Abstract
Approximately 40% of rectal cancers during initial diagnosis are identified as locally advanced rectal cancers (LARCs), for which the standardized treatment scenario is total mesorectal excision following neoadjuvant chemoradiotherapy (nCRT). nCRT can lead to discernible reductions in local relapse rate and distant metastasis rate in LARC patients, in whom previously inoperable tumors may potentially be surgically removed. However, only 4% to 20% cases can attain pathological complete response, and the remaining patients who are unresponsive to nCRT have to suffer from the side effects plus toxicities and may encounter poor survival outcomes due to the late surgical intervention. As such, employing potential biomarkers to differentiate responders from nonresponders before nCRT implementation appears to be the overarching goal. Well-defined competing endogenous RNA (ceRNA) networks include long noncoding RNA (lncRNA)-microRNA (miRNA)-mRNA and circRNA-miRNA-mRNA networks. As ceRNAs, lncRNAs, and circRNAs sponge miRNAs to indirectly suppress miRNAs downstream of oncogenic mRNAs or tumor-suppressive mRNAs. The abnormal expression of mRNAs regulates the nCRT-induced DNA damage repair process through pluralistic carcinogenic signaling pathways, thereby bringing about alterations in the nCRT resistance/sensitivity of tumors. Moreover, many molecular mechanisms relevant to cell proliferation, metastasis, or apoptosis of cancers (eg, epithelial-mesenchymal transition and caspase-9-caspase-3 pathway) are influenced by ceRNA networks. Herein, we reviewed a large group of abnormally expressed mRNAs and noncoding RNAs that are associated with nCRT resistance/sensitivity in LARC patients and ultimately pinpointed the backbone role of ceRNA networks in the molecular mechanisms of nCRT resistance/sensitivity.
Keywords
Introduction
Colorectal cancer is the third most common carcinoma and the second most common cause of cancer-related death in the world. 1 Approximately 30% of colorectal cancers are rectal cancers, of which 40% are definitively identified as locally advanced rectal cancer (LARC) during initial diagnosis.2,3 It is significant that rectal cancer has experienced a steadily reduced incidence in developed countries but has increased to a rate of 4.2% per year in mainland China, establishing it as the fifth leading cause of cancer-related mortality in this country.4,5 Currently, the standardized treatment paradigm for LARC patients is total mesorectal excision after neoadjuvant chemoradiotherapy (nCRT), which results in a resounding shrinkage of the local relapse rate and an increased pathological complete response (pCR) rate.6,7 In parallel, nCRT may provide an opportunity for radical surgery on previously inoperable tumors. Nevertheless, only 4 to 20% of postnCRT LARC patients can achieve pCR. 8 The remaining patients who are inert to nCRT not only suffer from side effects and toxicities but also have a high risk of poor prognosis due to late surgical interventions. Therefore, it is crucial that a distinction is made between the nCRT-sensitive cohort and the nCRT-resistant cohort by predicting which patients may benefit from nCRT. In this case, the sensitive cohort will receive nCRT directly, and the resistant cohort will be spared from nCRT and triaged to local resection or other mild therapeutic regimens.
nCRT resistance gives rise to tumor recurrence and metastasis and is a direct reason for polytherapeutic failure and the shortened overall survival (OS) of LARC patients; thus, there is an urgent need to overcome nCRT resistance and ameliorate nCRT sensitivity. After nCRT, tumor regression grade (TRG) is an independent factor influencing the long-term survival of LARC patients. 9 Mandard et al 10 in 1994 defined TRG, which was divided into 5 histologic grades: TRG1 (complete regression), no histologically recognizable residual tumor cells and fibrosis permeating all layers of the esophageal wall; TRG2, rare residual tumor cells scattered in the fibrosis; TRG3, presence of an increased number of residual cancer cells but still with a lower proportion than fibrosis; TRG4, residual cancer cells overgrowing fibrosis; and TRG5 (complete nonregression), absence of tumor regressions. Generally, postnCRT patients with TRG1-2 are considered to be the sensitive cohort, while those with TRG3-5 belong to the resistant cohort. 11
The growing appreciation of the molecular mechanisms in nCRT resistance/sensitivity and the investigation of potential biomarkers for differentiating sensitive cohorts from resistant cohorts have evolved through many clinical studies using the above definitive classifications. Herein, we reviewed relevant biomarkers from mRNA and noncoding RNA (ncRNA) domains within the present molecular mechanisms of nCRT resistance/sensitivity in LARC patients. In addition, we summarized or recalculated the prognostic performance of many potential biomarkers in distinguishing sensitive cohorts and resistant cohorts in terms of the original data from the included studies.
