Abstract
Introduction
While numerous putative signaling pathways have been proposed underlying tumorigenesis through cellular and animal studies over the years, their clinical relevance may only be established through investigations using human cancer tissues. However, in addition to ethical and privacy concerns, there are several technical challenges to utilizing human cancer tissues in biomedical research, especially at the protein level.
Human cancer tissues are mainly preserved in formalin fixed paraffin embedded (FFPE) format. The heavy crosslinking of the proteins in these tissues limits the usage of conventional protein techniques including enzyme linked immunosorbent assay and Western Blot analysis. 1 On the other hand, unlike in a typical laboratory setting where experimental materials are relative homogenous, human cancer tissues differ by age, sex, tumor size, tumor stage, and other clinicopathological factors. Therefore, a large number of human cancer tissue specimens need to be included in the study to eliminate potential bias. Evidently, a high-throughput protein technique is needed to meet this challenge.
In this regard, mass spectrometry (MS) may not be the best choice, as it is more suitable for analyzing a large number of proteins in a few tissue specimens, rather than a few proteins in a large number of tissue specimens. 2 The only high-throughput method to suit this purpose presently is reverse phase protein array. 3 However, the high equipment cost and stringent technical requirement associated with this method make it unlikely to be a standard part of a typical signaling transduction laboratory.
Recently, we introduced the quantitative dot blot (QDB) method to measure protein levels in fresh, frozen, and FFPE tissues absolutely and quantitatively.1,4-6 This is a high-throughput method requiring minimum equipment and training, adaptable in a regular laboratory setting. This method is well suited to investigate and validate findings from cellular and animal studies in human cancer tissues.
In the last 2 decades, extensive clinical studies led to the identification of 2 dominant signaling transduction pathways underlying tumorigenesis in human breast cancer: the estrogen receptor (ER)/progesterone receptor (PR) signaling pathway and human epidermal growth factor receptor 2 (Her2) pathway. Accordingly, surrogate assay has been developed to guide breast cancer patients for targeted therapies in daily clinical practice. 7 In surrogate assay, the activation statuses of these 2 pathways are monitored by examining the protein levels of key molecules in these pathways, including ER, PR, Her2, and Ki67, using immunohistochemistry (IHC). The success of surrogate assay in daily clinical practice suggests that we can monitor the activation statuses of ER/PR and Her2 pathways by following the expression levels of these key molecules in the FFPE specimens.
Yet, despite the success of targeted therapies in clinical practice, patients frequently developed resistance to these treatments afterward. Bidirectional talk between these 2 pathways is proposed to explain this acquired resistance, suggesting that ER/PR and Her2 signaling pathways are inversely related with each other.8,9 Suppression of one pathway through clinical intervention leads to the activation of the other pathway as an escape pathway
Indeed, activation of Her2 signaling pathway was found to lead to ER degradation, while inhibition of Her2 signaling pathway in preclinical studies led to the restoration of ER expression at both mRNA and protein levels. 11 Direct binding between Her2 and ER has also been shown in 293T cells when co-transfected with both ER and Her2 cDNA constructs. 12 Subsequently, co-administration of both endocrine and anti-Her2 therapies (combined therapy) was evaluated in several clinical trials as the solution to overcome acquired resistance among breast cancer patients. 10
In this study, we attempted to further explore the relationship between ER/PR and Her2 pathways by measuring ER, PR, Her2, and Ki67 protein levels as absolute and continuous variables in 852 FFPE breast cancer specimens using the QDB method, prior to any clinical intervention. Protein lysates were extracted from 2 × 15 μm FFPE slices, and the same lysates were used for quantitation of all 4 protein biomarkers to maximally avoid tumor heterogeneity at tissue level. Our results provide a visual support of an inherent bidirectional talk between ER/PR and Her2 pathways leading to breast cancer tumorigenesis.
Materials and Methods
Human Subjects and Human Cell Lines
The reporting of this study conforms to STROBE guidelines. 13 In this retrospective, descriptive, and observational study, a total of 852 FFPE breast cancer tissue specimens were provided consecutively and nonselectively together with documented IHC scores of 4 biomarkers from Yuhuangding Hospital at Yantai, China. The inclusion and exclusion criteria were specimens of female breast cancer patients administered sequentially and nonselectively from 2013 to 2017 prior to any clinical intervention. The ages of the patients were from 32 to 82 years, averaged at 53 years. All the details of the patients have been deidentified.
