Abstract
We previously showed that microRNA-182 (miR-182) might promote cell proliferation and migration in triple-negative breast cancer (TNBC). This study aimed to investigate circular RNAs (circRNAs) that interact with miR-182 and play important roles in TNBC. Thirty patients with TNBC were enrolled. One pair of tumor and adjacent tissue samples (control) were submitted for circRNA sequencing to establish the expression profile of circRNAs. Concomitantly, circRNAs aberrantly expressed between TNBC and control groups were identified, and these differentially expressed circRNAs (DEcircRNAs) were subjected to Gene Ontology and KEGG pathway enrichment analyses, as well as prediction of interactions with miRNAs. The expression levels of 5 circRNAs interacting with miR-182 were validated using qRT-PCR. Associations between the expression of circUSP42 and clinicopathological features and prognosis were evaluated. A total of 825 upregulated and 1127 downregulated DEcircRNAs were identified between tumor and control groups. Upregulated DEcircRNAs were significantly involved in proteoglycans in cancer, and endocytosis. Downregulated DEcircRNAs were involved in the pathway of resistance to EGFR tyrosine kinase inhibitors. Prediction of circRNA-miRNA interactions showed that hsa_circ_0002032, chr6:131973682-132047340+, hsa_circ_0005982, hsa_circ_0007823 (circUSP42), and hsa_circ_0001777 might act as miRNA sponges for miR-182. qRT-PCR showed consistent results with circRNA sequencing data (P < 0.05). Downregulation of circUSP42 was significantly associated with lymph node metastasis (P = 0.005) and advanced clinical stage (P = 0.032). Furthermore, Kaplan-Meier plots showed that low expression of circUSP42 was closely associated with poor outcome (log-rank test, P < 0.001). Our data suggested that dysregulation of circUSP42 might contribute to the development and progression of TNBC.
Introduction
Breast cancer is the most commonly diagnosed cancer among women worldwide. 1 In particular, triple-negative breast cancer (TNBC), which is defined as a tumor that lacks expression of estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2) is known to be the most aggressive breast cancer subtype. Approximately 15 to 20% of breast cancer cases are categorized as this subtype. 2 During the last few decades, there have been made outstanding advances in breast cancer treatment, resulting in a decreased overall mortality. 3,4 However, due to the absence of specific molecular targets, the prognosis for TNBC remains poor. 5,6 Therefore, it is significant to identify more anticancer molecular targets for the treatment of TNBC.
Recent advances in sequencing technology have revealed circular RNAs (circRNAs) as a new class of non-coding RNAs (ncRNAs). 7 circRNAs are a class of highly stable noncoding RNAs that are widely expressed in mammals. 8 Moreover, it has been confirmed that circRNAs have pre- and post-transcriptional regulatory functions in mammalian cells, 9,10 abundance changes of which have been reported to adversely influence gene expression. 11,12 With the development of RNA-Seq technology and bioinformatics, circRNAs have been demonstrated to be associated with disease progression, including cancers. 13 -15 However, the function of circRNAs in the progression of TNBC has been rarely reported. 4,16
Our previous study explored the function of miRNA-182 on the behavior of TNBC cells and demonstrated that miR-182 might promote cell proliferation and migration in TNBC via the downregulation of forkhead box F2 (FOXF2). 17 In this study, using tumor tissues obtained from patients with TNBC, we performed circRNA sequencing to establish the circRNA expression profile and identify circRNAs aberrantly expressed in TNBC. Accordingly, we identified circRNAs, including circUSP42 that interacts with miR-182, and verified their expression levels by qRT-PCR analysis. Further analysis using the target miRNA prediction software showed that circUSP42 might have a possible binding site for miR-182-5p at its 371-377 bp. Herein, we contemplated that the expression of tumor circUSP42 might be valuable in the management and prognosis estimation of patients with TNBC in a clinical setting. In this study, we further evaluated the associations between the expression of circUSP42 and clinicopathological features and prognosis.
