Abstract
Background and Aim:
There are an increasing number of studies indicating the important roles served by long non-coding RNAs (lncRNAs) in the development of different types of cancer. LINC00460 is a novel identified lncRNA that was found to be upregulated in colorectal cancer. However, the biological roles of LINC00460 in colorectal cancer have yet to be fully elucidated. This study was aimed to investigate the functions and molecular mechanisms of LINC00460 on colorectal cancer metastasis.
Methods:
Expression of LINC00460 and biglycan (BGN) in colorectal cancer tissues and cell lines were quantified by real time PCR or western blotting assay. Cell migration and invasion assays were performed to determine the effect of LINC00460 on tumor metastasis in vitro. The binding interaction between microRNA-149-5p and LINC00460 was revealed by luciferase reporter assay.
Results:
In the present study, lncRNA LINC00460 was shown to be upregulated in colorectal cancer tissues, and overexpression of LINC00460 significantly promoted metastasis of colorectal cancer in vitro. Furthermore, miR-149-5p interacted with LINC00460, and they negatively regulated expression of each other. Transfection of miR-149-5p mimics partially counteracted the tumor metastasis-promoting effects induced by LINC00460 overexpression. Finally, overexpression of LINC00460 upregulated the expression levels of biglycan, a target gene of miR-149-5p, which has also been identified as an oncogenic driver in colorectal cancer.
Conclusion:
Taken together, the present study demonstrated that LINC00460 promoted metastasis of CRC by sponging miR-149-5p and thereby affecting biglycan expression levels.
Introduction
Colorectal cancer (CRC) is the second leading cause of cancer-associated death in the United States, and accounts for ∼5 million deaths annually, worldwide. 1,2 Over the last few decades, the overall survival of patients with CRC has shown a slow, but steady increase. 3 The clinical tumor stage at diagnosis affects patient prognosis. 3 According to statistical data, the 5-year relative survival rate is 90% and 10% in patients with stage I and IV CRC, respectively. 3,4 The molecular mechanisms underlying development of CRC are clinically important as they are associated with the prognosis and treatment response of patients. 5,6 Therefore, improving our understanding of the basic molecular events in the tumorigenesis and progression of CRC may improve the clinical outcomes of patients through the development of improved therapeutics.
Long noncoding RNAs (lncRNAs) are non-coding RNAs >200 nucleotides in length. An increasing number of studies have demonstrated the central functions modulated by various lncRNAs in a variety of biological processes, and lncRNAs can regulate gene expression at different levels, including through chromatin organization, and at the transcriptional and post-transcriptional levels. 7 Data from several previous studies have shown that certain lncRNAs may function as competing endogenous RNAs (ceRNAs) to regulate mRNAs by binding with shared micro (mi)RNAs. 8 -12 LncRNAs exert important roles in various types of cancer, including CRC. 13,14 Studies have identified several CRC-associated lncRNAs, including HOTAIR, 15,16 H19, 17,18 PVT-1 19 and MALAT1. 20,21 However, the functions and molecular mechanisms of lncRNAs in CRC development are not yet fully understood. Recent studies have reported that a novel identified lncRNA LINC00460 was found to be up-regulated in several cancers, including CRC. 22 -25 It acted as a prognostic lncRNA signature in head and neck squamous cell carcinoma (HNSCC). 26 And it was also reported that elevated levels of LINC00460 was associated with poor clinical outcome of lung adenocarcinoma patients. 22 LINC00460 was found to be overexpressed in CRC. 27 whereas the functional roles of LINC00460 in CRC remains unclear and urgently need further investigation.
In the present study, we investigated relationship between LINC00460 high expression and CRC cell migration and invasion capacity. Moreover, we also studied the mechanism underlying the tumor-promoting functions of LINC00460 in vitro. We found that LINC00460 might functions as a ceRNA, and regulating expression of BGN by sponging miR-149-5p and promoting metastasis of CRC.
