Abstract
Background:
In the present study, we aimed to find an effective target for the treatment of tongue cancer using gene chip screening and signal pathway research.
Methods:
We used microarray screening and gene expression profile analyses to find important differentially expressed genes in tongue cancer. We constructed a protein-protein interaction network, and used enrichment analysis of the Kyoto Encyclopedia of Genes and Genomes to screen for important genes. We then silenced the genes of interest in SCC154 cells to study the relationship with the Phosphatidylinositol 3-kinase/Akt signal pathway. Western blot analyses, the 3-(4,5Dimethylthiazol-yl)-2,5Dimethylthiazol-2-yl)-2,5diphenyltetrazolium bromide (MTT) test, and immunofluorescence assays were used to compare the expression levels of Phosphatidylinositol 3-kinase/Akt signal pathway-related proteins, cell viability, and cell proliferation ability in normal SCC154 cells, Si-RNA SCC154 cells, and gene-silenced SCC154 cells. The scratch test, Transwell test, and western blotting were used to determine migration, invasion, and carcinogenesis.
Results:
Using GSE9844, GSE13601, and GSE31056 gene chips, we identified 93 upregulated genes and 76 downregulated genes in tongue cancer. Using the protein-protein interaction network and Kyoto Encyclopedia of Genes and Genomes enrichment analyses, we further identified 47 differentially expressed genes. Using Kaplan-Meier plotter online tools, we also identified 3 genes (SPP1, Recombinant Human Secreted Phosphoprotein 1; PLAU, plasminogen activator urinary; and APP, amyloid precursor protein). Compared with normal SCC154 cells and Si-RNA control SCC154 cells, the expressions of Phosphatidylinositol 3-kinase/Akt pathway proteins in si-SPP1 SCC154 cells were significantly decreased (*P < 0.05), and the protein activities and proliferation abilities were also significantly decreased (*P < 0.05), while the migration ability, invasion ability, and cancer forming ability were significantly increased (*P < 0.05).
Conclusion:
Inhibition of the SPP1 gene may have a therapeutic effect on tongue cancer, and could be an effective target for the treatment of this disorder.
Keywords
Introduction
Tongue cancer is a type of head and neck tumor, which occurs in the tongue. 1 The general clinical manifestations of tongue cancer are pain, erosion, ulcer, and mass. 2 Tongue cancer is more likely to occur in males, and females have a better prognoses than males. 3 The incidence of tongue cancer is presently increasing in many parts of the world, which is mainly related to the external environment and genes of the human body. The external causes of tongue cancer involve an unsanitary diet; often eating hot or irritating food; the influence of denture material; chewing of betel nuts; long-term exposure to cigarette butts; and other harmful substances. 4 Tongue cancer will continue to develop, and patients with advanced tongue cancer often relapse after surgery. The effects of radiotherapy and chemotherapy are not satisfactory, and the prognosis is poor, 5 so tongue cancer should be diagnosed and treated early.
The treatment of tongue squamous cell carcinoma mainly involves surgery, and adjuvant treatments include radiotherapy and chemotherapy, but recurrence is very high, 6 so the treatment of tongue cancer needs long-term follow-up observation and monitoring of the disease development. 7 The tongue is a common organ, which has a great influence on fasting, drinking water, and speaking. 8 There is therefore an urgent need to find more targets for the treatment of tongue cancer, as well as to develop more effective drugs and treatment methods. The analysis and summary of gene data is therefore the fastest and most effective way to identify effective targets. 9
Materials and Methods
Gene Chip Data
Gene chip data from the Gene Expression Omnibus (GEO) database were used. Tongue cancer samples were collected from data sets labeled GSE9844, GSE13601, and GSE31056. 10 Among them, the number of genes related to tongue cancer in GSE 9844 was 646, the number of genes related to tongue cancer in GSE 13601 was 1,857, and the number of genes related to tongue cancer in GSE 31056 was 2,070. These data were obtained using the Affymetrix human genome U133 Plus 2.0 array chip platform (Affymetrix, Santa Clara, CA, USA). 11
Standardization of the Original Data
For the original data from the chip, we used the Robust Multichip Average method to first normalize the normalized value, and then used the logarithm of log2FC (fold-change, log2FC = log2mean group1/group2) to generate the value as the standardized data, and used the standardized data for downstream analyses. 12
Differential Gene Analysis and Enrichment Analysis of the Kyoto Encyclopedia of Genes and Genomes (KEGG)
Gene differential expression analysis was based on the edger function package of R. For the GSE9844, GSE13601, and GSE31056 tongue cancer sample data sets, we divided the samples into a pair of healthy periodontal tissues and inflammatory periodontal tissues for differential gene analysis. 10 The absolute value of logFC > 1 and p-value ≤ 0.05 were used as the criteria to screen the upregulated differentially expressed genes. The absolute value of logFC < −1 and p-value ≤ 0.05 were used as the criteria to screen the downregulated differentially expressed genes. We used 3 databases to map upregulated genes and downregulated genes, and found that the upregulated genes and downregulated genes were shared in 3 gene chips. The differentially expressed genes were enriched and analyzed by the KOBAS (KEGG ontology based annotation system) database. When the p-value was < 0.05, the corresponding entries were considered to be significantly enriched. 13
Construction of the Protein-Protein Interaction (PPI) Network and Screening of Key Genes
For the differentially expressed genes, the PPI network was constructed by using the string database (https://string-db.org/cgi/input.pl). The PPI network was input into the software of Cytoscape, and key genes in the PPI network were screened based on the algorithm of Multiscale Curvature Classification (MCC) by using the cytoHubba plug-in of the software of Cytoscape. 14
Cell Culture, Treatment, and Transfection
