Abstract
Introduction
Breast cancer is one of the most common malignancies in women across the world.1,2 Although the great progress of early discovery and multiple treatment modalities, led to a decrease in incidence and mortality rates, the patients with advanced stages and metastasis had disappointing survival rates.3,4 Hence, more research is needed to investigate the pathological mechanisms of breast cancer progression and develop effective targets for breast cancer diagnosis and treatment.
Long non-coding RNAs (lncRNAs) are a class of transcripts greater than 200 nt, which lacks an effective open reading framework. 5 Recent studies have shown that lncRNAs exert multiple biological functions in various cancers including breast cancers. 6 For example, lncRNA, breast cancer-related transcript 1 (BCRT1), HOX transcript antisense RNA (HOTAIR), a regulator of phospholipid metabolism (ROPM), and NR2F1 antisense RNA 1 (NR2F1-AS1) have been reported to be associated with the occurrence and development of breast cancer.7–10 Long intergenic noncoding RNA 460 (LINC00460) is located in the chr13q33.2 region and has been demonstrated to be increased in many types of solid tumors, such as colorectal cancer, hepatocellular carcinoma, lung cancer, and renal cell carcinoma. 11 The levels of LINC00460 were upregulated in the breast cancer tissues and positively associated with lymphatic metastasis and poor overall survival of patients with breast cancer. 12 Moreover, LINC00460 was identified as an oncogenic gene and promoted breast cancer progression through the miR-489-5p/Fibroblast Growth Factor 7 (FGF7)/AKT axis. 12 However, the specific mechanisms of LINC00460 in the modulation of breast cancer remain undetermined.
As a kind of programmed cell death, ferroptosis, first coined by Dr Brent R Stockwell in 2012, is characterized by the accumulation of lethal lipid peroxides in an iron-dependent manner.13,14 Amounting evidence showed that ferroptosis plays an important role in the initiation, development, invasion, metastasis, and therapy resistance of cancer. 13 For instance, F-box and WD repeat domain-containing 7 (FBW7) regulated lipid peroxidation, promoted ferroptosis, and potentiated the cytotoxic effect of gemcitabine in pancreatic cancer cells through the nuclear receptor subfamily 4 group A member 1 (NRA41)-stearoyl-CoA desaturase (SCD1) axis. 15 Liu et al 16 noted that chronic 27-hydroxycholesterol (27HC) enhanced the tumorigenic and metastatic capability of breast cancer cells by promoting resistance to ferroptosis. Therefore, it is worth exploring the roles of ferroptosis in regulating cancer progression.
In our studies, we found that the expression of LINC00460 was significantly increased in breast cancer tissues and associated with the prognosis of patients with breast cancer. Furthermore, LINC00460 knockdown inhibited cell proliferation and promoted ferroptosis of breast cancer cells by downregulating the expression of myelin and lymphocyte protein 2 (MAL2), a lipid raft-resident protein of the MAL family. In addition, we found that LINC00460 modulated MAL2 expression by sponging miR-320a.
Materials and Methods
LncRNA Expression Profiling Data Analysis
The expression of LINC00460 in cancer patients was also obtained from the UALCAN database (http://ualcan.path.uab.edu/) and the GEPIA database (http://gepia.cancer-pku.cn/) with TCGA data. The LINC00460 expression in breast cancer was also analyzed using GSE39431 data through the lnCAR database (https://lncar.renlab.org/). For overall survival analysis, the UALCAN database, the TANRIC database (https://ibl.mdanderson.org/), and GSE16446 data from the Kaplan-Merier Plotter (http://kmplot.com/) database were used. Additionally, GSE16446 data from the Kaplan-Merier Plotter was also used to analyze the disease-free survival rate.
Cell Culture
Both MDA-MB-231 and BT-549 cells were obtained from American Type Culture Collection (ATCC, Manassas, VA, USA). MDA-MB-231 cells were cultured in Dulbecco's modified Eagle's medium (DMEM, Biological Industries, EitHaemek, Israel, #06-1170-87-1A), and BT-549 cells were grown in RPMI-1640 medium (Biological Industries, #01-104-1A). The medium was supplemented with 10% fetal bovine serum (FBS, Biological Industries, #04-001-1ACS), penicillin/streptomycin (1%, Beyotime, Shanghai, China, #C0222), L-glutamine (1%, sigma. Sigma-Aldrich, St. Louis, MO, USA, #56-85-9) and recombinant human EGF (20 ng/ml, Pepro Tec Inc, MD, USA, #AF-100-15).
