Abstract
Cancer stem cells, a special subgroup of cancer cells, have self-renewal capabilities and multidirectional potential, which may be reprogrammed from the dedifferentiation of cancer cells, contributing to the failure of clinical treatments. Esophageal adenocarcinoma grows in an inflammatory environment stimulated by deoxycholic acid, an important component of gastroesophageal reflux content, contributing to the transformation of esophageal squamous epithelium to the precancerous lesions of esophageal adenocarcinoma, that is, Barrett esophagus. In the present study, deoxycholic acid was used to investigate whether it could induce the expression of reprogramming factors Krüppel-like factor, OCT4, and Nanog; the transformation to cancer stem cells in esophageal adenocarcinoma; and the involvement of the interleukin-6/signal transduction and activation of transcription 3 inflammatory signaling pathway. OE33 cells were treated with deoxycholic acid (250 μM) for 0 hour, 3 hours, 6 hours, and 12 hours before evaluating the messenger RNA expression of Krüppel-like factor, OCT4, Nanog, interleukin-6, and Bcl-xL by reverse transcription-quantitative polymerase chain reaction. Interleukin-6 protein was detected by enzyme linked immunosorbent assay, while signal transduction and activation of transcription 3, phosphorylated signal transduction and activation of transcription 3, Krüppel-like factor, and OCT4 were detected by Western blot. Signal transduction and activation of transcription 3 small interfering RNA and human recombinant interleukin-6 were used to treat OE33 cells and to detect their effects on Krüppel-like factor, OCT4, Nanog, CD44, hypoxia-inducible factor 1-α, and Bcl-xL expression. Results showed that deoxycholic acid promotes the expression of reprogramming factors Krüppel-like factor and OCT4, which are regulated by the interleukin-6/signal transduction and activation of transcription 3 signaling pathway. Deoxycholic acid has a malignancy-inducing effect on the transformation of esophageal adenocarcinoma stem cells, improving the antiapoptotic ability of tumors, and increasing the malignancy of esophageal adenocarcinoma. Deactivating the regulatory signaling pathway of interleukin-6/signal transduction and activation of transcription 3 and neutralizing deoxycholic acid may be novel targets for improving the clinical efficacy of esophageal adenocarcinoma therapy.
Introduction
In recent years, an increasing incidence of gastroesophageal reflux disease (GERD) has been observed, accompanied by more reported cases of Barrett esophagus (BE) and esophageal adenocarcinoma (EAC). The incidence of esophageal cancer was ranked sixth in the world using the global burden of disease study estimation method of absolute years of life lost in both sexes between 2007 and 2017. 1 Barrett esophagus is commonly agreed upon by academic societies to be a precancerous lesion of EAC; it could be critically diagnosed by its characteristic pathological changes, wherein the distal esophageal squamous epithelial cells are replaced by intestinal columnar epithelial cells. Alarming data reported from a large clinical epidemiological study in 2011 showed that about 0.12% of BE patients would develop into EAC every year. 2 The current developmental disease model of EAC is described by multiple steps of progression, starting from normal squamous epithelium of the esophagus to GERD, BE, low-grade dysplasia (LGD), high-grade dysplasia (HGD), and finally EAC. Since GERD is an important initiator of the disease, exposure of the gastroesophageal reflux to esophageal cells is believed to be the major cause. In the past, the role of gastric acid reflux exposure has been overemphasized in the pathogenesis of EAC, until the use of proton pump inhibitors for the past few decades failed to decrease the increasing number of disease cases. Instead, it created a 6-fold increase in the incidence of EAC as reported in Western countries. 3 These observations suggest that acid reflux is not the only factor for the progression of GERD to EAC. The composition of gastroesophageal reflux is complex, containing mixtures of gastric acid, pepsin, bile acid, and so on. In this regard, the elucidation of the deterministic pathogenesis of EAC is urgent, as the cases of BE and EAC increase uncontrollably.