nCRT Resistance/Sensitivity-Related mRNAs
DNA Damage Repair Pathway
The most classical mechanism for tumorous CRT resistance is DNA damage repair (DDR), 12 which predominantly includes 2 mainstays, the nonhomologous end joining (NHEJ) pathway and the homologous recombination (HR) pathway. The DNA repair proteins Ku70 and Ku80 participate in the DDR process by the NHEJ pathway, which is crucial for the G1 to S phase. Concomitantly, the DNA repair proteins RAD23B and RAD51 are involved in the HR pathway, which plays a key role in the S to G2 phase and first needs to form a heterodimer with a variety of excision repair cross-complementing (ERCC) proteins (eg, ERCC1, ERCC2, XRCC2, XRCC3, and BRCA2) (Figure 1).

The backbone role of ceRNA networks in neoadjuvant chemoradiotherapy resistance/sensitivity of rectal cancers.
Dysregulated mRNA-Induced nCRT Resistance/Sensitivity via the DNA Damage Repair Pathway
Chromodomain helicase DNA-binding protein 4 (CHD4), an important subunit of the nucleosome-remodeling and histone deacetylation chromatin-remodeling complex, is involved in the DDR process and maintains genomic integrity and stability via the HR pathway. 13 In rectal cancer cells, CHD4 mRNA, in cooperation with DNA methyltransferases, can silence many tumor-suppressive genes (eg, MLH1, SFRP1, SFRP2, SFRP4, TIMP2, and TIMP3), thus driving the Wnt/β-catenin signaling pathway. 14 Findings indicated that upregulated CHD4 mRNA performed well in the prognosis of nCRT resistance in LARC patients (accuracy = 60%; Table 1), and CHD4 mRNA knockdown enhanced nCRT sensitivity. 19 The nucleotide excision repair (NER) pathway shows a major impact on nCRT resistance, which requires the participation of multiple ERCC proteins (eg, ERCC1 and ERCC2).32-34 Overexpressed ERCC1 mRNA in LARC patients can elicit nCRT resistance by accelerating the NER pathway and results in a poor prognosis, with reliable predictive performance of nCRT resistance (accuracy = 73%; Table 1). 22
Prognostic Performance of Overexpressed Biomarkers in Predicting nCRT Resistance and nCRT Sensitivity of Locally Advanced Rectal Cancer.

Abbreviations: PPV, positive predictive value; NPV, negative predictive value.