Materials and Methods
Human Tissue Samples and Cell Lines
A total of 40 pairs of CRC and matching adjacent non-tumor tissues were obtained from patients with CRC. The clinicopathological of characteristics of these 40 CRC patients was shown in Table 1. The present study was approved by the Ethics Committee at Wuxi People’s Hospital Affiliated to Nanjing Medical University (Wuxi, China; approval no. KS2019024). Human CRC cell lines HCT116, HT-29 cells, the normal colorectal mucosa cell line FHC, and 293 T cells were obtained from ATCC. All the cell lines were cultured in DMEM (Gibco; Thermo Fisher Scientific, Inc.) supplemented with 10% FBS (Gibco; Thermo Fisher Scientific, Inc.,), 100 units/ml penicillin and 100 mg/l streptomycin, and were incubated at 37˚C in humidified incubated with 5% CO2.
Clinicopathological Characteristics of 40 CRC Patients.

Construction of Stably Transfected Cell Lines
The lentiviral vectors pLKO.1-sh-NC, pLKO.1-sh-00460, pLKO.1-control and pLKO.1-OVE-00460 were prepared in the laboratory. The backbone of the lentiviral vectors was the pLKO.1-GFP plasmid. The LINC00460 short hairpin (sh)RNA target sequence was CGTGGGAAAGAAGACGCATTCTGAA. 293 T cells transfected with lentiviral vectors was used to produce lentiviral particles. After 72 h of transfection, supernatants containing the lentiviruses were collected and the cells were removed by concentration, centrifugation at 2000 rpm for 10 min at room temperature. For infection with lentiviruses, 106 HCT116 cells were seeded and cultured overnight. The lentiviral particle solution containing the virus, DMEM and Polybrene (10 µg/ml) (Sigma-Aldrich) was added to the cells for 24 h. Subsequently, the culture medium was replaced with fresh medium. Successfully infected cells were sorted using FACS according to GFP expression.
RNA Transfection
miR-149-5p mimics, small interfering (si)BGN and their corresponding negative controls were purchased form (Shanghai GenePharma Co., Ltd.). 4 ×105 Cells were transfected with miRNA or siRNA at final concentration of 50 nM using Lipofectamine® 3000 (Invitrogen; Thermo Fisher Scientific, Inc.). The si-BGN sequence was 5′-CCTTTGAGCAGAGAGGCTT-3′.
RNA Isolation and Reverse Transcription-Quantitative (RT-q) PCR
Total RNA was isolated from tissue samples or cultured cells using TRIzol® (Tiangen Biotech Co., Ltd.). Reverse transcription was performed using FastQuant RT Super Mix (Tiangen Biotech Co., Ltd.) according to the manufacturer’s protocol. qPCR was performed using SYBR Green PCR Master mix (Tiangen Biotech Co., Ltd.). Bugle-Loop™ miRNA qPCR (Guangzhou RiboBio Co., Ltd.) was used to quantify miR-149-5p expression. The primer sequences used for RT-qPCR were as follows: LINC00460 forward, 5′-ACAGCATGAGCCAGGACATC-3′ and reverse, 5′-GAAAGCTGCAACATGCTCCC-3′; BGN forward, 5′-GCCAAGCTGACTGGCATCC-3′ and reverse, 5′-AGTAGCGAAGCAGGTCCTCCA-3′, and GAPDH forward, 5′-AGCCACAATCGCTCAGACAC-3′ and 5′-GCCCAATACGACCAAATCC-3′. Expression was calculated relative to GAPDH.
Nuclear and cytoplasmic RNA were isolated separately using the Cytoplasmic & Nuclear RNA purification kit (Norgen Biotek) according to the manufacturer’s protocol.
