Abstract
Objective:
Breast cancer remains the most threatening triggers of cancer death in women. Drug resistance inevitably leads to the weakness of treatment for breast cancer. Macrophages, as one of the most abundant immune cells in tumor immune-infiltrating microenvironment, involves in cell survival, migration, and invasion of breast cancer.
Methods:
In this study, we compared the proportions of macrophages in patients with breast cancer with and without paclitaxel treatment, and investigated the targeted genes associated with macrophages for paclitaxel response. To explore the relationship between drug-related genes and breast cancer prognosis, survival analysis based on the drug-related genes were performed by website of Kaplan-Meier plotter with the threshold of significant P value < .05.
Results:
Compared to the normal samples, we revealed that paclitaxel significantly enhanced the ratio of macrophages in the tumor microenvironment. Furthermore, the expression of 3 drug-related genes (IFT46, PEX11A, and TMEM223) were significantly negatively associated with the proportions of macrophages. And it is worth to notice that PEX11A and TMEM223 were associated with better progression-free survival outcomes of patients with breast cancer. Moreover, PEX11A was associated with longer overall survival time of breast cancer.
Conclusion:
Taken all together, all the findings support to gain a better understanding to the development of more effective therapies targeted with paclitaxel.
Introduction
Breast cancer (BC) is the most common cancer among women worldwide, with a rising trend in aggressive neoplasm in young women. 1,2 In China, BC occupied 12.2% of all newly diagnosed and 9.6% of all cancer mortality of tumor patients, respectively. 3 Reproductive and lifestyle factors, such as age, exogenous hormone, diet, body fatness, and physical activities can influence the risk of BC. 4 -6 Over the past decade, a set of novel therapeutic strategies are developed for diagnosis to treatment 3 ; however, more accurate and effective treatments are still worthy of attention.
Paclitaxel, as an inhibitor of microtubule depolymerization, is an effective chemotherapeutic agent in the therapy for BC. 7 To strengthen the effect of the antibody therapy, combining paclitaxel with lumretuzumab (anti-human epidermal growth factor [EGF] receptor 3; anti-HER3) or pertuzumab (anti-human EGF receptor 2; anti-HER2) is used to treat patients with metastatic BC. Compared with paclitaxel alone treatment, combined solution (eg, paclitaxel plus bevacizumab) prolongs progression-free survival (PFS), but not overall survival (OS). 8 Although accelerating rates of the application of paclitaxel, the paclitaxel-evoked pain syndrome is inevitable, accompanying with paclitaxel-sensitivity and paclitaxel-resistance. 9 -11 Some functional pathways and related genes have investigated to regulate the validation process in paclitaxel-sensitive or paclitaxel-resistant cells. For example, IκBα super-repressor enhanced NF-κB pathway by inhibiting NF-κB DNA binding activity and Mn-SOD expression in paclitaxel sensitivity of BC cells. 12 Upregulating of AKT2 contributes to the increase of resistance to paclitaxel in BC cells. 13
Understanding tumor immune-infiltrating microenvironment is an important link between tumorigenesis mechanism and therapeutic solutions. 14 Many theoretical and experimental evidences have illustrated that the tumor-associated macrophages, as a major component of immune infiltrate, play a crucial role in tumor immune-infiltrating microenvironment. 15 -17 For instance, macrophages improve tumor cell migration and invasion via regulating EGF and vascular endothelial growth factor. 14,18 However, there is little evidence as to how to link the paclitaxel response and paclitaxel-evoked pain syndrome with the macrophages. In this study, we acquire gene expression and clinical data from GDAC FIREHOSE to examine the proportion of macrophages in patients with BC with and without paclitaxel treatment and to capture more characteristic the targeted genes associating with macrophages on paclitaxel response in paclitaxel-treated and without paclitaxel-treated patients with BC.
Materials and Methods
Data Collection
Gene expression and clinical data were acquired from the GDAC FIREHOSE database version 2016_01_28 (http://gdac.broadinstitute.org, RRID: SCR_016213). The bioinformatics method CIBERSORT (RRID: SCR_016955) 19 was used to estimate the proportions of immune cells in the tumor microenvironment. Combining with the immunophenotypic expression profiles provided by CIBERSORT, we selected 6 immune cell types (B cells, T cell CD8, T cells CD4, macrophages, dendritic, neutrophils) as subjects to calculate the proportions in each BC sample by using the R package {“EpiDISH”}.https://labs.genetics.ucla.edu/horvath/CoexpressionNetwork/, RRID: SCR_003302).