Materials and Methods
Patients and Samples
A total of 30 patients with TNBC who underwent modified radical resection in our hospital were enrolled in this study from January 2010 to December 2015. None of the included patients received radiotherapy or chemotherapy before surgery. Pairs of tumor tissue and adjacent tissue samples (control) were collected during surgery and confirmed by histopathology. The detailed patient characteristics are displayed in Supplementary Table 1. Samples were stored at -80°C until RNA extraction. Among them, circRNA sequencing was performed in one sample pair, whereas 30 pairs were used for circRNA validation, as well as for identifying correlations between the expression of circRNAUSP42 and TNBC clinic-pathological parameters and outcome. Our experiments were approved by the ethics committee of Shanghai Changning Maternity & Infant Health Hospital(Ethics approval number: CNFBLLQKW-2016003). Informed consent was obtained from all patients.
RNA Extraction
Total RNA was isolated from tissue samples. RNA concentration was measured using NanoDrop ND-1000 (Thermo Fisher Scientific, Waltham, MA, USA). The OD260/OD280 value was used as an indicator of the purity of RNA, with the OD260/OD280 value ranging from 1.8 to 2.1 being considered as qualified purity. The integrity and potential contamination of RNA samples were detected through agarose gel electrophoresis.
Library Preparation for circRNA Sequencing
High throughput sequencing was provided by CloudSeq Biotech Inc. (Shanghai, China). Briefly, 5 µg total RNA was pretreated to enrich circRNAs using the CircRNA Enrichment Kit (Cloud-seq Inc, USA). RNA libraries were constructed by using pretreated RNAs with the TruSeq Stranded Total RNA Library Prep Kit (Illumina, San Diego, CA, USA) according to the manufacturer’s instructions. Libraries were controlled for quality and quantified using the BioAnalyzer 2100 system (Agilent Technologies). Libraries were denatured as single-stranded DNA molecules, captured on Illumina flow cells, amplified in situ as clusters and finally sequenced for 150 cycles on an Illumina HiSeq 4000 Sequencer.
Quality Control and Genome Alignment
Paired-end reads harvested from sequencing were quality controlled using Q30. Then, 3′ adaptor-trimming and removal of low quality reads were performed using the cutadapt software (v1.9.3). 18 Reads were aligned to a reference genome through alignment of spliced transcripts to a reference (STAR) software. 19
Annotation and Bioinformatics Analyses of circRNAs
Detection and annotation of circRNAs were performed with the DCC software(https://github.com/dieterich-lab/DCC). 20 Then, the circBase database 21 and circ2Trait disease database 22 were used to annotate identified circRNAs. Raw junction reads for all samples were normalized by the number of total reads, and log2 transformed. Differentially expressed circRNAs (DEcircRNAs) between tumor and adjacent tissues were identified using thresholds of fold change ≥ 2.0, p-value ≤ 0.05. We used the Heatmap2 package of R for clustering analysis of DEcircRNAs based on the normalized junction reads. Gene Ontology (GO, http://www.geneontology.org) including molecular function (MF), biological process (BP) and cell component (CC), and KEGG pathway enrichment analyses were conducted for the host genes of the DEcircRNAs. A p-value ≤ 0.05 was taken as the threshold of significant enrichment. The circRNA-miRNA interactions were predicted using the miRanda, miRWalk, and Targetscan databases prediction software. 23
