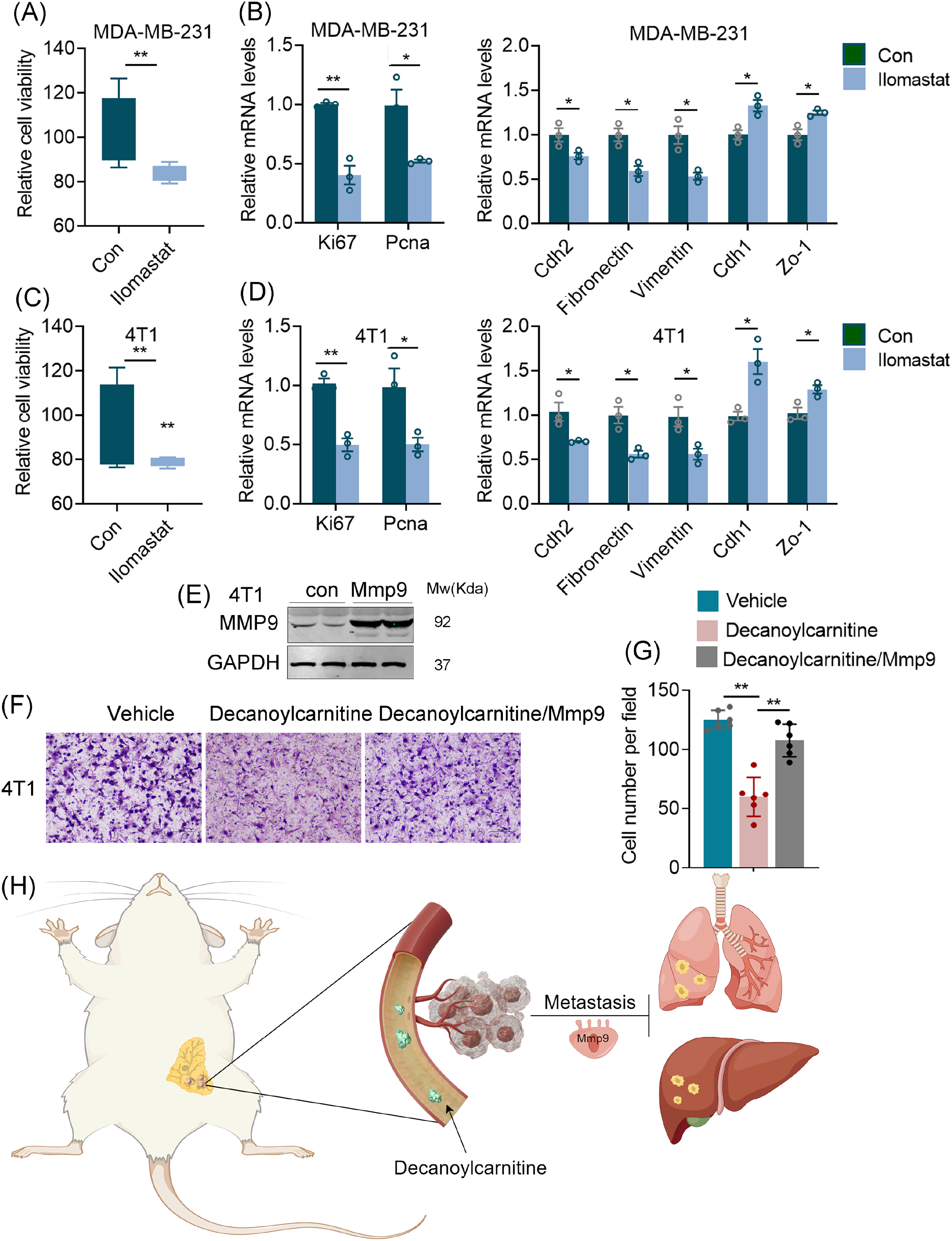
Mmp9-mediated inhibition of TNBC cell migration by decanoylcarnitine. (A–D) The CCK8 assay and qPCR analysis of cell proliferation, proliferative genes (Ki67, Pcna), mesenchymal genes (Cdh2, Vimentin, Fibronectin), and epithelial genes (Cdh1, Zo-1) in MDA-MB-231 (A, B) and 4T1 (C, D) cells treated with 10 nM Ilomastat for 24 hours. (E) Immunobloting assay to detect the expression of MMP9 in Mmp9 overexpressed 4T1 cells. (F) The Transwell assay to detect the effect of Mmp9 overexpression on decanoylcarnitine-inhibited cell migration. (G) The cell number per field of transwell assay. (H) Schematic depicting decanoylcarnitine's mechanism in TNBC inhibition. *P < 0.05; **P < 0.01. Scale bars, 50 μm.