Thymidylate synthase (TYMS) is a downstream targeted molecule of 5-fluorouracil (5-FU) chemotherapy and is essential for DNA synthesis. 35 Upregulation of TYMS mRNA maps to 5-FU resistance in LARC patients. 36 Furthermore, positive expression of TYMS mRNA in circulating tumor cells is an effective biomarker to predict nCRT resistance (accuracy = 83%; Table 1) and is not detected in any responders, 21 suggesting that TYMS-negative patients are sensitive to nCRT. RAD18 is an E3 ubiquitin-linked enzyme that maintains the integrity and stability of the genome through several DNA repair pathways, by which overexpressed RAD18 mRNA leads to CRT resistance in multiple human cancers.8,37 Increased RAD18 mRNA expression concurrently inhibits caspase-9-caspase-3-mediated apoptosis in rectal cancers, which further reinforces nCRT resistance (Figure 1). 8 A previous clinical study corroborated the good prognostic performance of overexpressed RAD18 mRNA in identifying nCRT resistance in LARC patients (accuracy = 65%; Table 1). 8
Epidermal growth factor receptor (EGFR) indirectly engages in the regulation of the DDR process. Specifically, EGFR can be activated by irradiation and then internalized into the nucleus to form a complex with DNA-dependent protein kinase (DNA-PK), which is associated with the promoted DDR process.38,39 Rab5C modulates the internalization process of EGFR and elevates the expression of Ku70 and Ku80, 40 thus enhancing CRT resistance in rectal cancers (Figure 1). In light of these findings, a clinical study found that EGFR mRNA-negative results combined with VEGF mRNA-positive results performed well in differentiating the nCRT resistant cohort from the nCRT sensitive cohort (accuracy = 65%; Table 1). 23
The PI3 K/AKT/mTOR signaling pathway changes the CRT resistance of cancer cells via direct regulation of the DDR process. 27 Coenzyme A (CoA) and its derivatives take part in multiple pathways of cell metabolism, including pyruvate oxidation, fatty acid synthesis, cell cycle processes, and cell death. 41 Human rectal cancer enriches CoA synthase (COASY), which causes an inferior response to CRT by activating the PI3 K/AKT/mTOR signaling pathway, 17 while the overexpression of COASY mRNA has value in predicting nCRT resistance in LARC patients (accuracy = 67%; Table 1). The oncogene Ha-Ras uses the PI3 K/AKT/mTOR signaling pathway to activate astrocyte elevated gene-1 (AEG-1) mRNA, which in turn prompts the activation of multiple carcinogenic signaling pathways, eg, PI3 K/AKT/mTOR, Wnt/β-catenin, and NF-kB (Figure 1).42-46 Overexpressed AEG-1 mRNA is an independent risk factor for disease-free survival and distant metastasis-free survival of postnCRT LARC patients, 47 meaning that it can be used as a potential biomarker for the prognosis of nCRT resistance.
There is unequivocal evidence that C-C motif chemokine receptor 6 (CCR6) mRNA facilitates the oncogenesis, progression, and metastasis of rectal cancer.48,49 Additionally, upregulated CCR6 mRNA maintains the integrity and stability of DDR direct participants (eg, DNA-PK and RAD51) by activating its downstream signaling molecules, that is, Akt and ERK, thus triggering nCRT resistance and attenuating nCRT efficacy in LARC patients (Figure 1). 16 Table 1 notes that overexpression of CCR6 mRNA was a reliable biomarker in the prognosis of nCRT resistance in LARC patients (accuracy = 71%).
Cyclooxygenase-2 (COX-2), a well-known inflammatory reaction factor, demonstrates critical functions in tumorigenesis, including progression, metastasis, and angiogenesis.50-53 Prostaglandin E2 (PGE2)-induced transformations extend the G1 phase by strengthening the expression of cyclin D1 (CCND1) and establishing the antiapoptotic function of PGE2. 54 COX-2 fosters PEG2 production; thus, the inhibition of apoptosis occurs. 55 COX-2 mRNA overexpression is positively associated with the poor prognosis of post-nCRT LARC patients, 56 and accordingly, there is a postulation that the administration of COX2 inhibitors prior to nCRT may reduce the distant metastasis rate and prolong OS.
N-myc downstream-regulated gene 1 (NDRG1) maintains the function of E-cadherin and β-catenin, which are both involved in cell adhesion and localization and are thought to be potential tumor metastasis suppressors. 57 Interestingly, NDRG1 mRNA overexpression gives rise to elevated expression levels of CRT resistance-related proteins, eg, MDR, LRP-1, and MRP-1. 58 Gene silencing of NDRG1 mRNA makes rectal cancer cells sensitive to CRT by creating more DNA double-strand breakages. 59 Epithelial-mesenchymal transition (EMT) constitutes the molecular mechanism of tumorous CRT resistance, in which nucleic β-catenin accumulates, E-cadherin is reduced, miR-200c is underexpressed, and tumor budding begins to advance.29,60 Nucleic β-catenin was significantly increased in the nCRT-resistant cohort of LARC patients relative to the nCRT-sensitive cohort of those patients (57.6% vs 16.7%, P < .001), and utilizing it as a single biomarker to predict nCRT resistance showed an effective performance (accuracy = 83%; Table 1). 15 OCT4 is a key transcription factor in embryonic stem cells that can render LARC patients resistant to nCRT by accelerating the EMT process. In detail, OCT4 mRNA upregulates the expression of an EMT-related transcription factor, zinc finger E-box binding homeobox 1 (ZEB1), and gene silencing of ZEB1 in turn can reverse OCT4 mRNA-induced nCRT resistance. 61 Collectively, OCT4 mRNA is ZEB1-dependent in its elevation of nCRT resistance in LARC patients.