The human SCC154 cell line was purchased from American Type Tissue Collection (Manassas, VA, USA) (product number: crl-3241 TM). Approximately 1.5 × 10 4 cells per well were cultured in MEM medium 11090081 (Gibco, Gaithersburg, MD, USA), fetal bovine serum 900-108 Gemini, and penicillin 091670249 MP. After 24 h, 20 pmol Si-SPP1 (siRNA) was dissolved in 50 UL Opti MEM serum-free medium. One µL lipofectamine 2000 was dissolved in 50 µL Opti MEM serum-free medium, mixed well, incubated for 5 min at room temperature, and the above 2 tubes of solution were further incubated for 20 min. During transfection, the 24 well plate medium was replaced by 400 µL per well of serum-free medium. The mixed solution was added to corresponding wells of a 24-well plate, and the serum-containing medium was replaced at 4–6 hours. 15 The si-SPP1 was purchased from Proteintech (Chicago, USA), and had the following sequences: si-SPP1-1, 5′-CGAUCGAUAGUGCCGAGAAGC-3′ and si-SPP1-2, 5′-AGCUAGUCCUAGACCCUAAGA-3′. 16
Western Blot Analysis
SCC154 cells and SCC154 cells transfected with si-SPP1 were extracted using cold lysis buffer to obtain total protein. The concentrations of total protein were determined using the bicinchoninic acid protein concentration assay kit (Pierce, Rockford, IL, USA). Protein samples were resolved using sodium dodecyl sulfate polyacrylamide gel electrophoresis and transferred onto polyvinylidene fluoride membranes. The membranes were incubated with primary antibody overnight at 4°C. The antibodies used in this study were as follows: anti-IκBα (1:1,000, 4812; Cell Signaling Technology, Danvers, MA, USA), anti-SPP1 (1:1,000, 40672 S; Cell Signaling Technology), anti-p-PI3 K (1:1,000, 17366 S; Cell Signaling Technology), anti-PI3 K (1:1,000, 4249 T; Cell Signaling Technology), anti-p-Akt (1:2,000, 4060 T; Cell Signaling Technology), anti-Akt (1:2,000, 2920 S; Cell Signaling Technology), anti-vimentin (1:1,000, 5741 T; Cell Signaling Technology), anti-E-cadherin (1:1,000, 14472 S; Cell Signaling Technology), and anti-cytokeratin (1:5,000, ab9377; Abcam, Cambridge, UK). The membranes were then incubated with horseradish peroxidase (HRP)-labeled goat anti-rabbit IgG (1:5,000, ab205718; Abcam) secondary antibody or HRP-labeled goat anti-mouse IgG (1:5,000, ab205719; Abcam) secondary antibody for 1 h at 37°C. Finally, the protein bands were visualized by enhanced chemiluminescence exposure solution, and quantified using a gel imaging system (UVP, Upland, CA, USA). Anti-glyceraldehyde 3-phosphate dehydrogenase (1:1,000, 5174 T; Cell Signaling Technology) was used as the internal reference. 17
MTT Detection Assay
SCC154 cells in log phase were adjusted to a concentration of 50,000 cells/mL, and 100 µL of the cell suspension were added to each well of a 96-well plate. The cells were cultured in the incubator, followed by addition of the drug. The drug was first prepared using an Eppendorf tube, then the medium of the 96-well plate was discarded, followed by 2 washes and addition of the drug. The cells were cultured 24 h. Twenty microliters of MTT solution (5 mg/mL) was added to each well and cultured for 3–4 h. The MTT was purchased from Solarbio (Beijing, China). The medium was then carefully removed from each well, followed by addition of 150 µL of dimethyl sulfoxide (DMSO), followed by incubation for 10 min at 37°C with shaking at low speed for 10 min. The absorbance at 490 nm was then measured. The zeroing well (serum-free medium, MTT, and DMSO) and the control well (cells, maximum concentration of drug-dissolving medium, serum-free medium, MTT, and DMSO) were then measured. The cell viability was calculated using the following formula: cell viability of the control = (absorbance of the drug − absorbance of the zeroing well)/(absorbance of the control well—absorbance of the zeroing well) × 100%. 18
Immunofluorescence Detection of Cell Proliferation
Cover slips were placed into a 12-well plate, and SCC154 cells and the SCC154 cells transfected with si-SPP1 were then added. After the cells adhered, the slides that had adhered cells were washed with phosphate-buffered saline (PBS) 3 times for 3 min each time, then fixed with 4% paraformaldehyde for 15 min, followed by washing the slides with PBS 3 times for 3 min each time, followed by addition of 0.5% Triton X-100 in PBS for 20 min at room temperature. The slides were then washed with PBS 3 times for 3 min each time. The PBS was then absorbed with absorbent paper, normal goat serum was added for 30 min at room temperature, the solution was removed without washing, and diluted primary antibody to which Ki67 was added to each slide, which was then placed in a wet box and incubated overnight at 4°C. Fluorescence secondary antibody was then added, and the slides were washed 3 time for 3 min each time. After removing the solution with absorbent paper, diluted fluorescence-labeled secondary antibody was added and the slides were incubated in a wet box for 1 h at 20–37°C, then the slides were washed with PBST for 3 times for 3 min each time. From the time of adding the fluorescent secondary antibody, all the following steps were conducted in the dark place as much as possible. To stain the nucleus, 4′,6-diamidino-2-phenylindole was added and incubated in the dark for 5 min, rinsed using PBST 4 time for 5 min each time and absorbent paper was used to dry the liquid. Fluorescence quenching agent was then added and the cells were examined using a fluorescence microscope. 19
Scratch Test
Before the assay, a ruler and marker pen were sterilized by irradiation with ultraviolet light for 30 min on a sterilized table. The marker pen was then used at the back of the 6-well plate, with a ruler to draw a horizontal line, about every 0.5–1 cm across the well. At least 5 lines were drawn for each well. Approximately 5 × 10 5 cells were added to the wells (the number varied with different cells), to ensure that they could adhere to the well and fully spread. The next day, a gun head was used to compare with the ruler, to be perpendicular to the horizontal line scratch on the back. The gun head was vertical and not inclined. The cells were washed with PBS 3 times to remove excess cells, followed by addition of serum-free medium. The cells were then incubated at 37°C in a 5% CO2 incubator. 20 Photographs were taken at 0 and 72 h using an inverted AE2000 fluorescence microscope (Motic, Richmond, Canada).