Cell Transfection and Infection
LINC00460 siRNA-1 (5′-AGACCTAATAGCCAATAAG-3′) and LINC00460 siRNA-1 (5′-CCATGTGAAGTGTAGAACA-3′) were purchased from RiboBio Life Science Co., Ltd (Guangzhou, China, #siG161024092500-1-5 and #siG161024092509-1-5). miRNA mimic negative control (NC), miR-320a mimic, miRNA inhibitor NC, and miR-320a inhibitor were obtained from Genepharma (Suzhou, China). The MAL2 overexpression plasmids were purchased from Nanjing Ruizhen Biotechnology Co., Ltd (Nanjing, China). Transfections were performed using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA, #11668) accordingly. The LINC00460 shRNA lentivirus (LV-LINC00460 siRNA-1) carrying the sequence of LINC00460 siRNA-1 (5′-AGACCTAATAGCCAATAAG-3′) was established and packaged by Genepharma. For lentivirus infection, LV-LINC00460 siRNA-1 was used to infect MDA-MB-231 and BT-549 cells at a multiplicity of infection (MOI) of 50.
RNA Extraction and Real-Time Quantitative PCR
Total RNA was isolated using RNA isolater Total RNA Extraction Reagent (Vazyme, Nanjing, China, #ER501-01). To detect the gene expression, 1 μg of total RNA was reverse-transcribed into cDNA with a HiScript III First Strand cDNA Synthesis Kit (+gDNA wiper) kit (Vazyme, #R312). Then, the cDNA was used for real-time quantitative PCR (RT-qPCR) by using the AceQ Universal SYBR qPCR Master Mix kit (Vazyme, #Q511) according to the manufacturer's instructions. GAPDH was used as an internal control. To detect the expression of miRNA, total RNA (1 μg) was reverse-transcribed with a miRNA First Strand cDNA Synthesis Kit (by stem-loop) kit (Vazyme, #MR101), and the RT-qPCR was performed using the miRNA Universal SYBR qPCR Master Mix kit (Vazyme, #MQ101). U6 was used as an internal control. The primer sequences for RT-qPCR are described in Supplemental Table S1.
Cell Counting Kit-8 Analysis
Cells (1 × 103/well) were grown in 96-well plates in quintuplicate. At designated time points (0, 24, 48, and 72 h), cell counting Kit-8 (CCK-8) reagent (10 µL, Dojindo, Kumamoto, Japan, #CK04) was added to each well. After incubation for 2 h at 37 °C, the optical density at 450 nm was measured.
Colony Formation Assay
Cells (1 × 103/well) were seeded into 6-well plates in triplicate and cultured for 2 weeks. The cell colonies were washed with PBS, fixed with 4% paraformaldehyde (Sangon Biotech, Shanghai, China, #E672002), and stained with 1% crystal violet (Sigma-Aldrich, #C3886).
MDA, Reduced GSH, and Iron Assay
To explore the effects of LINC00460 on ferroptosis, the cells were treated with RAS-selective lethal 3 (RSL3, 3 µM) for 24 h. The MDA content was measured by the malondialdehyde (MDA) kit (Cominbio, Suzhou, China, #MDA-1-Y). The reduced glutathione (GSH) levels were detected with a reduced GSH assay kit (Njjcbio, Nanjing, China, #A006). And the iron content was measured with an iron assay kit (Shyuanye, Shanghai, China, #R22185) according to the manufacturer's instructions.
RNA Sequencing
MDA-Mb-231 cells infected with LV-LINC00460 siRNA-1 or LV-NC were seeded into 6-well plates and cultured for 24 h. Total RNA was isolated and used for RNA sequencing (RNAseq) analysis. The RNAseq analysis was performed by GENEWIZ Biotech Co. (Suzhou, China).
Western Blot
The total protein was obtained from cells using RIPA lysis buffer (Beyotime, #P0013D) containing protease inhibitors and phosphatase inhibitors (Beyotime, #P1045) and was quantified with a BCA protein assay kit (Beyotime, #P0010). 30 µg total protein was separated by 10% SDS-PAGE (Beyotime, #P0012AC) as previously described. 17 The primary antibodies anti-MAL2 (Abacm, #ab75347, 1:1000) and anti-GAPDH (Abacm, #ab8245, clone 6C5, 1:5000) were incubated with the membranes.
Luciferase Activity Assay
The luciferase reporter plasmids containing wide type (WT) LINC00460 sequence, LINC00460 sequence containing the mutant miR-320a-binding site, WT MAL2 3′untranslated region (UTR) sequence or MAL2 3′UTR sequence containing the mutant miR-320a-binding site were obtained from Nanjing Ruizhen Biotechnology Co., Ltd For luciferase reporter assay, MDA-MB-231 cells were co-transfected with luciferase reporter plasmids and miRNA mimic NC or miR-320a mimic using Lipofectamine 2000. After 24 h, the luciferase activity was measured with a dual-luciferase reporter assay system (Promega, Fitchburg, USA, #E1910) according to the manufacturer's protocol.
Statistical Analysis
GraphPad 6.0 statistical software packages were used for statistical analyses. The statistically significant differences were analyzed by Student's t-test or one-way ANOVA. *P < .05, **P < .01, and ***P < .001 was considered statistically significant in all cases.
