Abstract
Objectives:
To develop a model to predict biochemical recurrence (BCR) after radical prostatectomy (RP), using artificial intelligence (AI) techniques.
Patients and Methods:
This study collected data from 7,128 patients with prostate cancer (PCa) who received RP at 3 tertiary hospitals. After preprocessing, we used the data of 6,755 cases to generate the BCR prediction model. There were 16 input variables with BCR as the outcome variable. We used a random forest to develop the model. Several sampling techniques were used to address class imbalances.
Results:
We achieved good performance using a random forest with synthetic minority oversampling technique (SMOTE) using Tomek links, edited nearest neighbors (ENN), and random oversampling: accuracy = 96.59%, recall = 95.49%, precision = 97.66%, F1 score = 96.59%, and ROC AUC = 98.83%.
Conclusion:
We developed a BCR prediction model for RP. The Dr. Answer AI project, which was developed based on our BCR prediction model, helps physicians and patients to make treatment decisions in the clinical follow-up process as a clinical decision support system.
Keywords
Introduction
Prostate cancer (PCa) is the most common cancer in men, and its treatment options vary widely. Among the treatment options, radical prostatectomy (RP) is the surgical removal of the prostate gland. RP may be used to treat PCa that has not spread beyond the prostate or has not spread very far. However, PCa can recur after radical prostatectomy.
Physicians determine the risk of cancer recurrence after RP based on prostate-specific antigen (PSA) level and cancer stage. Biochemical recurrence (BCR) is used when assessing the outcome of RP. BCR denotes an increase in the PSA level of patients who have received surgery and/or radiotherapy for PCa. BCR is used as an endpoint to assess treatment success. 1 That is, BCR may indicate that PCa has recurred; this is also termed PSA failure or biochemical relapse. BCR has important implications for the treatment of PCa and predicting BCR has been a topic of interest for a considerable period. 1 -3 We developed a model to predict BCR following RP, based on artificial intelligence (AI) techniques.
Recently, there has been a trend to introduce AI technology into the urology field. The application of AI has shown promise; AI has been reported to be excellent for outcome prediction and diagnosis. 4 -7 In South Korea, the PROMISE CLIP Project and the Dr. Answer AI have been ongoing since 2018. 8 The Dr. Answer AI project is South Korea’s largest project to develop software using AI technology. The project focuses on 8 diseases: cardiocerebrovascular disease, cardiac disorder, breast cancer, colorectal cancer, PCa, dementia, epilepsy, and childhood genetic and rare diseases (http://dranswer.kr). 9 PROMISE CLIP is project to develop intelligent software to solve medical demands related to PCa. The PROMISE CLIP registry consists of clinical, imaging, and pathology data for PCa based on multicenter EMR data.
Here, we developed a model to predict BCR following RP, using clinical data from the PROMISE CLIP project registry. We developed Dr. Answer AI software for PCa based on this prediction model.
Patients and Methods
BCR Definition
We developed a model to predict BCR after RP. BCR was defined as an increase in the blood level of PSA in patients with PCa after treatment with surgery or radiation, namely a patient whose PSA value exceeded 0.2 ng/mL at any point after RP. 2 This BCR definition was defined by discussion between 2 clinicians.
PCa Data for BCR Prediction Model
We collected data from 7,128 patients with PCa who received RP at 3 tertiary hospitals: Seoul St. Mary’s Hospital of the Catholic University, Samsung Medical Center, and Asan Medical Center. The participating hospitals are in Seoul and the Gyeonggi province (capital area). We utilized 6,755 cases out of 7,128 available, after excluding 369 for which relapse data were missing and 4 cases with a T-stage value of TX.
Input Variables for BCR Prediction Model
We used 16 variables to develop a model to predict BCR: age at diagnosis, BMI, marital status, education, smoking, drinking, family history of PCa, initial PSA, Gleason group, max positive core count, core ratio, neoplasm high risk malignant, extracapsular extension (ECE), seminal vesicle invasion (SVI), lymph node metastasis (LM), and T staging (Table 1). Neoplasm high risk malignant is a state neoplasm exists under epithelial tissue, not invading the stromal tissue into the basement membrane. In the Table 1, N is number of missing values and % is the missing values percentile. We filled the missing values with the mode for the categorical features and the median for the continuous feature grouped by age group and T stage.