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
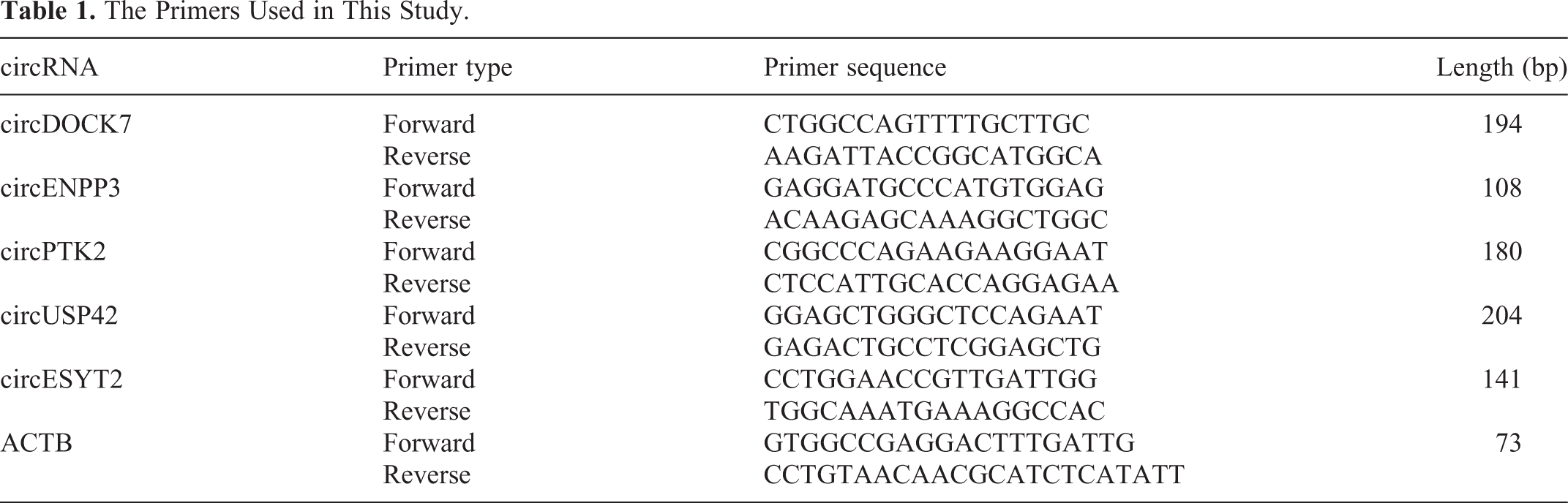
To verify the expression levels of several key DEcircRNAs we performed qRT-PCR analysis. Briefly, total RNA was extracted from tissues of 2 groups using TRIzol® reagent (Invitrogen, USA). cDNA was synthesized using the Superscript II reverse transcriptase (Invitrogen). Amplification was performed on a ViiA 7 Real-time PCR System (Applied Biosystems, USA) using a qRT-PCR SYBR Green master mix (Cloudseq, Shanghai, China). Actin beta (ACTB) was used as the internal control. The relative expression level of circRNA was calculated using the 2−ΔΔCq method. Primers used are shown in Table 1.
The Primers Used in This Study.
Statistical Analysis
Data are shown as mean ± standard deviation and were analyzed using the SPSS 22.0 software. Normality test was conducted using the One-Sample Kolmogorov-Smirnov Test for quantitative data. Normal distribution data were expressed as mean ± standard deviation (SD), whereas non-normal distribution data were expressed as median (interquartile range). Comparisons between 2 groups were conducted by Student’s t test for normal distribution data or Mann-Whitney U test for non-normal distribution data. Qualitative data were expressed as n (%) and compared by Chi-square test. Comparison of the expression of circUSP42 between cancer and normal tissues was analyzed by paired t-test. Kaplan-Meier plot and log-rank test were applied to determine the correlation of the expression of circUSP42 with disease-free survival (DFS) and overall survival (OS). P < 0.05 indicated statistical significance.
Results
Quality Control and Differential Expression Analysis
The Q30 for tumor and control groups was shown to be 90.53% and 90.66% respectively. Following differential expression analysis with thresholds of fold change ≥ 2.0 and p-value ≤ 0.05, we identified a total of 825 upregulated and 1127 downregulated DEcircRNAs between tumor and control groups. A heatmap of DEcircRNAs is shown in Figure 1. Samples in the same group were clustered in a branch.

Profiling of circRNAs in tumor and adjacent tissues in patients with TNBC. (A) Heatmap of differentially expressed circular RNAs. Red represents high expression, whereas green represents low expression. (B) Number of novel circRNAs in upregulated circRNAs and downregulated circRNAs. (C) Distribution of differentially expressed circRNAs (DEcircRNAs) based on their length of nuclear acids. (D) Distribution of DEcircRNAs based on their location on human chromosomes. (F) Counts of DEcircRNAs based on their categories of circle components.