Results
LINC00460 was Increased in Breast Cancer Tissue Specimens
Initially, we analyzed the expression of LINC00460 using TCGA and GEO data. The levels of LINC00460 were significantly upregulated in breast cancer according to the UALCAN and GEPIA databases (Figure 1A and B). GSE29431 microarray data from the lnCAR website also showed that the LINC00460 expression was markedly increased in breast cancer tissues compared with normal tissues (Figure 1C). Moreover, we observed that compared with normal tissues, the expression of LINC00460 was significantly elevated in cancer tissues of patients with luminal breast cancer, human epidermal growth factor receptor 2 (HER2) positive breast cancer, and triple-negative breast cancer (TNBC) (Figure 1D). Patients with high LINC00460 expression had a longer overall survival time based on the data from UALCAN, TANRIC, and GSE16446 data from the Kaplan-Merier Plotter database (Figure 1E to G). In addition, the GSE16446 data showed that high LINC00460 expression represented a longer disease-free survival rate (Figure 1H).

LINC00460 was overexpressed in breast cancer tissue specimens and associated with the survival of breast cancer patients. (A, B) Expression of LINC00460 in breast cancer tissue samples from TCGA data based on UALCAN (A) and GEPIA (B) databases. (C) Expression of LINC00460 in breast cancer tissue samples from GSE29431 microarray data based on the lnCAR website. (D) Expression of LINC00460 in the cancer tissues of patients with luminal breast cancer, HER2 positive breast cancer, and TNBC. (E, F) Relationship between the LINC00460 expression and the overall survival of breast cancer patients from UALCAN (E) and TANRIC (F) database. (G, H) Relationship between the LINC00460 expression and the overall survival (G) and recurrence-free survival (H) of breast cancer patients from GSE16446 data based on Kaplan-Merier Plotter database.
LINC00460 Knockdown Inhibited Breast Cancer Cell Proliferation and Promoted Ferroptosis
To explore the effects of LINC00460 on cellular function, we reduced the expression of LINC00460 in breast cancer cells using LINC00460 siRNAs. As shown in Figure 2A, both LINC00460 siRNA-1 and siRNA-2 markedly downregulated the levels of LINC00460 in MDA-MB-231 and BT-549 cells. Then, we used the sequence of LINC00460 siRNA-1 to establish the lentivirus (LV-LINC00460 siRNA-1), which resulted in a decrease in LINC00460 expression in MDA-MB-231 and BT-549 cells (Figure 2B). The results of CCK-8 and colony formation assay showed that the knockdown of LINC00460 significantly suppressed cell growth of MDA-MB-231 and BT-549 cells (Figure 2C and D). Moreover, LINC00460 silencing led to an increase in MDA levels in MDA-MB-231 and BT-549 cells after RSL3 treatment (Figure 2E). Compared with the control group, the GSH levels were markedly reduced in the LINC00460 knockdown group (Figure 2F). Additionally, the depletion of LINC00460 also elevated the iron content in MDA-MB-231 and BT-549 cells treated by RSL3 (Figure 2G). These observations suggest that LINC00460 plays an important role in the regulation of cell proliferation and ferroptosis of breast cancer cells.

LINC00460 knockdown modulated the proliferation and ferroptosis of breast cancer cells. (A) Transfection with LINC00460 siRNA-1 and siRNA-2 decreased LINC00460 levels in MDA-MB-231 and BT-549 cells. (B) Infection with lentivirus with LINC00460 siRNA-1 (LV-LINC00460 siRNA-1) decreased LINC00460 levels in MDA-MB-231 and BT-549 cells compared with infection with LV-NC. (C) CCK-8 assay of MDA-MB-231 and BT-549 cells after infection with LV-LINC00460 siRNA-1. (D) Colony formation assay of MDA-MB-231 and BT-549 cells after infection with LV-LINC00460 siRNA-1. (E-G) The MDA (E), GSH (F), and Iron (G) levels in MDA-MB-231 and BT-549 cells after infection with LV-LINC00460 siRNA-1. Three samples were analyzed per condition, and the experiments were performed in triplicate. *P < .05, **P < .01, and ***P < .001.
LINC00460 Silencing Suppressed MAL2 Expression
To investigate the molecular mechanisms about how LINC00460 contributed to cell proliferation and ferroptosis of breast cancer cells, we performed RNA-seq analysis to identify the differentially expressed genes (DEGs) in MDA-MB-231 cells infected with LV-LINC00460 siRNA-1 or LV-NC. As shown in Figure 3A and B, there were 190 upregulated and 59 downregulated DEGs (P < .05) in LINC00460 knockdown MDA-MB-231 cells compared with control cells. Among these, we identified 78 DEGs with the logarithm of fold change (|logFC|) > 1.5 (Figure 3C and Supplemental Table 2). Then, we observed that MAL2, a member of the MAL proteolipid family, has been demonstrated to be involved in cancer progression.18–20 Therefore, we hypothesized that LINC00460 may exert its biological function through the MAL2. As shown in Figure 3D and E, LINC00460 silencing obviously decreased the mRNA and protein expression of MAL2 in MDA-MB-231 and BT-549 cells. Furthermore, the expression of MAL2 was higher in breast cancer tissues than that in normal tissues according to the UALCAN and GEPIA databases (Figure 3F and G). Additionally, compared with normal tissues, the MAL2 expression was obviously increased in the cancer tissues of patients with luminal breast cancer, HER2-positive breast cancer, and TNBC (Figure 3H).