Sixteen Input Variables for BCR Prediction Model.

Random Forest for BCR Prediction Model
We used a random forest (RF) to develop the BCR prediction model. RF has the advantage of addressing small sample sizes, high dimensional feature spaces, and complex data structures in a healthcare context. 10 The RF is nonparametric and interpretable, highly accurate, and efficient for diverse types of data. RF approaches have been used in a variety of healthcare fields, including PCa research. 11 -15
There existed a class imbalance problem in this study. The problem of data imbalance can be solved by 1) oversampling, 2) undersampling, and 3) combining oversampling and undersampling. We applied a random forest with diverse sampling methods. We used several undersampling methods such as One side selection (OSS), condensed nearest neighbor (CNN), edited nearest neighbors (ENN), and neighborhood cleaning rule (NCR). OSS is an undersampling technique that combines tomek links and the CNN rule. Tomek links are ambiguous points on the class boundary and are identified and removed in the majority class. The CNN method is then used to remove redundant examples from the majority class that are far from the decision boundary. Another undersampling method is ENN which is deleting the closest k data among multiple class data if not all or multiple classes. NCR is an undersampling technique that combines both the CNN rule to remove redundant examples and the ENN rule to remove noisy or ambiguous examples. Like OSS, the CSS method is applied in a 1-step manner, then the examples that are misclassified according to a k-nearest neighbor (KNN) classifier are removed, as per the ENN rule. Unlike OSS, less of the redundant examples are removed and more attention is placed on “cleaning” those examples that are retained.
We used oversampling methods such as random oversampling, adaptive synthetic sampling (ADASYN), synthetic minority oversampling technique (SMOTE), synthetic minority oversampling ENN, and synthetic minority oversampling tomek. Random Oversampling is replacing the data of a minority class over and over again. It is similar to increasing the weight. ADASYN method is a method of creating virtual fractional class data on a straight line between the fractional class data and data randomly selected from among the k-numbered fractional class data closest to the data. SMOTE is an oversampling method in which a sample of a class with a small number of data is taken and a new sample is created by adding a random value to the data. Synthetic minority oversampling ENN is a mixture of SMOTE method and ENN method. SMOTE Tomek is a mixture of SMOTE method and Tomeklink method.
Oversampling refers to a method of increasing the fractional class of data, including random sampling, ADASYN, and SMOTE. Undersampling is a method in which only a part of multiple class data is used. Methods include Tomek’s link method, ENN, and neighborhood cleaning rule. Additionally, it is possible to combine oversampling and undersampling, such as in SMOTE + ENN and SMOTE + Tomek. We applied several techniques to solve the class imbalance problem. Finally, we developed a new model that considers the influence of variables in general by using several ensemble sampling techniques: random forest with SMOTE + Tomek, ENN, and random sampling.
The data were divided into training datasets and a test dataset: 9 training datasets were matched to 1 test dataset. When developing predictive models, studies often use a dataset ratio of 8 to 2 or 7 to 3. As mentioned earlier, we used several undersampling and oversampling techniques to address class imbalance problems. In such techniques, the data used to train models contain values of matched characteristics as compared to the raw data, to improve model sensitivity. Accordingly, we considered it desirable to increase the size of the training dataset rather than increase the reliability of verification by increasing the size of the test dataset. Therefore, we used the aforementioned 9:1 ratio of training to test datasets.
Weka 3.8.3 and Python 3.7 were used as the machine learning programs. This study used the 5 Python libraries to develop the BCR prediction model: Pandas, NumPy, Sklearn, Pickle, and Imblearn. Finally, we achieved BCR prediction performance over 10-fold cross validation.
Ethics
The study was performed in accordance with the Declaration of Helsinki and was approved by 3 Institutional Review Boards: Catholic University (IRB number: KC18RNDI0509), Samsung Medical Center (IRB number: SMC201807069001), and Asan Medical Center (IRB number: 2018-0963).