All the studies were carried out in accordance with the Declaration of Helsinki and approved by the Medical Ethics Committees of Yuhuangding hospital [2017(76)] on August 31, 2017. The informed consent form requirement was waived for archived specimens.
MCF-7 and BT474 cell lysates were obtained from the Cell Bank of Chinese Academy of Sciences (Shanghai, China), and used as controls for all 4 biomarkers.
General Reagents
Recombinant human Her-2/ErbB2 protein (cat# 10004-H20B1) was purchased from Sino Biological Inc. ER, PR, and Ki67 recombinant proteins were purified in the house. QDB plate was purchased from Quanticision Diagnostics, Inc (RTP). Anti-PR (clone 1E2, cat# 790-4296) rabbit monoclonal primary antibody was purchased from Roche Diagnostics GmbH. Rabbit anti-ER (clone SP1, cat# ab13370) antibody was purchased from Abcam Inc. Rabbit anti-Her2 antibody (clone EP3, cat# ZA-0023) and mouse anti-Ki67 (clone MIB1, cat# ZM-0167) were purchased from ZSGB-BIO (www.zsbio.com). HRP labeled Donkey Anti-Rabbit IgG secondary antibody was purchased from Jackson Immunoresearch lab (West Grove). BCA total protein quantification kit was purchased from Thermo Fisher Scientific Inc (Calsband).
Purification of Protein Standards for ER, PR, and Ki67
DNA sequences corresponding to the 1162-1254AA of human Ki67 (NCBI #: NM_002417.4), 455-595AA of human ER-α (NCBI #: NM_000125.3), and 310-417AA of human PR isoform B (NCBI #: NP000917.3) were inserted into pET-32a ( + ) expression vector, respectively, and expressed in
Preparation of FFPE Tissue and Cell Lysates
Two breast cancer tissue FFPE slices (2 × 15 μm) were put into 1.5 ml Eppendorf tubes, and deparaffinized before they were solubilized using solubilization buffer (50 mM HEPES, 137 mM NaCl, 5 mM EDTA, 1 mM MgCl2, 10 mM Na2P2O7, 1%TritonX-100, 10% glycerol, pH 7.6). The recovery rate of the current extraction method was estimated by adding known amounts of formalin fixed Ki67 recombinant protein to microtubes containing a slice of FFPE breast cancer specimen with minimum Ki67 protein expression. The total protein was extracted as described and used for QDB measurement. The recovery rate was calculated as the percentage of measured amount of Ki67 over the amount added to the microtube. We were able to achieve over 80% recovery rate using our extraction method (Supplemental Table 1).
Cells (MCF-7& BT474) were lysed in the solubilization buffer (50 mM HEPES, 137 mM NaCl, 5 mM EDTA, 1 mM MgCl2, 10 mM Na2P2O7, 1%TritonX-100, 10% glycerol, pH 7.6), supplemented with protease and phosphatase inhibitors (100 mM NaF, 0.1 mM phenylmethylsulfonyl fluoride, 5 μg/mL pepstatin, 10 μg/mL leupeptin, 5 μg/mL aprotinin). The supernatants were collected after centrifugation, and the total amount of protein was measured using BCA protein assay kit by following manufacturer's instructions.
Interpretation of IHC Scores
All IHC scores were collected from patients’ medical records. For scoring of ER and PR, specimens with less than (<) 1% positively stained tumor cell nuclei are considered negative, while those with equal or more than (≥) 1% tumor cells with positively stained nuclei are considered positive. Her2 is categorized by following ASCO/CAP guidance “0 as either no staining is observed, or membrane staining that is incomplete and is faint/barely perceptible and within ≤10% of tumor cells; 1 + as incomplete membrane staining that is faint/barely perceptible and within >10% of tumor cells; 2 + as circumferential membrane staining that is incomplete and/or weak/moderate and within >10% of tumor cells or complete and circumferential membrane staining that is intense and within ≤10% of tumor cells and 3 + as circumferential membrane staining that is complete, intense, and within >10% of tumor cells.” 14 The scoring of Ki67 level is to calculate the percentage of positively stained nuclei of tumor cells by counting at least 500, preferably 1000 tumor cells in 3 field of views under high magnification (×400). 15 Staining of any level is considered positive.
QDB Analysis
Sample pools were prepared by mixing tissue lysates from 4 FFPE tissue specimens with an IHC score of 3 + for Her2, and IHC score of 90% for ER, PR, and Ki67 to define the linear range of QDB assay respectively (Supplemental Fig. S1). The pooled lysates were serially diluted side by side with the recombinant proteins for defining the standard curves of QDB analysis.