LINC00460 knockdown reduced MAL2 expression in breast cancer cells. (A) RNA-seq analysis showed the number of DEGs (59 downregulated and 190 upregulated) in MDA-MB-231 cells after infection with LV-LINC00460 siRNA-1 or LV-NC. (B) RNA-Seq volcano plot showing logarithm of fold change in DEGs in MDA-MB-231 cells after infection with LV-LINC00460 siRNA-1 or LV-NC. (C) RNA-Seq heatmap showing the DEGs in MDA-MB-231 cells after infection with LV-LINC00460 siRNA-1 or LV-NC. (D, E) The mRNA (D) and protein (E) levels of MAL2 in MDA-MB-231 and BT-549 cells after infection with LV-LINC00460 siRNA-1 or LV-NC. (F, G) Expression of MAL2 in breast cancer tissue samples from TCGA data based on UALCAN (F) and GEPIA (G) databases. (H) Expression of MAL2 in the cancer tissues of patients with luminal breast cancer, HER2 positive breast cancer, and TNBC. *P < .05 and **P < .01.
LINC00460 Regulated MAL2 Expression via the miR-320a
Next, we explored how LINC00460 regulated MAL2 expression. Recently, amounting evidence noted that lncRNAs may act as miRNA sponges, hence regulating the expression of downstream target genes.21,22 Therefore, we used the publicly available prediction tools starBase 3.0 to search for miRNAs, which could modulate both LINC00460 and MAL2. As shown in Figure 4A, there were 3 miRNAs (miR-320a, miR-320b, and miR-654-3p), which could bind to both LINC00460 and MAL2. Previous studies indicated that miR-320a was downregulated in breast cancer tissues and functioned as a tumor suppressor gene in breast cancer.23–25 Hence, we inferred that LINC00460 may modulate MAL2 expression in breast cancer cells via the miR-320a. The hybridization models between LINC00460 and miR-320a was shown in Figure 4B. Moreover, transfection with miR-320a mimic significantly reduced the luciferase activity of the WT-LINC00460 luciferase plasmids but failed to reduce that of the Mutant-LINC00460 luciferase plasmids (Figure 4C). In addition, LINC00460 knockdown upregulated the miR-320a levels in MDA-MB-231 and BT-549 cells (Figure 4D). These data suggest that LINC00460 can act as ceRNA, binding with miR-320a. Furthermore, we showed the hybridization models between miR-320a and MAL2 (Figure 4E). Luciferase reporter assay verified that miR-320a could directly combine with MAL2 (Figure 4F). Importantly, we observed that miR-320a expression was higher in breast cancer tissues than that in normal tissues (Figure 4G). Moreover, compared with normal tissues, the expression of miR-320a was markedly increased in the cancer tissues of patients with luminal breast cancer, HER2-positive breast cancer, and TNBC (Figure 4H).