Results
After data preprocessing, we finally used the data of 6,755 patients with PCa who received radical prostatectomy: 1,719 cases from Seoul St. Mary’s Hospital of the Catholic University (25%), 2,383 cases from Samsung Medical Center (35%), and 2,653 cases from Asan Medical Center (39%; Table 2). Among the 6,755 patients with PCa, 2,200 experienced BCR 200 (33%). Online Appendix A shows correlations among continous predictors resampling.
Basic Information.

Most patients were between 60 and 74 years of age at diagnosis (n = 5,024, 74.50%). Nearly all patients were married (n = 5,165; 97.40%), most had a high school education or lower (n = 3,604; 89.50%), 55.40% were non-smokers (n = 3,667), and 70.70% drank alcohol (n = 4,687). Most patients had no family history of PCa (n = 4,798; 87.60%). Among the patients, 36% had a Gleason score of 7 (n = 2,269), 77.30% had neoplasm high-risk malignancy (n = 5,084), 66.10% had no extracapsular extension (n = 4,312), 87.20% had no seminal vesicle invasion (n = 5,725), and 75.40% had no lymph node metastasis (n = 4,708). Stage 2 classification applied to 66.50% of patients (n = 3,286). The average BMI was 34.53, average initial PSA was 9.72, and the average maximum positive core count was 46.05. The average core ratio was 0.46 (Table 3).
Descriptive Statistics for 16 Input Variables.
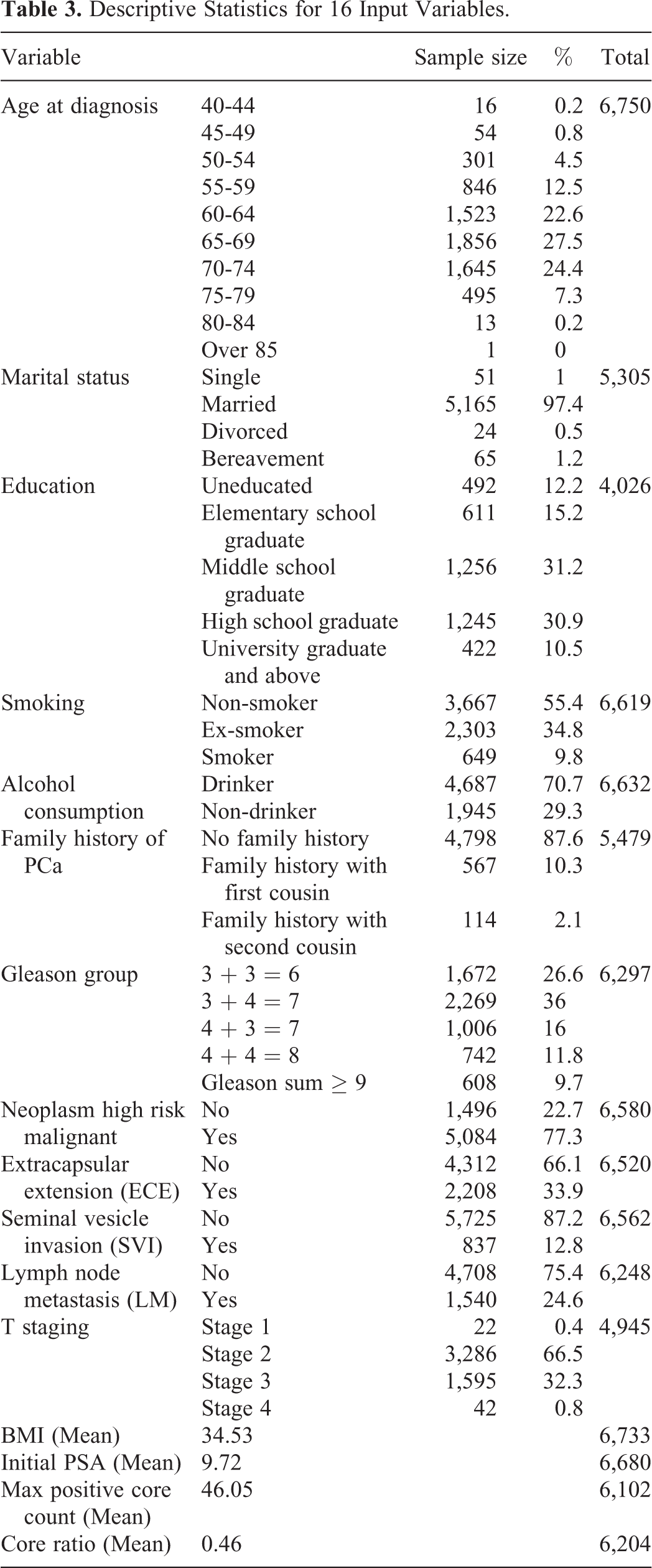
Table 4 compares the performance of other algorithms with that of the final algorithm. The random forest with SMOTE + Tomek, ENN, and random sampling was chosen based on comparison with other algorithms. The random forest achieved good performance: accuracy = 96.59%, recall = 95.49%, precision = 97.66%, F1 score = 96.59%, and ROC AUC = 98.83%.