It is currently suggested that bile acid, not gastric acid, is an important component of reflux leading to the formation and progression of BE and EAC. Bile acids, with the function of fat emulsification, are structurally classified into 2 groups: free bile acids, including cholic acid, deoxycholic acid (DCA), goose deoxycholic acid, and a small amount of licholic acid; the other includes the conjugated products of bile acid with glycine or taurine, called conjugated bile acids. Bile acid concentration from the reflux of patients with BE usually ranges from 3 to 820 μM, with a median of 180 μM, 4 but in some BE patients, it may reach as high as 6400 μM. 5 In the rat reflux model, it was found that gastric acid alone could not induce the formation of BE; on the contrary, bile acid alone could produce it. 6 Shen et al 7 successfully used a DCA-induced immortal esophageal squamous epithelial cell line et-1a to undergo reprogramming to express CDX2 and MUC2, markers of intestinal epithelial cells. Huo et al 8 treated BE cell lines with DCA and ursodeoxycholic acid (UDCA) and found that DCA caused oxidative stress reaction and subsequent cell DNA damage, while hydrophobic UDCA did not damage the cell DNA. Therefore, unconjugated bile acid DCA may be an important determinant in bile components that cause the transformation of esophageal squamous epithelial cells into BE and eventual carcinogenesis. 9 Studies have shown that DCA can activate the BMP4 pathway to induce esophageal squamous epithelial cells to reprogram and transform into BE. 10 Although DCA was shown to have a certain malignant induction effect on esophageal cells, the specific mechanism by which DCA induces the malignant transformation process of esophageal squamous epithelial cells to form BE and EAC remains unclear. It is speculated that DCA induces the malignant transformation of EAC through the activation of stem cell–associated gene expression. Because stemness marker genes, to some extent, can reflect the existence and degree of activity of stem cell–associated genes, several stem cell–associated genes were chosen in this experiment to detect the existence and activity of cancer stem cells (CSCs).
Octamer-binding transcription factor 4 (OCT4) is a member of the pit, oct, unc (POU) transcription factor family, that is, mainly expressed in embryonic stem cells, reproductive stem cells, and undifferentiated embryonal carcinoma. It plays an important role in maintaining embryonic stem cell pluripotency and self-renewal. In fact, OCT4 also plays a role in maintaining the pluripotency of adult stem cells, which is involved in its self-renewal ability and a sign of their pluripotency. Moreover, as a transcription factor, OCT4 is also necessary for somatic reprogramming. Octamer-binding transcription factor 4 directly binds to SOX2 (sex-determining region Y-box 2) to trans-activate target genes. OCT4 and SOX2 target their enhancer regions by combining the composite element SOX-OCT, thus regulating their own transcription activities. 11 These stemness transcription factors OCT4, SOX2, and Nanog can jointly occupy promoter regions of about 350 genes in the genome, among which OCT4 occupies more than 90% of these regions, indicating that OCT4-SOX2-Nanog is a key cascade of stemness and OCT4 occupies an important core position. Studies have found that downregulating the expression of OCT4 can reduce the tolerance of pancreatic cancer to chemotherapy drugs and block the signal transduction of stem cells in pancreatic cancer stem cells. 12 In liver cancer, it has also been found that inhibition of OCT4 can reduce both tumor resistance to chemotherapy drugs and the stemness of tumor cells. 13 These aforementioned facts indicate that OCT4 is an important regulator of CSCs and is crucial in the process of cell reprogramming.