Cancer stem cells (CSCs) exhibit an inherent antiapoptotic nature and robust CRT resistance due to their intrinsic DDR capability. 62 CD133 mRNA is a specific biomarker for multiple phenotypes of CSCs, and its overexpression implies the probable presence of nCRT resistance in LARC patients (accuracy = 72%; Table 1). 18 Since retinoblastoma binding protein 6 (RBBP6) demonstrates the capability to bind with the tumor suppressors p53 and Rb, it may be involved in the cell cycle, proliferation, apoptosis, and CRT resistance of tumor cells.63-65 Overexpressed RBBP6 mRNA arrests the G2 to M phase and regulates the apoptosis pathways, which leads to CRT resistance of rectal cancer cells. In contrast, inhibiting cellular RBBP6 expression predisposes rectal tumors to increased CRT sensitivity. 66 Pim-3 is a member of the Pim family that is recognized as an anti-apoptotic oncogene and causes tumor growth by synergistically working with the proto-oncogene Myc. 20 Studies have shown that the expression of Pim-3 in colorectal cancer tissues was significantly increased compared to that in normal colon tissues (32.6% vs 0.02%; P < .001). 67 The pCR rate in LARC patients with overexpressed Pim-3 mRNA was significantly lower than that in patients with underexpressed Pim-3 mRNA (P = 0.001), 20 indicating good prognostic performance of the overexpressed Pim-3 mRNA in differentiating the nCRT-resistant cohort from the nCRT-sensitive cohort (accuracy = 63%; Table 1).
Direct Evidence of Dysregulated mRNA-Induced nCRT Resistance/Sensitivity
In addition to the aforementioned mRNAs involved in the molecular mechanisms of nCRT resistance, a large body of studies have identified a direct correlation between many upregulated mRNAs68-82 or other downregulated mRNAs72,83,84 and the nCRT resistance of LARC patients (Table 2), indicating that they can be used as potential biomarkers to predict postnCRT prognosis. For example, overexpression of KRAS, PDPK, PPP2R5C, and YES1 mRNAs and underexpression of PPP2R1B mRNA are responsible for the poor prognosis of postnCRT LARC patients via activation of the PI3 K/AKT/mTOR signaling pathway. 72
Direct Evidence of Abnormal RNA Expression-Induced nCRT Resistance in Locally Advanced Rectal Cancer.