Among the 1952 DEcircRNAs, 411 (21.1%) circRNAs were identified for the first time as novel circRNAs, whereas the other 1541 (78.9%) circRNAs were previously identified (Figure 1B). The majority of DEcircRNAs had a length less than 2000 nucleotides (Figure 1C). These DEcircRNAs were observed to be located on all human chromosomes, with the largest number of DEcircRNAs being found on chromosome 2 (chr2) (Figure 1D). DEcircRNAs were mainly divided into 5 categories, including antisense circRNAs, exonic circRNAs, intergenic circRNAs, intronic circRNAs, and sense overlapping circRNAs. In particular, 83% of all identified DEcircRNAs were demonstrated to be originating from exonic regions (Figure 1E).
GO and KEGG Pathway Analyses
We further performed functional analyses for the host genes of DEcircRNAs. GO analysis showed that the upregulated DEcircRNAs were significantly associated with the BP of organelle organization, protein localization, and cellular component organization, CC of intracellular part, and MF of binding, ion binding, and Ras GTPase binding (Figure 2A). Concomitantly, the downregulated DEcircRNAs were shown to be significantly enriched in the BP related to organelle organization, and cellular macromolecule metabolic process, CC related to intracellular part, and MF related to catalytic activity, and binding (Figure 2B).

Enrichment of the molecular function (MF), biological process (BP), and cell component (CC) terms by upregulated (A) and downregulated (B) differentially expressed circular RNAs.
KEGG pathway enrichment analysis revealed that the upregulated DEcircRNAs were significantly involved in ubiquitin mediated proteolysis, phosphatidylinositol signaling system, proteoglycans in cancer, and endocytosis (Figure 3A), whereas the downregulated DEcircRNAs were involved in pathways of lysine degradation, RNA transport, and EGFR tyrosine kinase inhibitor resistance (Figure 3B).

Enrichment of pathways by upregulated (A) and downregulated (B) differentially expressed circular RNAs.
circRNA-miRNA Interaction Prediction
It has been reported that some circRNAs can act as miRNA sponges to regulate the expression of other related RNAs through miRNA response elements. At present, we identified the interaction of circRNAs and miRNAs. Respectively, miRNA binding sites on DEcircRNAs are listed in Supplementary Table S1. We noted that there were many target sites in circRNAs for a specific miRNA. The top 5 predicted miRNA targets of each DEC are shown in Supplementary Table S2.
Validation of the Expression Levels of Key circRNAs by qRT-PCR Analysis
In this study, we focused on circRNAs that interacted with miR-182, including 3 upregulated circRNAs, hsa_circ_0002032, chr6:131973682-132047340+, and hsa_circ_0005982, as well as 2 downregulated circRNAs, hsa_circ_0007823, and hsa_circ_0001777. We performed qRT-PCR to analyze the expression of these 5 circRNAs in 10 pairs of TNBC and adjacent tissues. In accordance with the results of our bioinformatics analysis, the selected circRNAs were demonstrated to be significantly upregulated or downregulated in TNBC compared with adjacent tissues (Figure 4A, P < 0.05 or P < 0.01). To further investigate the correlation between the expression of circRNAs and clinical information, hsa_circ_0007823 was selected as a candidate because it exhibited the most significant difference. Briefly, hsa_circ_0007823 is 451 bp in length and located at chr7:6150756-6155154+ in the USP42 gene; therefore, we labeled it as circUSP42.