Western Blot Analysis
Total protein was extracted from cells and tissues using RIPA lysis buffer (50 mM Tris-HCl pH 8.0, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, and 0.1% SDS) containing 1 mM PMSF and protease inhibitor cocktail (Roche Diagnostics). Protein quantification was performed by using BCA protein assay kit (Beyotime). Then proteins were loaded on to an SDS-gel and resolved using SDS-PAGE and subsequently electrophoretically transferred onto a PVDF membrane (EMD Millipore). Blots were incubated in blocking buffer [(5% non-fat-dried milk, TBS with 0.5% Tween-20 (TBST)] at room temperature for 1 h and subsequently incubated with diluted anti-BGN monoclonal antibody (Santa Cruz Biotechnology, Inc.) overnight at 4˚C, anti-human GAPDH polyclonal antibody (Santa Cruz Biotechnology, Inc.) was used as the loading control. Blots were washed with TBST and incubated with horseradish peroxidase-conjugated secondary antibody at room temperature for 1 h. Signals were visualized suing enhanced chemiluminescence reagent (Tiangen Biotech Co., Ltd.). Densitometry analysis was performed using ImageJ 1.48v (National Institutes of Health).
Luciferase Reporter Assay
Luciferase reporter plasmids pGL3-LINC00460-wild-type (WT), pGL3-LINC00460-mutant (MUT) and pGL3-BGN-3′ untranslated region (UTR) were prepared in the laboratory. Briefly, the LINC00460-WT, LINC00460-MUT, 3′UTR of BGN were cloned into pGL3 plasmids, downstream of the firefly luciferase gene. 293 T cells were seeded in 24-well plates at a density of 4 × 104 cells/well and cultured for 24 h to reach a confluency of 70% to 80%. Subsequently miRNA mimics (200 ng) were co-transfected with the indicated pGL3 firefly luciferase vector (200 ng) and pGL3 Renilla luciferase vector (20 ng) into 293 T cells. After 48 h, luciferase activity was measured using the dual luciferase reporter assay system (Promega, Corporation) and activity was normalized to Renilla luciferase.
Cell Migration and Invasion Assays
Cell invasion assays were performed using BD BioCoat™ Matrigel™ invasion chamber with 8 µm pores (BD Biosciences). Briefly, HCT116 cells at a density of 8 × 104 in 200 µl DMEM (without FBS) were placed in the upper chamber, whereas DMEM containing 10% FBS was added to the lower wells. After 24 h of culture, the invasion chambers were processed according to the manufacturer’s protocol and the number of cells which had invaded were stained with Hoechst at room temperature for 5 min, and then counted using a fluorescence microscope (Eclipse 80i; Nikon Corporation). Migration assays were performed using the same protocol as the invasion assay except uncoated BD Falcon Chambers were used (BD Biosciences).
Statistical Analysis
All data are presented as the mean ± standard error of the mean of the indicated number of repeats. One-way ANOVA test or the Student’s t-test was used to analyze the data. P < 0.05 was considered to indicate a statistically significant difference.
Results
LINC00460 Expression Is Significantly Upregulated in CRC Tissues and Cell Lines
The expression levels of LINC00460 in 40 pairs of CRC tissues and their corresponding adjacent non-tumor tissues were detected using RT-qPCR. The results showed that LINC00460 expression was upregulated ∼5.25-fold in CRC tissues (Figure 1A). In addition, the expression levels of LINC00460 were significantly higher in the CRC cell lines, HCT116 and HT-29, compared with the normal human colorectal mucosa cell line, FHC (both P < 0.001; Figure 1B). To observe the potential clinical value of LINC00460 in CRC, we analyzed the relationship between LINC00460 expression and the disease free survival of patients with colon adenocarcinoma or rectum adenocarcinoma using an online tool called GEPIA 28 from TCGA data. The results showed that patients with higher LINC00460 expression had poorer disease free survival rate than those with lower expression (Figure 1C and D).

LINC00460 expression is significantly upregulated in CRC tissues and cell lines. A, mRNA expression levels of LINC00460 in 40 human CRC tissues and adjacent non-tumor tissues. ***P < 0.001. B, Relative expression of LINC00460 in HCT116 and HT-29 CRC cell lines and the human normal colorectal mucosa cell line FHC. ** *P < 0.001 vs. FHC. C and D, Results of TCGA data (GEPIA) showed that high LINC00460 expression predicted poor disease free survival of patients with colon adenocarcinoma (C) and rectum adenocarcinoma (D). E, RT-qPCR showed the cellular distribution of LINC00460 in HCT116 cell and HT29 cell, the paired t test was used to analyze the data. CRC, colorectal cancer.