Krüppel-like factor (KLF4) is a member of the Spl/Kruppel zinc finger transcription factor family and plays an important role in the process of gastrointestinal morphogenesis, intestinal mucosa formation, and other related functions. Krüppel-like factor is also involved in the physiological processes of cell proliferation, differentiation, and the cell cycle. The process of adult cell reprogramming to form induced pluripotent stem cells (iPSCs) is usually accompanied by the overexpression of 4 Yamanaka transcription factors: KLF4, OCT4, SOX2, and MYC. Moreover, it has been found that these 4 Yamanaka transcription factors have successfully induced osteosarcoma cells and breast cancer cells into osteosarcoma stem cells and breast CSCs, 14 -16 respectively. Krüppel-like factor is a stem cell gene involved in inducing adult cells for reprogramming into iPSCs and promoting the transformation of tumor cells into CSCs. On the other hand, KLF4 is also thought to be associated with inflammation. Studies have found that KLF4 can be activated by bile acids and KLF4 can increase the transcriptional activity of MUC2 and CDX2, indicating the potential role of KLF4 in the development of BE. 17 Other studies have shown that after inhibiting the Notch1 signaling pathway in esophageal squamous epithelial cells, upregulation of KLF4 can induce the transformation of esophageal squamous epithelium into BE-like metaplasia. 18
Nanog, a member of the antennapedia (ANTP) homologous gene family, is a gene specifically expressed in embryonic stem cells. Its main function is not only to help the self-renewal and maintenance of the undifferentiated state of embryonic stem cells but also to promote their proliferation. Nanog was found to be expressed in cells of seminoma, breast cancer, human fetal germ cells, testicular carcinoma in situ, and germ cell tumors. The higher the tumor grade, the stronger the Nanog expression, and the worse the clinical prognosis. Moreover, high-grade tumors also showed characteristics of embryonic stem cells. The insulin-like growth factor (IGF) pathway is linked to the self-renewal potential of cells, where its components are downstream molecules of Nanog. In Nanog+ cancer stem cells, IGF2 and IGF receptors are upregulated, while silencing Nanog inhibits IGF1 receptor expression. 19 Nanog has been reported as a transcription factor of pluripotent stem cells in many studies on CSCs and it was also recently discovered that it can reprogram somatic cells into a state similar to that of embryonic stem cells.
CD44 is an important adhesive molecule, which is a transmembrane protein of the cell membrane and is widely distributed on the surface of lymphocytes, monocytes, endothelial cells, and other cells. 20 Under normal conditions, CD44 expression on the cell surface is in a relatively static state, but it is overexpressed in tumor cells. The extracellular functional region of CD44 can degrade the extracellular matrix hyaluronic acid, so that tumor cells could get rid of the intracellular and stromal domains. At the same time, the intracellular domain could connect with cytoskeletal proteins to improve the activity of tumor cells, thus promoting tumor invasion and metastasis. Therefore, tumor cells with a high expression of CD44 also had a high potential for invasion and migration. In addition, as a marker of many tumor stem cells, CD44 was often used to classify tumor stem cells. Recently, it has been found in liver cells that activation of the signal transducer and activator of transcription 3 (STAT3) pathway could upregulate CD44, which could activate the AKT pathway, and finally remove the inhibition of P53 gene on stem cells, thereby promoting liver cancer formation. 21 Overall, CD44 is a key marker in CSCs. 22
Hypoxia-inducible factor 1-alpha (HIF-1α) is a regulatory factor of hypoxia response, especially the hypoxia response switch, enabling the body to start and adapt to a series of hypoxia reactions. Because of its high dependency on hypoxia, HIF-1α is rapidly degraded under constant oxygen conditions. Along with the reduction in oxygen concentration, the expression and activity of HIF-1α could increase with the enhancement of cell viability in the case of local hypoxia and ischemia, with the subsequent generation of a large number of blood vessels. 23 Hypoxia exists widely in solid tumors; HIF-1α plays an important role in the process of tumor formation and development by enabling tumor cells to survive, adapt, and proliferate in that stressful state. In addition to regulating cellular adaptation to hypoxia, HIF-1α could affect the growth of tumor cells by controlling the activity downstream genes and altering the metabolism of tumor cells by accelerating vascular endothelial growth factor transcription or by improving glucose transport and glycolytic enzyme activities. Furthermore, as a hallmark of tumors, HIF-1α can induce gene expression of stem cells, regulate tumor stem cell renewal, and moderate multipotent signaling pathways. 24 Researchers have found that the activation of glioma stem cells is involved in the activation of HIF-1α, whose activation could inhibit the differentiation of stem cells. However, the knockout of HIF-1α eliminated the proliferation of anoxia-mediated glioma stem cells. 25 The above findings indicate the important modulating function of HIF-1α in tumor stem cells, which made it a key marker of tumor stem cells.