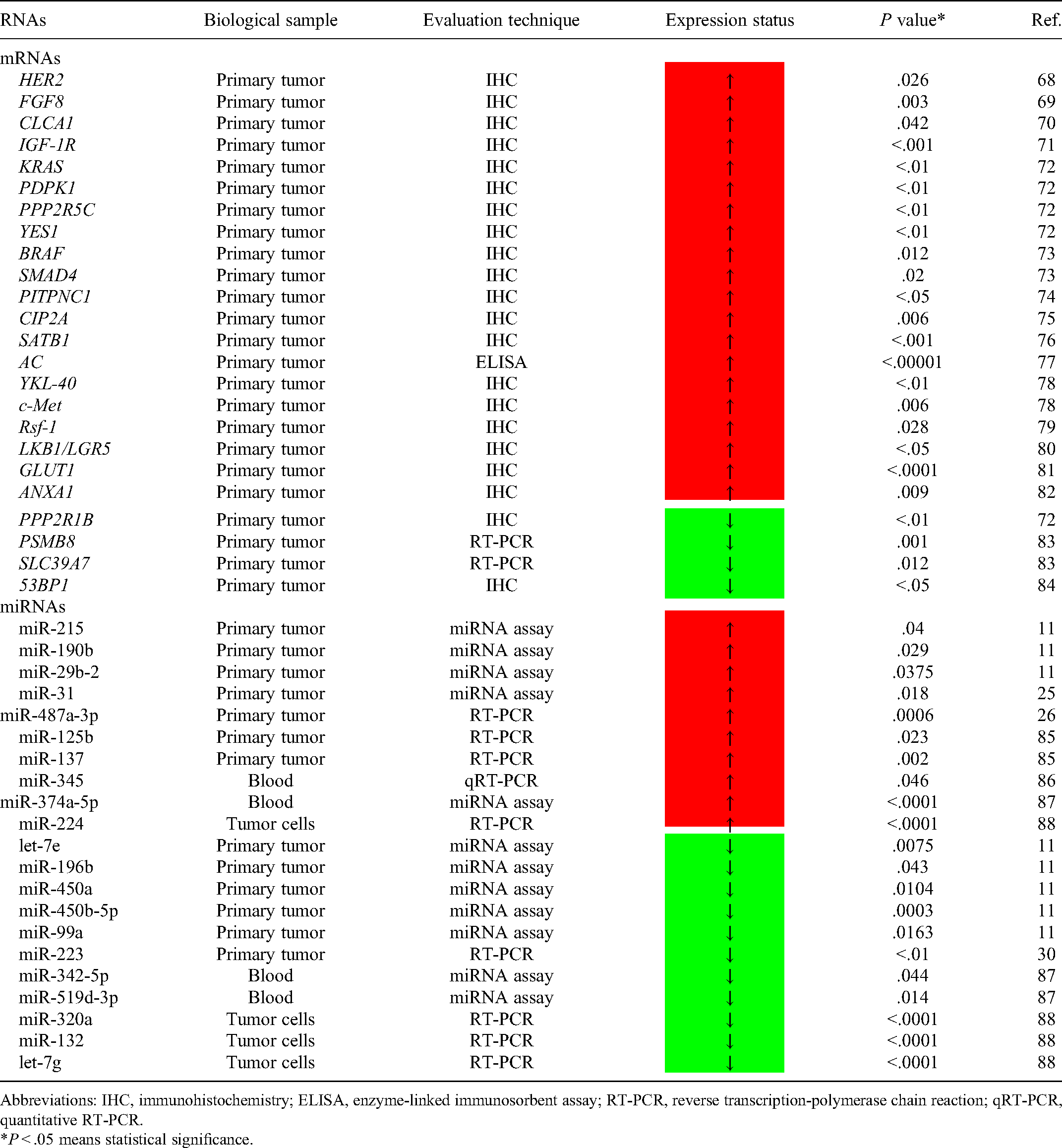
Abbreviations: IHC, immunohistochemistry; ELISA, enzyme-linked immunosorbent assay; RT-PCR, reverse transcription-polymerase chain reaction; qRT-PCR, quantitative RT-PCR.
*P < .05 means statistical significance.
nCRT Resistance/Sensitivity-Related miRNAs
Dysregulated miRNA-Induced nCRT Resistance/Sensitivity Against Downstream Targeted mRNAs
Several studies have only evaluated the association between dysregulated miRNAs and the abnormal expression of downstream targeted mRNAs and CRT resistance in rectal cancers. As previously reviewed, XRCC3 mRNA encodes one of the kernel proteins in the HR pathway; the expression level of XRCC3 mRNA in rectal cancers can be increased by the downregulation of miR-185, 89 and CRT resistance is thereby enhanced. Rectal cancers treated with CRT overexpress miR-95, which suppresses its downstream targeted sphingolipid phosphatase SGPP1 mRNA, resulting in weakened tumor necrosis and reinforced cell proliferation. 90 These results highlight the core role of miR-95 in nCRT resistance in LARC patients.