To determine the subcellular localization of LINC00460, the cytoplasmic and nuclear RNA from cultured cells were isolated. And the RT-qPCR data demonstrated that LINC00460 is predominantly located in the cytoplasm of CRC cells (Figure 1E). Therefore, LINC00460 is a cytoplasmic lncRNA and may function in a post-transcriptional manner.
LINC00460 Regulates Metastasis of CRC Cells in Vitro
To determine the biological effects of LINC00460 in CRC, HCT116 were used to establish cell lines with stable knockdown or overexpression of LINC00460. LINC00460 expression was assessed using RT-qPCR (Figure 2A and B). Subsequently, the effects of LINC00460 knockdown and overexpression on tumor metastasis were determined using transwell migration and invasion assays. Knockdown of LINC00460 in HCT116 cells significantly decreased cell motility and inhibited cell invasion, whereas overexpression of LINC00460 promoted these effects (all P < 0.01; Figure 2C and D).

LINC00460 regulates metastasis of CRC cells. A, HCT116 cells were infected with lentiviral shRNA-00460 to establish a stable LINC00460 knockdown HCT116 cell line. RT-qPCR was used to assess expression of LINC00460. **P < 0.01. B, HCT116 cells were infected with lentiviral OVE-00460 to establish a stable LINC00460 overexpression HCT116 cell line. RT-qPCR was used to assess expression of LINC00460. **P < 0.01. C, Cell migration and invasion were examined in control HCT116 cells and cells transfected with sh-NC, sh-00460 or OVE-00460. D, Quantification of the number of migrated or invaded cells in the Transwell assays. **P < 0.01. All the quantitative data are presented as the mean ± standard error of the mean of at least 3 replicates. Statistical analysis was performed using a Student’s t-test or a one-way ANOVA test. CRC, colorectal cancer; RT-qPCR, reverse transcription-quantitative PCR; sh, short hairpin; NC, negative control; sh-00460, shRNA targeting LINC00460; OVE-00460, LINC00460 overexpression vector.
miR-149-5p Interacts With and Negatively Regulates LINC00460 Expression
Including miRDB 29 and miRcode 30 were used to predict miRNAs that may interact with LINC00460. The results of the alignment suggested that miR-149-5p was potentially one of the miRNAs specific for LINC00460, and there were 2 predicated miR-149-5p binding sites in the LINC00460 sequence (Figure 3A). Transfection of miR-149-5p mimics in HCT116 cells resulted in a reduction of LINC00460 expression in a concentration-dependent manner (P < 0.01; Figure 3B). Furthermore, LINC00460 interacted with miR-149-5p directly, as determined using a luciferase reporter assay (Figure 3C). To investigate whether this binding activation was dependent on the predicated miR-149-5p binding sites, a pGL3-LINC00460-MUT plasmid was constructed containing mutations in the recognition sites, and the luciferase assay results confirmed that the mutant LINC00460 was no longer downregulated by miR-149-5p mimics (Figure 3C).