The QDB process was described elsewhere with minor modifications.1,4,6 In brief, the final concentration of the FFPE tissue lysates were adjusted to 0.25 μg/μL for Her2 and Ki67 and 0.125 μg/μL for ER and PR, and 2 μL/unit was used for QDB analysis in triplicate. The QDB plate was then dried for 1 h at RT and then blocked in 4% nonfat milk for 1 h. Next, it was put into a 96-well microplate with 100 μL primary antibody. Dilution used for antibodies were Her2 1:1500 in blocking buffer; ER 1:250 in blocking buffer; PR 1:8 in PBS; Ki67 1:1000 in blocking buffer. Then the plate was incubated overnight at 4 °C. Afterward, the plate was rinsed twice with TBST and washed 3×10 min. It was then incubated with either a donkey anti-rabbit or donkey anti-mouse secondary antibody for 4 h at RT, rinsed twice with TBST, and washed 4 × 10 min. Finally, the QDB plate was inserted into a white 96-well plate prefilled with 100 μL/well ECL working solution for 3 min. The chemiluminescence signal of the combined plate was quantified by using the Tecan Infiniti 200pro Microplate reader with the option “plate with cover.”
For the 852 FFPE specimens, each sample was measured 3 times, each time in triplicate. To ensure the accuracy of the results, the consistency of the experiments was maintained by including BT474 and MCF-7 cell lysates with predocumented biomarker levels in all the experiments. The result was considered valid when the calculated biomarker levels of control cells were within 25% of the predocumented biomarker levels. The absolute biomarker levels were determined based on the dose curve of protein standard, with those specimens with chemiluminescence reading of less than 2 fold over blank were defined as nondetectable, and entered as 0 for data entry. The reliability of the QDB method was evaluated by performing Spike-and-recovery assay using Ki67 recombinant protein in lysates extracted from FFPE breast cancer tissues as shown in Supplemental Table 2.
Data Analysis
All the data were presented as Mean ± SEM. All the 3D analyses of biomarkers were performed using Origin pro 9.1 software from Originlab Corp (Northampton). All the statistical analyses, including the unpaired two-tailed Student's
Results
In this study, all the specimens were collected from local hospital sequentially, nonselectively, and anonymously as 2 × 15 μm FFPE slices prior to any clinical intervention. These specimens were used for the preparation of total tissue lysates by deparaffinization and solubilization with a lysis buffer. All 4 biomarkers were measured using the same lysate prepared from each FFPE specimen, with intra- and inter-CVs below 25% when measured 3 times, each time in triplicate.
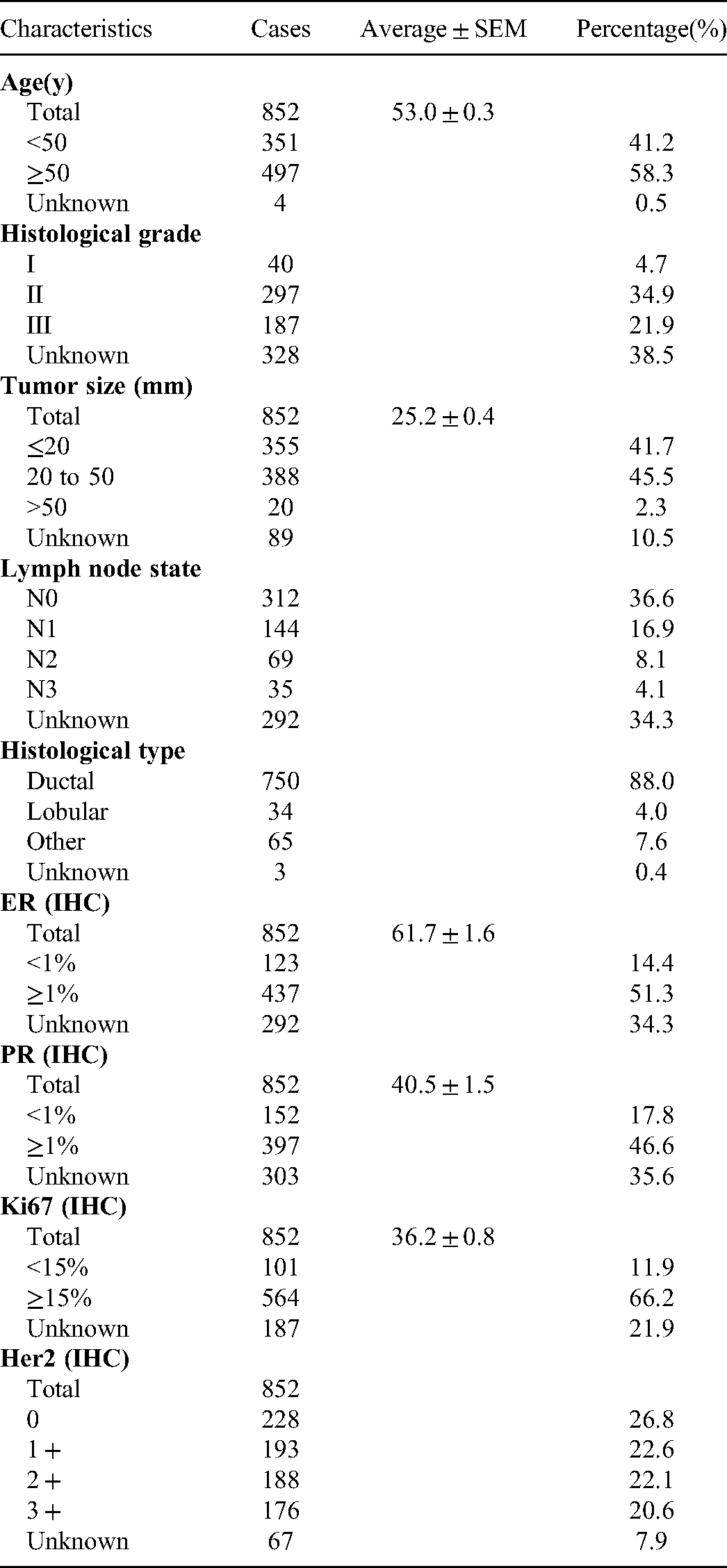