In contrast, some studies have revealed that overexpressed miRNAs increase the CRT sensitivity of rectal tumors by regulating their downstream targeted mRNAs. miR-205 suppresses its downstream targets ZEB1 mRNA and Ubc13 mRNA to blockade the DDR process. 91 miR-130a hampers the DDR process by directly targeting SOX4 mRNA and dramatically reverses the EMT phenotype of rectal cancer cells. 92 Consequently, there is a likelihood that miR-205/miR-130a may function as a CRT sensitizer in the treatment of rectal cancers and be utilized as a potential therapeutic target to improve post-CRT prognosis. miR-451a is expressed at significantly higher levels in the nCRT-sensitive cohort of LARC patients than in the nCRT-resistant cohort of those patients, whereas its downstream targets CAB39 mRNA and EMSY mRNA tend to be expressed at lower levels, 93 suggesting that miR-451a-induced nCRT sensitivity is based on the suppression of CAB39 mRNA and EMSY mRNA. The overexpression of ARID4B mRNA is reported to be positively correlated with tumorigenesis, invasion, and metastasis of breast cancers. 94 miR-519b-3p directly bonds to the 3′-UTR of ARID4B mRNA, giving rise to ARID4B mRNA underexpression that improves CRT sensitivity; 28 thus, the upregulation of miR-519b-3p has an excellent predictive performance in identifying nCRT sensitivity in LARC patients (accuracy = 92%; Table 1). An effective prognostic performance in defining the nCRT sensitivity of LARC patients is also exhibited by miR-21-5p (accuracy = 89%; Table 1), which increases tumorous nCRT sensitivity predominantly by targeting ASTB1 mRNA. 27 Overexpressed miR-194 indirectly curbs the Wnt/β-catenin signaling pathway by suppressing the expression of its downstream target TRAF6 mRNA, leading to elevated nCRT sensitivity in LARC patients. 95
Although the abovementioned articles report that many miRNAs targeting their downstream targeted mRNAs are closely related to the varied nCRT resistance of LARC patients, there is less understanding of the upstream targets of these miRNAs, that is, lncRNAs and circRNAs. This presents challenges in biologically constructing nCRT resistance-related ceRNA networks in LARC patients and increases the difficulty of developing inhibitors against potential biomarkers.
Direct Evidence of Dysregulated miRNA-Induced nCRT Resistance/Sensitivity
Several studies have only investigated the expression difference of miRNAs between the nCRT-resistant cohort and the nCRT-sensitive cohort. The overexpression of miR-21 (accuracy = 83%), miR-31 (accuracy = 72%), and miR-487a-3p (accuracy = 64%) showed good to effective performance in predicting nCRT resistance in LARC patients,24-26 while the overexpression of miR-1246 (accuracy = 70%), miR-1290-3p (accuracy=74%), miR-205-5p (accuracy = 63%), miR-200c (accuracy = 79%), and miR-223 (accuracy = 90%) revealed good to excellent predictive performance in ascertaining the nCRT sensitivity of LARC patients (Table 1).27,29,30 Compared to the nCRT-sensitive cohort, many phenotypes of miRNAs are upregulated11,25,26,85-88 or downregulated11,30,87,88 in tumors or biological fluids (ie, blood, saliva, and urine) of the nCRT-resistant cohort (Table 2), and these dysregulated miRNAs in turn could be used to discern nCRT sensitivity or nCRT resistance.
nCRT Resistance/Sensitivity-Related ceRNAs
Competing Endogenous RNAs Networks
The classifications of ncRNAs include short ncRNAs (sncRNAs), long ncRNAs (lncRNAs), and circRNAs; sncRNAs are further divided into Piwi-interacting RNAs, small interfering RNAs, tRNAs, rRNAs, snoRNAs, and microRNAs (miRNAs).96,97 miRNAs are highly conserved, with lengths of 18 to 25 nucleotides, and regulate gene expression at the posttranscriptional level by degrading and/or translationally repressing their downstream targeted mRNAs. 98 A recent study suggested that miRNAs mediate the expression level of many oncogenes, anti-oncogenes, and CRT resistance-related genes. 99 lncRNAs, endogenous ncRNAs with a length over 200 nucleotides, frequently do not encode proteins and have the following major biological contributions: transcriptional regulation, encoding modulation, and organization of nuclear domains.100,101 Studies have revealed that the dysregulation of lncRNAs is associated with carcinogenesis and CRT resistance. 102 circRNAs are endogenous ncRNAs lacking 5′- and 3′-untranslated regions (UTRs) which are produced by backsplicing of precursor mRNAs. 103 They have a circular structure, evolve conservatively, and are highly stabilized, thereby showing inherent resistance to RNase but can exhibit abnormal expression when they are mediated in cancer progression.