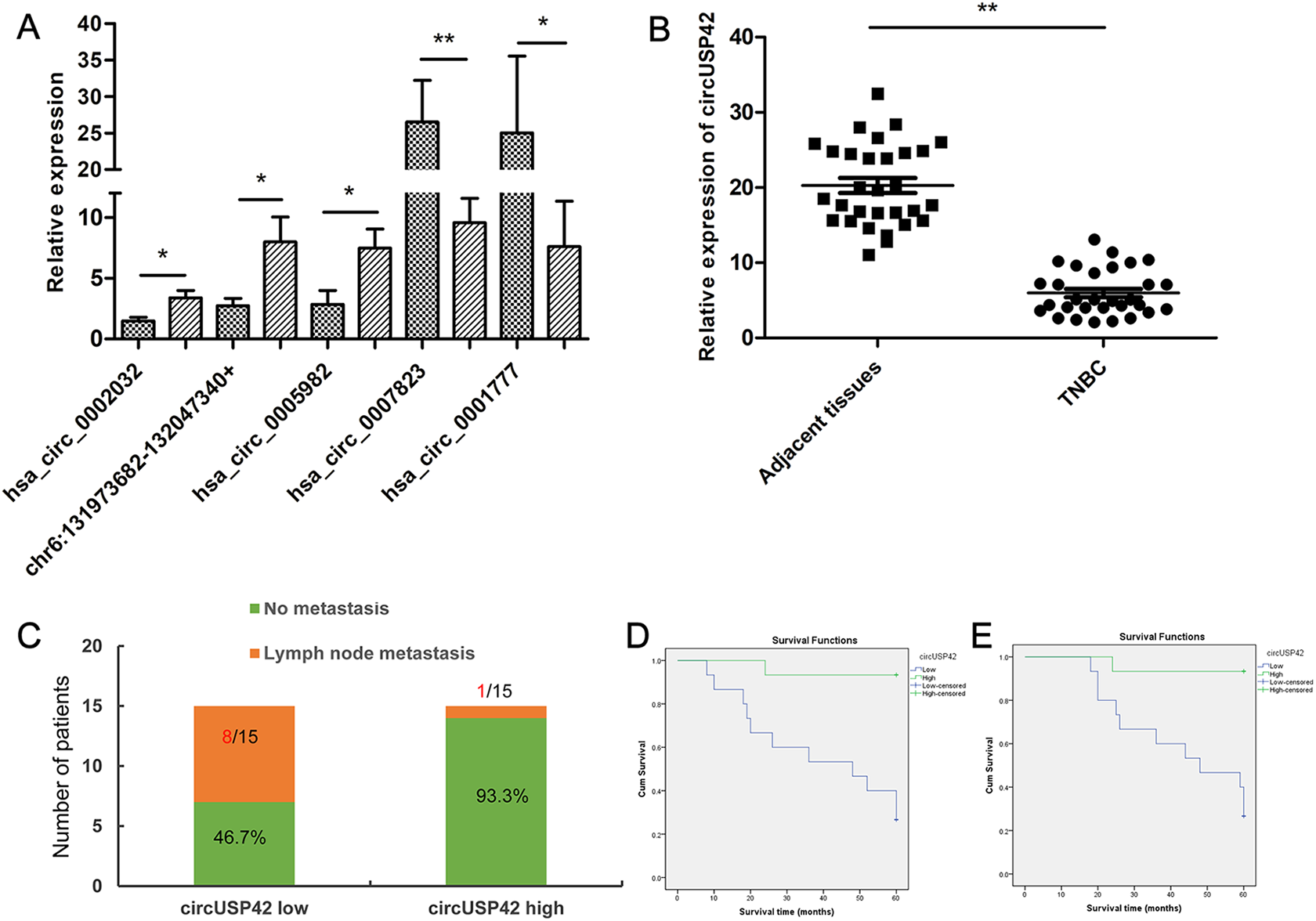
Expression of circUSP42 in TNBC. (A) qRT-PCR analysis of the expression of 5 DEcircRNAs in 10 pairs of TNBC and adjacent normal tissues. (B) qRT-PCR analysis of the expression of circUSP42 in 30 pairs of TNBC and adjacent normal tissues. (C) Analysis of the expression of circUSP42 in patients with TNBC with or without lymph node metastasis. The median circUSP42 value was used as cutoff. (D) Kaplan-Meier survival curves of disease-free survival of patients with TNBC exhibiting low and high expression of circUSP42. The median circUSP42 value was used as cutoff. (F) Kaplan-Meier survival curves of overall survival of patients with TNBC exhibiting low and high expression of circUSP42. The median circUSP42 value was used as cutoff.
circUSP42 Was Decreased in TNBC and Predicted Poor Prognosis
Expression of circUSP42 was validated in 30 pairs of TNBC and adjacent tissues (including the RNA sequencing sample and 10 previous cases). Accordingly, qRT-PCR analysis showed that circUSP42 was significantly downregulated in TNBC tissues compared with adjacent normal tissues (Figure 4B, 20.28 ± 5.44 vs. 5.97 ± 3.08, P < 0.01). Correlation between clinicopathological parameters and the expression of circUSP42 is displayed in Table 2. We observed that downregulation of circUSP42 was significantly associated with lymph node metastasis (P = 0.005) (Figure 4C) and advanced clinical stage (P = 0.032) (Table 2). Furthermore, Kaplan-Meier plots showed that low expression of circUSP42 was closely associated with poor DFS (log-rank test, P < 0.001) (Figure 4D) and poor OS (log-rank test, P < 0.001) (Figure 4F). These data indicated that dysregulation of circUSP42 might contribute to the development and progression of TNBC.