The clinicopathological parameters of these specimens are listed in Table 1. To ensure the consistency of the method, the absolute levels of both Her2 and Ki67 of the first 336 specimens were measured with 2 IHC antibodies independently. The correlations of the measured results were all above 0.96 when analyzed with Pearson's correlation analysis.1,6
Clinicopathological Characteristics of 852 FFPE Breast Cancer Samples.
The distributions of all 4 biomarkers are shown in Figure 1a to 1d. The correlations of QDB results with provided IHC scores are shown in Figure 1e to 1h. Our results were found to be highly correlated, with IHC results for Her2 (

Distribution of Her2, estrogen receptor (ER), progesterone receptor (PR), and Ki67 levels as absolute and continuous variables among 852 FFPE specimens (
The expression levels of these 4 biomarkers were also measured in the breast tissues of other pathological states other than cancer (nontumor tissues). We found that both Ki67 and Her2 levels in nontumor tissue specimen were undetectable (Supplemental Fig. S3a). Meanwhile, there were significantly lower expressions of ER and PR in nontumor tissue specimens than those of tumor tissue specimens when analyzed using an unpaired two-tailed Student
Next, we explored the putative relationships among ER, PR, and Her2 by creating 3-dimensional scatterplot using their protein levels as X, Y, and Z coordinates (Figure 2a). We observed the specimens segregating naturally into 3 distinctive subgroups based on their spatial distributions. We named the first group of specimens the hormone receptor (HR) group as they spread flat on the ER–PR plane, representing specimens with dominant expression of hormone receptors and minimum Her2 expression. The second group we named the Her2 group as they were found wrapping the Her2 axis, representing specimens with strong Her2 expression and minimum hormone receptors expression. The third group accumulated at the intersections of ER, PR, and Her2 axes, representing specimens lacking strong expressions of ER, PR, and Her2. We named this group the corner group. Few specimens were found floating in the ER–PR–Her2 space, indicating the lack of specimens with strongly expression of all 3 biomarkers simultaneously.

Three-dimensional distribution of 852 FFPE specimens based on the absolute and quantitative levels of ER, PR, and Her2. (
This unique pattern persisted until we narrowed the 3D view into a small block of ER <0.2 nmol/g, PR <0.8 nmol/g and Her2 <0.3 nmol/g, where we began to find specimens distributed randomly inside (Figure 2b). Therefore, we used these values as cutoffs to separate specimens as 264/588 (31.0%/69.0%) for ER, 301/551 (35.3%/64.7%) for PR, and 220/632 (25.8%/74.2%) for Her2. We also assigned 357 specimens into HR group (41.9%), 220 into Her2 group (25.8%), and 275 into corner group (32.3%).