Interactome Analysis of Targeted Gene on Paclitaxel Response
To investigate the targeted genes on paclitaxel response, the drug-related response data were obtained from Genomics of Drug Sensitivity in Cancer (GDSC) database (http://www.cancerrxgene.org/, RRID: MGI: 5909258). There are only 12 cell lines (HCC1187, HCC1599, HCC2157, HCC2218, COLO-824, DU-4475, EVSA-T, MRK-nu-1, OCUB-M, MFM-223, CAL-148, BT-474) involved in paclitaxel drug response from GDSC. Furthermore, these cell lines were classified into drug-resistance and drug-sensitivity using SL Forward-Backward methods. The brief description of procedures was as follow: (1) To randomly classify one cell line into drug-resistant cells. If fold change (FC) value of the average 50% inhibitory concentration (IC50) between the drug-resistant and drug-sensitive cell line was of > 2, it proceeded to the next step, otherwise randomly restarted. (2) To randomly capture a drug-sensitive cell line into the drug-resistant cell line, and then calculate the FC value of the IC50 between the 2 cell lines. If the FC value was greater than that in step (1), we moved to the next; otherwise, returned to the drug-sensitive cell line. (3) To repeat step (2) for screening all the drug-sensitive cell lines. (4) To randomly selected a drug-resistant cell line into sensitive cell lines, and then calculate the FC value of the IC50 between 2 cell lines. If the FC value was greater than that in step (1), it moved to the next; otherwise, returned to the drug-resistant cell line. (5) To repeat step (4) for screening all the drug-sensitive cell lines. After clarifying the drug-resistant and drug-sensitive cell lines, we screened all drug-related genes in expression profiles of these cell lines from GDSC and calculated the FC values and P values of t test. Finally, the genes with P value < .05 and (FC) > 2 or FC < 1/2 were considering as the potential-targeted gene on paclitaxel response.
The Effect of Macrophages on Drug-Related Genes
To further evaluate the effect of macrophages on drug-related genes, Pearson correlation coefficients (PCC) between drug-related gene expression and macrophage ratio were calculated by the {cor} and {cor.test} functions in the R environment. And the genes with PCC < −0.5 or PCC > 0.5 were considered as associating with macrophages.
Prognostic Analysis of Drug-Related Genes
To explore the relationship between drug-related genes and BC prognosis, survival analysis based on the drug-related genes were performed by website of Kaplan-Meier plotter (https://kmplot.com/analysis/index.php?p=service&cancer=breast) with the statistically significant P value< .05. The Kaplan-Meier plotter is an online database based on expression data and clinical data from BC, gastric cancer, lung cancer, and ovarian cancer. The patient samples were divided into 2 groups according to the median expression of the gene. Then, Kaplan-Meier survival curve was plotted to obtain the prognostic value of drug-related genes. Briefly, the drug-related genes were uploaded into the database respectively to get the Kaplan-Meier curve.
Results
Paclitaxel Upregulated Tumor-Infiltrating Macrophages
To investigate the effects of paclitaxel on the immune-infiltrating microenvironment of patients with BC, we obtained the frequency of 6 immune cells (Figure 1) in 13 BC-related patients who received paclitaxel in the GDAC FIREHOSE database (Table 1) and one normal control nontumor sample (TCGA-BH-A0B3-10).
Details of Patients With BC With Paclitaxel.

Abbreviation: BC, breast cancer.

The frequency of 6 immune cells in BC-related tissues. All 13 BC-related tissues and control (TCGA-BH-A0B3-10) are obtained from the GDAC FIREHOSE. BC indicates breast cancer.
To further clarify the ratio variations of immune-related cells, we randomly selected 12 adjacent nontumor samples in patients with BC without taking paclitaxel treatments to match the cohort size of 13 tumor samples. The proportion of macrophages was upregulated in paclitaxel-treated patients with BC. The detailed results showed that mean proportion in patients was 0.6168 but in normal samples was 0.4957 (Table 2, Figure 2A). On the other hand, the mean proportion of other immune cell types was not changed so much, such as B cells in patients was 0.0609 and in normal samples was 0.0670; T cells CD4 even reduced, in patients it was 0.2318 but in normal samples it was 0.2555. Besides, we randomly selected 50 BC tissues and 50 adjacent nontumor samples in patients with BC without paclitaxel-treatment. No obvious difference was detected (Figure 2B). As a consequence, we may deduce that paclitaxel-induced macrophage infiltration in the tumor microenvironment of BC. The results showed that paclitaxel upregulated the proportion of macrophages, compared to the control.