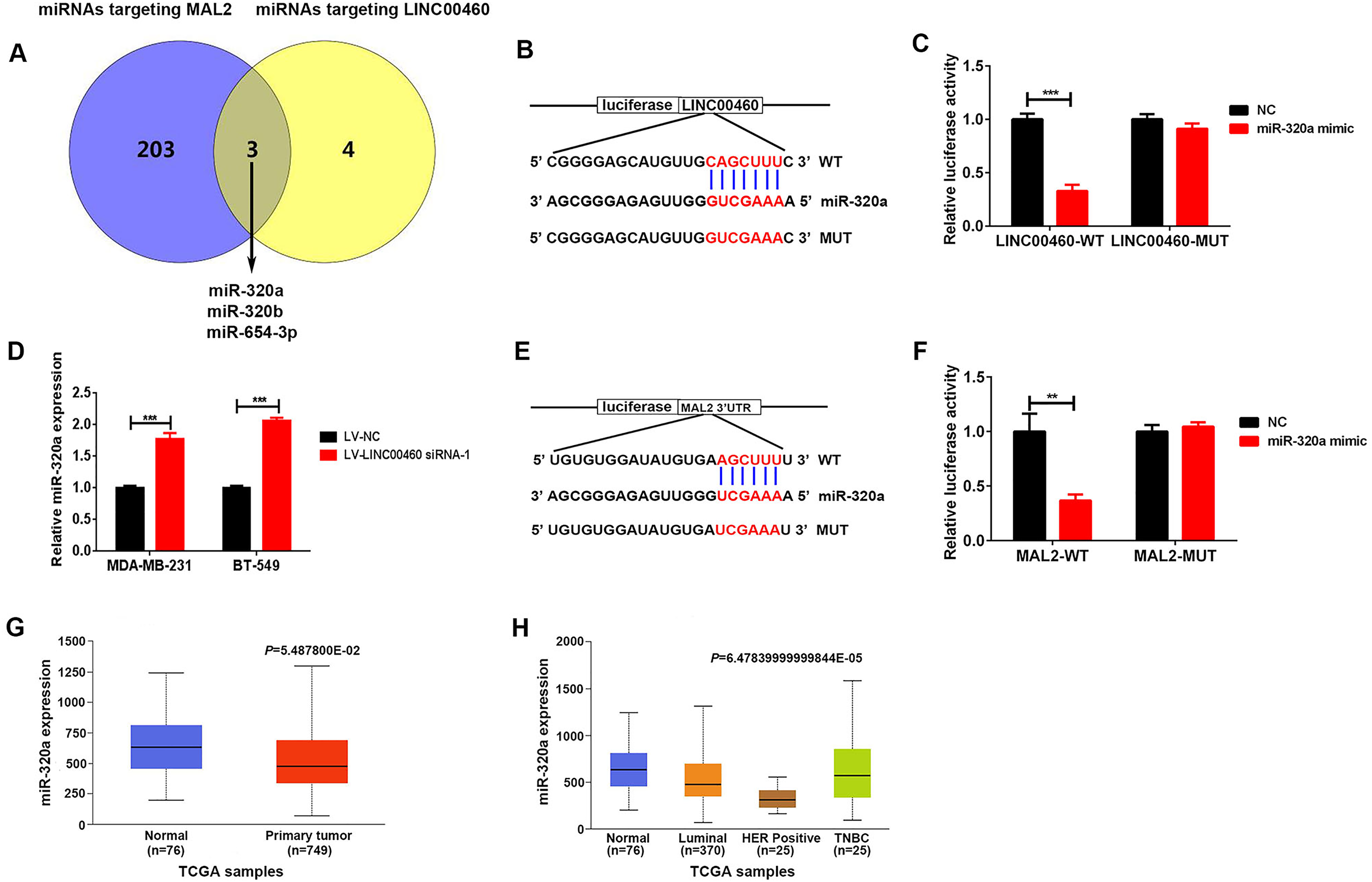
miR-320a targeted LINC00460 and MAL2. (A) Three putative miRNAs targeting both MAL2 and LINC00460 were predicted by StarBase 3.0. (B) Hybridization models between LINC00460 and miR-320a. (C) Luciferase reporter assay showed the binding of miR-320a and WT LINC00460 but not mutant (MUT) LINC00460. (D) The miR-320a expression in MDA-MB-231 and BT-549 cells after infection with LV-LINC00460 siRNA-1 or LV-NC. (E) Hybridization models between miR-320a and MAL2 3′UTR. (F) Luciferase reporter assay showed the binding of miR-320a and WT MAL2 3′UTR but not mutant (MUT) MAL2 3′UTR. (G) Expression of LINC00460 in breast cancer tissue samples from TCGA data based on UALCAN databases. (H) Expression of miR-320a in the cancer tissues of patients with luminal breast cancer, HER2 positive breast cancer, and TNBC.
Then, we investigated whether LINC00460 regulated MAL2 expression in breast cancer cells via miR-320a. As shown in Figure 5A, transfection with miR-320a mimic led to a significant increase in the miR-320a levels in MDA-MB-231 and BT-549 cells. Besides, miR-320a overexpression significantly downregulated the protein levels of MAL2 in MDA-MB-231 and BT-549 cells (Figure 5B and C). Treatment with miR-320a inhibitor downregulated the miR-320a levels, which increased in LINC00460 knockdown MDA-MB-231 and BT-549 cells (Figure 5D). Importantly, treatment with miR-320a inhibitor significantly abolished the inhibitory effects of LINC00460 silencing on MAL2 expression in MDA-MB-231 and BT-549 cells (Figure 5E and F). Together, our results suggest that LINC00460 regulated MAL2 expression in breast cancer cells via the miR-320a.

LINC00460 promoted MAL2 expression by sponging miR-320a. (A) The expression of miR-320a in MDA-MB-231 and BT-549 cells after transfection with miR-320a mimics. (B) The MAL2 protein expression in MDA-MB-231 and BT-549 cells after transfection with miR-320a mimics. (C) The western blot band and (B) densities were quantified using ImageJ software. (D) The expression of miR-320a in MDA-MB-231 and BT-549 cells after treatment with LV-LINC00460 siRNA-1 and miR-320a inhibitor. (E) The MAL2 protein expression in MDA-MB-231 and BT-549 cells after treatment with LV-LINC00460 siRNA-1 and miR-320a inhibitor. (F) The western blot band and (E) densities were quantified using ImageJ software. *P < .05, **P < .01, and ***P < .001.
