Abstract
Glycosylation has a clear role in cancer initiation and progression, with numerous studies identifying distinct glycan features or specific glycoproteoforms associated with cancer. Common findings include that aggressive cancers tend to have higher expression levels of enzymes that regulate glycosylation as well as glycoproteins with greater levels of complexity, increased branching, and enhanced chain length1. Research in cancer glycoproteomics over the last 50-plus years has mainly focused on technology development used to observe global changes in glycosylation. Efforts have also been made to connect glycans to their protein carriers as well as to delineate the role of these modifications in intracellular signaling and subsequent cell function. This review discusses currently available techniques utilizing mass spectrometry-based technologies used to study glycosylation and highlights areas for future advancement.
Introduction to Glycosylation
Protein glycosylation is a post-translational modification (PTM) in which individual monosaccharides or longer polysaccharides (also called carbohydrates, sugars, or glycans) decorate asparagine (N-glycan), serine (S) or threonine (T) (O-glycan),1,2 and, rarely, cysteine (S-glycan) 3 or tryptophan (C-mannosylation) 4 residues. Along with protein glycosylation, lipid-glycan conjugates, extended free poly or oligosaccharides such as hyaluronan, and recently identified RNA-glycan conjugates are all essential for the regulation of healthy cellular processes. 2 This review focuses on an overview of mass spectrometry-focused workflows to study N- and O-glycoproteoforms present in mammalian cells, but all forms of glycosylation have been shown to be perturbed in various cancer models. 5 These modifications are vital to protein folding, solubility, and protein-protein interactions including those regulating innate immunity and infection. Protein glycosylation is challenging to predict in silico as it is not encoded genetically but is regulated through complex combinations of aptamer amino acid sequence, larger protein folds interacting with specific glycosyltransferases, proximity of glycosyltransferases and acceptor proteins, the presence or absence of additional glycan modifying enzymes (glycosidases, acetyl-, phospho- or sulfo-transferases), and the availability of activated glycan substrates. The fundamental building block of glycan PTMs is glucose, which can be activated and modified through the hexosamine biosynthetic pathway (HBP) into monosaccharide subunits described below. 2 Although all cells can synthesize activated glycans to varying extents, these building blocks are often salvaged by the action of glycosidases (enzymes which degrade oligosaccharides) during the trimming of larger oligosaccharide modifications in the endoplasmic reticulum (ER) and Golgi, or during lysosomal degradation of misfolded proteins. 2 The presence or absence of glycosylation at specific amino acids (macro-heterogeneity), differences in the structures of glycans at individual sites (micro-heterogeneity), as well as the sum of variation in occupied glycosylation sites across a protein (meta-heterogeneity) can result in biological and technical challenges to link specific glycans to potential health versus pathological function. 6
Oligosaccharide Structure and Biosynthesis
Monosaccharides and combinations of oligosaccharides (2 +monosaccharides in combination) differ across the kingdoms of life, with the deepest understanding being in mammalian cells and tissue. Originally termed carbohydrates, or hydrates of carbon, they follow the molecular formula of Cx(H2O)n. Although they generally exist in a conformationally stable ring formation, often drawn as a chair, the structures have various linear form equilibriums which can be trapped for downstream chemical analyses including mass spectrometry (Figure 1). Each carbon in the glycan ring (C1-C5) is a chiral center which can have substituents at the α or β position (Figure 1). Nomenclature of glycan linkages follows the rule of orientation (α or β) followed by the carbon atoms of the sugar rings being linked in the bond. The complexity of oligosaccharides arises from the α or β bonds which can be formed on each chiral carbon, creating a large combinatorial number of potential oligosaccharides with identical chemical compositions but completely different structural characteristics. A classic example is the polysaccharide structures of cellulose versus amylose, which are both formed from glucose. The β(1-4) linkage in cellulose makes a strong large fiber while the α(1-4) linkages in amylose are compact and easily digested. 7 Individual monosaccharides are often simplified in symbolic notation as shown in Figure 1. 8 If the precise stereochemistry of a monosaccharide is unknown, the associated symbol can be left with no color and a generic designation such as hexose. In humans, there are 9 typical monosaccharides which create combinations of linear and branched glycoconjugates (Hexoses: D-glucose (Glc), N-Acetyl-D-glucosamine (GlcNAc), D-galactose (Gal), N-Acetyl-D-galactosamine (GalNAc), D-Mannose (Man), D-Glucoronic acid (GlcA); Pentoses: D-Xylose (Xyl), L-fucose (Fuc); Nonosulonic or sialic acids: N-Acetylneuraminic acid (Neu5Ac)).

(A) Chair and (B) Fisher projection representations of D-Glucose indicating chiral carbons. (C) Representative nomenclatures of structures of N- and O-glycans including the location of cleavage by commonly utilized endoglycosidases. 137
Although protein glycosylation was identified in the 1800s and utilized for common assays like blood typing by the early 1900s, detailed studies of the biosynthesis of glycans and glycan chains have required the development of analytical tools capable of defining monosaccharide linkage orders as well as the stereochemistry at each anomeric center. 9 Complex glycosylated proteins are mostly thought of as localizing to the cell surface and facilitating cell-cell, immune response, or infection defense responses. 2 On extracellular and excreted proteins, both N- and O- linked glycosylation often occurs during processing in the ER and Golgi prior to sorting and transport through secretory pathways.10–12 Monosaccharides must be in an activated format such as -UDP, -GDP, or -CMP in order to be transferred to a growing oligosaccharide, lipid, or directly to a protein, but these activated sugars cannot easily pass membranes and must be actively transported into the ER/Golgi, creating another point of potential glycosylation dysregulation.10,13 Both N -and O- glycan modifications have a “core” structure attached to an amino acid side chain which can then be further extended/modified by specific glycosyltransferases. N-linked glycans have a core glycan moiety of two N-acetylglucosamine (GlcNAc) and three mannose residues are generally associated with the amino acid backbone consensus sequences (N-X-S/T, Figure 1).