Interestingly, we identified that 169 out of 220 (76.8%) specimens from Her2 group were within both the ER and PR cutoffs. For the remaining 51 specimens, 42 out of 51 (82.4%) were within either the ER or PR cutoffs. In other words, 211 out of 220 (95.9%) specimens in the Her2 group were with either ER <0.2 nmol/g or PR <0.8 nmol/g. Among the 9 outliers, 6 specimens were at the vicinity of ER or PR cutoffs. The other 3 specimens were also the only ones with medium to strong expressions of all 3 biomarkers simultaneously, in agreement with our observation that few specimens were floating in the ER–PR–Her2 space.
The Ki67 levels of these 3 groups were evaluated in Figure 3a. We found the averaged Ki67 levels were 4.00 ± 0.39 nmol/g, 4.35 ± 0.26 nmol/g, and 2.81 ± 0.16 nmol/g for corner, Her2, and HR groups, respectively. There were statistical differences between the corner and HR groups (
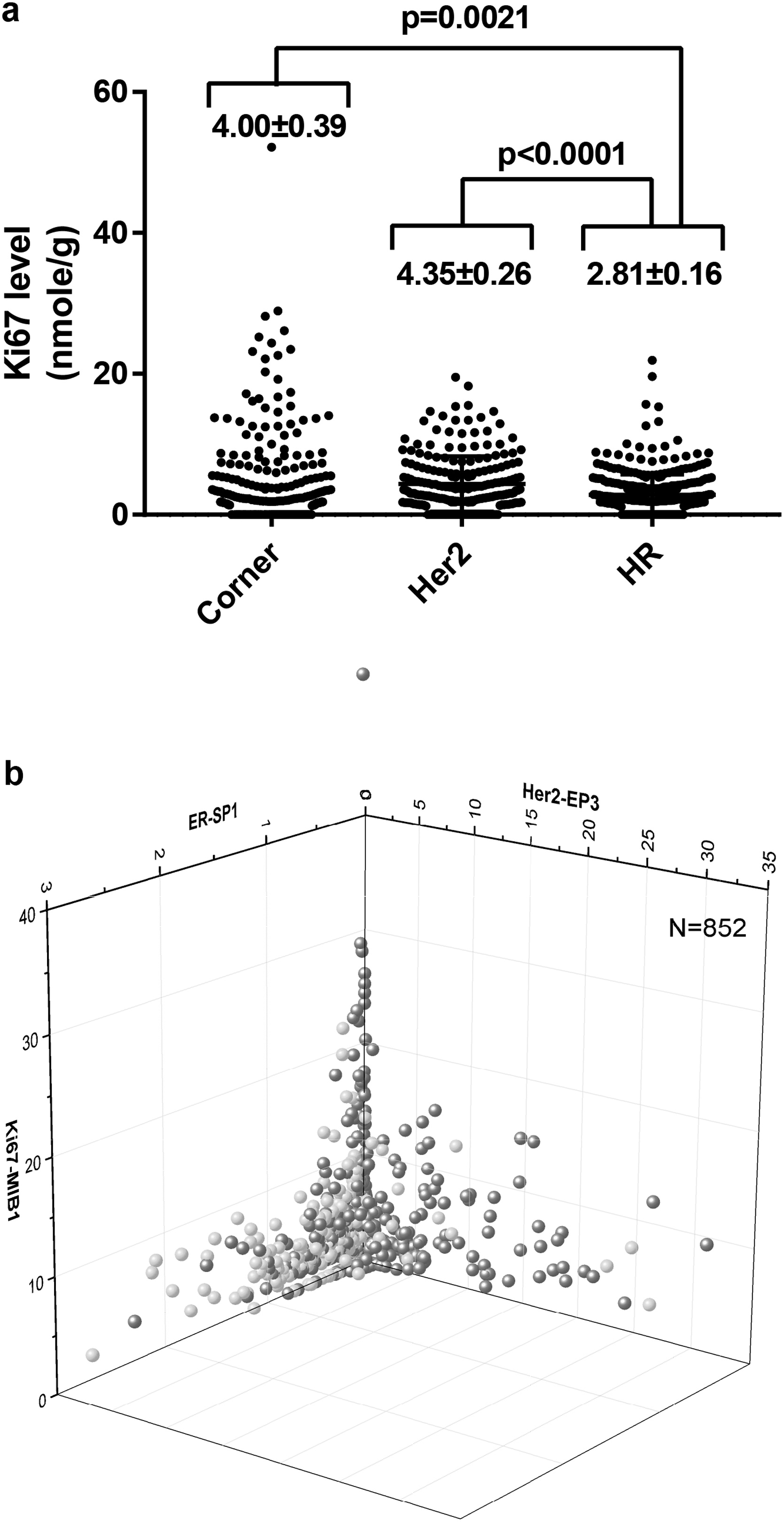
Evaluation of Ki67 levels of specimens grouped by their spatial distribution. (
However, when observing the 3D scatterplots of ER–Her2–Ki67 (Figure 3b), ER–PR–Ki67, and PR–Ki67–Her2 (Supplemental Fig. S4), we found that specimens with the highest Ki67 levels were at the intersection of ER, PR, and Her2. We managed to include PR information in the 3D scatterplot of ER–Her2–Ki67 by assigning specimens with PR <0.8 nmol/g as red, and PR≥0.8 nmol/g as blue in Figure 3b. Specimens with the highest Ki67 levels were found exclusively in red in this picture.