Discussion
Over the years, IF has been extensively researched for its positive impact on various metabolic pathways, including insulin sensitivity, 19 reduce inflammation, 20 and promote autophagy, 21 all crucial physiological responses for overall health and disease prevention. Notably, recent studies suggest that IF may regulate the tumor microenvironment to mitigate the progression of TNBC,22,23 adding another layer to the array of benefits associated with this dietary regimen. Though increasing studies investigate the benefits of IF diet in cancer cells, very few have considered this from a systematic metabolic view and address how metabolites integrate signaling pathway to slow down cancer progression. Our study is groundbreaking as it sheds light on the positive impact of metabolites of IF mouse model on pulmonary metastasis of TNBC in obese mice, emphasizing the pivotal role of dietary patterns related metabolites change in the progression of TNBC. Our study illuminates the positive impact of metabolites from the IF (5:2 diet) mouse model on cancer progression of TNBC in obese mice.
Our research provides hitherto undocumented evidence that decanoylcarnitine, a carnitine ester with decanoic acid widely present in the human body, can influence the invasive ability of breast cancer cells. It exhibits a significant decrease and can serve as a biomarker for lung metastasis in lung tumors, suggesting its potential as a biomarker for lung metastasis with high diagnostic accuracy. 24 Additionally, its application in diagnosing oral squamous cell carcinoma in cases of oral erosive lichen planus has been identified, 25 while in renal cell carcinoma, it is notably more abundant in cancer tissue and urine compared to normal samples. 26 The heterogeneity of decanoylcarnitine in different tumors underscores the need for a comprehensive exploration of different cancer types. Notably, decanoylcarnitine levels in plasma significantly decrease in patients with esophageal squamous cell carcinoma and show notable recovery following radiation and chemotherapy. This highlights its potential not just in diagnosis but also in monitoring treatment efficacy. 27 The heterogeneity of decanoylcarnitine in various tumors underscores the need for further studies, particularly larger clinical trials focusing on breast cancer, to validate its clinical utility. Although significant research has been devoted to exploring the diagnostic potential of decanoylcarnitine in clinical settings, there is a notable gap in understanding how it influences tumor progression. Our study, employing transcriptome sequencing has revealed that decanoylcarnitine markedly inhibits Mmp9 expression. Mmp9, known for its ability to degrade collagen and facilitate basement membrane degradation, plays a critical role in promoting cancer cell migration, invasion, and metastasis.28,29 This finding positions decanoylcarnitine as a pivotal factor in affecting TNBC outcomes, evident in its capacity to curb cell proliferation and migration and impact extracellular matrix-related proteins, such as Mmp9. These insights offer a glimpse into how IF could modulate the cancer microenvironment.
While our study contributes valuable insights, it is essential to acknowledge its limitations. Firstly, the exploration of decanoylcarnitine's role in the progression of TNBC primarily centers around TNBC cell lines. To strengthen the robustness and applicability of our findings, future research endeavors should prioritize the inclusion of in vivo data, allowing for a more comprehensive understanding of decanoylcarnitine's impact in a physiological context. Secondly, the direct functional mechanism of decanoylcarnitine remains a subject that warrants further detailed investigation. While our study provides initial insights, a more in-depth examination is necessary to unravel the intricacies of decanoylcarnitine's specific actions and interactions within the cellular milieu. This area of research presents an opportunity for future investigations to delve deeper into the molecular mechanisms underpinning the influence of decanoylcarnitine in the context of TNBC. Addressing these research gaps will contribute to a more nuanced understanding of decanoylcarnitine's role and its potential implications for TNBC therapeutic interventions.
While these results are promising, further research is necessary to comprehensively understand the molecular intricacies and validate these findings in a broader population. Despite encouraging preclinical data, the lack of extensive clinical trials assessing the impact of IF on breast cancer across diverse populations poses a significant challenge.9,30 Since robust clinical evidence is essential to form conclusive insights. Further clinical trials are crucial to substantiate and expand upon these preliminary findings.
Conclusion
In summary, the hypothesis that IF might mitigate TNBC progression enriches the ongoing discourse about the health benefits of this dietary practice. Persistent research efforts are vital to uncover the underlying mechanisms and evaluate their wider implications for human health and disease management. Our current understanding is primarily based on targeted metabolomics and transcriptome sequencing data, with limited in further validation. Future research will aim to thoroughly demonstrate and validate the role of decanoylcarnitine in metabolic enhancement and breast cancer progression inhibition.
Footnotes
Authors’ Contributions
The results presented in this article have not been previously published in whole or in part. The experiments were conceived and designed by Ninghan Feng and Dongmei Wang. The main experiments were conducted by Yifan Tang, Shuai Chen, while data analysis was performed by Yifan Tang, Shuai Chen, Saijun Wang, Kun Zhang, Ke Xu. The results were discussed and guided by Ninghan Feng and Dongmei Wang. The manuscript was written by Ninghan Feng, Dongmei Wang, Yifan Tang, and Shuai Chen. The article was reviewed and approved by all the authors.
Authors’ Note
There are no human subjects in this article and informed consent is not applicable. This project does not encompass experiments involving humans; hence, there is no necessity for an informed consent form. Animal experiments were conducted with the primary objective of minimizing the quantity of animals involved and alleviating their distress (Approval no. IACUC-2308039).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from National Natural Science Foundation of China (82273232), Changzhou Medical Center of Nanjing Medical University Program (CZKY102RC202301, JX10205000).