During N-glycan biosynthesis, a multi-enzyme pathway begins on the cytoplasmic side of the ER and flips to the lumen, eventually creating a 14-sugar oligosaccharide on a dolichol phosphate anchor. This oligosaccharide is then transferred to a nascent protein, and, following successful protein folding, the oligosaccharide is trimmed to a core 11-unit oligomannose structure (Figure 1, oligomannose or high mannose structure). As the protein matures through the ER and into Golgi compartments, this core oligosaccharide structure may be trimmed, extended, and modified by sulfation or phosphorylation, eventually creating hybrid and complex glycoproteins for secretion or display on the cell surface. 2
Core O-GalNAc glycans (4 basic cores shown in Figure 1) are also synthesized in the ER/Golgi; initiated by the transfer of GalNAc in an α linkage to serine (S) or threonine (T) by one of several polypeptide-N-Acetylgalactosaminyltransferases. This initial GalNAc is then expanded by additional glycosyltransferases within the Golgi to one of 4 di- or tri-saccharide cores in a cell type-specific manner. 2 Extended glycosylation, along with antennae fucosylation and sialylation, can be performed by some of the same glycosyltransferases that extend complex N-glycans. These O-GalNAc glycans can be extremely diverse and complex, and are responsible for self-recognition, preventing infection, and the definition of blood groups. Adding to the challenge of analysis, some heavily O-glycosylated glycoproteins, known as mucins, have dense regions of Proline/Serine/Threonine domains with extensive, branching O-glycan structures creating a characteristic “bottlebrush” 3D structure. 14
O-GlcNAcylation is extremely common, generally underappreciated, and catalyzed within the cytoplasm by a single enzyme, O-GlcNAc transferase (OGT). 15 This modification appears to have fairly rapid on/off rates and competes with phosphorylation at the same or adjacent sites creating a means of intracellular signaling. 16
In addition to these canonical glycosylation pathways, recent studies have identified glycosyltransferases and glycoproteins in unexpected cellular locations such as the mitochondria, 17 so it is clear that there is still much to learn about when, where, and how proteins are glycosylated in healthy cells. Although prior knowledge of glycan PTMs can be incorporated into glycobiology experiments, researchers should both be open to unexpected findings and demand strong analytical confirmation of unusual modifications.
Glycosylation in Disease
As expected from the heterogeneity and ubiquitous presence of glycan modifications, they are vital to health; genetic disruptions in glycan biosynthesis are often embryonic lethal but lead to severe neurological defects if present.18–20 Acquired diseases related to glycosylation run the gamut and include cardiac, 21 neurological, 22 immune-related, 23 gastrointestinal, 24 and metabolic disorders. 25 A mid-year 2022 PubMed.gov search of glycosylation + cancer results in nearly 13,000 scientific articles. Cancer-specific glycan biomarkers have been implicated in breast,26–28 ovarian,29,30 lymphoma, 31 colon, 32 myeloma, 33 pancreatic, 34 and prostate35,36 cancers. 1 Research in cancer progression and glycosylation clearly indicates cancer cells often have modified glycan profiles due to aberrant increases or decreases of glycotransferases and glyco-modifying enzymes. Additionally, saccharide subunit and oligosaccharide biosynthesis occur through a branch of the glycolysis pathway, important in cell metabolism and often disregulated in cancer.37,38 Furthermore, cancer cells evolve to have mechanisms that promote survival, circumventing the severe loss of survivability imposed by the consequences of aberrant glycosylation. Some studies have identified either global changes in glycan structures associated with specific cancers or specific glycan/glycoproteoforms that directly lead to increased cancer. Both may lead to potential therapeutic targets and continued studies using the mass spectrometry workflows described in Figure 2 are expected to create deeper connections between glycobiology and cancer. Below we highlight some common forms of glycosylation dysregulation identified in cancer.