Details of Microenvironment Between BC Patients’ Tissues With Paclitaxel and Normal Samples Without Paclitaxel.

Abbreviation: BC, breast cancer.

The effect of paclitaxel treatment on immune cells. A, Comparison of breast cancer tissue and adjacent nontumor samples in patients with BC with paclitaxel treatments. B, Comparison of breast cancer tissue and adjacent nontumor samples in patients with BC without paclitaxel treatments.
Potential Drug-Related Genes
According to gene expression profiles between cell lines of drug-resistant (HCC2157, HCC2218, COLO-824, DU-4475, EVSA-T, MFM-223, CAL-148, BT-474) and drug-sensitive (HCC1599, HCC1187, MRK-nu-1, OCUB-M) (Table 3), we screened 62 potential drug-related genes. Further, the correlation between gene expression level and the ratio of macrophages within each sample allow us to obtain drug macrophages–related genes. Three targeted genes were found, including intraflagellar transport 46 (IFT46), peroxisome biogenesis factor 11A (PEX11A), and transmembrane protein 223 (TMEM223). The expression of these genes were significantly negatively associated with the proportions of macrophages (γIFT46 = −0.604; γPEX11A = −0.698; γTMEM223 = −0.616; Figure 3).
Details of GDAC Cell Lines.
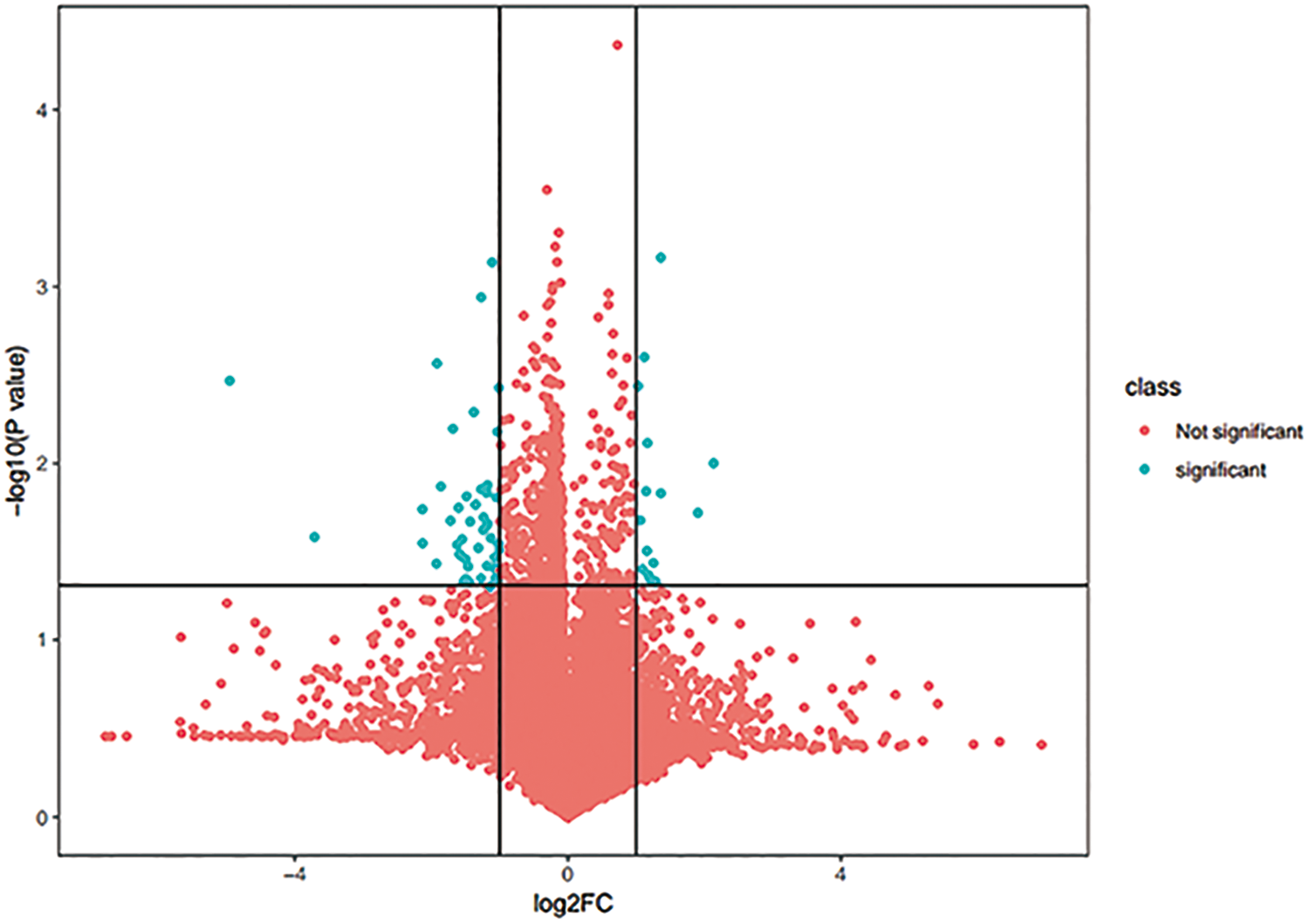
Potential targeting genes on paclitaxel response. Three targeted genes were found, including intraflagellar transport 46 (IFT46), peroxisome biogenesis factor 11A (PEX11A), and transmembrane protein 223 (TMEM223). The expression of these genes were significantly negatively associated with the proportions of macrophages (γIFT46 = −0.604; γPEX11A = −0.698; γTMEM223 = −0.616).