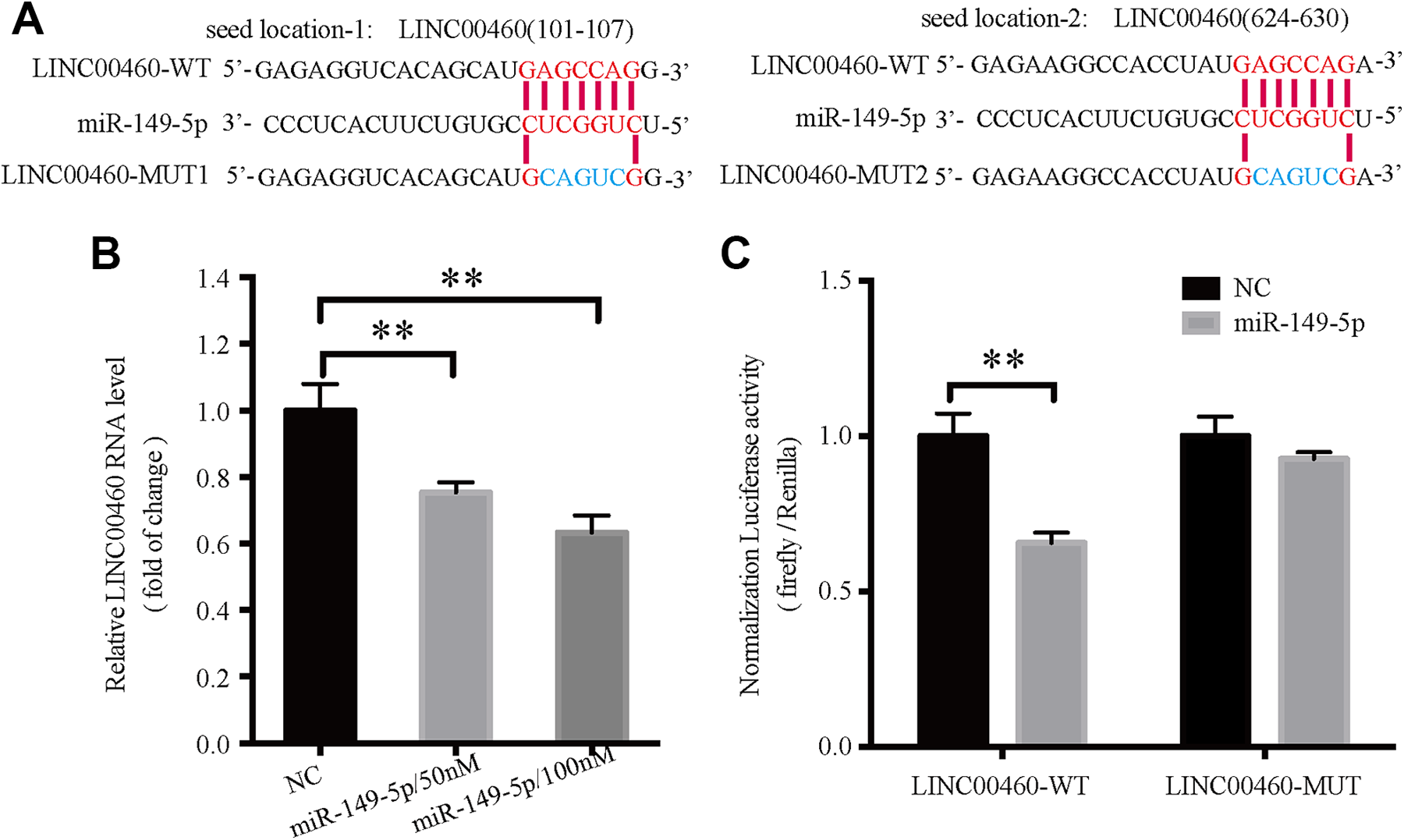
miR-149-5p interacts and regulates expression of LINC00460. A, Sequence alignment of miR-149-5p with predicated binding sites to the LINC00460-WT or LINC00460-MUT. B, LINC00460 expression in HCT116 cells following transfection with 50 or 100 nM miR-149-5p mimics. **P < 0.01. C, Luciferase reporter assays showed that transfection of miR-149-5p mimics significantly reduced the luciferase activity of LINC00460-WT vector, whereas it had no effect on the LINC00460-MUT vector. **P < 0.01. All the quantitative data are presented as the mean ± standard error of the mean of at least 3 replicates. Statistical analysis was performed using a Student’s t-test or a one-way ANOVA. miR, microRNA; WT, wild-type; MUT, mutant.
LINC00460 Functions as a ceRNA by Sponging miR-149-5p in CRC Cells
miR-149-5p was significantly downregulated in CRC tissues compared with the matching non-tumor tissues (P < 0.001; Figure 4A). To determine whether LINC00460 regulated miR-149-5p levels in the CRC cell lines, miR-149-5p levels were detected using RT-qPCR in HCT116 cells with stable knockdown or overexpression of LINC00460. The results showed that overexpression of LINC00460 resulted in a 39% reduction of miR-149-5p expression (P < 0.01), whereas knockdown of LINC00460 resulted in a significant increase in miR-149-5p levels (P < 0.001; Figure 4B).
