Abstract
Background
Esophageal cancer is a severe malignant disorder originating in the esophagus, and the sixth most common cause of cancer-related death worldwide. Patients with esophageal cancer have a low 5-year survival rate, and the disease has an incidence that significantly differs with geographical location.1,2 Patients with esophageal cancer usually experience difficulty in swallowing, chest pain, weight loss, worsening indigestion, coughing, and other severe symptoms.3,4 Recent epidemiological investigations have revealed that esophageal cancer development is closely associated with multiple risk factors such as genetic mutations, smoking, alcohol abuse, obesity, frequent drinking of hot beverages, radiation, as well as pathogenic conditions including gastroesophageal reflux disease, Barrett's esophagus, and achalasia.1,5,6 The pathogenesis of esophageal cancer is mediated by an accumulation of oncogene activation and tumor suppressor gene inactivation, which results in increased cell proliferation, decreased apoptosis, as well as increased cell migration capacity and angiogenesis, which contribute to esophageal cancer metastasis.1,7 While chemotherapeutic agents such as cisplatin (cis-diaminodichloroplatin [DDP]) have been widely used as first-line therapy in treating esophageal cancer, the fast-developing resistance to available chemotherapeutic agents presents a major clinical problem. 8 A more thorough understanding of the molecular mechanisms that control angiogenesis and chemotherapy resistance would facilitate the prevention and management of esophageal cancer.
ATP7A (ATPase copper transporting alpha), alternatively known as MNK (Menkes protein), is one member of the heavy metal transporting P-type ATPase family of proteins that is characterized by having multiple metal binding domains in the N-terminal region. 9 It is well documented that similar to other P-type ATPase members, ATP7A can function as a major transporter capable of transferring copper ions out of mammalian cells by binding with copper ions and initiating copper-induced intracellular trafficking.9,10 Under normal physiological conditions, the ATP7A gene is widely expressed in the trans-Golgi network of various tissue types. As cellular copper content increases, ATP7A is transferred into cytoplasmic vesicles that subsequently fuse with the plasma membrane to allow an efflux of copper ions needed to maintain copper homeostasis.9,11 A mutation in the ATP7A gene and resultant dysregulation of copper homeostasis can result in various metabolic disorders such as Menkes disease, which is characterized by a general copper deficiency due to inadequate transportation of copper ions from enterocytes into the blood system.9,12 Moreover, ATP7A has been implicated in cancer pathogenesis due to its ability to promote cancer cell invasiveness by providing sufficient copper ions for oncogenic enzymes. 10 Importantly, ATP7A proteins have also been identified as major contributors to cancer cell resistance to platinum (Pt)-based chemotherapy drugs such as cisplatin.9,10 For instance, ATP7A was found to be highly expressed in DDP-resistant esophageal squamous cell cancer (ESCC) cells, and capable of enhancing ESCC cell resistance to cisplatin treatment. 13 However, little is currently known about the mechanisms which control changes in ATP7A expression that are associated with cisplatin resistance.
Epigenetic regulation mediated by noncoding RNAs has been established as an essential mechanism for altering the expression of genes involved in various biological and pathogenic processes. 14 MiRNAs comprise a large family of small single-stranded noncoding RNAs that commonly contain 22 nucleotides. 15 Extensive research in past decades has shown that microRNAs are expressed in various human cell types and help to regulate distinct biological events by modulating gene expression.15,16 MicroRNAs inhibit gene expression mainly by preventing translation by base-paring with the 3′-UTR (3′-untranslated region) of their target genes or causing mRNA destabilization and degradation.15-17 Due to their potent effects on cell cycle progression, differentiation, apoptosis, migration, inflammation, angiogenesis, and other tumorigenesis-related cellular processes, microRNAs have been established as critical regulators of cancer development and progression in humans.17-19 For instance, the proliferation and migration capabilities of esophageal cancer cells were shown to be repressed by microRNA-124 by its ability to inhibit expression of the NRP1 (Neuropilin 1) gene. 20 Moreover, another microRNA (miR-495) was found to enhance the susceptibility of gastric and ovarian cancer cells to doxorubicin and Taxol treatments by inhibiting MDR1 (multiple drug resistance 1) gene expression. 21 However, the roles played by miR-495 in the development of chemotherapy resistance in esophageal cancer cells remain largely unknown.
In the present study, we investigated how miR-495 might mediate cisplatin resistance and angiogenesis in esophageal cancer cells, and whether it might also regulate ATP7A expression. Our results provide new insights into the molecular mechanisms by which noncoding RNAs mediate chemotherapy resistance in esophageal cancer, and could serve as a basis for reversing cancer drug resistance by targeting microRNA expression.