Algorithm Performance Results.

Online Appendix B compares undersampling performance which shows mostly lower than 70% accuracy. It is poor performance compare to the oversampling performance.
The random forest model’s accuracy summary, after running the model 30 times with different seeds selected, shows minimum 0.9491, maximum 0.9704, and mean 0.9613 (Online Appendix C).
Software for BCR Prediction Model
We developed the Dr. Answer AI for PCa software based on the BCR prediction model for radical prostatectomy. Figure 1 shows the prediction software main screens. The SW receives the individual patient’s unique values and outputs changes before and after compared with previously entered data. The SW allows patients with PCa to recognize and prevent negative prognoses in advance, and facilitates the patient’s understanding of their current condition. The “Pathology” table summarizes the pathological results after surgery. Patients with PCa can become aware of their prognosis and risk of BCR in advance by referring to the results. The yellow bar on the right indicates the BCR prediction. The gray value on the left represents the most recent blood test value after surgery.
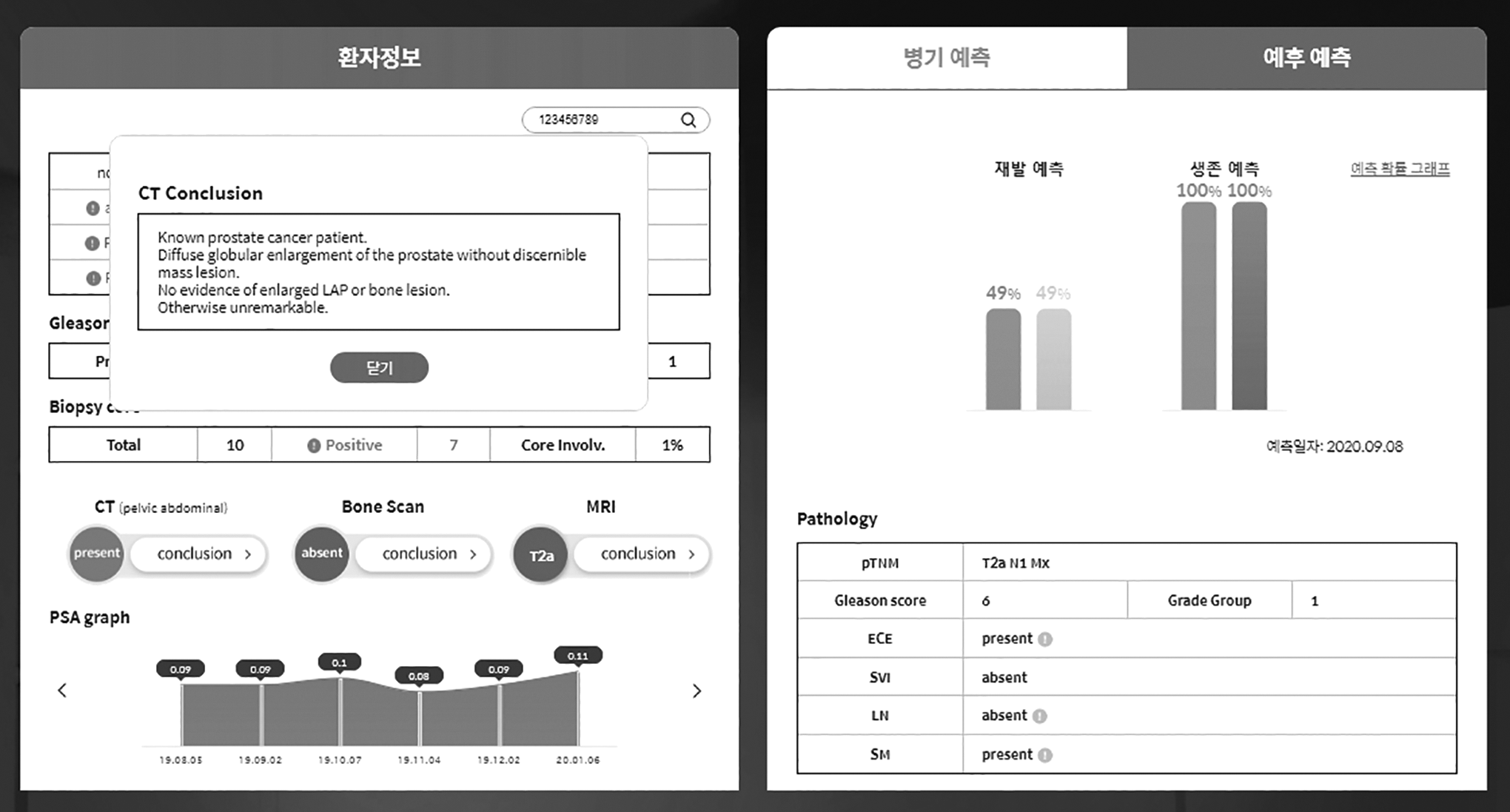
Dr. Answer AI software for BCR prediction model.jpg.
Discussion
We developed a model to predict BCR following radical prostatectomy. The model achieved good performance (accuracy: 96.59%, AUC: 98.83%) for predicting BCR based on AI techniques. BCR represents an important clinical prognosis; previous studies have assessed treatment and target features. 16 -19 It has been reported that machine learning can achieve good performance in predicting BCR. In robot-assisted prostatectomy, the accuracy of K-nearest neighbor, random forest tree, and logistic regression were reported as 0.976, 0.953, and 0.976, respectively. 17 Zhang et al (2016) indicated that an imaging-based approach using support vector machine (SVM) classification was superior at predicting PCa outcome(sensitivity = 93.3%, specificity = 91.7%, accuracy = 92.2%). 20 The performance of our model was significantly better than model performance reported in previous studies. These models offer a variety of individualized treatment options for the clinical follow-up process. The Dr. Answer AI software helps physicians and patients to make treatment decisions during the clinical follow-up process. The BCR prediction model following RP could be have meaningful clinical used, given its good performance.
In recent years, there has been a trend toward building AI-based predictive models in the field of urology. 21 AI is effective for application to PCa, which is complex in diagnosis and treatment. 4 However, although AI technology is playing a growing role in the urology field, practical implementation of AI models is limited. 6 We developed the Dr. Answer AI software based on a random forest. It has been reported that random forests have high accuracy and efficiency when applied to diverse types of data. Random forests have been used in various aspects of prostate cancer research. 11 -15 The current software represents the response to one of the 8 diseases considered in the Dr. Answer AI project. 9 The intention is to use the Dr. Answer AI software in hospitals from 2020, upon obtaining a software license from the Ministry of Food and Drug Safety. The Dr. Answer AI software has the advantage that it is technology that can be applied to actual medical practice.