The clinical relevance of the suggested block of ER <0.2 nmol/g, PR <0.8 nmol/g, and Her2 <0.3 nmol/g was explored next. We hypothesized that these cutoffs might represent the cutoffs to identify specimens with overexpressed biomarkers. We tested this hypothesis with Her2 first, as it is one of the best-standardized protein biomarkers for breast cancer patients.17,18 The recommendations from American Society of Clinical Oncology/College of American Pathology (ASCO/CAP) was followed to differentiate Her2 + specimens from Her2− specimens based on IHC results, and receiving operative characteristic (ROC) analysis was performed with measured Her2 levels. As expected, we confirmed Her2 at 0.3 nmol/g as the optimized cutoff value to achieve the best sensitivity and specificity at 84.8% and 97.2% respectively with IHC results, with overall concordance rate at 93.8% with IHC analysis (Figure 4a and b).
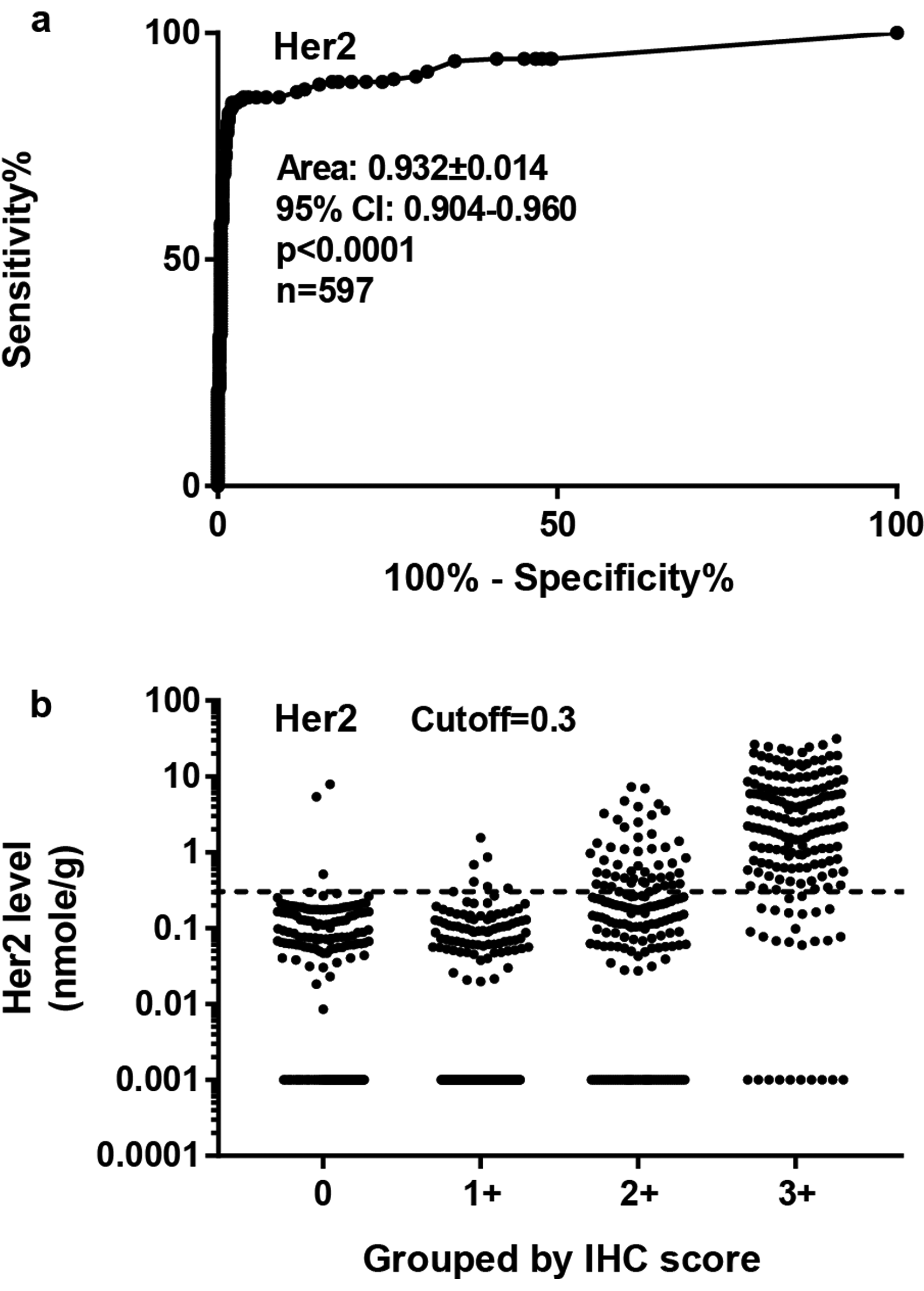
Evaluating the sensitivity and specificity of QDB method using receiver operative characteristics (ROC) analysis based on provided Her2 IHC scores from local hospitals. (
However, when cutoffs were developed based on the recommended IHC score at 1% for both ER and PR by ASCO/CAP using ROC analysis, we achieved optimized sensitivity and specificity at 0.045 and 0.47 nmol/g for ER and PR, respectively (data not shown), significantly different from our proposed 0.2 and 0.8 nmol/g for ER and PR. We also failed to observe any distributional differences graphically around these values (0.045 or 0.47 nmol/g) in our 3D scatterplot.
Discussion
In this study, we were able to unprecedentedly measure the protein levels of ER, PR, Her2, and Ki67 absolutely and quantitatively in over 800 FFPE human breast cancer specimens using the QDB method. These specimens covered tumors of all sizes, grades, and node statuses to be a true reflection of the real world situation. We also showed the first 3D scatterplot based on absolutely quantitated ER, PR, and Her2 protein levels of these specimens. Their unique distribution pattern in the ER–PR–Her2 space provided direct visual support of bidirectional talk between ER/PR and Her2 pathways in breast cancer.