Materials and Methods
Clinical Sample Collection and Cell Culture
Tissues collected between January 2017 and December 2018 from a total of 20 patients who were diagnosed with esophageal squamous cancer and scheduled to undergone a surgical resection operation in the Department of Medical Oncology, Affiliated Dongguan People's Hospital, Southern Medical University (Dongguan, China) were enrolled in this study. The study protocol was preapproved by the Ethics Committee of the Affiliated Dongguan People's Hospital, Southern Medical University, and each patient provided their written consent for study participation. The esophageal cancer tissues and corresponding adjacent noncancerous esophageal tissues (3 cm from site of tumor) were surgically collected at the same time and then stored in liquid nitrogen for use in further studies. The human esophageal cancer cell lines Eca-109 and TE1, as well as human umbilical vein endothelial cells, were obtained from the Cell Bank of the Typical Culture Preservation Committee, affiliated with the Chinese Academy of Science (Shanghai, China). The esophageal cancer cells were cultured in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% fetal bovine serum (Thermo Fisher Scientific) and 1% penicillin/streptomycin (Sigma Aldrich) in a 37 °C humidified chamber containing 5% CO2.
Cell Transfection and Resistance Induction
MiR-495 mimics were synthesized by GenePharma Company. The CDS sequences of the ATP7A gene were amplified by real-time polymerase chain reaction (RT-PCR) and then ligated onto a pcDNA 3.0 plasmid. Next, the sequences and recombinant plasmids were transfected into esophageal cancer cells by using Lipofectamine 3000 reagent (Thermo Fisher Scientific) according to the manufacturer's instructions. Eca-109 + DDP and TE1 + DDP cell lines with significant resistance to cisplatin (DDP) were established by incubating the cells with increasing drug concentrations. Briefly, Eca-109 and TE1 cells in logarithmic growth phase were cultured in DMEM containing 0.02 μg/mL DDP (Sigma Aldrich) for 24 h; after which, they were cultured in normal medium until complete recovery of cell growth. Subsequently, the cells were cultured in DMEM containing slightly higher DDP concentrations for 24 h, until the cells could stably grow in DMEM containing 1 μg/mL DDP.
Quantitative RT-PCR
Total RNA was extracted from esophageal cancer tissues or cultured esophageal cancer cells using an EasyPure® RNA Kit (#ER101-01; Transgen Biotech) according to the manufacturer's instructions. RNA concentrations were measured with a NanoDrop microvolume spectrophotometer. Next, ∼ 2 μg of RNA from each sample was reverse transcribed into cDNA using a Fast Quant RT Super Mix kit (#KR108; Tiangen) as instructed by the manufacturer. The levels of target mRNA or microRNA were subsequently detected by quantitative polymerase chain reaction (qPCR) assays that were performed using a TransStart Green qPCR Super Mix kit (#AQ101-01; Transgen) according to the manufacturer's instructions. Finally, the relative levels of mRNA or microRNA expression were calculated by the 2−△△Ct method with GAPDH or U6 serving internal standards for mRNA and microRNA, respectively. The primers sequences used in the study are shown in Table 1.
Primer Pairs Used for Quantitative Real-Time Polymerase Chain Reaction (RT-PCR) Analysis.

Western Blotting
The total proteins from esophageal cancer tissues or cultured esophageal cancer cells were extracted using a Total Protein Extraction kit (#C006225; Sangon Biotech) according to the manufacturer's instructions. Next, ∼25 μg of protein from each extract was added to a protein loading buffer and boiled at 100 °C for 5 min. The protein samples were then separated by 12% SDS-PAGE (sodium dodecyl sulphate–polyacrylamide gel electrophoresis; Sangon Biotech), and the protein bands were transferred onto PVDF (polyvinylidene fluoride) membranes, which were subsequently blocked with 5% lipid-free milk solution. The membranes were then incubated overnight with diluted primary antibodies at 4 °C, washed 3 times with TBST, incubated for 1 to 2 h with diluted secondary antibodies at room temperature, and finally developed with ECL Western Blotting Substrates (#32109; Thermo Fisher Scientific) as instructed by the manufacturer. The primary antibodies used for western blotting were anti-ATP7A (#ab13995; Abcam), anti-MRP1 (#14685; CST), anti-MRP1 (#14685; Abcam), anti-ABCG1 (#ab218528; Abcam), anti-ABCA1 (#ab18180; Abcam), anti-VEGF (#2463; CST), anti-Caspase 3 (#ab2302; Abcam), and anti-GAPDH (#5174; CST).
Cell Proliferation and IC50 Determination
The proliferation rates of esophageal cancer cells were measured by using a Cell Counting Kit 8 (WST-8) (#ab228554) according to the manufacturer's instructions. Briefly, cultured cancer cells in logarithmic growth phase (3000 cells/well) were seeded in a 96-well plate (Corning) and then cultured under normal conditions another 24 h; after which, they were mixed with 10 µL WST-8 solution and incubated under normal conditions for yet another 2 to 3 h at 37 °C. The proliferation rates of cancer cells were finally determined by measuring their absorbance at 450 nm (OD450) with a microplate reader. The IC50 (half-inhibitory concentration) values for DDP in esophageal cancer cells were determined by the CCK-8 method after the cancer cells had been treated for 48 h with a DDP concentration gradient ranging from 1 to 100 mM.