Transwell Assay Detection of Cell Invasion
The SCC154 cells and SCC154 cells transfected with si-SPP1 were starved for 12–24 h by removing the serum. After digestion, the cells were centrifuged and the culture medium was discarded, washed with PBS for 1–2 times, and resuspended with serum-free medium. A cell suspension (200 µL) was added to the Transwell inserts purchased from Chemicon International (Temecula, CA, USA). Generally, 600 µL culture medium containing 15% fetal bovine serum was added to the lower chamber of a 24-well plate. It should be noted that bubbles were often generated between the lower culture medium and the chamber. Once the bubbles were generated, chemotaxis of the lower culture medium decreased or even disappeared. Special attention was therefore made when planting the plate. Once the bubbles appeared, the chamber was lifted to remove the bubbles, and then the chamber was placed into the culture plate, followed by routine culturing for 24 h. The Transwell chamber was then removed, the culture solution was discarded, and cells were washed twice with calcium-free PBS. The cells in the upper layer of non-migrating cells were then removed with a cotton swab, fixed with methanol or formaldehyde for 30 min, and air-dried. A 0.1% Crystal Violet stain was then added and incubated for 30–60 min, followed by washing with PBS for 3 times. The moisture in the upper chamber was then removed using a cotton swab. Using a magnification of 400×, the cells were observed and counted in 5 visual fields. 21
Statistical Analysis
All experiments were performed at least 3 times, and all experimental data were presented as the mean ± standard deviation. A p-value of < 0.05, was considered statistically significant. 22 For comparisons between 3 or more groups, 1-way analysis of variance was used, followed by Tukey’s multiple comparison tests. Statistical analysis was performed using the Prism software, version 5.0 (GraphPad Software, San Diego, CA, USA) and SPSS statistical software for Windows, version 18.0 (SPSS, Chicago, IL, USA).
Results
Identification of Differentially Expressed Genes in Tongue Cancer
After checking the gene chip data from the GEO database, we found that the number of genes related to tongue cancer in GSE 9844 was 646, the number of genes related to tongue cancer in GSE 13601 was 1857, and the number of genes related to tongue cancer in GSE 31056 was 2070. Figure 1A shows the shared relationships of upregulated genes. Among the tongue cancer related genes of these 3 gene chips, 93 were upregulated in common (Table 1). Figure 1B shows the shared relationships of downregulated genes. Among the tongue cancer related genes of these 3 gene chips, 76 were downregulated in common (see Table 1). LogFC = log (mean group1/group2); p-value ≤ 0.05.

Identification of differentially expressed genes in tongue cancer. The number of genes related to tongue cancer in GSE 9844 was 646, the number of genes related to tongue cancer in GSE 13601 was 1,857, and the number of genes related to tongue cancer in GSE 31056 was 2,070. A, Shared relationships of upregulated genes. Among the tongue cancer related genes of these 3 gene chips, 93 were upregulated in common. B, Shared relationships of downregulated genes. Among the tongue cancer related genes of these 3 gene chips, 76 were downregulated in common. LogFC = log (mean group1/group2); p ≤ 0.05.
All 216 Commonly Differentially Expressed Genes (DEGs) Were Detected From 3 Profile Datasets, Including 93 Upregulated Genes and 76 Downregulated Genes in Tongue Cancer Tissues Compared to Normal Tongue Tissues.