Bcl-xL is an antiapoptotic protein, a member of the Bcl-2 family of proteins. Members of this family of proteins are key regulatory factors for the apoptosis and survival of cells. They regulate the permeability of the mitochondrial membrane, balance the release of cytochrome C from mitochondria into the cytoplasm, and control the release of apaf-1 into the cytoplasm. The caspase recruitment domain (CARD) domain and C-terminal domain of apaf-1 can combine with caspase-9 and cytochrome C to form apoptotic bodies, respectively. When caspase-3 is activated by apoptosis, the caspase cascade reaction is activated, which in turn initiates cell death. Therefore, proteins of the Bcl-2 family are particularly important for the regulation of apoptosis. Members of this family of proteins are divided into antiapoptotic and pro-apoptotic groups, according to their functions. The antiapoptotic proteins of the Bcl-2 family mainly include Bcl-2, Bcl-xL, Bcl-w, and Mcl, while the pro-apoptotic proteins mainly include Bax, Bak, Bad, Bid, and Bim. Tumor cells generally express antiapoptotic proteins of the Bcl-2 family at high levels and reduce the expression of pro-apoptotic proteins of the Bcl-2 family, which attenuate the levels of cytochrome C and apaf-1 entering the cytoplasm and reduce the inhibition of apoptotic body formation. These actions prevent the cells from entering the cascade to activate the caspase reaction and inhibit the initiation of programed apoptosis, thus playing an antiapoptotic role and increasing the survival of tumor cells.
It is well known that EAC is associated with inflammatory tumors, and the inflammatory microenvironment plays an important role in the formation and development of BE and EAC. Signal transduction and activation of transcription 3 is a key molecule related to inflammation, such as nuclear factor κ-light chain enhancer of activated B cells (NF-κB), which functions in protein tyrosine phosphorylation signal transduction pathways. Cytokine interleukin-6 (IL-6) can activate downstream JAK kinase, which further phosphorylates STAT3 to form the active form phosphorylated-signal transducer and activator of transcription 3 (pSTAT3; Tyr705). The pSTAT3 dimer enters the nucleus and targets the expression of relevant downstream genes, playing a role in regulating cell proliferation, differentiation, apoptosis, immune escape, and other functions. 26 In addition, the STAT3 pathway plays an important role in maintaining the pluripotency of iPSCs and embryonic stem cells by mediating inflammation and tumorigenesis. Immunohistochemical results showed that BE and EAC tissues express more pSTAT3 located in the nucleus than normal esophageal squamous epithelial tissues. 27 Under stimulation with DCA, BE cell lines of high malignancy express more pSTAT3 and its downstream antiapoptotic protein Mcl than that of BE cell lines with low malignancy. 28 Therefore, the activation of the STAT3 pathway may promote the malignant progression of esophageal squamous epithelia to BE and EAC.
In summary, CSCs have a special existence in cancers, which may be the origin of tumorigenesis or the cause of the current therapeutic failure to tumors. OCT4, KLF4, Nanog, CD44, and HIF-1α are typical markers of CSCs. At present, the source of CSCs is controversial, but the development, growth, and survival of any cell cannot be separated from its microenvironment. Moreover, EAC is caused by the long-term progression of GERD, BE, LGD, and HGD. Thus, gastroesophageal reflux is an important environmental inducing factor, among which the reflux of DCA has also attracted attention in recent years. Therefore, in this study, we first explored the expression of STAT3 and other stemness-related genes in normal esophageal and EAC cell lines under the stimulation of DCA. This study aimed to explore whether DCA can affect the expression of reprogramming factors KLF4, OCT4, and Nanog and whether it can induce the malignant transformation of EAC cells into EAC stem cells. Specifically, whether IL-6/STAT3, an inflammatory signaling pathway, acts as a signal bridge in EAC is also worth further study.
Materials and Methods
Cell Culture
Human EAC, OE33, and normal esophageal HEEC cell lines were purchased from Guangzhou Jenniobio Biotechnology Co Ltd. The complete culture medium included 90% Dulbecco’s Modified Eagle Medium (DMEM) basic (1X; GIBCO, Thermo Fisher Scientific), 10% fetal bovine serum (TBD), and 1% penicillin and streptoMYCin (Tianjin Hao Yang Biological Manufacture CO). Fetal bovine serum was filter-sterilized using a 0.22-µm syringe-driven filter (JET BIOFIL), and the cells were cultured in an incubator (MEMMERT) at 37 °C, 95% humidity, and 5% CO2.