Cell Apoptosis
The apoptosis of esophageal cancer cells was measured by flow cytometry using a Dead Cell Apoptosis Kit containing Annexin-V FITC and PI (#V13242; Thermo Fisher Scientific) as instructed by the manufacturer. Briefly, cultured esophageal cancer cells were harvested by centrifugation, washed with phosphate buffered saline (PBS) solution, and then incubated with 5 μL of FITC Annexin V and 1 μL of 100 μg/mL PI working solution for 13 min at room temperature. The percentages of apoptotic esophageal cancer cells were determined by flow cytometry at fluorescence emissions of 530 and 590 nm.
Enzyme linked immunosorbent assay
The levels of TNF-α (tumor necrosis factor alpha) and VEGF (vascular endothelial growth factor) in the culture medium of esophageal cancer cells were measured by enzyme linked immunosorbent assays (ELISAs) that were performed using a Human TNF-α ELISA Kit (#PT518; Beyotime Biotech) and Human VEGF ELISA Kit (#PV963; Beyotime Biotech), respectively, according to the manufacturer's instructions.
Tube Formation Assay
The angiogenic capability of human umbilical vein endothelial cells (HUVECs) cells was evaluated using the tube formation assay. Briefly, after being cocultured with esophageal cancer cells as designated, the HUVEC cells in complete culture medium were seeded into the wells of 24-well plates (Corning) which had been precoated with 0.1 mL of Matrigel Matrix (standard formulation; Corning). The formation of capillary-like structures was observed and photographed under a light microscope under normal conditions for 24 h at 37 °C.
Immunofluorescence
The in-situ expression of NLRP3 (Nod-like receptor protein 3) proteins in esophageal cancer cells was analyzed by immunofluorescence (IF). Briefly, slides with esophageal cancer cells were first treated with 4% paraformaldehyde solution for 12 min at room temperature, and then washed 3 times (5 min per wash) with prechilled PBS solution. The slides were then blocked with 10% bovine serum albumin (BSA) solution for 25 min at room temperature and subsequently incubated with primary antibodies against NLRP3 (#ab4207; Abcam) and anti-goat IgG antibodies conjugated with FITC (#ab47846; Abcam). After being counterstained with 4′,6-diamidino-2-phenylindole (DAPI), the esophageal cancer cells were observed using a confocal microscope.
Dual Luciferase Reporter Assay
The binding of miR-495 to ATP7A gene sequences in both Eca-109 and TE1 cells was verified by dual luciferase assays that were conducted using a pmirGLO vector combined with the Dual-Luciferase Reporter NanoDLR Assay (Promega) according to the manufacturer's instructions. Briefly, Eca-109 and TE1 cells expressing the wide type (WT) or mutant (MUT) sequences of the ATP7A gene were first established and then simultaneously transfected with miR-495 mimics or the negative control sequences, respectively. The relative luciferase activities of esophageal cancer cells were finally measured using a GloMax 20/20 Luminometer (Promega).
Statistics
Quantitative results are based on data obtained from at least 3 replicate experiments and are presented as a mean value ± standard deviation. Differences between two groups were analyzed by Student's t-test, and one-way analysis of variance was used for differences analysis between three or more groups. A P-value < .05 was regarded as being statistically significant.
Results
Elevated ATP7A Expression and Decreased miR-495 Expression Were Detected in Esophageal Cancer Patients
To study the potential pathogenic involvement of miR-495 and ATP7A expression in esophageal cancer, we first detected the expression of both miR-495 and ATP7A in cancerous tissues collected from 20 esophageal cancer patients. Clinicopathological characteristics of miR-495 to 3p and ATP7A in esophageal cancer patients were shown in Table 2. There was significant difference between tumor size and expression of ATP7A, lymph node metastasis, and expression of miR-495 to 3p (Table 2). Quantitative RT-PCR studies showed that the levels of ATP7A mRNA in the esophageal cancer tissues were significantly higher than those in the corresponding adjacent noncancerous esophageal tissues (Figure 1A). In contrast, we also found that the levels of miR-495 expression in the cancerous tissues of esophageal cancer patients were significantly lower than those in the adjacent esophageal tissues (Figure 1B). The great enhancement of ATP7A expression and decrease in miR-495 expression in esophageal cancer tissues were also verified by subsequent western blotting assays (Figure 1C). Further analysis showed the expression of ATP7A and miR-495 to 3p presented negative correlations (Figure 1D). Those remarkable and opposite alterations of miR-495 and ATP7A expression in cancerous tissues from esophageal cancer patients suggest their potential roles in esophageal cancer development and chemotherapy resistance.