While this observation had been hinted by Konecny
The success of surrogate assay in daily clinical practice clearly suggested that we might use the expression levels of ER, PR, and Her2 to monitor the activation statuses of ER/PR and Her2 signaling pathways. Thus, based on their spatial distribution, we may consider HR group as specimens dominated by ER/PR signaling pathway, the Her2 group as those dominated by Her2 signaling pathway, and the corner group as those with neither pathways being activated. In addition, among 852 specimens used in this study, only 3 were with both pathways activated simultaneously. Thus, for the majority of specimens, there was only one pathway activated at a time, representing a “seesaw” relationship between these 2 pathways.
More importantly, we observed this “seesaw” relationship in specimens prior to any clinical intervention. Thus, our observations challenge the escape theory underlying the acquired resistance of breast cancer patients.8,10,11 Rather, they suggest that this regulation existed as part of an intrinsic signaling network.
The “seesaw” hypothesis might be used to explain the seemingly disappointing results of several clinical trials of combined therapy. 10 While both overall response rate and clinical benefit rate were improved significantly with combined therapy, these improvement did not translate into improved overall survival (OS) of patients. 10 Considering only one pathway was activated at any given time, it is possible the added side effects from combined therapy overshadowed their putative benefit over time. Thus, we propose to alternate endocrine therapy with ant-Her2 therapy periodically to take full advantage of the clinical benefits of these therapies. Of course, the timing of their alternation would pose a new challenge to clinicians in the future.