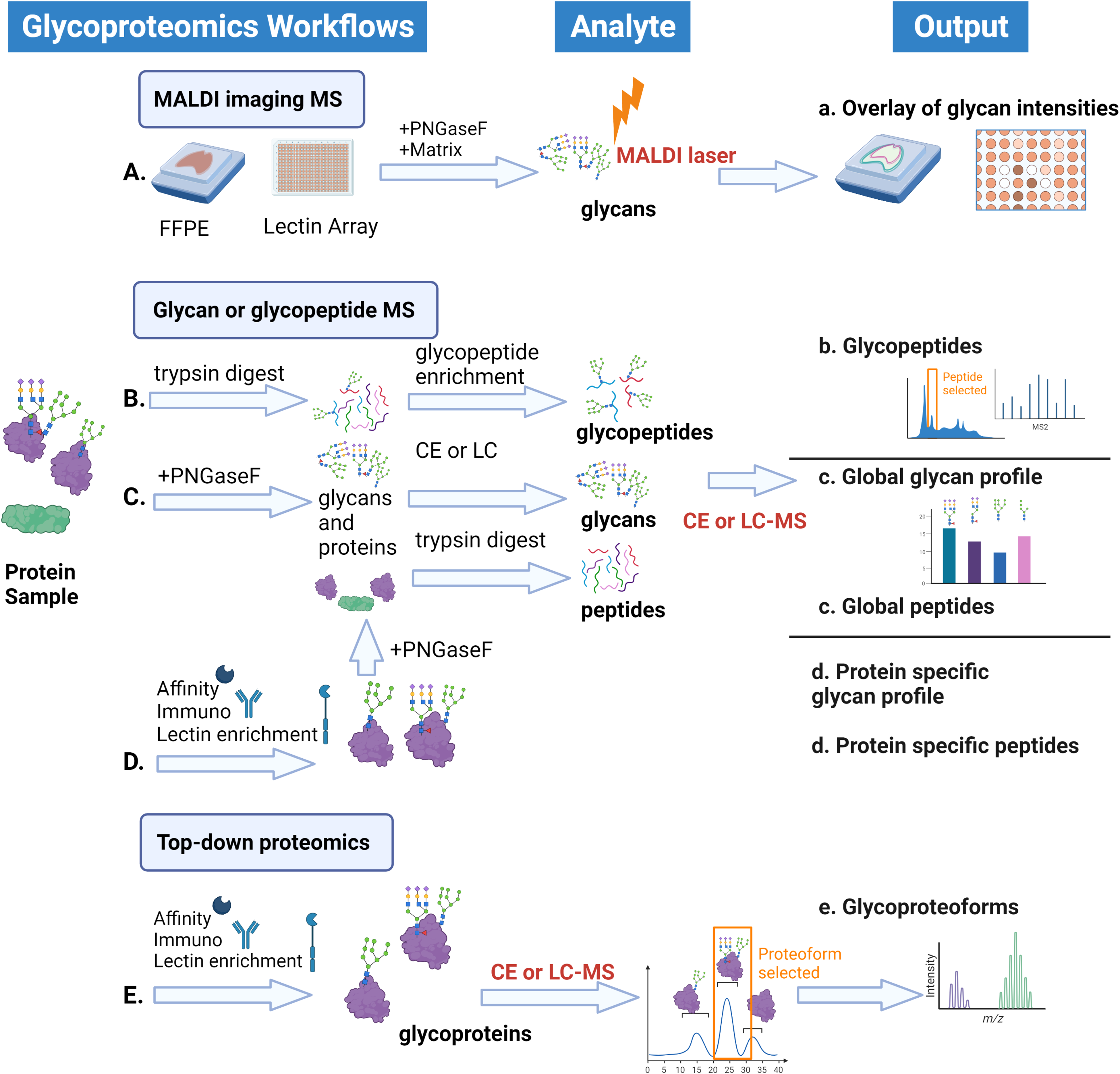
Glycoproteomics workflows. Based on sample type, available technology, and final goals some or all of these may be combined to define glycan micro and macro heterogeneity in a sample. (A). MALDI imaging mass spectrometry utilizes a solid phase to create glycan intensity overlays directly from tissue samples or from spotted slides of antibodies or lectins. (B–D). Separations-based glycan or glycopeptide mass spectrometry utilizes traditional bottom-up mass spectrometry in which proteins are digested with a protease, usually trypsin, to create peptides. (B). Glycopeptides can be enriched and analyzed for glycan and peptide structure in tandem or (C). Glycans can be removed by endoglycosidase treatment for separate mass spec runs of glycans and peptides. (D). Specific proteins or sets of glycans be enriched first to examine the glycan profiles for variation across conditions, as in biological manufacturing. (E). Top-down proteomics utilizes enrichment of a specific protein of interest and state-of-the-art mass spectrometry to identify glycoproteoforms—or the sum of protein sequence and macroheterogeneity of glycan modifications on the protein.