Another advantage of the current model is that we used several techniques to overcome the class imbalance problem. We collected data from 7,128 patients with PCa following RP, and used the data of 6,755 cases; there were 2,200 patients with PCa patients who experienced BCR (33%). The problem of data imbalance is caused by the different number of datapoints in each class. There are several sampling techniques by which to address class imbalances. Most techniques overweight problems or overgeneralize certain variables in the model. The current study considered the influence of variables in general via several ensemble sampling techniques: RF with SMOTE + Tomek, ENN, and random oversampling. SMOTE + Tomek combines the sampling methods of the SMOTE and Tomek techniques. SMOTE randomly selects K-nearest neighbors of the minority class. Tomek is a data cleaning technique that defines a Tomek link as a pair neighbors with minimal Euclidian distance. SMOTE + Tomek utilizes both methods’ advantages, which consist of obtaining balanced data sets with a clear boundary between the majority and minority classes. 22 ENN represents an undersampling method that discards a sample if it is misclassified followed a nearest neighbors classification. ENN addresses performance issues with nearest neighbor classification due to the presence of minority class samples. ENN performance approaches that of the Bayes classifier. 23
In contrast, random oversampling represents a method of repeatedly replacing imbalanced classes for analysis. Random oversampling constructs data sets with both an expected average and standard deviation equal to those of the original minority class data. 24 We were able to improve the accuracy of our model by applying several techniques to address class imbalance problems.
A further advantage of the current study is that it used the data of 6,755 patients with PCa from 3 centers. Previous studies infrequently exceeded a sample size of 1,000. 4 We developed a model to predict BCR following radical prostatectomy using clinical data from the PROMISE CLIP registry. In multi-center studies such as this, there are differences in the data formats used by each hospital. It required effort to format each institution’s data because the data were not organized into a common data model (CDM). In addition, to ensure data security, the data of each institution were uploaded to the Naver Cloud Platform, 25 data were deleted after the project, and the researchers could not access individual datasets. When conducting large-scale research that involves multiple institutions, a CDM for data sharing is important. In South Korea, many hospitals are converting data into a CDM, but it takes time for multi-center research based on a CDM to become feasible. Thus, difficulties associated with multi-center research existed in this study but were addressed successfully. Accordingly, a suitable algorithm and associated software were developed.
Although the current findings are meaningful, the study was subject to some limitations. First, BCR referred to a patient whose PSA value was greater than 0.2 ng/mL at least once following RP. 2 However, other studies have proposed 0.4 ng/m as a suitable criterion for BCR. 1,3 That is, criteria deemed appropriate differ, depending on the researcher. These differences in criteria can change model performance. Future research could consider different definitions of BCR. Second, the data of the hospitals that participated in this project were not converted into a CDM, although the project was carried out successfully. Conversion of hospital data to the same standard and integration of the data required considerable effort. If hospital data were converted to a CDM, this would increase the efficiency of future large-scale multi-center studies.
PCa is one of the most complex cancers to diagnose and treat. Therefore, it is very important to accurately predict BCR and formulate treatment plans based on this knowledge. The BCR prediction model will be useful for the aggressive treatment of PCa. The Dr. Answer AI software helps physicians and patients to make treatment decisions during the clinical follow-up process as a clinical decision support system (CDSS).
Supplemental Material
Supplemental Material, sj-pdf-1-tct-10.1177_15330338211024660 - Dr. Answer AI for Prostate Cancer: Predicting Biochemical Recurrence Following Radical Prostatectomy
Supplemental Material, sj-pdf-1-tct-10.1177_15330338211024660 for Dr. Answer AI for Prostate Cancer: Predicting Biochemical Recurrence Following Radical Prostatectomy by Jihwan Park, Mi Jung Rho, Hyong Woo Moon, Jaewon Kim, Chanjung Lee, Dongbum Kim, Choung-Soo Kim, Seong Soo Jeon, Minyong Kang and Ji Youl Lee in Technology in Cancer Research & Treatment
Footnotes
Authors’ Note
Jihwan Park and Mi Jung Rho contributed equally to this work. The study was performed in accordance with the Declaration of Helsinki and was approved by 3 Institutional Review Boards: Catholic University (IRB number: KC18RNDI0509), Samsung Medical Center (IRB number: SMC201807069001), and Asan Medical Center (IRB number: 2018-0963).
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Authors Jaewon Kim, Changjung Lee, and Dongbum Kim are employed by LifeSemantics, which developed the Dr. Answer AI software for prostate cancer, and which owns the full rights to use the software for commercial purposes, as seen fit by the company. The co-first authors, Mi Jung Rho and Jihwan Park, are married couples. They participated in this project at the time of the research.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Institute for Information & communications Technology Promotion(IITP) grant funded by the Korea government (MSIT) (2018-0-00861, Intelligent SW Technology Development for Medical Data Analysis).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