PPI Network Construction Using the STRING Online Database and Cytoscape Software
We constructed the differentially expressed gene PPI network complex image. The nodes denoted proteins and the edges denoted the interactions of proteins (Figure 2A). The clusters of 1–4 for PPI were obtained by module analysis using Cytoscape software (degree of cutoff = 2, node score cutoff = 0.2, k-core = 2, and maximum depth = 100). The first 47 genes with high importance in the PPI network were
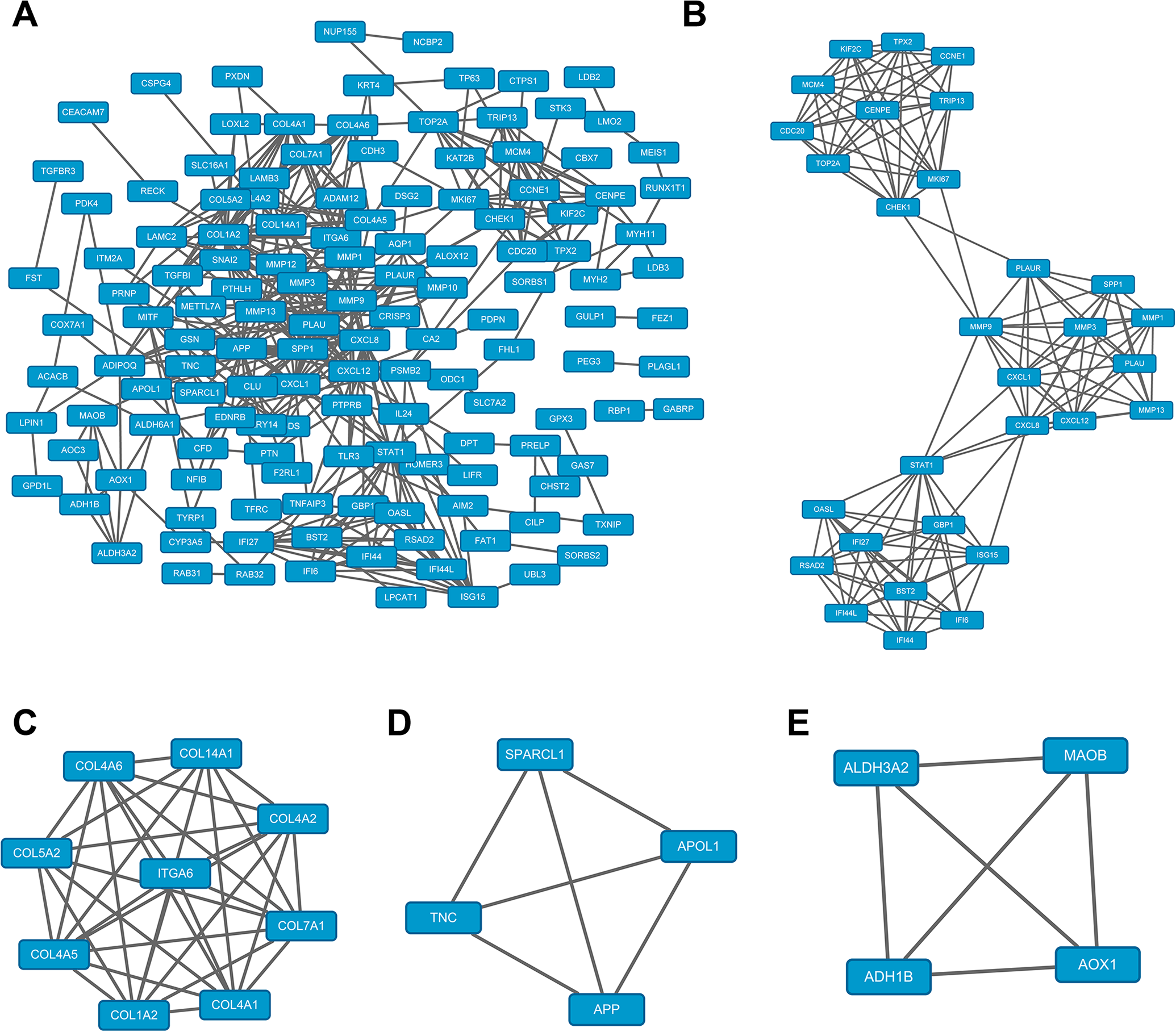
The protein-protein interaction (PPI) network constructed using the STRING online database and Cytoscape software. A, The differentially expressed gene PPI network complex. The nodes denote proteins, and the edges denote the interaction of proteins. B-E, The clusters 1–4 for PPI were obtained by module analysis using the Cytoscape software (the degree of cutoff = 2, node score cutoff = 0.2, k-core = 2, and maximum depth = 100). The first 47 genes with high importance in the PPI network were screened out from B to E. They are
KEGG Pathway Analysis of 47 Commonly Differentially Expressed Genes in Tongue Cancer.

ECM = extracellular matrix.
SPP1 Gene in Human Tongue Cancer of GEO DataSets.

SPP1: secreted phosphoprotein 1.
The Prognostic Information of the 47 Core Genes
We used Kaplan-Meier plotter online tools to identify the prognostic information of the 47 core genes, with only 4 of 47 genes having a significantly worse survival rate (p < 0.05). The survival time of patients with high expressions of

The prognostic information of the 47 core genes. Kaplan-Meier plotter online tools were used to identify the prognostic information of the 47 core genes and only 4 of 47 genes had a significantly worse survival rate (p < 0.05). HNSC: head-and-neck squamous cell carcinoma. A, The effect of SPP1 expression levels on HNSC patient survival. B, The effect of PLAU expression levels on HNSC patient survival. C, The effect of CXCL1 expression levels on HNSC patient survival. D, The effect of APP expression levels on HNSC patient survival.