LINC00460 Modulated Breast Cancer Cell Proliferation and Ferroptosis Through the miR-320a/MAL2 Axis
We further examined whether the LINC00460 modulated cell proliferation and ferroptosis of breast cancer cells through the miR-320a/MAL2 signaling axis. The CCK-8 assay showed that treatment with miR-320a inhibitor or MAL2 overexpression plasmids counteracted the decreases in the cell proliferation rate of MDA-MB-231 and BT-549 cells infected with LV-LINC00460 siRNA-1 (Figure 6A and B). Furthermore, miR-320a knockdown or MAL2 overexpression reversed the effects of LINC00460 silencing on RSL3-induced ferroptosis, evidenced by the detection of MDA, GSH, and Iron (Figure 6C to H).

LINC00460 modulated the proliferation and ferroptosis of breast cancer cells through the miR-320a/MAL2 axis. (A, B) CCK-8 assay of MDA-MB-231 (A) and BT-549 (B) cells after treatment with LV-LINC00460 siRNA-1 and miR-320a inhibitor or MAL2 overexpression vectors. (C, D) The MDA levels in MDA-MB-231 (C) and BT-549 (D) cells after treatment with LV-LINC00460 siRNA-1 and miR-320a inhibitor or MAL2 overexpression vectors. (E, F) The GSH levels in MDA-MB-231 (E) and BT-549 (F) cells after treatment with LV-LINC00460 siRNA-1 and miR-320a inhibitor or MAL2 overexpression vectors. (G, H) The Iron levels in MDA-MB-231 (G) and BT-549 (H) cells after treatment with LV-LINC00460 siRNA-1 and miR-320a inhibitor or MAL2 overexpression vectors. *P < .05, **P < .01, and ***P < .001.
Discussion
It has been reported that LINC00460 is increased in diverse cancer tissues and is associated with clinicopathological characteristics of cancer patients. 11 For example, the expression levels of LINC00460 were increased in colorectal cancer tissues and colorectal cancer patients with high LINC00460 expression showed poor 5-year overall survival and disease-free survival. 26 In prostate cancer, LINC00460 was reported to be increased in cancer tissues, and high LINC00460 expression was correlative to tumor size and high Gleason score. 27 Zhu et al 12 also found that LINC00460 expression was much higher in breast cancer tissues than that in benign breast tissues. In the current study, we found that LINC00460 was significantly elevated in breast cancer tissues based on the TCGA and GSE39341 data. In addition, we also observed that the expression of LINC00460 was significantly elevated in cancer tissues of patients with luminal breast cancer, HER2-positive breast cancer, and TNBC, suggesting that LINC00460 plays a crucial role in the progression of breast cancer. A study showed that high LINC00460 expression was positively associated with lymphatic metastasis and poor overall survival. 12 However, we found that patients with high LINC00460 expression had a longer overall survival and disease-free survival rate. We inferred that the cohorts from different research teams had different clinicopathological characteristics such as patients’ race, estrogen receptor (ER) status, HER2 status, and mRNA expression of 50 genes (PAM50) subtype, which may result in inconsistent prognostic outcomes for breast cancer patients with high LINC00460 expression. Hence, it is important to collect a large number of tissue samples from breast cancer patients for analyzing the relationship between LINC00460 expression and survival.
Previous studies have noted that LINC00460 is involved in multiple cell biological processes of tumor cells including cell proliferation, epithelial-mesenchymal transition, metastasis, apoptosis, and drug resistance. The overexpression of LINC00460 sufficiently induced the epithelial-mesenchymal transition of colorectal cancer cells and enhanced cell proliferation, migration, and invasion in vitro and tumor growth and metastasis in vivo. 26 LINC00460 knockdown suppressed the growth of A549 cells, a kind of lung cancer cells, but promoted their apoptosis. 28 Furthermore, the LINC00460-mediated miR-149-5p/miR-150-5p-mutant p53 feedback loop elevated the resistance to oxaliplatin in colorectal cancer cells. 29 Herein, our results also showed that silencing of LINC00460 led to a decrease in the proliferation rate of breast cancer cells. As far as we know, we first investigated the effect of LINC00460 on ferroptosis. Our data indicated that silencing of LINC00460 promoted the ferroptosis of breast cancer cells treated by RSL3. Collectively, LINC00460 was a key regulator of cell proliferation and ferroptosis in breast cancer. However, the expression and biological function of LINC00460 in other cells (eg, immune cells, fibroblast, and endothelial cells) in the tumor microenvironment remain unclear, which may affect the relationship between LINC00460 and the prognosis of patients with breast cancer. Further investigation is needed in the future.