We further hypothesized that the activation of ER/PR and Her2 pathways required a common factor (Figure 5). Access of this factor by ER/PR pathway would deny its access by Her2 pathway (Figure 5a). Likewise, access of this factor by Her2 pathway denied its access by ER/PR pathway (Figure 5b). On the other hand, Figure 5c represented the state when none of these 2 pathways were able to get access to this common factor. Perceivably, this putative common factor, or factors, would be a promising drug target to shut down both pathways without incurring added side effects of combined therapy.

The “seesaw” hypothesis. The unique distribution patterns of the breast cancer specimens in the ER–PR–Her2 space suggest a “seesaw” relationship, with predominant activation of ER/PR pathway in the HR group (
The spatial distribution of individual specimens in 3D space also suggest that absolutely quantitated protein biomarker levels may be used collectively as a “fingerprint” to differentiate individual specimen at population level. 19 The current prevailing IHC-based categorized results are clearly insufficient to distinguish a specimen accurately at population level. Their results are not proportional variables. They are also plagued with subjectivity and inconsistency inherently associated with IHC method. In contrast, absolutely quantitated protein biomarkers provide the much needed precision to distinguish individual specimen, as reflected convincingly in the ER–PR–Her2 space described in current study. Admittedly, there remains a portion of specimens gathered at the corner of this space. However, we can always add more protein biomarkers to separate these specimens effectively at population level. For example, we may use ER, PR, Her2, Ki67, and cyclinD1 to further separate the specimens accumulating in the corner. Thus, the protein levels of a set of biomarkers, combined with the traditional clinicopathological factors, maybe used to generate a unique “fingerprint” for every breast cancer specimen available. 19
When large number of FFPE specimens with known outcomes are associated with their unique “fingerprints,” we will have a powerful tool to evaluate the clinical outcomes of these specimens individually at population level. We foresee that patients will no longer be limited by 5 intrinsic subtypes, but rather can be grouped in as many subgroups as necessary based on the similarity of their “fingerprints.” With a sufficient number of FFPE specimens available, this platform should provide clinicians/patients an unprecedented opportunity to evaluate clinical outcomes from various angle based on their “fingerprints” at population level. 19
However, caution is warranted when interpreting this study for 2 reasons. First, this was a retrospective observational study. It remains to be seen if this is a universal phenomenon in breast cancer specimens. Nevertheless, the large number of the specimens of all types in this study should compensate for this weakness to a certain degree. Second, our hypothesis is very speculative, lacking much needed clinical evidences other than what was observed
Conclusion
By using the QDB method to unprecedentedly measure ER, PR, Her2, and Ki67 as absolute and continuous variables in 852 FFPE breast cancer specimens, we provided visual support of a “seesaw” relationship between ER/PR and Her2 pathways. Its universal presence, regardless of the tumor sizes, grades, and intrinsic subtypes, suggests that it is part of intrinsic signaling network underlying breast cancer tumorigenesis. The spatial distribution of individual specimens in ER–PR–Her2 also suggested that absolutely quantitated protein biomarkers may be used as a “fingerprint” to distinguish individual specimen at population level, which may serve as the foundation for “big data”-supported cancer research in the near future. More importantly, our results showed that QDB method may be used as an effective tool to explore, evaluate, and validate the signaling network underlying tumorigenesis in archived human cancer specimens.
Supplemental Material
sj-docx-1-tct-10.1177_15330338211065603 - Supplemental material for 3D Visualization of the Dynamic Bidirectional Talk Between ER/PR and Her2 Pathways
Supplemental material, sj-docx-1-tct-10.1177_15330338211065603 for 3D Visualization of the Dynamic Bidirectional Talk Between ER/PR and Her2 Pathways by Jiahong Lyv, Guohua Yu, Yunyun Zhang, Yan Lyv, Wenfeng Zhang, Jiandi Zhang and Fangrong Tang in Technology in Cancer Research & Treatment
Footnotes
Author Contributions
JL, YZ, WZ, and FT performed all the assays and data analysis; GY provided clinical samples; YL performed all the statistical analysis; JZ designed and supervised the overall study and drafted the manuscript; JL, GY, and YZ contributed to data interpretation and edited the manuscript.
Declaration of Conflicting Interests
The authors disclosed receipt of the following potential conflict for the research, authorship, and/or publication of this article: JL, YZ, WZ, JZ, and FT are employees of Yantai Quanticision Diagnostics, Inc., a division of Quanticision Diagnostics, Inc., which owns the patent for the QDB method. GY declared no conflict of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study is funded by Yantai Quanticision Diagnostics, Inc.
Ethics Statement
Supplemental Material
Supplemental material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