Correlations Between circUSP42 Expression and Clinical Characteristics in Breast Cancer Patients (n = 30).

Discussion
In this study, we established the expression profile of circRNAs in TNBC through circRNA sequencing, and identified circRNAs that were differentially expressed between TNBC tumor and normal tissues. A total of 825 upregulated and 1127 downregulated DEcircRNAs were identified between tumor and control groups. Upregulated DEcircRNAs were shown to be significantly involved in proteoglycans in cancer, and endocytosis. In contrast, downregulated DEcircRNAs were observed to be involved in the pathway of resistance to EGFR tyrosine kinase inhibitors. Moreover, prediction of circRNA-miRNA interactions showed that hsa_circ_0002032, chr6:131973682-132047340+, hsa_circ_0005982, hsa_circ_0007823, and hsa_circ_0001777 might act as miRNA sponges for miR-182.
Proteoglycans are molecular effectors of cell surface and pericellular microenvironments, with multiple functions in angiogenesis and cancer through their ability to interact with ligands and receptors that regulate neovascularization and neoplastic growth. 24 During tumor development, the expression of proteoglycans is markedly modified in the tumor microenvironment, affecting cancer cell signaling, survival, growth, as well as cell adhesion and migration. 25 Proteoglycans are known to be composed of a specific core protein with one or more covalently linked glycosaminoglycan chains, resulting in a high degree of structural and functional complexity. 26 The levels of proteoglycan/glycosaminoglycan expression have been associated with breast cancer growth, invasion, and metastasis. Therefore, proteoglycans have been reported to act as potential pharmacological targets of breast cancer. 27
Endocytosis entails selective packaging of cell-surface proteins, such as receptors of cytokines and receptors of adhesion components, in cytoplasmic vesicles. 28 It has been reported that the endocytic pathway taken by surface receptors could influence the activity of downstream signaling pathways and thereby control cellular responses. 29 On this note, the crosstalk between endocytosis and signaling has been implicated in cancer progression, since alterations in proliferative, survival, and migratory signals have been reported to be necessary for metastasis. 28 In this study, we showed that the upregulated circRNAs were involved in the endocytosis pathway, suggesting that this pathway might be associated with the progression of TNBC.
In contrast, the downregulated circRNAs were shown to be associated with the resistance to epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors. Tyrosine kinase inhibitors have been widely used for the treatment of cancers harboring aberrant activation of EGFR or EGFR mutations. 30 Some cancers have been reported to initially be sensitive to EGFR tyrosine kinase inhibitors; however resistance has been shown to invariably develop later on. 31 There have been various mechanisms suggested to be associated with the resistance to EGFR-tyrosine kinase inhibitors, such as the activation of alternative pathways, such as hepatocyte growth factor (HGF), MET proto-oncogene, receptor tyrosine kinase (c-Met), AXL receptor tyrosine kinase (AXL), impairment of the EGFR- tyrosine kinase inhibitors-mediated apoptosis pathway, ATP binding cassette transporter effusion, and so on. 30 In this study, we speculated that altered expression of some circRNAs might be also implicated in the resistance to EGFR tyrosine kinase inhibitors in TNBC.