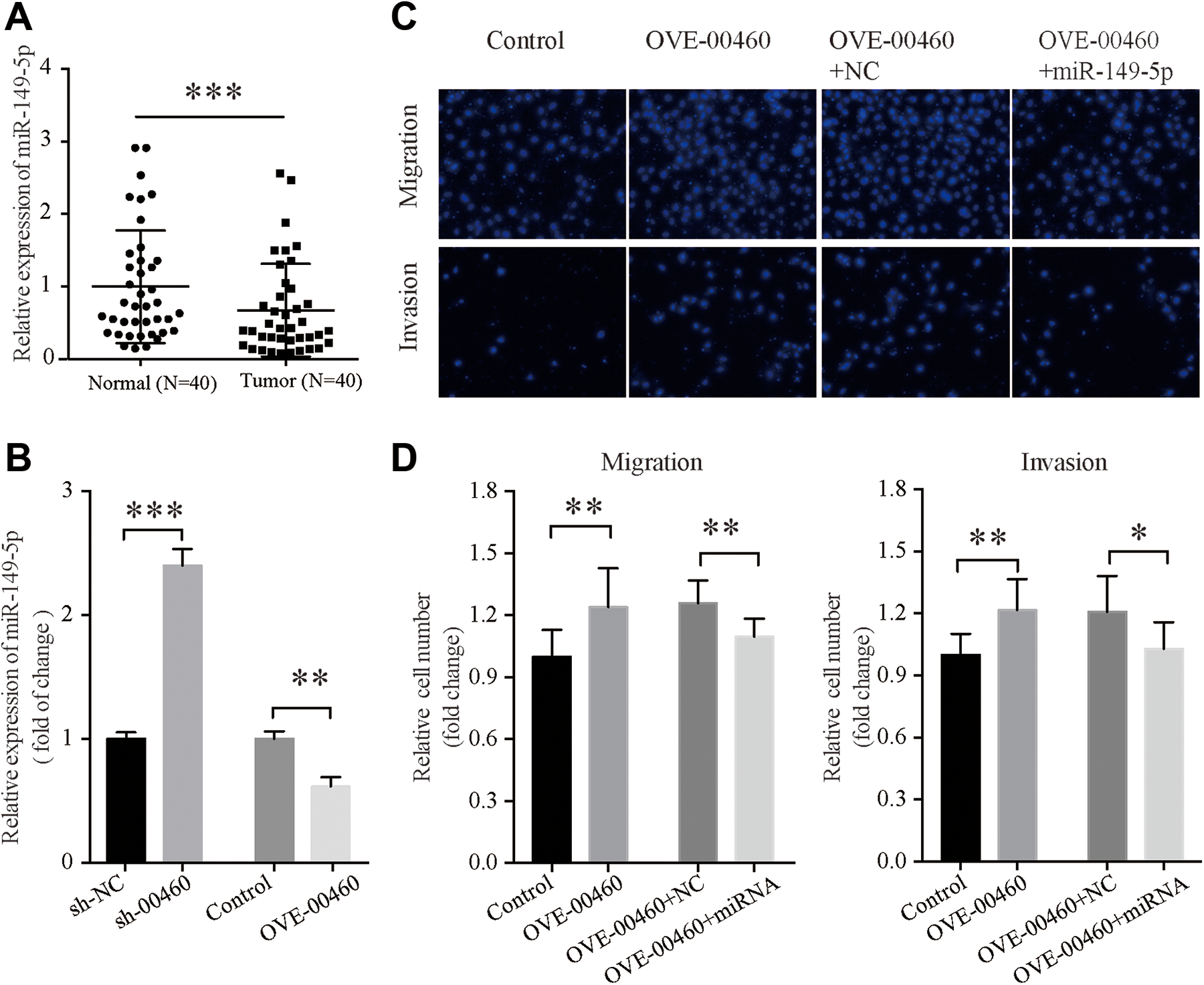
LINC00460 functions as a competing endogenous RNAs by sponging miR-149-5p in CRC. A, Expression levels of miR-149-5p in 40 pairs of CRC tissues were significantly decreased compared with the adjacent non-tumor tissues, the paired t test was used to analyze the data, ***P < 0.001. B, Expression levels of miR-149-5p in the LINC00460 knockdown HCT116 cells or LINC00460 overexpressing HCT116 cells. C, Effects of miR-149-5p overexpression on Transwell migration and invasion. miR-149-5p was overexpressed in HCT116 cells already overexpressing LINC00460. D, Quantification of the number of cells which had successfully migrated or invaded. The metastasis promoting effects induced by LINC00460 were partially but significantly reversed by transfection of miR-149-5p. *P < 0.05, **P < 0.01. All the quantitative data are presented as the mean ± standard error of the mean of at least 3 replicates. Statistical analysis was performed using a Student’s t-test or a one-way ANOVA test. miR, microRNA; CRC, colorectal cancer; sh, short hairpin; NC, negative control; sh-00460, shRNA targeting LINC00460; OVE-00460, LINC00460 overexpression vector.