As a lipid raft-resident protein of the MAL family, MAL2 has been demonstrated to be involved in cancer progression.18–20 For example, MAL2, upregulated in pancreatic cancer tissues, enhanced the proliferation and invasion capabilities of pancreatic cancer cells by interacting with IQ motif-containing GTPase-activating protein 1 (IQGAP1). 30 Bhandari et al 31 showed that MAL2 was remarkably overexpressed in breast cancer tissues and MAL2 silencing inhibited cell proliferation, migration, and invasion of breast cancer cell lines through the regulation of epithelial-mesenchymal transition. Herein, to investigate the potential molecular mechanism of why LINC00460 regulated cell proliferation and ferroptosis in breast cancer, we performed RNAseq and found that the expression of MAL2 was remarkedly decreased in breast cancer cells after LINC00460 silencing. Moreover, the expression of MAL2 was markedly increased in breast cancer tissues. Importantly, MAL2 overexpression reversed the effects of LINC00460 knockdown on cell proliferation and ferroptosis in breast cancer cells. Taken together, our results suggest that LINC00460 promotes cell proliferation and suppressed ferroptosis in breast cancer cells via the MAL2. In addition, we observed that neuronal growth regulator 1 (NEGR1) was the most upregulated gene among the DEGs (Figure 3A to C). Given that NEGR1 was involved in the progression of cancers such as thyroid cancer, neuroblastoma, and colon adenocarcinoma,32–34 it is worth to investigate the effect of LINC00460/NEGR1 axis in the cell proliferation and ferroptosis in breast cancer. Further investigation is needed in the future.
The competitive endogenous RNAs (ceRNA) hypothesis states that some RNAs can regulate the expression of downstream mRNA by combining shared miRNAs, which is an important mechanism of lncRNA-mediated biological functions.35,36 Previous studies have demonstrated that LINC00460 modulates various cellular progress through the ceRNA mechanism.28,37 A study by Ruan et al 37 reported that LINC00460 sponged miR-149-5p, and promoted biglycan (BGN) expression, leading to the elevation of metastasis of colorectal cancer cells. Additionally, LINC00460 functioned as a ceRNA and upregulated p21 (RAC1) activated kinase 1 (PAK1) expression by sponging miR-485-5p in hepatocellular carcinoma cells. 36 In the current study, we hypothesized that LINC00460 might modulate MAL2 expression through sponging a miRNA. We used the publicly available prediction tools starBase 3.0 and found that miR-320a, miR-320b, and miR-654-3p could bind to both LINC00460 and MAL2. miR-320b was significantly increased in breast cancer tissues compared to the normal tissue based on the UALCAN database (Supplemental Figure S1A). Although the expression of miR-654 was decreased in breast cancer tissues (Supplemental Figure S1B), there were few literature that reported the biological function of miR-645-3p in the progression of breast cancer. MiR-320a has been identified as a tumor suppressor in various cancers, including breast cancer. 24 MiR-320a could suppress breast cancer cell proliferation, migration, and invasion by targeting Rab14. 25 In addition, miR-320a overexpression sensitized tamoxifen-resistant breast cancer cells to tamoxifen by negatively regulate cAMP-regulated phosphoprotein 19 (ARPP-19) and estrogen-related receptor γ (ERRγ). 38 Therefore, we inferred that miR-320a could combine both LINC00460 and MAL2. Our results showed that miR-320a knockdown significantly downregulated the levels of MAL2 and LINC00460 in breast cancer cells. Moreover, the expression of miR-320a was markedly decreased in breast cancer tissues. Importantly, the miR-320a inhibitor significantly abolished the inhibitory effects of LINC00460 silencing on MAL2 expression, cell proliferation, and ferroptosis in breast cancer. These results indicated that LINC00460 promoted MAL2 expression by sponging miR-320a. Herein, we focused on the miR-320a, which could combine LINC00460 and MAL2. We can’t exclude that LINC00460 exerts roles in regulating the progression of breast cancer through other miRNAs. For example, Zhu et al 12 showed that LINC00460 promoted cell viability, migration, and invasion of breast cancer cells through the miR-489-5p/FGF7/AKT axis.
Besides the ceRNA mechanism, lncRNAs also mediated their biological functions through interactions with proteins, DNA, or a combination of these. 39 Therefore, we could not exclude that LINC00460 modulated cell proliferation and ferroptosis in breast cancer through other mechanisms. Further investigation was needed.
Conclusion
In summary, our work demonstrated that LINC00460 was overexpressed in breast cancer tissues and associated with the prognosis of patients with breast cancer. Moreover, the knockdown of LINC00460 inhibited cell proliferation and promoted ferroptosis of breast cancer cells through the miR-320a/MAL2 signaling pathway (Figure 7), therefore providing potential targets for breast cancer diagnosis and therapy.