Oligomannose or high mannose N-glycans (Figure 1C), arising from dysregulation of glycosidases and glycosyltransferases in the N-glycan biosynthetic pathway, have been identified as increased in aggressive cancer.39–44 N-glycans impact protein stability and solubility and folding, and in healthy cells improper glycosylation should lead to protein degradation. 13 In addition, N-glycans also regulate extracellular and physiological functions including cell-cell adhesion, maintenance of mucosal barriers, and immune cell function.2,45 Oligomannose N-glycans terminate in unsubstituted mannose residues which are typically trimmed during ER-to-Golgi protein processing (Figure 1), but, in cancer, these glycoproteins avoid lysosomal degradation and are sent to the cell surface or excreted leading to both immune responses as well as potential biomarkers with this glycan modification.46–48,49
Truncated O-GalNAc glycans, containing a single O-GalNAc (Tn antigen) or a sialylated O-GalNAc (sTn) are also common in cancers.50–52 Where healthy cells have mature linear or branched O-GalNAc glycans, cancer cells have decreased expression of upstream glycosyltransferases and chaperones creating membrane-bound and secreted glycoproteins bearing only Tn or sTn. Sialyl-Tn is currently being targeted with several Car-T type therapies in development.51,53
Core N-glycan fucosylation (shown in complex and hybrid structures in Figure 1) is associated with several kinds of aggressive cancers.26,29,39,54 Mass spectrometry in combination with specific lectin and glycosidases has been vital for identifying this cancer biomarker and differentiating it from outer arm fucosylation (Figure 1). 36 Some of these core-fucosylated N-glycoproteins are known and understood, for example, alpha-fetoprotein (AFP) and L-AFP, when core-fucosylated, are associated with poor cancer prognosis. 55
Increased or changed sialylation (Neu5Ac) profiles on both mature N- and O-glycans have been identified as cancer biomarkers (Figure 1).5,56 Since many of the glycosyltransferases responsible for the antennae modifications are capable of acting on either N- or O-glycans, perturbation in this pathway can potentially impact all complex glycosylated proteins. For example, increases in glycotransferases in pancreatic cancers create an extra sialylated N- or O-glycan known as C19-9 which is found in the serum of 75% of pancreatic cancer patients. 34 Non-human glycan N-glycolylneuraminic acid (Neu5Gc), which differs by one oxygen from Neu5Ac, has been found in tumors and malignant tissues and is suspected to be incorporated into glycoproteins by scavenging from dietary sources combined with increased glycosyltransferase activity.57,58
O-GlcNAcylation is generally increased in human cancers,59–61 directly impacting several nucleocytoplasmic signaling pathways.16,32,62–78 An example of a key cancer-related process impacted by O-GlcNAc is epithelial to mesenchymal transition (EMT). TGFβ activates Smad transcription factors leading to the transcription of EMT-related genes. SMAD2 and SMAD4 have been reported to be modified by O-GlcNAc,62,79 and, in the case of SMAD4, this modification prevents proteasomal degradation by blocking interaction and phosphorylation of SMAD4 by glycogen kinase-3β (GSK3β), increasing SMAD4 half-life and transcriptional activity. 80
Often one can find various combinations or all forms of aberrant glycosylation on specific proteins important for proper cell growth and death. 81 Recently, there has been a rapid development of technologies to analyze glycosylation. The enzymes needed for glycosylation are ubiquitous and often essential across organ systems. Therefore, it is imperative to determine which tumor glycoforms drive aggressive disease. The development of technologies to study glycosylation has often overshadowed the need for connection to the functional consequences of the specific glycosylation modifications on individual proteins. Only once these connections are made can potential novel cancer therapies target glyco-modifications and glycoproteins. 82 This review discusses current technologies used to study glycosylation as well as the strengths and limitations of these technologies.
Chemical Biology Toolkit
Since the discovery of glycosylated proteins in the late 1800s scientists have made use of natural biological systems which create, degrade or bind glycan structures. The proteins and enzymes described below are regularly used with traditional methods such as western blotting, immunofluorescence, and UV or fluorescence liquid chromatography (LC) separations as well as with mass spectrometry. While these tools have greatly increased our knowledge about glycan profiles of specific cell and tissue types, alone, they lack analytical precision to identify specific glycan structures. Due to the isobaric and isomeric nature of oligosaccharide modifications, it is often necessary to utilize complementary analytical techniques including various affinity or LC, lectin binding, mass spectrometry and Nuclear Magnetic Resonance (NMR) to fully annotate glycan linkages and isomeric status. Advances in protein level NMR as well as the accessibility of techniques such as Cryo-electron microscopy are additional means of determining the 3D structure and potential molecular interactions of glycoproteins which will not be discussed here.
Glycan Binding Proteins and Antibodies
The most common classes of naturally occurring glycan-binding proteins are known as lectins. These proteins are found throughout nature and bind carbohydrates, either as the soluble forms of oligosaccharides or conjugated to glycoproteins of interest.2,83 First isolated in the late 1800s when it was noted that plant seed extracts tended to agglutinate human blood, novel lectins are constantly being reported in the literature especially as isolated from plants and fungi. Lectins can be purified from their natural sources or recombinantly expressed, and then chemically bound to fluorophores for imaging/immunohistochemistry or to various solid phases for use as affinity purification reagents in studying specific oligosaccharide motifs (Workflows Figure 2). 84 The range of specificity/selectivity of these proteins can vary significantly and can be used to cross-correlate glycan changes.27,42,48,85–87 Lectins can be used individually or in combination to obtain more confidence about glycan structures and have been historically utilized in combination with tools such as HPLC which provide some information about glycan structure but not the precision of precise mass or hydrogen/carbon interactions which can be obtained from techniques like mass spectrometry or NMR respectively. A disadvantage of lectin-based technologies is that often the full specificity of a lectin (or lack thereof) is not perfectly understood and multiple glycoforms will be bound and purified/imaged.
A common reagent, Concanavalin A (ConA) is a lectin originally isolated from jack bean and is now recombinantly expressed and commercialized (Pierce™, Sigma-Aldrich(R), Thermo Fisher ScientificTM, CytivaTM, Vector Laboratories, GlycoMatrixTM, and others) as a glycoprotein purification reagent which preferentially binds α-linked mannose and glucose residues. 88 However, ConA has an affinity for several N-glycan types including oligomannose-type and hybrid-type N-glycans, which bind with high affinity, and biantennary complex-type N-glycans, which bind weakly. Other examples of useful lectins in cancer research include those targeting highly branched, fucosylated, or sialylated (complex) N-glycans. These include Aleuria aurantia lectin (AAL, isolated from orange peel fungus) which binds fucose 89 and Maackia amurensis Lectin (MAL, isolated from seed from the Amur tree) 90 and Sambucus nigra lectin (SNA isolated from elderberry bark) which bind different linkages of sialic acid.91,92 More recently, recombinant engineering of lectins has been a useful approach to improving specificity. One example is the development of a mutant form of AAL with a point mutation at asparagine (N) 224 changing this residue to a glutamine (Q) resulting in N224Q AAL, which has enhanced binding for core fucose, a key cancer biomarker. 89
Development of oligosaccharide-specific antibodies as analytical reagents and biotherapeutics is of great interest but has limited specific success due to the challenges of obtaining specific immune responses only to a glycoform of interest. Glycan, lectin, and antibody arrays are still extremely useful as diagnostic tools and even more powerful when combined with mass spectrometry.93,94
Glycosidases
Endo or exoglycosidases are enzymes that break saccharide bonds to release oligo or monosaccharides respectively. Glycosidases are often derivatives of natural enzymes which break specific or general glycan linkages. These enzymes can be utilized combinatorially with downstream western blots, lectin purifications, HPLC, or mass spectrometry analysis to sequentially break down and identify glycan profiles.