Three Genes Were Significantly Expressed in Head and Neck Squamous Cell Carcinoma (HNSC) Patients When Compared to Healthy Controls
Four genes, which were related with poor prognoses, including
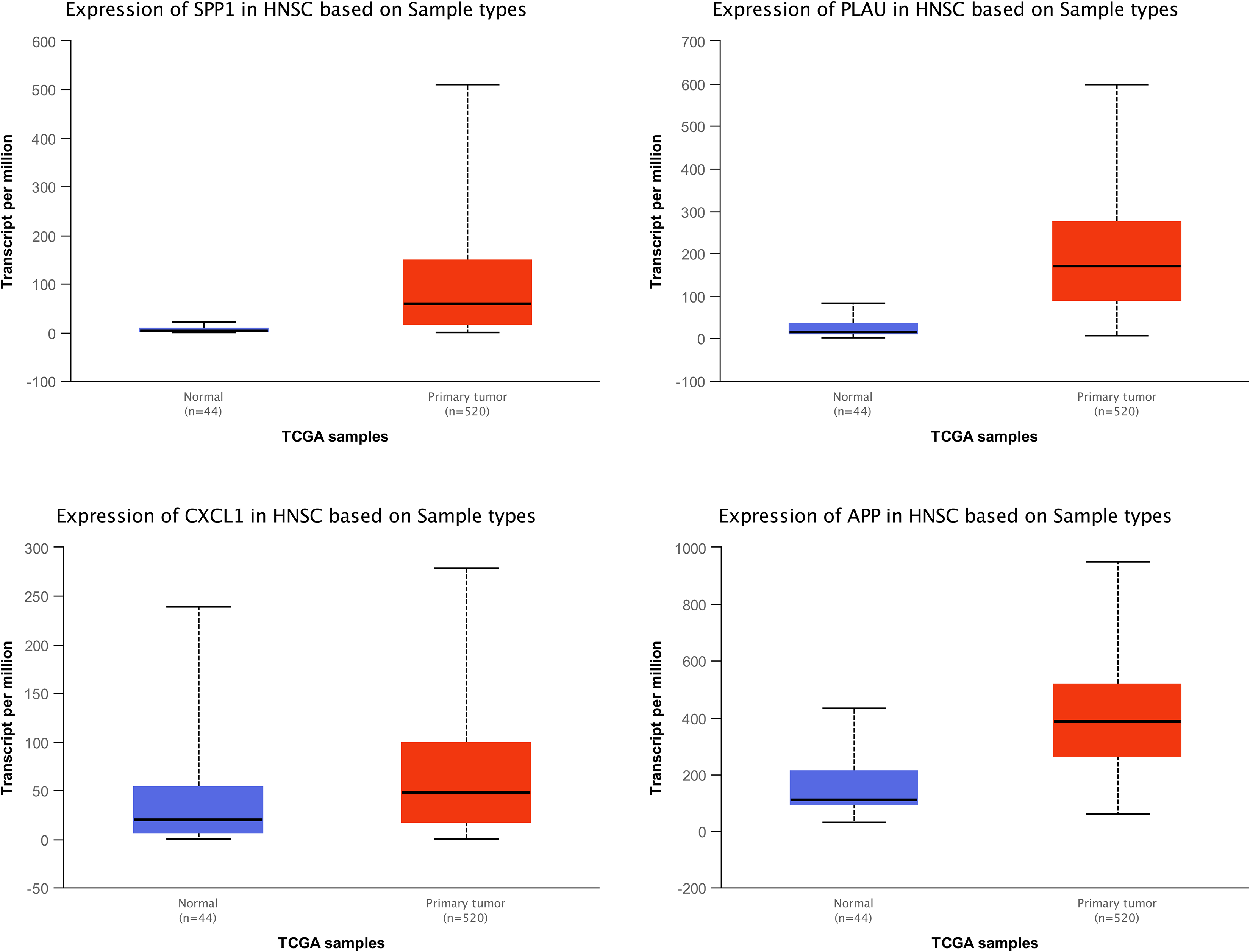
Three significantly expressed genes in head-and-neck squamous cell carcinoma (HNSC) patients compared to controls. Four genes, which were related with poor prognosis, including
Silencing of SPP1 Suppresses PI3K/Akt Signaling Pathway and Proliferation of Tongue Cancer Cells
We used the WB method to detect the protein expressions in the PI3K/Akt signaling pathway in tongue cancer control cells and si-SPP1 cancer control cells. The proteins included SPP1, p-PI3 K, PI3 K, p-Akt, and Akt. The proteins in the PI3K/Akt signaling pathway of si-SPP1 SCC154 cells were decreased (Figure 5A). We used the MTT assay to detected cell viability assay in tongue cancer control cells and si-SPP1 cancer control cells. The protein levels of the Si-SPP1 group were significantly lower than that of the normal control group (p < 0.01, Figure 5B). Compared with the si-SPP1 SCC154 cells, tongue cells from the tongue cancer control group and the tongue cancer control group of the si-RNA control had stronger proliferative ability (p < 0.05, Figure 5C).