Prognostic Analysis of Drug-Related Genes
Kaplan-Meier plotter was performed to explore the relationship between drug-related genes and BC prognosis. The results revealed that PEX11A and TMEM223 were associated with longer PFS of patients with BC, and the log-rank test P value were 3.3e-08 and 1.8e-07. Peroxisome biogenesis factor 11A was associated with longer OS of BC and the log-rank test P value was .002. However, the gene IFT46 had not been found in Kaplan-Meier plotter online database (Figure 4).
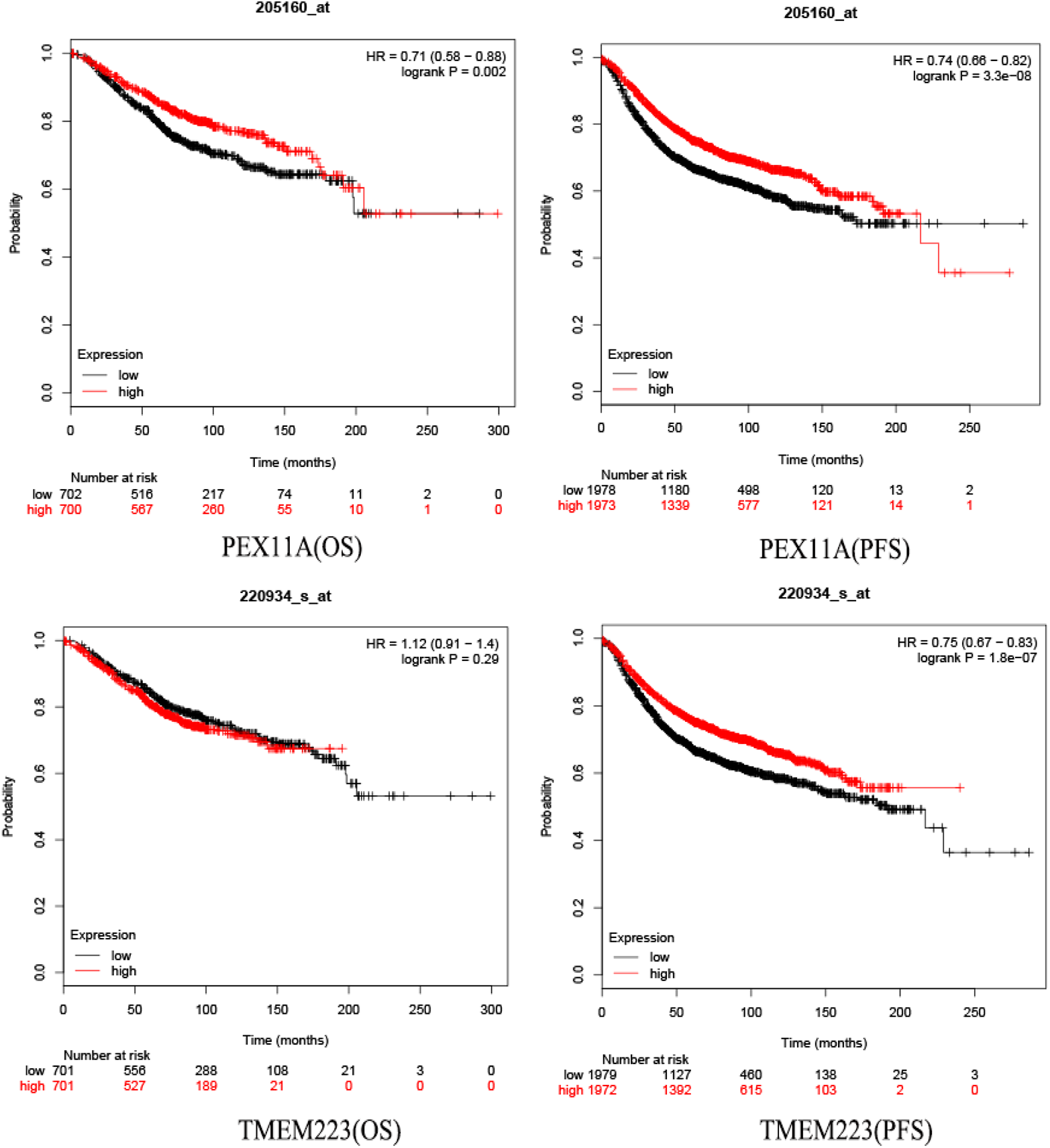
Prognostic analysis of drug-related genes. Kaplan-Meier plotter database prognostic analysis of PEX11A, TMEM223 (overall survival [OS]; progression-free survival [PFS]). PEX11A indicates peroxisome biogenesis factor 11A; TMEM223, transmembrane protein 223.
Discussion
Paclitaxel is a frontline antineoplastic agent widely applied in the treatment of BC, ovarian cancer, gastric cancer, and lung cancer. 20 -23 In this study, compared with nonpaclitaxel-treated patients with BC, paclitaxel can upregulate the proportion of macrophages in the tumor-infiltrating cells of tumor microenvironment in paclitaxel-treated patients with BC. Similar results have shown that paclitaxel may restore tumor-induced macrophages production of proimmune factors (eg, immunostimulatory cytokine IL-12) in tumor-bearing host. 24 Low-dose paclitaxel suppresses the M2 macrophages by acting on Toll-like receptor 4, but induces the M1 macrophages in gastric cancer. 22
However, paclitaxel-induced peripheral sensory neuropathy is well established and is a serious problem that requires chemotherapy in combination with paclitaxel.
9,25
Previous study have shown that the paclitaxel-induced mechanical allodynia is due to the increase of macrophages induced by high-dose paclitaxel, an