To determine the effects of miR-149-5p on metastasis of CRC cells, HCT116 cells stably expressing LINC00460 were transfected with miR-149-5p mimics. Results of the transwell migration and invasion assays showed that LINC00460 promoted cell metastasis, but transfection with the miR-149-5p mimics partially reversed the tumor-promoting effects induced by LINC00460 (Figure 4C and D).
Based on the above results, miR-149-5p may mediate the functions of LINC00460 in the metastasis of CRC cells and LINC00460 may act as a ceRNA by sponging miR-149-5p in CRC.
LINC00460 Regulates Expression of BGN
BGN was predicated to be a key target gene of miR-149-5p by miRDB. miR-149-5p was confirmed to bind directly to the 3′UTR of BGN using a luciferase reporter assay (Figure 5A). Furthermore, BGN expression was significantly upregulated in CRC tissues (P < 0.001; Figure 5B). To further determine the association between LINC00460 and BGN, BGN expression levels were determined in the HCT116 cell lines with stable knockdown or overexpression of LINC00460 using RT-qPCR and western blotting. The results showed that knockdown of LINC00460 resulted in a decrease in BGN expression, and overexpression of LINC00460 resulted in an increase in BGN expression (both P < 0.01; Figure 5C and E). To verify whether LINC00460 regulated BGN expression by sponging miR-149-5p, LINC00460 overexpressing HCT116 cells were transfected with miR-149-5p mimics. The results suggested that overexpression of LINC00460 upregulated BGN expression, but miR-149-5p mimics partially reversed the increase in BGN expression (Figure 5D and E). To examine the role of BGN on the biological function of LINC00460, si-BGN was used to knockdown BGN expression in HCT116 cells stably overexpressing LINC00460 (Figure 5F). The results showed that LINC00460 increased the metastatic behaviors of cells, and si-BGN partially, but significantly, reversed the effects on migration and invasion (both P < 0.05; Figure 5G and H).

LINC00460 overexpression promotes metastatic behavior in CRC cells, partially by regulating expression of BGN. A, Luciferase reporter assays showed that miR-149-5p significantly reduced the luciferase activity of the reporter vector containing the 3’UTR of BGN. **P < 0.01. B, mRNA expression levels of BGN in 40 pairs of CRC and adjacent non-tumor tissues, the paired t test was used to analyze the data, ***P < 0.001. C, mRNA expression levels of BGN in the LINC00460 knockdown HCT116 cells or LINC00460 overexpressing HCT116 cells. GAPDH was used as the internal control. **P < 0.01. D, mRNA expression levels of BGN in cells overexpressing LINC00460 alone or overexpressing both LINC00460 and miR-149-5p. BGN expression was upregulated in HCT116 cells overexpressing LINC00460 compared with the control, whereas LINC00460 overexpressing HCT116 cells transfected with miR-149-5p mimics resulted in a partial reversal of the LINC00460-mediated increase in BGN expression. *P < 0.05, **P < 0.01. E, BGN protein expression levels were determined in the indicated cells by western blotting. F, si-BGN knockdown efficiency was determined by RT-qPCR. ***P < 0.001. G, Effects of si-BGN on Transwell migration and invasion. Transfection of HCT116 cells overexpressing LINC00460 with si-BGN resulted in a decrease in both invasion and migration. *P < 0.05. H, Quantification of the number of cells which had successfully migrated or invaded. The metastasis promoting effects induced by LINC00460 were partially but significantly reversed by transfection of si-BGN. *P < 0.05. All the quantitative data are presented as the mean ± standard error of the mean of at least 3 replicates. Statistical analysis was performed using a Student’s t-test or a one-way ANOVA test. miR, microRNA; CRC, colorectal cancer; si, small interfering; sh, short hairpin; NC, negative control; sh-00460, shRNA targeting LINC00460; OVE-00460, LINC00460 overexpression vector; BGN, biglycan.