Changes in ATP7A and miR-495 expression in esophageal cancer patients. (A) The elevated levels of ATP7A expression in cancerous tissues from 20 esophageal cancer patients. ATP7A mRNA levels in cancerous and noncancerous esophageal tissues were analyzed by quantitative RT-PCR. (B) Decreased miR-495 expression in esophageal cancer tissues from 20 esophageal cancer patients. MiR-495 expression was analyzed by quantitative RT-PCR. (C) The levels of ATP7A protein in both cancerous and adjacent noncancerous tissues collected from esophageal cancer patients. The ATP7A protein levels in esophageal tissues were detected by western blotting. (D) Correlations between the expression of ATP7A and miR-495 to 3p.
Clinicopathological Characteristics of miR-495 to 3p and ATP7A in Esophageal Cancer Patients.

*Statistical significance (P < 0.05).
MiR-495 Repressed Cisplatin Resistance in Esophageal Cancer Cells
To investigate the roles of miR-495 in modulating the chemotherapy resistance of esophageal cancer, we created cisplatin-resistant Eca-109 and TE1 cells as described in the Material and Methods section. When compared with the cisplatin-sensitive cells, both the cisplatin-resistant Eca-109 and TE1 cells exhibited greatly decreased levels of miR-495 expression (Figure 2A). To further confirm the function of miR-495, we forced the overexpression of miR-495 in both Eca-109 and TE1 cells with resistance to cisplatin and subsequently verified that such overexpression was present in both cell lines (Figure 2B). Subsequent CCK-8 assays showed that the IC50 values of cisplatin in the Eca-109 and TE1 cells with miR-495 overexpression were remarkably elevated when compared with the negative controls (Figure 2C). Moreover, we found that the proliferation rates of cisplatin-resistant Eca-109 and TE1 cells were significantly repressed by miR-495 overexpression, when compared with the negative controls (Figure 2D). In contrast, the apoptosis rates of cisplatin-resistant Eca-109 and TE1 cells were greatly increased by overexpression of miR-495, when compared with the negative control cells (Figure 2E). Those results showed that miR-495 serves as a negative regulator of resistance to cisplatin treatment in esophageal cancer cells.
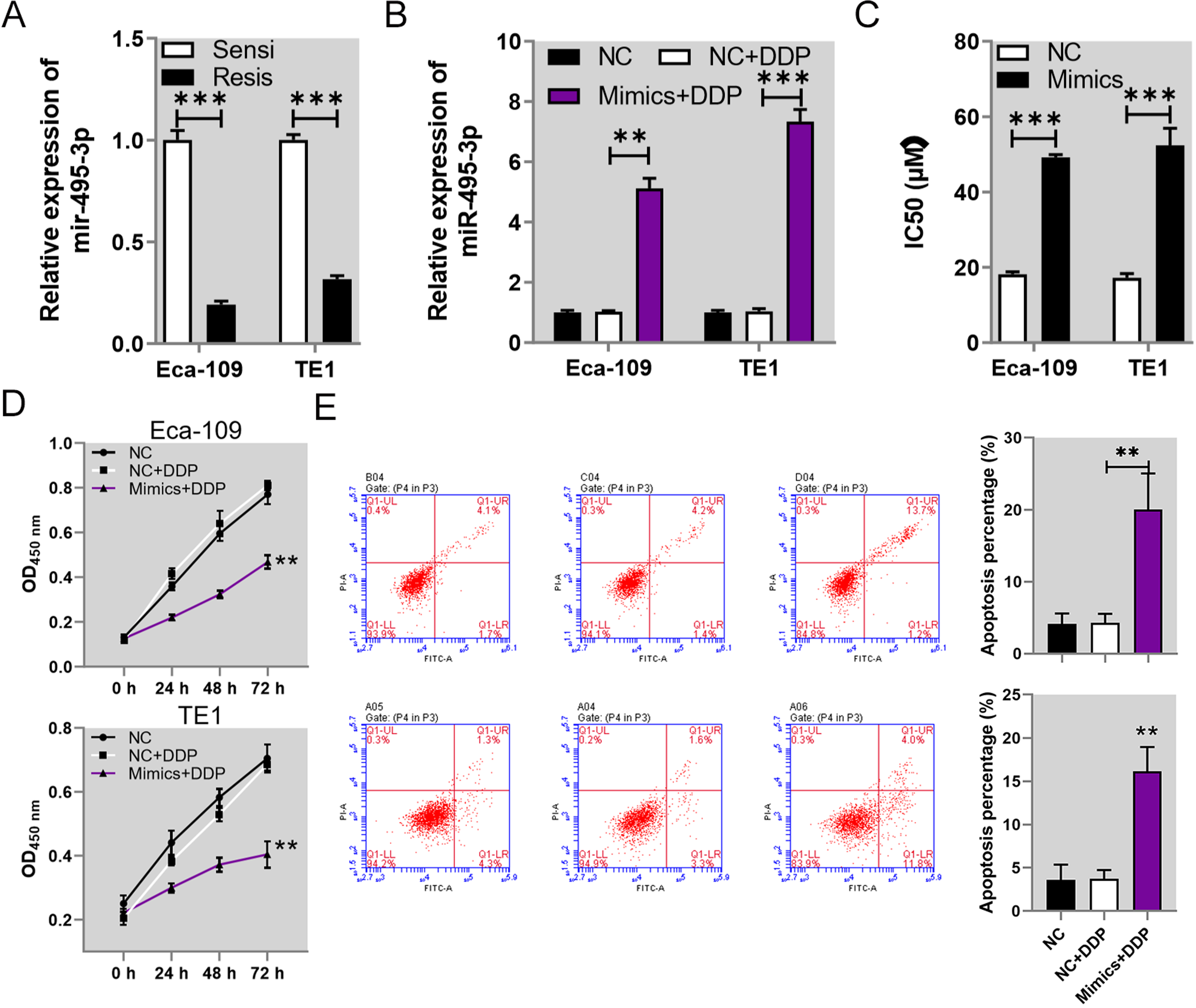
Suppression of cisplatin resistance in esophageal cancer cells by miR-495. (A) The relative levels of miR-495 expression in cisplatin-sensitive and -resistant esophageal cancer cells. The levels of miR-495 expression in Eca-109 and TE1 cells were evaluated by quantitative RT-PCR. (B) The increases in miR-495 expression in Eca-109 and TE1 cells transfected with specific mimics. (C) The IC50 values of cisplatin in both Eca-109 and TE1 cells that overexpressed miR-495. The CCK-8 method was used to determine IC50 values in esophageal cancer cells. (D) The proliferation rates of cisplatin-resistant Eca-109 and TE1 cells with miR-495 overexpression. Cell proliferation was measured using the CCK-8 assay. (E) The increased apoptosis rates of cisplatin-resistant Eca-109 and TE1 cells that overexpressed miR-495. The apoptosis of esophageal cancer cells was detected by flow cytometry.