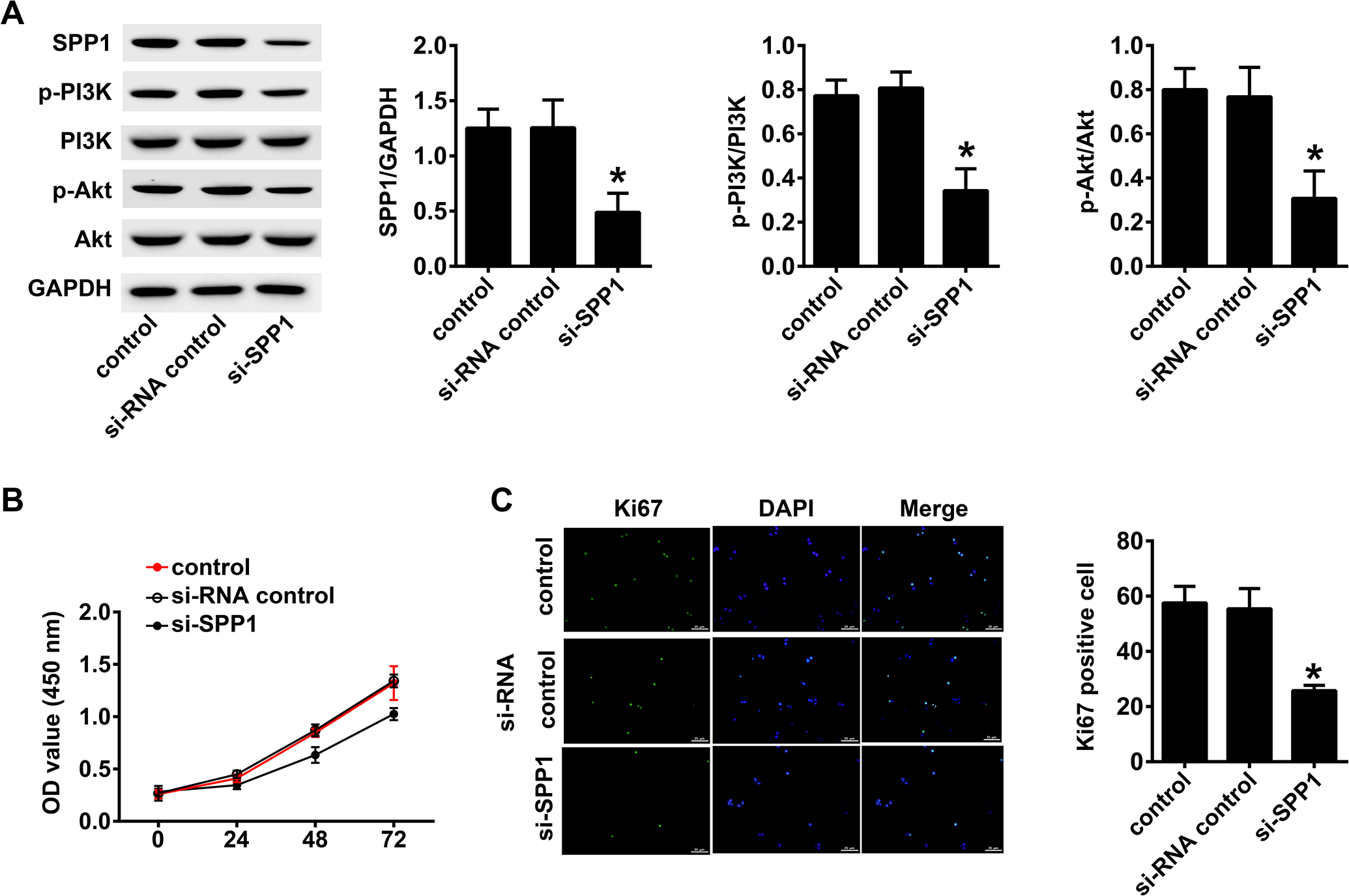
Silencing of SPP1 suppresses the PI3K/Akt signaling pathway and proliferation of tongue cancer cells. A, The change in expression of each protein of the PI3K/Akt signaling pathway detected by western blotting in control SCC154 cells and si-SPP1 SCC154 cells. The proteins included SPP1, p-PI3 K, PI3 K, p-Akt, and Akt. The proteins in the PI3K/Akt signaling pathway of si-SPP1 SCC154 cells were decreased. B, The detection of cell growth activity using the MTT assay of tongue cancer control and si-SPP1 SCC154 cells. The Si-SPP1 group was significantly lower than that of the normal control group (p < 0.01). C, Immunofluorescence detection of cell proliferation. Compared with the si-SPP1 SCC154 cells, the tongue cells in the tongue cancer control and the tongue cancer control groups of si-RNA had stronger proliferative ability (p < 0.05).