Taken together, these results suggested that LINC00460 regulates metastasis partially through increasing BGN expression.
Discussion
Accumulating evidence has shown that lncRNAs serve important roles in the tumorigenesis and progression of different types of cancer. In the present study, it was demonstrated that expression of LINC00460 was upregulated in CRC. This result was consistent with previous studies. 22,27 Furthermore, recent studies have demonstrated that LINC00460 expression was significantly increased in a number of different types of tumors. 22 -24 Upregulated expression of LINC00460 was associated with poor prognosis and poor overall survival of patients with lung adenocarcinoma. 22 Furthermore, Transwell migration and invasion assays demonstrated that knockdown of LINC00460 in HCT116 cells significantly reduced metastatic behaviors of cells, whereas overexpression of LINC00460 increased these behaviors. Based on these results, it was hypothesized that LINC00460 may serve an important role in the tumorigenesis and progression of CRC.
It has been previously been demonstrated that lncRNAs, mRNAs, transcribed pseudogenes and circRNAs can act as sponges of miRNAs. 31 -34 Consistent with these findings, recent studies reported that RNA transcripts function as ceRNAs to regulate expression by binding with a shared miRNA. 8 Therefore, one aim of the present study was to determine whether LINC00460 functioned as a sponge of a miRNA. Bioinformatics analysis and luciferase reporter assays showed that miR-149-5p could directly interact with recognition sites in LINC00460. Furthermore, miR-149-5p was able to suppress LINC00460 expression and LINC00460 also reduced the levels of miR-149-5p.
Recently, several studies have reported that miR-149-5p served as a tumor suppressor, and was found to be significantly downregulated in a number of different types of cancer, such as gastric cancer, 35 breast cancer, 36 lung cancer 37 and CRC. 38,39 In the present study, miR-149-5p was downregulated in CRC. In addition, expression of BGN, a target gene of miR-149-5p, was upregulated. LINC00460 regulated BGN expression through binding and attenuating the effects of miR-149-5p. LINC00460 overexpression increased the expression levels of BGN, whereas knockdown of LINC00460 had the opposite effect.
The BGN gene encodes a member of the small leucine-rich proteoglycans family, 40 which is a critical regulator of bone growth, 41 muscle development, 42 inflammation and innate immunity. 43,44 Recent studies reported that expression of BGN was elevated in tumor tissues compared with non-tumor adjacent tissues in several types of cancer, including pancreatic cancer, 45 CRC 46 and esophageal squamous cell carcinoma. 47 Upregulated expression of BGN was associated with advanced stage cancer and a poor prognosis. 47 -51 BGN is involved in the development of different types of cancer. 51,52 In the present study, the oncogenic role of BGN was confirmed, as overexpression LINC00460 promoted cell migration and invasion, whereas silencing BGN suppressed these behaviors.
In conclusion, upregulation of LINC00460 promoted metastasis of CRC cells, at least in part, by acting as a ceRNA for miR-149-5p, which resulted in upregulation of BGN.
Footnotes
Authors’ Note
This study was approved by the Ethics Committee at Wuxi People’s Hospital Affiliated to Nanjing Medical University (Wuxi, China; approval no. KS2019024).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The present study was supported by The Scientific Research Programs of Wuxi People’s Hospital Affiliated to Nanjing Medical University (Wuxi, China; grant no. RKA201811).