Cell Treatment
Cells were grown in culture wells to reach 80% density before they were treated with DCA (Yuanye bio-technology). The treatment protocols were designed considering the conditions used in previous in vitro cell tests to avoid cell lysis. Previous studies indicated treatment of immortalized esophageal squamous epithelial cell lines with 200-μM DCA for 2 to 12 hours did not affect the cell viability, but it was decreased by 38% and 51% at 18-hour and 24-hour treatment, respectively. 7 Treating the Bar-t cell line of BE for 12 hours with a DCA concentration less than 200 μM did not affect cell activity, but treatment at 300 μM slightly inhibited cell activity by less than 20%. 29 Therefore, in this study, the DCA concentration was prepared at 250 μM with a maximum treatment time of 12 hours to minimize the influence on cell activity and apoptosis. In the first part, HEEC and OE33 were treated with STAT3 small interfering RNA (siRNA) and DCA. In the second part, OE33 cells were stimulated with DCA for 0 hour, 3 hours, 6 hours, and 12 hours before RNA extraction, after which protein extraction was performed. In addition, 10 ng/mL and 100 ng/mL concentrations of recombinant human 4IL-6 (Beyotime) were prepared to stimulate OE33 cells for 24 hours. Cells treated with DMEM for 24 hours were used as a blank control group for RNA extraction and protein extraction.
Gene Silencing Using siRNA
Transfection procedures were performed according to the instructions provided by the manufacturer (RIBOBIO). The transfection agents used were riboFECTTMcp buffer and riboFECTTMcp reagent, also according to the manufacturer’s instructions. Small interfering RNA STAT3 (RIBOBIO; si-h-STAT3_001: GATACGACTGAGGCGCCTA) was used to knock down the expression of STAT3 while the cells are in the logarithmic growth stage, at a cell density of 50%. This treatment lasted for 48 hours.
RNA Extraction and Reverse Transcription-Quantitative Polymerase Chain Reaction
The RNAsimple Total RNA Kit (TIANGEN) was used to lyse cells and extract the total RNA content. The extraction procedure was conducted according to the operation instructions provided by the manufacturer (TIANGEN). The reaction system was carried out using a fluorescence quantitative polymerase chain reaction instrument (BIONEER). Primer information is shown in Table 1.
Primer Information.

Abbreviations: HIF-1α, hypoxia-inducible factor 1-α; IL-6, interleukin-6; Krüppel-like factor; STAT3, signal transducer and activator of transcription 3.
Detection of IL-6 via ELISA
After OE33 cells were treated with DCA for 0 hour, 3 hours, 6 hours, and 12 hours, 500 µL of culture medium was taken and centrifuged at 4 °C and 6000 rpm for 10 minutes for the detection of IL-6 using enzyme linked immunosorbent assay (ELISA) according to the instructions of IL-6 ELISA Kit provided by Jiangsu Enzyme-linked. The IL-6 content in the supernatant was detected in a 96-well plate.
Total Protein Extraction
Radioimmunoprecipitation assay Lysis Buffer (strong; CWBIO CW2333S) was mixed with protease inhibitor cocktail (100×; CWBIO CW2200) in a ratio of 1:99 for the preparation of 1× extraction reagent for protein extraction. After the treated cells were rinsed with PBS, 1× protein extraction reagent was placed on ice. The lysates were transferred to new centrifuge tubes and incubated on ice for 20 minutes. They were then centrifuged at 14 000×g for 10 minutes before the supernatants were transferred to new tubes. The final extracted samples were kept at −80 °C.
Western Blot
Western blotting was used to detect the target protein in the sample. The total protein content in the sample was detected in a 96-well plate using a BCA protein concentration determination kit (Dingguochangsheng). Protein tracer sample buffer (reduction, 5×; CWBIO) was mixed with protein samples in a ratio of 1:4. The mixtures were then placed in a boiling water bath for 3 minutes. The samples were cooled to room temperature and centrifuged at 13 000×g at 4 °C for 30 seconds. Denatured proteins were directly loaded to a sodium dodecyl sulfate-page gel, and conventional electrophoresis (concentration gel voltage 60 V, separation gel voltage 120 V) and membrane transfer (current 200 mA) were performed using a Bio-Rad protein imprinting device. After completion, the transferred films were incubated at 4 °C overnight. The rabbit primary antibodies used were STAT3 (Proteintech), pSTAT3 (Tyr705; ABclonal), KLF4 (Proteintech), OCT4 (Proteintech), and β-actin (Proteintech). Samples were incubated with HRP-labeled goat anti-rabbit secondary antibody (Proteintech) at room temperature for 1 hour, washed with 1× TBST 3 times for 10 minutes each, and finally ECL development was performed.