By August 2011, Salmena et al 104 presented a “competing endogenous RNAs (ceRNAs)” hypothesis that communication across all types of RNA transcripts uses the letters of a new language, called miRNA recognition elements (MREs). This communicated information will ultimately be “heard” and translated by using an increasing body of updated experimental techniques. lncRNAs and circRNAs are affiliated with ceRNAs, mechanically called miRNA sponges that indirectly regulate miRNAs downstream of targeted mRNAs by sponging miRNAs and eventually influencing the occurrence, proliferation, and metastasis of cancer cells. 105 The ceRNA networks mainly comprise lncRNA-miRNA-mRNA networks and circRNA-miRNA-mRNA networks. Functionally, overexpression of oncogenic lncRNAs/circRNAs against the expression of tumor-suppressive miRNAs can promote tumorous cell proliferation and metastasis, and overexpression of tumor-suppressive lncRNAs/circRNAs restrains the expression of oncogenic miRNAs and thus inhibits carcinogenesis (Figure 1). 105
Techniques for Identifying lncRNAs and circRNAs
Previous techniques (eg, RNA microarray and mRNA-Seq) used to identify mRNA relied on poly(A) tails and tended to favor the detection of overexpressed transcripts; as such, they have challenges in identifying and analyzing lncRNAs due to their low expression level and poly(A) tail-free properties. In this context, many innovative techniques have been developed for this purpose (Table 3).106-114 Among them, RNA sequencing, especially transcriptome-wide RNA sequencing, is one of the most commonly used methods.
Techniques for Identifying lncRNAs and circRNAs.

Abbreviations: RNA-Seq, RNA sequencing; CaptureSeq, capture sequencing; RIP, RNA immunoprecipitation; ChIRP, chromatin isolation by RNA purification; FISH/ISH, fluorescence in situ hybridization; RNA-3C, RNA-chromosome conformation capture; ASO, antisense oligonucleotides; RPD, RNA pull-down assay; RPA, RNase protection assay; EMSA, electrophoretic mobility shift assay.
The special circular structure of circRNAs gives rise to significantly fewer enriched binding sites of RNA binding proteins than the corresponding linear mRNAs. 119 Therefore, historical nucleotide sequence-based approaches may not be suitable for identifying circRNAs and analyzing circRNA-protein interactions. RNA pull-down assays and RNA immunoprecipitation are the mainstays in detecting circRNAs and circRNA-protein interactions (Table 3).115,116 RNase protection assays, 117 fluorescence in situ hybridization techniques, 115 and electrophoretic mobility shift assays 118 are also used to determine, profile, and understand the biogenesis and functions of circRNAs.