A wide variety of recombinant enzymes have been optimized and purified by companies such as New England BioLabs, Inc(R), Promega, QABio, and others. In the case of N-glycosylation, PNGaseF is a common commercially available reagent which cleaves N-glycan chains from the asparagine side chain, resulting in free glycans and deamidated asparagine residues (Figures 1 and 2).95,96 Endo H, a different recombinant endoglycosidase, cleaves at the core GlcNAc of oligomannose and some complex N-glycans, leaving a GlcNAc-asparagine residue. 97 For O-glycans, O-glycosidase (Endo-α-N-Acetylgalactosaminidase) hydrolyzes the removal of core 1 and core 3 disaccharides from glycoproteins. 98 Other glycosidases hydrolyze bonds at branching or terminal portions of the glycoform. Both linear and branched sialic acid residues can be removed from N-glycan antennae structures using neuraminidase. 99 Various fucosidases exist which cleave at the core or terminal fucose residues within N-glycan structures. 100 Galactosidase hydrolyzes terminal β1-3 and β1-4 linked galactose residues from oligosaccharides. 101 Various recombinant commercial or newly discovered natural glycosidases will have mixed specificities for various glycan core/antenna/and glycan linkages so the product notes should be carefully studied prior to experimental design.
After the removal of part or the full glycan, separations and mass spectrometry techniques can be tailored more specifically for glycans and peptides. 102 Combinatorial usage of lectins and glycosidases in the workflows shown in Figure 2 can enable more confident identification of glycan structures and pinpoint classes of perturbed glycosylation patterns in disease.83,103
Mass Spectrometry Techniques
Mass spectrometers are analytical instruments which allow for the measurement of a mass-to-charge ratio (m/z) of ionized species such as small molecules, lipids, peptides, and proteins which have been given positive or negative charge(s). They have been useful in the analysis of oligosaccharide structure determination since the late 1950s, although the instrument availability, modes of ionization and fragmentation have changed significantly in the last 70 years. 104 Although gas chromatography-mass spectrometry is still a useful tool for glycomics, here we will focus on the current common ionization and fragmentation techniques used in glycoproteomics.
Utilizing a high-resolution, precision technique such as mass spectrometry in order to identify specific glycoforms and modification sites on a protein may seem obvious, but numerous technological hurdles remain to be overcome in order to standardize mass spectrometry-based glycoproteomics.1,105 Isomeric branched glycan structures may have identical mass-to-charge ratios making them indistinguishable without additional separation (retention time) or chemical modifications. The fundamentally different physiochemical properties of glycans versus peptides or proteins mean that varying separation, ionization techniques, and fragmentation methods within the mass spectrometer are required for optimally identifying glycans versus protein or peptide sequences (Figure 2). 106
Adding to the overall complexity of mass spectrometry-based glycoproteomics, N-linked versus O-linked glycopeptides also require different cleavage enzymes, workflows for the identification of sites and glycan structures, enrichment strategies, and even mass spectrometer instrument settings. 106
The mass spectrometry workflows described in Figure 2 use either matrix-assisted laser desorption ionization (MALDI) or electrospray ionization (ESI) of the prepared glycan, glycopeptide, or glycoprotein to create gas phase ions which are directed into the mass spectrometer. In MALDI, a solid phase matrix, such as α-Cyano-4-hydroxycinnamic acid (CHCA), absorbs energy at the wavelength of the pulsed laser leading to ablation from the solid surface and proton transfer creating gas phase sample ions. 107 In electrospray ionization, electrical voltage applied directly to a liquid-liquid junction or emitter creates charged droplets of solvent and sample which desolvate into gas phase sample ions. Once these gas phase ions have been created, they are manipulated and isolated by the use of a voltage gradient, ion optics, and an increasing vacuum. 108 Several common mass analyzers include low resolution (±1 Dal measurements) quadrupoles and ion traps, medium resolution time of flight (ToF), and high resolution Fourier-transform ion cyclotron resonance magnets (FTICR) and orbitraps. Most mass spectrometers also have the ability to fragment ions using collisions of ions with each other as well as with neutral gases (collision induced dissociation (CID) and higher energy collision-induced dissociation (HCD), anion radicals (electron transfer or capture dissociation, ETD/ECD), or the application of UV light (UV photon dissociation). The set of m/z fragments can then be used to reassemble a larger structure. 108
One of the challenges of utilizing mass spectrometry to assign either glycan (removed from protein) or glycopeptide structures is that differing fragmentation techniques and energies are necessary to create fragments of cross-ring, internal glycan, glycan-peptide, and peptide or protein sequences.106,109