MiR-495 Inhibited the Angiogenic Capability of Cisplatin-Resistant Esophageal Cancer Cells
To gain additional insight into the mechanisms by which miR-495 regulates cisplatin resistance, we examined how miR-495 influences the angiogenesis of esophageal cancer cells. ELISA studies showed that the levels of 2 angiogenic factors (TNF-α and VEGF) found in the culture media of cisplatin-resistant Eca-109 and TE1 cells were greatly downregulated by overexpression of miR-495 (Figure 3A and B). Results from tube formation assays and studies in which esophageal cancer cells were cocultured with HUVECs revealed that miR-495 overexpression significantly impaired the angiogenic capabilities of both cisplatin-resistant Eca-109 and TE1 cells (Figure 3B). Moreover, western blot studies showed that the levels of MRP1 (multidrug resistance protein 1), ABCG1 (ATP-binding cassette transporter-G1), ABCA1 (ATP-binding Cassette Transporter A1), and VEGF proteins in cisplatin-resistant Eca-109 and TE1 cells were substantially reduced by miR-495 overexpression, while C-Cas 3 (cleaved Caspase-3) expression was significantly increased (Figure 3D). We also found that the levels of NLRP3 (NLR family, pyrin domain-containing 3) protein expression in the cisplatin-resistant Eca-109 and TE1 cells were markedly downregulated by miR-495 overexpression (Figure 3E). These results indicated that miR-495 could effectively repress angiogenesis in esophageal cancer cells with cisplatin resistance.

Inhibition of cisplatin resistance in esophageal cancer cells by miR-495. (A,B) The relative levels of angiogenic factors in the culture media of cisplatin-resistant Eca-109 and TE1 cells that overexpressed miR-495. TNF-α (A) and VEGF (B) levels in cell culture media were measured by ELISA methods. (C) The impaired angiogenic capability of HUVECs cocultured with cisplatin-resistant esophageal cancer cells that overexpressed miR-495. HUVEC angiogenesis was detected by the tube formation assay. (D) The relative levels of MRP1, ABCG1, ABCA1, VEGF, and C-Cas3 proteins in cisplatin-resistant esophageal cancer cells that overexpressed miR-495. (E) In-situ expression of NLRP3 protein in cisplatin-resistant esophageal cancer cells that overexpressed miR-495. NLRP3 expression in esophageal cancer cells was determined by immunofluorescence. Abbreviations: TNF-α, tumor necrosis factor alpha; VEGF, vascular endothelial growth factor; NC, negative control; DDP, cisplatin; MRP1, multidrug resistance protein 1; ABCG1, ATP-binding cassette transporter-G1; ABCA1, ATP-binding Cassette Transporter A1; C-Cas 3, cleaved Caspase-3; GAPDH, glyceraldehyde-3-phosphate dehydrogenase; NLRP3, NLR family, pyrin domain-containing 3; RT-PCR, real-time polymerase chain reaction; **P < .01; ***P < .001.