Dysregulated lncRNA-Induced nCRT Resistance/Sensitivity by Sponging Downstream miRNAs
RNA microarray analysis has revealed the close correlation between some lncRNA-miRNA-mRNA networks and nCRT resistance in LARC patients. 72 The expression levels of 3 lncRNAs (ie, lncRNA-KLF7-1, lncRNA-MAB21L2-1, and LINC00324) in the nCRT-sensitive cohort of LARC patients outperformed those in the nCRT-resistant cohort of LARC patients; applying them as the variable subset to predict nCRT sensitivity showed excellent prognostic performance (accuracy = 90%; Table 1). 31 In rectal cancer tissues and cells, upregulating lncRNA-ROR can increase neoplastic CRT resistance by negatively regulating the activity of the p53/miR-145 pathway; conversely, gene knockdown of lncRNA-ROR decreases cell vitality and promotes apoptosis, creating a high CRT sensitivity within tumors. 120 Radiotherapy-activated lncRNA-OIP5-AS1 upregulates DYRK1A mRNA by suppressing miR-369-3p, where the remission of cell viability and the promotion of apoptosis also occur, and finally improves the CRT sensitivity of rectal cancer cells. 121 lncRNA-p21 is underexpressed in rectal cancer tissues and cells but can be overexpressed by radiotherapy. 122 Upregulated lncRNA-p21 leads to increased CRT sensitivity in colorectal cancers by blocking the Wnt/β-catenin signaling pathway and inducing the overexpression of the apoptotic gene Noxa. 122 The overexpression of lncRNA-EGOT in rectal cancers induces ErbB4 mRNA expression by targeting miR-211-5p, therefore hampering apoptosis and facilitating cell proliferation, which are both critical to increased CRT resistance. 123 Unfortunately, there are few studies investigating the correlation between circRNA-miRNA-mRNA networks and nCRT resistance in LARC patients, which is a breakthrough worth anticipating.
Conclusions
To date, studies investigating the involvement of transcriptome RNAs (ie, mRNAs and ncRNAs) in variation-related molecular mechanisms for nCRT resistance/sensitivity in LARC patients are still limited. Specifically, dysregulation inherent in rectal cancer or CRT-induced mRNA dysregulation both influence multiple DDR pathways, thus either driving or curbing the DDR process, which plays a central role in changing nCRT resistance/sensitivity in LARC patients. The expression level of these mRNAs is regulated by their upstream targeted ncRNAs; as such, the ceRNA networks constituted by mRNAs plus ncRNAs appear to have a backbone role in affecting the DDR process and the resultant alteration of nCRT resistance/sensitivity. The presently acknowledged lncRNA-miRNA-mRNA networks were identified by RNA microarray analysis on known RNA sequences, so they are confined and imperfect. Additionally, circRNA-miRNA-mRNA networks warrant further investigation since it is still unclear how they impact nCRT resistance/sensitivity in LARC patients. In order to shed more light on the correlation between ceRNA networks and nCRT resistance in LARC patients, it may be worth applying a myriad of novel biological and experimental technologies.
Search Strategy and Inclusion Criteria
Articles published in English were searched in the PubMed database using the search terms (radiotherapy) OR (radiation therapy) OR (radiosensitive) OR (radiosensitivity) OR (radioresistant) OR (radioresistance) AND (rectal OR rectum) AND (cancer OR tumor OR tumor OR carcinoma OR neoplasm) AND (RNA OR gene). The publications were retrieved on May 28, 2021. Clinical studies that assessed the dysregulation of mRNAs/ncRNAs affecting nCRT resistance/sensitivity in local advanced rectal cancer patients or experimental articles that investigated the association between dysregulated mRNAs/ncRNAs and CRT resistance/sensitivity in rectal cancer cells met the inclusion criteria.
Statistical Methods
All articles present sensitivity, specificity, and the number of sensitive cases and resistant cases for calculating the prognostic performance of biomarkers (ie, overexpressed ceRNAs, miRNAs, and mRNAs). A 4 × 4 contingency table with affiliated formulas (shown in Table 4) was constructed to compute the accuracy of biomarkers in the prognosis of neoadjuvant chemoradiotherapy resistance in locally advanced rectal cancer.
A 4 × 4 Contingency Table With Affiliated Formulas.

Sensitivity = A/(A + C); Specificity = D/(B + D); Positive predictive value = A/(A + B); Negative predictive value = D/(C + D); Accuracy = (A + D)/(A + B + C + D).
Footnotes
Abbreviations
Authors’ Contributions
LH: writing manuscript, table and figure drawing; HC: writing manuscript; YQ: writing manuscript, supervision; BZ: writing manuscript, data collection; QS: conception/design; final approval of manuscript. All authors reviewed and approved the manuscript prior to submission.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