Ion Mobility
Ion mobility is an analytical technique where gas-phase ions are separated as they move through an applied electrical field. It can be considered analogous in concept to the “separations based techniques” described below, however it is now often integrated within the mass spectrometer. 110 After sample ionization, ion mobility separates ions based on size and 3D shape around an electrode (field asymmetric ion mobility) or in a drift tube (traditional linear ion mobility) in the presence of an inert buffer gas, often nitrogen. This adds an analytical determinant known as collision cross section (CCS) which can be used to assist in precise identification of glycans and glycopeptides precisely.111,112 Since glycan isomers move differently, especially if the hydroxyls have been modified with methyl, acetyl, or larger substituents, this technique can provide additional confidence in terms of identification of glycan, glycopeptide, or glycoprotein structure. Although classically a technique that was only available in specialized, commercial mass spectrometers incorporating ion mobility are now available from several vendors (BrukerTM (timsTOF), WatersTM (Cyclic ion mobility), SciexTM (SelexIon), MOBILion (MOBIE(R)) and Thermo Fisher ScientificTM (FAIMSpro)). Integrated ion mobility mass spectrometry can be used in any of the techniques described below and has successfully been used in combination with high-resolution precursor m/z and cross-ring fragment ions to confidently elucidate N-glycan isomer structures. 113
MALDI Imaging Mass Spectrometry
MALDI is a form of mass spectrometry where a sample on a solid support can be mixed with a matrix and ionized. In the context of glycosylation, this technique can give a spatial resolution of glycan patterns and is known as either imaging mass spectrometry (IMS) or mass spectrometry imaging (Workflow Figure 2A). Pioneered by the Drake lab in 2013, this technique is especially insightful in the analysis of tissue slices, formalin-fixed paraffin-embedded (FFPE) tissue blocks, and tissue microarrays which are commonly used for clinical samples in cancer.27,114,115 An endoglycosidase such as PNGaseF is applied to release glycans, which are ionized from the solid phase and identified in the mass spectrometer with an output of overlays of intensities of specific glycan forms (Figure 2). This can lead to a deeper understanding of spatial changes to glycans within tissue or tumor biopsies, for example, this technique specifically identified high mannose N-glycans localizing with tumors. 116 Recent mass spectrometry instrumentation advances and the use of laser-induced post-ionization with negative mode detection have the potential to increase the sensitivity of N-glycan tissue imaging by 10´. 115
In addition to direct tissue glycan imaging, the use of the MALDI solid phase allows for profiling of low sample amounts of glycoproteins applied to microspots of lectins, specific antibodies, or affinity tag reagents.28,35,117–120 This is especially useful for potential biomarker discovery and subsequent screening as biofluid samples can be applied to slides and quickly analyzed for glycan patterns.117,121
IMS has the advantage of potentially minimal extra sample processing, no LC-associated issues, and fast total acquisition time per sample (depending on spatial resolution). As in most techniques described here, large amounts of data are created, but specific to IMS, appropriate statistical modeling of spatial resolution must occur on large project scales in order to determine clinically relevant changes. Process convolution R packages are now available to assist with spatial information 122 ; additionally, other bioinformatics teams are beginning to investigate the use of machine learning to improve the initial raw data processing of these complex data sets.123,124
Separations Based Bottom-up Mass Spectrometry
The following techniques utilize methods of separating glycopeptides, glycans, or peptides such as capillary electrophoresis (CE) or LC prior to mass spectrometry detection (Figure 2 workflows B-D). Although mass spectrometry can provide a high-resolution intact glycopeptide, glycan, or peptide m/z with fragment ions, the isomeric and isobaric structure of glycan structures (ie, numerous combinatorial oligosaccharide structures can be formed with the same intact mass with the same subunits in different order and configurations, Figure 1) means that separations such as CE or LC provide orthogonal characteristics (in addition to the potential built-in ion mobility) necessary to provide analytical precision and confidence to isomeric glycoform identifications.
The workflows in this section consist of bottom-up or “shotgun” mass spectrometry in which proteins are digested by proteases into peptides. The most common protease in mass spectrometry workflows is trypsin, which cleaves after lysine and arginine residues (Figure 2). In the study of O-glycans, alternative proteases may be required due to the absence of lysine and arginine residues in regions of high glycosylation.125–127 Large glycan structures can also inhibit efficient digestion by any protease so release of glycans prior to proteolysis can be advantageous (Workflows 2C and 2D).