MicroRNAs (miRNAs) are known to be posttranscriptional regulators of gene expression, playing specific roles by direct base pairing to target sites within the noncoding region of mRNAs. 32 Recently, it has been reported that some circRNAs might act as miRNA sponges to regulate the expression levels of other related RNAs through miRNA response elements. 33 It has been shown that circRNAs could block the inhibitory effect of miRNAs on the expression of their target genes by making miRNA response elements bind to these miRNAs. 12 Our previous study demonstrated that miR-182 promoted cell proliferation and migration in TNBC. 17 A previous study also reported that miR-182 was upregulated in TNBC tissues and cells, and could promote the proliferation and invasion of TNBC cells. The present study identified several target binding sites for miR-182 in circRNAs, and showed that the expression levels of 5 circRNAs (circDOCK7, circENPP3, circPTK2, circUSP42, and circESYT2) were validated by qRT-PCR analysis. Taken together, we speculated that these 5 circRNAs might act as sponges for miR-182 in regulating the levels of expression of related RNAs, thus playing crucial roles in the development of TNBC. Furthermore, we found that circUSP42 was downregulated in TNBC tumor tissues, and the expression of circUSP42 was negatively correlated with the N and TNM stages. High expression of circUSP42 was revealed to be associated with favorable DFS and OS. Furthermore, circUSP42 high expression was an independent predictor for favorable DFS. In our opinion, there are several possible causes for these results. CircUSP42 might be able to repress tumor progression via the suppression of tumor-like behaviors of TNBC cells, such as inhibition of cancer cell invasion and migration, as well as via interaction with miRNAs, such as miR-182 identified in previous studies. Herein, circUSP42 was demonstrated to probably impede the disease progression or recurrence in patients with TNBC through the inhibition of tumor growth by regulating cancer cell functions, resulting in more favorable survival in patients. This preliminarily observation has demonstrated the value of circUSP42 as a potential biomarker for disease surveillance and prognosis in patients with TNBC.Our study inevitably had some limitations. First, we performed circRNA sequencing using only one case sample. This limitation might have led to research bias in our study, considering the selection of the target circular RNA. However, the results of the differential expression of the predicted circular RNA by sequencing could be further verified by PCR. Therefore, we collected 5 samples of cancer and paracancerous tissue, mixed them and then took a sample for sequencing. In order to ensure the reliability of the study, we further verified our results by PCR analysis, and determined that the expression of circUSP42 was significantly low in 30 samples of TNBC cancer tissues, partially compensating for this limitation in our study. Second, the sample size of the verification dataset was relatively small. Only 30 pairs of samples were included. Third, the downstream signaling of miR-182 and circUSP42 were not investigated in this study. Therefore, the significance of circUSP42 in TNBC and its molecular mechanism still warrant further study.
In conclusion, based on circRNA sequencing of TNBC tissues, thousands of DEcircRNAs were identified between TNBC and control groups. The pathways of proteoglycans in cancer, endocytosis, and resistance to EGFR tyrosine kinase inhibitors were suggested to potentially play important roles in the progression of TNBC through regulation by these circRNAs. Moreover, hsa_circ_0002032, chr6:131973682-132047340+, hsa_circ_0005982, hsa_circ_0007823 (circUSP42), and hsa_circ_0001777 were considered that they might act as miRNA sponges for miR-182. Downregulation of circUSP42 was significantly associated with lymph node metastasis, advanced clinical stage, and closely associated with poor outcome. Our data suggested that dysregulation of circUSP42 might contribute to the development and progression of TNBC.
Supplemental Material
TCRT-19-0477.R1_Supplementary_Table_S2 - circUSP42 Is Downregulated in Triple-Negative Breast Cancer and Associated With Poor Prognosis
TCRT-19-0477.R1_Supplementary_Table_S2 for circUSP42 Is Downregulated in Triple-Negative Breast Cancer and Associated With Poor Prognosis by Jinling Yu, Weida Shen, Jinping Xu, Bo Gong, Beimin Gao and Jiangfan Zhu in Technology in Cancer Research & Treatment
Footnotes
Authors’ Note
Our experiments were approved by the ethics committee of Shanghai Changning maternal and child health care hospital. Informed consent was obtained from all patients.
Acknowledgments
We thank Cloud-Seq Biotech Ltd. Co.(Shanghai, China) for the CirRNA-Seq service, special thanks to Dr. Jinsong Lu for the help during the study.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Highlights
Up- (825) and downregulated (1127) DEcircRNAs identified between tumor and control Five circRNAs, such as circUSP42, might act as miRNA sponges for miR-182 circUSP42 downregulation was significantly associated with lymph node metastasis circUSP42 downregulation was significantly associated with advanced clinical stage
Supplemental Material
Supplemental material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