Silencing of SPP1 Suppresses Migration, Invasion, and the Epithelial-to-Mesenchymal Transition (EMT) of Tongue Cancer Cells
For detection of cell migration in tongue cancer we used the scratch test. The scratch test was conducted on tongue cancer control cells during the logarithmic phase of si-RNA tongue cancer control cells and si-SPP1 SCC154 cells growth cycles. The cell migration was determined 72 h later. The results showed that si-SPP1 cells had the longest migration distance and the strongest migration ability (p < 0.05, Figure 6A). We used the Transwell assay to determine the invasive ability of SCC154 and si-SPP1 SCC154 cells. The invasive ability of si-SPP1 SCC154 cells was stronger than that of normal SCC154 and Si-RNA SCC154 cells (p < 0.05, Figure 6B). The western blot results indicated that vimentin in the si-SPP1 SCC154 cells was significantly lower than that of normal SCC154 and Si-RNA control SCC154 cells (p < 0.05). However, E-cadherin and cytokeratin in the si-SPP1 SCC154 cells were significantly higher than those in normal SCC154 and Si-RNA control SCC154 cells
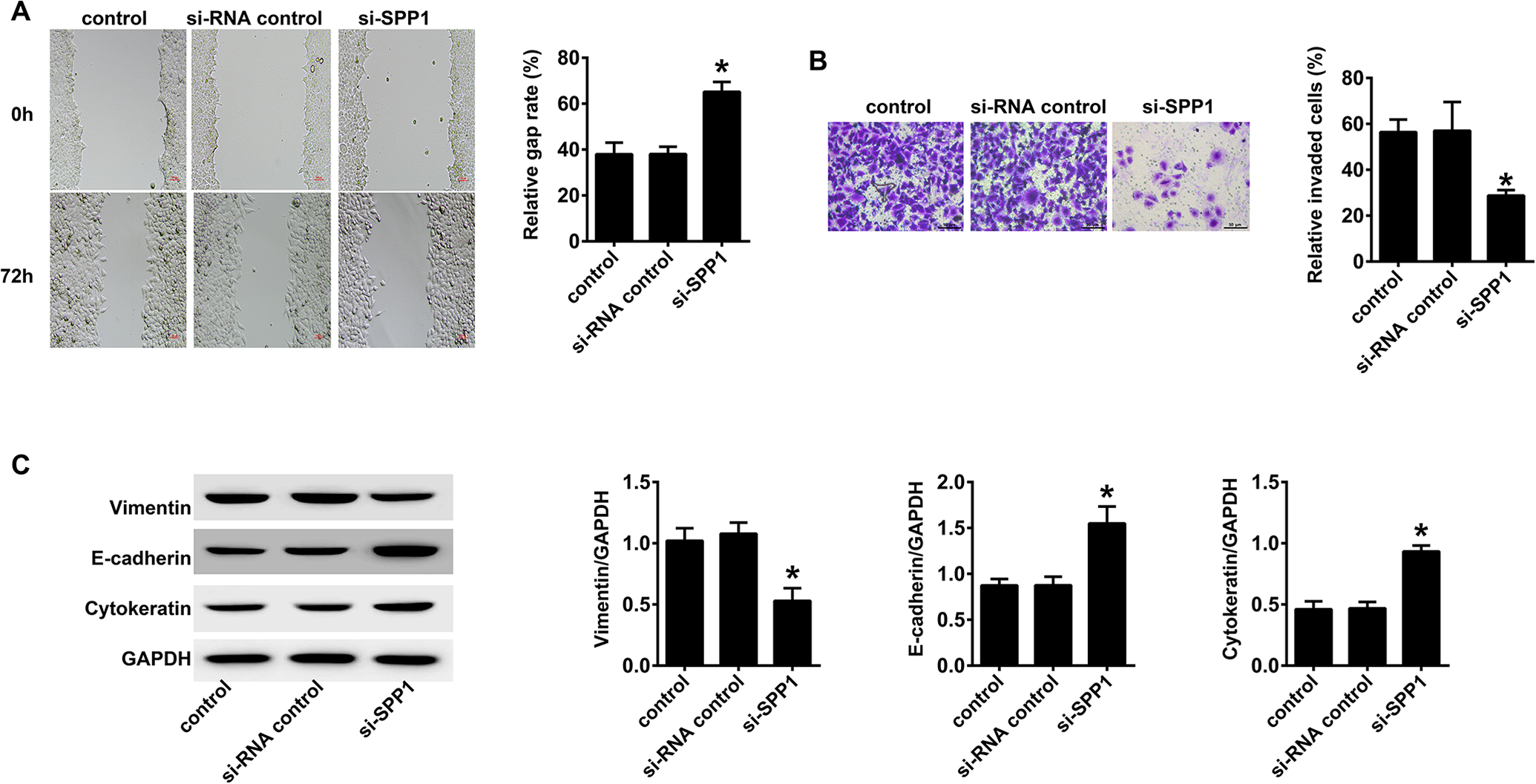
Silencing of SPP1 suppresses migration, invasion, and the epithelial-to-mesenchymal transition (EMT) of tongue cancer cells. A, The detection of cell migration in tongue cancer using the scratch test. The scratch test was conducted on tongue cancer control cells as well as Si-RNA tongue cancer control and si-SPP1 SCC154 cells during the logarithmic phase of cell growth. The cell migration was measured 72 h later. The si-SPP1 cells had the longest migration distance and the strongest migration ability (p < 0.05). B, The Transwell assay for the invasive ability of SCC154 cells. The cells were cultured in a co-culture system and stained with a fluorescent dye after 36 h. The invasive ability of si-SPP1 SCC154 cells was stronger than that of normal SCC154 and Si RNA SCC154 cells (
Discussion
Tongue cancer, or squamous cell carcinoma of the tongue, is one of the most common malignant tumors of the head and neck, as well as being a common oral cancer.
23
Tongue cancer can be divided into 2 types: tongue body cancer in the anterior 2/3 part and the tongue root cancer in the posterior 1/3 part.
24
At present, surgical treatment, radiotherapy, chemotherapy, and targeted treatment are mainly used to treat tongue cancer. However, the malignant degree of tongue cancer is high, the prognosis is poor, and the recurrence probability is high. Patients need long-term follow-up observation.
25
There are few specific targets for tongue cancer, mainly EGFR and EGFR/Akt.