Similar to the analysis of other PTMs by mass spectrometry, modified proteins or peptides may be enriched by various strategies and analyzed as intact glycopeptides or separately, as glycans and deglycosylated peptides (Figure 2). A thorough 2021 review by Riley et al summarizes the use of various chemical, immuno-, and affinity enrichments for successful mass spectrometry-based N- and O-glycoproteomics. 109 Briefly, a protein of interest may be enriched using in vivo tags (His, FLAG, HA), natural binding protein partners, antibodies/nanobodies, or lectins known to bind glycans on that protein (Figure 2 workflows D and E). Glycopeptides may also be enriched using chromatographic methods such as hydrophobic interaction (HILIC), electrostatic repulsion-hydrophilic interaction, or porous graphitic carbon chromatography (Figure 2 workflow B). Numerous chemical biology techniques utilizing non-natural sugar substrates for the destructive enrichment of glycoproteins or glycopeptides also exist.84,128
Glycopeptide Mass Spectrometry
Successful glycopeptide mass spectrometry (Figure 2B) will result in an identified peptide, glycan structure, and localized modification site. In the most common workflow, following digestion of a pool of proteins to create peptides, glycopeptides are enriched using hydrophilic interaction chromatography (HILIC) followed by LC or CE-mass spectrometry (Workflow Figure 2B). Since glycopeptides are generally more hydrophilic/polar than unmodified peptides they preferentially bind these solid phases. This enrichment strategy can be combined with upfront protein/lectin chromatography for increased success in purifying specific subsets of glycosylated peptides. Successful fragmentation of both the glycan chain and peptide is essential, and new improvements have led to either stepped fragmentation or use of combinations of fragmentation techniques when characteristic glycan fragment ions are identified by the mass spectrometer.106,126 Glycopeptide studies have led to the ability to characterize the glycan occupancy and heterogeneity of modification sites within a cellular milieu.31,109,129 Permutations of this standard workflow have identified many aberrant glycoforms and glycopeptides in various forms of cancer, for example, increased high-mannose N-glycans and sialylated O-glycans in aggressive ovarian cancers 130 and specific core fucosylated glycopeptides associated with liver tumors. 131
Both instrumentation and computational challenges exist in this approach as both a peptide and glycan must be fragmented and correctly identified. Due to the common core structure and consensus amino acid background sequence, there is typically only one N-glycan site per tryptic peptide making the study of these modifications slightly more straightforward than O-linked glycosylation. In contrast, O-linked glycosylation can and does occur on multiple serine or threonine sites per tryptic peptide, meaning extensive fragmentation of the glycopeptide is necessary to localize the glycan. However, the advantage of identifying both glycan site and structure in a single experiment has spurred the field forward in the last ten years. Studies indicate that stepped fragmentation energies can allow for accurate fragmentation of peptide and glycan separately. 106
Even after successful enrichment, detection and fragmentation of both glycan chain and peptide, software to successfully deconvolute these data has been a source of constant improvement in the field of glycobiology. Software was initially developed to identify either glycan or peptides from mass spectrometry-based data, but the field has seen rapid development in options for the analysis of intact glycopeptides improving the ability to identify site and glycan isomer in the same study and now numerous commercial and academic software are available (Byonic, Protein Prospector, GlycoPAT, GPQuest, pGlyco, MSfragger-Glyco, StrucGP to name a few105,118,132–135). An emphasis on appropriate controls and data transparency should be used as the glycoproteomics community is moving towards standards in data reporting. Recent reviews indicate that N-glycan moiety and site specificity can vary widely based on user and software and O-glycan identification and localization can lack reproducibility across laboratories. 136 However, computational focus on the appropriate incorporation of false discovery rates for confident identification of both peptides/glycan sites and glycan structures should begin to move the field towards more consistent and reproducible data analysis. 133
Glycomics (Glycan and Peptide) Profiling
Classic glycomics interrogates glycan profiles of cells or specific proteins by removing and analyzing glycans and protein/peptides using separate mass spectrometry runs (Figure 2C and 2D). N-glycans are typically cleaved with an endoglycosidase such as PNGaseF, while O-glycans are removed by O-glycosidases or β-elimination. 137 Separate experimental designs to analyze glycan and amino acid moieties allow for simplifications in the mass spectrometry and LC/CE separations. Once released, glycans can be purified by class offline on anion exchange columns followed by either direct injection or detection on the mass spec or separation on reverse phase or porous graphite column.138,139 Traditional glycomics commonly involves the modification of free hydroxyl groups on the released glycans with acetyl groups (examples: permethylation, peracetylation) making the released molecules more hydrophobic in order to improve chromatographic resolution. Other chemical modifications to the reducing end of the glycan, such as 2-aminobenzamide, 2-aminobenzoic acid, or 2-aminopyridine, add fluorescence, modify chromatography and can result in improved ionization for mass spectrometry. 140 O-GalNAc glycans can be unstable in alkaline conditions and are better analyzed following β-elimination and reduction of the hemiacetal group. Although complex glycans may have identical precursor ion m/z, CE, LC retention time and potential ion mobility of modified glycans and comparison with standards allows for rapid characterization of the overall glycan profiles of purified proteins. 141 All new researchers in the field should review the recommended Minimum Information Required for a Glycomics Experiment (MIRAGE) for field standard recommendations on designing experiments and reporting results. 142 Peptides can then be analyzed separately using typical reverse phase-LC/MS/MS instrumentation settings and software.