MiR-495 Repressed ATP7A Expression in Esophageal Cancer Cells via Direct Binding
To explore the relationship between miR-495 and ATP7A expression, we tested the potential interaction between miR-495 and ATP7A gene sequences in esophageal cancer cells. MiR-495 expression in the Eca-109 and TE1 cell lines was greatly increased by transfection with the miR-495 mimics when compared with its expression in the negative control groups (Figure 4A). Western blot results showed that overexpression of miR-495 resulted in a significant downregulation of ATP7A protein levels in both the Eca-109 and TE1 cell lines (Figure 4B). Subsequently, our dual luciferase reporter assays showed that miR-495 mimics induced remarkable decreases in luciferase activity in the Eca-109 and TE1 cell lines expressing WT ATP7A gene sequences, but not in those cells expressing the mutant (MUT) versions of ATP7A gene sequences (Figure 4C). These assays showed that miR-495 could directly bind with the ATP7A gene sequences to repress ATP7A expression in esophageal cancer cells.

Suppression of ATP7A expression by miR-495 binding in esophageal cancer cells. (A) The relative levels of miR-495 expression in esophageal cancer cells transfected with miR-495 mimics. MicroRNA expression was detected by quantitative RT-PCR. (B) The suppression of ATP7A protein levels in esophageal cancer cells that overexpressed miR-495. ATP7A protein levels were detected by western blotting, with GAPDH serving as an internal standard. (C) The direct binding of miR-495 to ATP7A gene sequences in esophageal cancer cells. The direct binding of miR-495 to ATP7A sequences was verified by dual luciferase reporter assays. Abbreviations: NC, negative control; ATP7A, ATPase copper transporting alpha; GAPDH, glyceraldehyde-3-phosphate dehydrogenase; WT, wild type; MUT, mutant; RT-PCR, real-time polymerase chain reaction; **P < .01; ***P < .001.
MiR-495 Suppressed Cisplatin Resistance in Esophageal Cancer Cells by Targeting ATP7A Expression
To elucidate the roles played by ATP7A in mediating the inhibition of cisplatin resistance by miR-495, we next simultaneously overexpressed miR-495 and the ATP7A gene in cisplatin-resistant esophageal cancer cells. We first showed that the levels of miR-495 expression in cisplatin-resistant Eca-109 and TE1 cells were greatly increased by the miR-495 mimics, but not by ATP7A overexpression (Figure 5A). However, the ATP7A mRNA levels in cisplatin-resistant Eca-109 and TE1 cells were significantly repressed by the miR-495 mimics, but greatly increased by transfection with the ATP7A-overexpressing vectors (Figure 5B). Furthermore, ATP7A mRNA levels in the cisplatin-resistant Eca-109 and TE1 cells transfected with miR-495 mimics were greatly increased by ATP7A overexpression (Figure 5B). Consistent with that finding, the levels of ATP7A protein expression in the above cells showed changes that were similar to changes in the ATP7A mRNA levels (Figure 5C). Moreover, we observed that the proliferation rates of cisplatin-resistant Eca-109 and TE1 cells transfected with miR-495 mimics were greatly increased by overexpression of the ATP7A gene (Figure 5D). In contrast, the increases in apoptosis of cisplatin-resistant Eca-109 and TE1 cells induced by miR-495 mimics were significantly repressed by the combined overexpression of the ATP7A gene (Figure 5E). These results proved that the regulation of esophageal cancer cell cisplatin resistance by miR-495 was mediated by its suppression of ATP7A gene expression.

ATP7A mediated miR-495-regulated cisplatin resistance in esophageal cancer cells. (A,B) The relative levels of miR-495 and ATP7A expression in cisplatin-resistant esophageal cancer cells that overexpressed miR-495 and ATP7A. The levels of miR-495 (A) and ATP7A (B) expression were measured by qRT-PCR, with GPADH serving as an internal standard. (C) Changes in ATP7A protein levels in cisplatin-resistant esophageal cancer cells caused by overexpression of miR-495 and ATP7A. Relative protein levels were detected by western blotting, with GAPDH serving as an internal standard. (D) The enhanced proliferation of cisplatin-resistant esophageal cancer cells transfected with miR-495 mimics and overexpressing ATP7A. Esophageal cancer cell proliferation was evaluated by the CCK-8 assay. (E) The decreased percentages of apoptotic cisplatin-resistant esophageal cancer cells transfected with miR-495 mimics and overexpressing ATP7A. Flow cytometry was used to measure cell apoptosis. Abbreviations: NC, negative control; DDP, cisplatin; ATP7A, ATPase copper transporting alpha; GAPDH, glyceraldehyde-3-phosphate dehydrogenase; OD450, absorbance at 450 nm; qRT-PCR, quantitative real-time polymerase chain reaction; *P < .05; **P < .01; ***P < .001; #P < .05 and ##P < .01 (compared with the Mimics + DDP group).