26
It is therefore important to find a specific target for tongue cancer. In our study, we determined the differential genes of SCC154 cells and normal tongue cells, and found 3 highly specific genes,
In our study, the expression levels of SPP1, PLAU and APP genes in head and neck squamous cell carcinoma patients were significantly higher than that of normal people, and the prognosis of patients with low expression of SPP1, PLAU and APP genes was significantly higher than that of patients with high expression. SPP1 is a secreted multifunctional glycoprotein. Its functions include bone metabolism, immune regulation, wound healing, cell survival and tumor progression. SPP1 is a member of the small integrin binding ligand N-linked glycoprotein family, which is expressed in bone, but also in other tissues. As a cytokine, SPP1 is involved in promoting the production of interferon gamma and interleukin-12 and reducing the production of interleukin-10. It is crucial in the pathway leading to type I immunity. SPP1 interacts with a variety of cell surface receptors widely expressed, which makes it play an active role in many physiological and pathological processes such as wound healing, bone transformation, tumorigenesis, inflammation and ischemia. 27 Urokinase type plasminogen activator (PLAU) gene is located in the 2-band 2 subzone (10q22.2) of the long arm of chromosome 10. Its coding product, urokinase type plasminogen activator (uPA), plays a key role in the process of β-amyloid protein degradation, Urokinase type plasminogen activator (uPA) can promote the conversion of plasminogen into plasmin to degrade aggregated and non aggregated β-amyloid proteins. Conversely, β-amyloid can also induce plau gene to express urokinase type plasminogen activator, thus increasing the plasma content of plasmin, thus blocking the neurotoxicity of β-amyloid protein. 28 App is a multifunctional transmembrane protein, which is highly conserved in the whole evolutionary process. Its regulatory role in development is consistent with the formation of synapses. At present, many studies have shown that app has potential functions in blood coagulation, platelet activation, immune cell response to injury and infection, and has a certain role in tumor formation and development. 29 CXCL1 was initially found in melanoma. CXCL2 plays a role by specifically binding with CXCR2 receptor, which has the function of chemotactic neutrophils, as well as promoting angiogenesis, tumor formation and development. 30 However, in this study, there was no significant difference in CXCL1 gene expression between head and neck tumors and normal head and neck tissues. Although the functions of these 3 genes SPP1, PLAU and APP are different, from the research results, they all have promoting effect on tongue cancer. These 3 genes are likely to be effective targets for the treatment and prevention of tongue cancer.
Cell signaling plays an important role in tumor development. The PI3K/Akt pathway is considered to be the primary pathway for cancer cell survival, which is called the “anti-apoptosis pathway.”
31
PI3 K is an intracellular phosphopeptide kinase discovered by Ito et al. (2010), which is related to the products of the
We used KEGG pathway analysis to identify the most important 47 differentially expressed genes in tongue cancer, and found that
Our results showed that SPP1 was an effective target of tongue cancer, and inhibition of SPP1 may inhibit the PI3K/Akt signaling pathway, which may be an effective treatment for tongue cancer. However, there are still some limitations in our research, such as the limited amount of data in the database of the gene chip, which likely caused the omission of differential genes. We only analyzed 3 tongue cancer gene databases. If we want to get the more accurate results, we should analyze as many tongue cancer gene chips as possible to obtain the differentially expressed genes. In fact, we should analyze the effect of SPP1 gene on the prognosis and survival of tongue cancer patients in as many GEO databases as possible, so as to determine the positive role of SPP1 gene in tongue cancer from the perspective of bioinformatics. In the follow-up study, we will include all the tongue cancer gene chips, and collect as much as possible the information of tongue cancer cases in hospitals to make a systematic analysis. For the simplification of enrichment analysis, we only used KEGG enrichment analysis instead of GO enrichment analysis, 35 which may lead to errors or omissions in the relationship between effective factors and signal pathways between SPP1 and the PI3K/Akt pathways. However, the relationship between SPP1 and PI3K/Akt has not been fully studied. In the KEGG pathway analysis, SPP1 was also related to EMC receptor interaction and focal adhesion. At present, there are few studies on the relationship between SPP1 gene and EMC receptor interaction. The KEGG pathway analysis in this study confirmed that SPP1 is related to EMC receptor interaction. The density and composition of EMC receptor interaction affect the progress of tumor. The rigidity and degeneration of EMC receptor interaction can lead to the proliferation, migration, invasion and angiogenesis of cancer cells. 36 This study also confirmed that the high expression of SPP1 gene was negatively correlated with the prognosis of tongue cancer. Therefore, the high expression of SPP1 in tongue cancer can promote the invasion and poor prognosis of tongue cancer. In recent years, there are many studies and reports about the relationship between SPP1 gene and focal adhesion, mainly in the field of tumor research. Zhu et al. 37 Analyzed the gene bank of endometrial cancer and found that SPP1 gene was related to focal adhesion. Qiu et al. 38 Analyzed the gene pool of gastric adenocarcinoma, and found that SPP1 gene was significantly up-regulated in gastric adenocarcinoma, which promoted the adhesion of gastric cancer cells and shortened the survival time of patients with gastric adenocarcinoma. These provide the directions for further study of SPP1 gene.
Research involving gene chips has become very popular and should be further used in the future. The large number of cases in the gene database enabled us to identify specific target genes. Tongue cancer is a type of oral cancer, but the extent of research is presently not large. In the future, we need to conduct additional studies of tongue and oral cancer. Finding the specific target gene of tongue cancer can provide a more accurate direction for drug development and treatment research.
Footnotes
Abbreviations
Authors’ Note
Our study did not require an ethical board approval because it did not contain human or animal trials.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