These methods are routinely used for quality control of biotherapeutics including those used in the treatment of cancer (Figure 2D), but glycan profiles of complex mixtures of proteins can also be monitored for a higher-level view of changes across a system (Figure 2C). Recent studies have combined IMS with glycomics to analyze which families of N-glycans are increased in metastatic breast cancer, identifying a specifically increased glycan with poor outcomes. 27 This technique can result in high confidence ratios of specific glycan moieties, but site localization of the glycan groups is lost. The cleavage of the glycan group from the amine asparagine creates a deamidated residue which can be used to infer sites of glycosylation; however, deamidation can and does occur within solution. To use deamidation as a confident identifier of N-glycan sites, multiple controls, and potentially orthogonal endoglycosidases should be used. 143 Separate techniques utilize stringent enrichment of glycosylated peptides (cell-surface capture technology) which may or may not destroy the structure of the glycans themselves, followed by release of the peptides by PNGaseF and identification/quantification by LC-MS. 33
Mass Spectrometry of Intact Glycoproteins
Top-down mass spectrometry is the study of full-length protein sequences to define biologically relevant proteoforms, or specific combinations of protein sequences and PTMs. These analyses used to be limited to labs with large and expensive FTICR mass spectrometers and specialized knowledge for full deconvolution of high m/z species. Recent instrumentation advances have made the techniques more accessible, and in 2018/2019 the Heck research group defined varying glycoproteoforms of engineered erythropoietin 144 and fetuin. 145 Since the interaction of glycoproteins is a key step of viral infection, a byproduct of the SARS-Cov2 pandemic has created a major push for deeper mass spectrometry-based analysis of glycoproteins interactions of viral and human host proteins. By analyzing full-length proteins, the sum of various modifications can be determined—information which is lost when a protein is digested into peptides. Traditionally, glycan analysis of intact proteins is still technically challenging from both instrumentation and computational perspectives. Recent advances by Kelleher et al, however, have proven the technique capable of quantifying glycan patterns on IgGs isolated from Covid-infected patients. 146 Researchers used magnetic beads with recombinantly expressed SARS-Cov-2 spike protein Receptor Binding Domain (RBD) to pull down serum antibodies followed by Individual Ion mass spectrometry. 147 A 2022 study by Wilson et al examined optimizing LC parameters, settling on HILIC for successful optimal resolution of glycoproteoforms of the RBD protein. 148 Although these workflows can be carried out on commercially available orbitrap mass spectrometers, specialized techniques and laborious data analysis are still hurdles to regular glycoproteoforms mass spectrometry. At the time of publication of this review, there were no published reports of intact glycoproteoforms from clinical cancer samples, but numerous groups have taken up the challenge. The commercialization of software and ultra-high mass range instrumentation developed in collaboration with many of the labs discussed above should enable greater community implementation of intact glycoproteomics in any case with well-defined affinity or immunopurification. Ideally, the ability to quantify ratios of specific glycoproteoforms, above and beyond specific glycopeptides, will provide the next level of connectivity between protein and glycosylation changes which occur within cancer.
Challenges and Future Directions
It is clear that the current discoveries surrounding glycosylation in cancer generated from mass spectrometry-based techniques are the tip of the iceberg for understanding these important PTMs.30,35 We expect that continued advances in specific glycan-specific enrichments, including the development of nanobodies 149 and the use of nanopores, 150 combined with standardized protocols and user-friendly software strategies will allow more laboratories to adopt discovery glycoproteomics. Deep visual proteomics combines the capabilities of workflow A and C in Figure 2 to allow for the spatial resolution of both glycans and deep global proteomics.151,152 The application of isobaric tags, in which sets of isotopologues modify peptides with the same exact mass, allows for multiplexing of global peptide and glycopeptide samples for consistent quantitation of glycopeptides across samples. Recent use of a “boost” channel (a sample with a higher quantity of the glycopeptides of interest) with the isobaric labeling system N,N-dimethyl leucine (DiLeu) as a modification of workflow 2B, indicates the potential for improved quantitation and accurate comparisons of glycopeptides across samples. 153 Advancement of the combination of imaging and low quantity or single-cell proteomics will improve the overall spatial resolution and cell type-specific understanding of glycoproteoforms. 154 As a complement, IMS of low abundant proteoforms (resulting in an output of combined 2A and 2E type data) shows real promise for connecting glycoproteoforms across various tissue types. 155
The field of mass spectrometry is also rapidly integrating data-independent acquisition for low-abundance peptides and PTMs, and this is also the case for glycoproteomics. 135 With standardized protocols for glycopeptide sample preparation, data acquisition, and analysis created by consortiums such as the human glycome project (https://human-glycome.org/), Consortium for functional glycomics (http://www.functionalglycomics.org/), Clinical Proteomic Tumor Analysis Consortium (CPTAC https://proteomics.cancer.gov/programs/cptac/), more labs will be able to implement these techniques for successful biomarker discovery.30,136,156 As these techniques are standardized and made more available to researchers and clinicians, we expect more focus on glycosylation as a biomarker or therapeutic target. 156 User-friendly interfaces such as Glycomics@ExPASy, 157 GlyConnect, 158 GlyGen, 159 GlyCosmos Portal, 160 and Glyco.me 161 have been developed, allowing researchers without access to mass spectrometry resources of their own to interrogate differential glycosylation patterns in their samples and probe the underlying pathways.
Comprehensive and user-friendly technologies and databases will continue to drive our collective understanding of glycosylation-regulated systems biology, and the study of these pathways has high potential to be just as fulfilling as the focus on kinase and phosphatase pathways over the last two decades. An understanding of the glycosylation status of cancer therapeutic targets as well as glycan-specific cancer therapeutics holds promise for the treatment of challenging malignancies.162,163 This review emphasizes that glycoproteomics is a challenging but potentially rewarding field with rapid technological advancement.
Footnotes
Abbreviations
Acknowledgments
Thanks to Whitney Smith-Kinnaman for reviewing this manuscript. Figures were created with BioRender.com.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics Statement
Our study did not require ethical board approval because it did not contain human or animal trials.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article. This work was supported by funds provided to ESY by Indiana University and the Simon Comprehensive Cancer Center. Work in the IUSM Proteomics Core was supported, in part, by the Indiana Clinical and Translational Sciences Institute which is funded by Award Number UL1TR002529 from the National Institutes of Health, National Center for Advancing Translational Sciences, Clinical and Translational Sciences Award.