ATP7A Overexpression Abrogated the miR-495-Induced Suppression of Angiogenesis in Esophageal Cancer Cells
To gain more insight into the effects of ATP7A on esophageal cancer cell functions regulated by miR-495, we examined how ATP7A overexpression influences the angiogenic capability of esophageal cancer cells. By coculturing esophageal cancer cells with HUVECs and then performing tube formation assays, we showed that ATP7A overexpression in both cisplatin-resistant Eca-109 and TE1 cells transfected with miR-495 mimics enhanced the tube formation by HUVECs, when compared with HUVECs cocultured with Eca-109 and TE1 cells transfected with miR-495 mimics alone (Figure 6A). Also, the reductions in levels of TNF-α and VEGF in the culture media of cisplatin-resistant Eca-109 and TE1 cells treated withmiR-495 mimics were significantly recovered by combined overexpression of miR-495 and ATP7A (Figure 6B). Similarly, the decreases in MRP1, ABCG1, ABCA1, and VEGF proteins and increase in C-Cas3 proteins in cisplatin-resistant Eca-109 and TE1 cells induced by miR-495 mimics were all significantly mitigated by overexpression of the ATP7A gene (Figure 6C). Moreover, the decreases in NLRP3 protein expression in cisplatin-resistant Eca-109 and TE1 cells transfected with miR-495 mimics were also remarkably recovered by the simultaneous overexpression of miR-495 and ATP7A (Figure 6D). When taken together, these results revealed that the inhibition of angiogenesis in esophageal cancer cells by miR-495 was mediated by its repression of ATP7A expression.

ATP7A mediated the miR-495-induced inhibition of angiogenesis in esophageal cancer cells. (A) Recovery of the angiogenic capabilities of HUVECs cocultured with cisplatin-resistant esophageal cancer cells overexpressing miR-495 as result of ATP7A overexpression. HUVEC angiogenesis was evaluated by the tube formation assay. (B,C) Levels of angiogenic factors in the culture media of cisplatin-resistant Eca-109 and TE1 cells overexpressing miR-495 and the ATP7A gene. ELISA methods were used to analyze the TNF-α (B) and VEGF (C) levels in cell culture media. (D) The levels of MRP1, ABCG1, ABCA1, VEGF, and C-Cas3 proteins in cisplatin-resistant esophageal cancer cells as altered by miR-495 and ATP7A overexpression. (E) The recovered expression of NLRP3 protein in cisplatin-resistant esophageal cancer cells transfected with miR-495 mimics and overexpressing ATP7A. NLRP3 protein expression in esophageal cancer cells was measured by immunofluorescence. Abbreviations: NC, negative control; ATP7A, ATPase copper transporting alpha; DDP, cisplatin; TNF-α, tumor necrosis factor alpha; VEGF, vascular endothelial growth factor; MRP1, multidrug resistance protein 1; ABCG1, ATP-binding cassette transporter-G1; ABCA1, ATP-binding Cassette Transporter A1; C-Cas 3, cleaved Caspase-3; GAPDH, glyceraldehyde-3-phosphate dehydrogenase; NLRP3, NLR family, pyrin domain-containing 3; ELISA, enzyme linked immunosorbent assay; *P < .05; **P < .01.
Discussion
Chemotherapy agents such as cisplatin (DDP) and other platinum drugs have been widely used as the main anticancer drugs for treating solid tumors, including esophageal cancer; however, their efficacy is frequently impaired by the development of drug resistance in cancer cells.8,9 Reductions in drug accumulation caused by alterations in drug transport systems substantially contribute to the insensitivity of cancer cells to platinum-based chemotherapies,9,12 and the underlying mechanisms for that resistance remain poorly understood. In the present study, we found that microRNA miR-495, whose expression was significantly repressed in esophageal cancer, served as an essential inhibitor of cisplatin resistance and angiogenesis in esophageal cancer. We also found that the regulation of esophageal cancer cell resistance to cisplatin treatment was mediated by its repression of 1 copper-translocating P-type ATPase (ATP7A) that could directly bind to ATP7A gene sequences. Overexpression of the ATP7A gene effectively mitigated the inhibition of cisplatin resistance and angiogenesis in esophageal cancer cells transfected miR-495. These investigations provide novel insights into the development of cisplatin resistance that is mediated by epigenetic alterations based on noncoding RNAs and the downstream regulation of transport systems.
Due to their potent regulation of gene expression at the posttranscriptional level, microRNAs have been established as critical players in the induction of cancer resistance to major chemotherapies.22,23 For instance, the sensitivity of many types of cancer cells to 5-fluorouracil and docetaxel treatments can be greatly altered by several microRNAs such as miR-21, miR-98, miR-155, and miR-194. 22 Cisplatin has long been used as a first-line chemotherapeutic agent for treating solid cancers. In has been reported that the efficacies of other chemotherapeutic agents are also severely impaired when treating tumors with cisplatin resistance associated with microRNA-induced epigenetic regulation of gene expression.24,25 Moreover, the cisplatin resistance of esophageal cancer cells may also accompanied by substantial changes in microRNA profiles, which have been suggested as potential biomarkers for predicting chemotherapeutic efficacy and toxicity in esophageal cancer patients. 26 In the present study, we showed that miR-495 expression was greatly downregulated in esophageal cancer tissues, and its overexpression resulted in the effective suppression of cisplatin resistance and angiogenesis in esophageal cancer cells, as evidenced by decreased cell proliferation and increases in apoptosis and Caspase 3 cleavage. These findings provide direct evidence for the ability of miR-495 to suppress cisplatin resistance in esophageal cancer cells, and are also consistent with previous reports showing the involvement of miR-495 in chemotherapy resistance in other cancer types such as gastric cancer and nonsmall cell lung cancer.27,28 The prevalent implication of miR-495 in chemotherapy resistance suggests that targeting of miR-495 might be a promising strategy for recovering cancer chemo-sensitivity.