The 3 drug-related genes (IFT46, PEX11A, TMEM223) screened were significantly negatively correlated with macrophages in BC. Besides, PEX11A and TMEM223 were associated with longer PFS of patients with BC. Peroxisome biogenesis factor 11A is associated with prolonged OS in BC. The intraflagellar transport (IFT) system has been well established to connect kinesin motor protein (linked by IFTB) and dynein motor protein (linked by IFTA). 31 The transmembrane transport protein IFT46 is a member of IFTB complex and is involved in the biosynthesis in cell organelles, and the maintenance and transport of IFT proteins. 31,32 However, few studies have investigated the relationship between IFTB complex and tumor-infiltrating macrophages. This is the first study that IFT46 has been linked to paclitaxel-responsive macrophages of BC. Peroxisomes are highly involved in cellular functions (eg, reactive oxygen species [ROS] metabolism). 33 Peroxisome biogenesis factor 11A is a factor of peroxisomes, which plays a key role in the regulation of peroxisome proliferator-activated receptor alpha in ROS metabolism and lipid metabolism. 33,34 Reactive oxygen species is involved in regulating the occurrence of tumor-associated macrophages and alternatively activating macrophages (M2) differentiation. 35 Therefore, PEX11A may regulate M2 macrophages via ROS metabolism under the combination of chemotherapy and paclitaxel. Transmembrane protein 223 as a transmembrane protein, we assumed that it may be involved in the phagocytosis of macrophages. 36,37 However, there is not enough experimental evidence or literature to evaluate the response of drug-related genes to macrophages in the tumor microenvironment.
In summary, our preliminary evidence suggested that paclitaxel can increase the proportion of tumor-infiltrating macrophages in BC, which may mediate the regulation of drug-related genes associated with macrophages. This study provides a potential basis of alleviating paclitaxel-induced peripheral sensory neuropathy for chemotherapy.
Footnotes
Abbreviations
Authors’ Note
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China (No. 81972453, No. 81972597, No. 81602471 and No. 81672729), Zhejiang Provincial Natural Science Foundation of China under Grants (No. LY19H160055, LY19H160059).