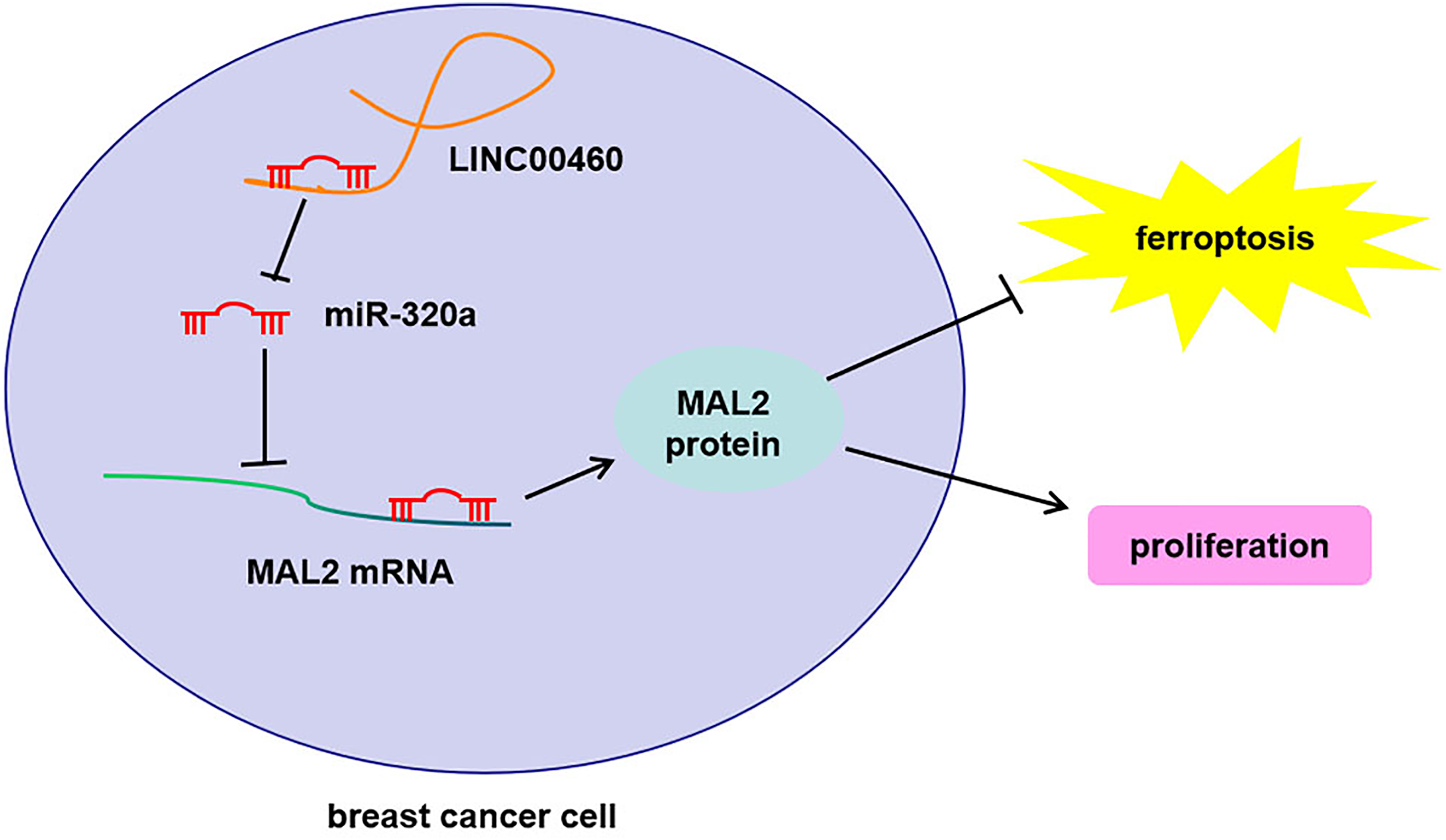
The outlined mechanism underlying LINC00460 facilitating cell proliferation and inhibits ferroptosis in breast cancer.
Supplemental Material
sj-docx-1-tct-10.1177_15330338231164359 - Supplemental material for LINC00460 Facilitates Cell Proliferation and Inhibits Ferroptosis in Breast Cancer Through the miR-320a/MAL2 Axis
Supplemental material, sj-docx-1-tct-10.1177_15330338231164359 for LINC00460 Facilitates Cell Proliferation and Inhibits Ferroptosis in Breast Cancer Through the miR-320a/MAL2 Axis by Chuanqiang Zhang, Liang Xu, Xiaowei Li, Yueqiu Chen, Tongguo Shi and Qiang Wang in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-3-tct-10.1177_15330338231164359 - Supplemental material for LINC00460 Facilitates Cell Proliferation and Inhibits Ferroptosis in Breast Cancer Through the miR-320a/MAL2 Axis
Supplemental material, sj-docx-3-tct-10.1177_15330338231164359 for LINC00460 Facilitates Cell Proliferation and Inhibits Ferroptosis in Breast Cancer Through the miR-320a/MAL2 Axis by Chuanqiang Zhang, Liang Xu, Xiaowei Li, Yueqiu Chen, Tongguo Shi and Qiang Wang in Technology in Cancer Research & Treatment
Footnotes
Acknowledgments
Not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics Statement
We analyzed the expression of LINC00460, miR-320a, and MAL2 in human samples using UALCAN, GEPIA, lnCAR, TANRIC, and Kaplan-Merier Plotter data. Therefore, the ethical approval of this article was not required.
Funding
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: This research was funded by Wujiang Science & Education Revitalizing Health Project (grant number WWK201921), Suzhou Clinical Team Introduction Project (SZYJTD201826), and Suzhou Key Discipline Project (SZXK202126).
Supplemental Material
Supplemental material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