ATP-binding cassette (ABC) transporters play essential roles in the development of resistance to various chemotherapeutic agents.29,30 ATP7A is one member of a heavy metal transporter family responsible for transporting copper ions and platinum-based drugs across cell membranes.9,10 Accumulating evidence indicates that elevated levels of ATP7A expression are correlated with poor clinical outcomes for cancer patients undergoing platinum-based therapy. 9 While a high level of ATP7A gene expression contributes to cisplatin resistance in esophageal cancer, 13 the underlying mechanisms for development of that resistance remain poorly understood. In this study, we found that ATP7A expression could be suppressed by the binding of miR-495 to ATP7A gene sequences in esophageal cancer cells, and that ATP7A overexpression could partially abrogate that suppressed proliferation and promote apoptosis and Caspase 3 cleavage in cisplatin-resistant esophageal cancer cells treated with miR-495 mimics. These findings increase our understanding of alterations that occur in a transporter system associated with cisplatin resistance. In addition, multiple other efflux pumps in the ABC transporter family (eg, MRP1, ABCG1, and ABCA1) have also been characterized as important regulators of cancer chemotherapy resistance.30-32 Here, we showed that the expression levels of all these transporters were greatly suppressed by miR-495 mimics in esophageal cancer cells, but their expression could be effectively recovered by ATP7A overexpression in esophageal cancer cells transfected with miR-495 mimics. These results verify the roles of transporter expression regulation by microRNAs in the development of cisplatin resistance.
The formation of new vessels (angiogenesis) is critical for the metastasis of solid cancers; an area which has been widely targeted by various chemotherapies and emerging new therapeutic strategies.33,34 Angiogenesis is also involved in the development of cancer resistance to chemotherapies, 35 which is mediated by formation of a tumor microenvironment niche via crosstalk between tumor cells and stroma favoring drug resistance. 36 In this study, we found that HUVEC angiogenesis by cisplatin-resistant esophageal cancer cells was significantly repressed by transfection with miR-495 mimics, while ATP7A overexpression recovered the angiogenic capabilities of cisplatin-resistant esophageal cancer cells transfected with miR-495 mimics. Moreover, we showed that TNF-α and VEGF contributed to tumor growth partially by enhancing angiogenesis progression. 37 We also found that the levels of TNF-α and VEGF in cisplatin-resistant esophageal cancer cells were decreased by miR-495 mimics, while ATP7A overexpression significantly mitigated the decreases in TNF-α and VEGF levels in cisplatin-resistant esophageal cancer cells transfected with miR-495 mimics. In addition, the NLRP3 inflammasome pathway has also been reported to regulate the angiogenesis process in certain contexts. 38 Here, we showed that NLRP3 expression in cisplatin-resistant esophageal cancer cells was inhibited by miR-495 mimics, and those decreases in expression could largely be recovered by simultaneous ATP7A overexpression. When taken together, these results clearly demonstrated role played by suppression of miR-495-induced ATP7A expression in mediating angiogenesis during cisplatin resistance in esophageal cancer.
In summary, this study revealed that miR-495, whose expression was greatly downregulated in esophageal cancer tissues, reduced the cisplatin resistance and angiogenesis of esophageal cancer cells by directly targeting and suppressing ATP7A gene expression. These investigations broadened our understanding of the molecular mechanism that underlies epigenetic regulation of transporter pathways during development of chemotherapy resistance in esophageal cancer, and also provide a basis for developing new therapeutic regimens that reverse cancer chemotherapy resistance. However, in this study, no significant difference was found from other analysis of clinicopathological characteristics, and this might be due to number of included sample was not enough. Further investigation would focus on this issue.
Author Contributions
Z.H.L., S.W.L., and K.J.L. designed the study. Z.H.L. and S.W.L. performed the experiment, collected the data, and drafted the manuscript. Y.Q.W. and J.T.C. analyzed the data and visualized the results. K.J.L. and J.J. revised the paper. All authors read and approved the final manuscript.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Research and Development Fund Project of Dongguan People's Hospital.
Ethics Approval and Consent to Participate
This study was approved by the Ethics Committee of Dongguan People's Hospital (KYKT2020-023), and informed consents were obtained from all patients.