Statistical Analysis
Each independent experiment was repeated 3 times for each group. The target gene mRNA change ratio was calculated using the 2(−△△Ct) method. Student t test was used to compare the data of the 2 groups, and P < .05 was considered statistically significant.
Results
Signal Transducer and Activator of Transcription 3 Induced the Initiation and Progression of EAC by Regulating the Expression of Stem Cell–Associated Genes
Figure 1 shows that after siRNA silencing of STAT3 mRNA in HEEC, the relative mRNA expression of STAT3 was reduced to 13% compared to that of the negative control, indicating that the silencing efficiency of STAT3 in HEEC was 87% (efficiency = 100% − Mean fold change %, P < .01). Since the silencing efficiency of STAT3 was statistically significant, these cells could be used to study the expression of other related genes. After STAT3 mRNA expression was silenced in HEEC, the mRNA expression of the stem cell–related genes OCT4, HIF-1α, Nanog, and CD44 were significantly reduced (P < .01), with expression reduction efficiencies of 84%, 89%, 86%, and 95%, respectively. Significant siRNA silencing of STAT3 mRNA was also seen in OE33, where the relative expression of mRNA was 17% of the control, with a silencing efficiency of 83% (P < .01). Since this silencing efficiency was statistically significant, it could also be used to study the expression of related genes. After STAT3 silencing in OE33, the relative mRNA expressions of OCT4, HIF-1α, and Nanog decreased (P < .01), but the relative expression of CD44 increased (P < .05). After the knockdown of STAT3, Bcl-xL gene expression in the HEEC and OE33 cell lines were reduced by 31% (P < .01) and 29% (P < 0.05), respectively, showing significant differences (Figure 1).

Mean fold change of stem cell–related genes of the HEEC (A) and OE33 (B) with STAT3 silencing under stimulation of DCA (3 hours), *P < .05; **P < .01, compared with the control. DCA indicates deoxycholic acid; STAT3, signal transducer and activator of transcription 3.
In this experiment, the relative expression of Bcl-xL mRNA was detected after silencing STAT3 by siRNA to explore the correlation between STAT3 and Bcl-xL in HEEC and OE33 cells. The expression of the antiapoptotic Bcl-xL protein was downregulated in the STAT3-silenced HEEC and a similar phenomenon was observed in OE33 cells (Figure 2).

Protein expression of related genes after STAT3 silencing in HEEC and OE33. STAT3 indicates signal transducer and activator of transcription 3.
Deoxycholic Acid Promoted the Expression of Stemness-Related Reprogramming Factors KLF4 and OCT4
In this study, 250-μM DCA was used to stimulate OE33 EAC for 0 hour, 3 hours, 6 hours, and 12 hours to investigate whether the gastroesophageal reflux constituent DCA has a stemness induction effect on EAC. On the one hand, our results showed that relative mRNA expression levels (fold change) of reprogramming factor KLF4 were 2.08 (P = .0585), 6.78 (P < .01), and 4.53 (P < .05), after 3 hours, 6 hours, and 12 hours of DCA stimulation, respectively, compared to 0 hour of DCA stimulation, that is, without DCA stimulation. Krüppel-like factor expression was statistically different after DCA stimulation for 6 hours and 12 hours, indicating that the expression of KLF4 mRNA (Figure 3) and protein (Figure 4) in EAC increased after DCA stimulation, reaching a peak at 6 hours. At the same time, the relative mRNA expression levels of cellular reprogramming factor OCT4 were 1.43 (P = .133066), 1.92 (P < .05), and 4.51 (P < .01), after 3 hours, 6 hours, and 12 hours of DCA stimulation, respectively. The expression levels of OCT4 were also statistically different after 6 hours and 12 hours of DCA stimulation. The mRNA and protein expression of OCT4 in EAC showed a gradually increasing trend under DCA stimulation (Figure 3). In addition, the mRNA expression level of Nanog after 3-hours stimulation with DCA was 1.75 times higher than that at 0 hour (P < .05), showing a significant difference, while it has not significantly changed after 6 hours and 12 hours of DCA stimulation (Figure 3).
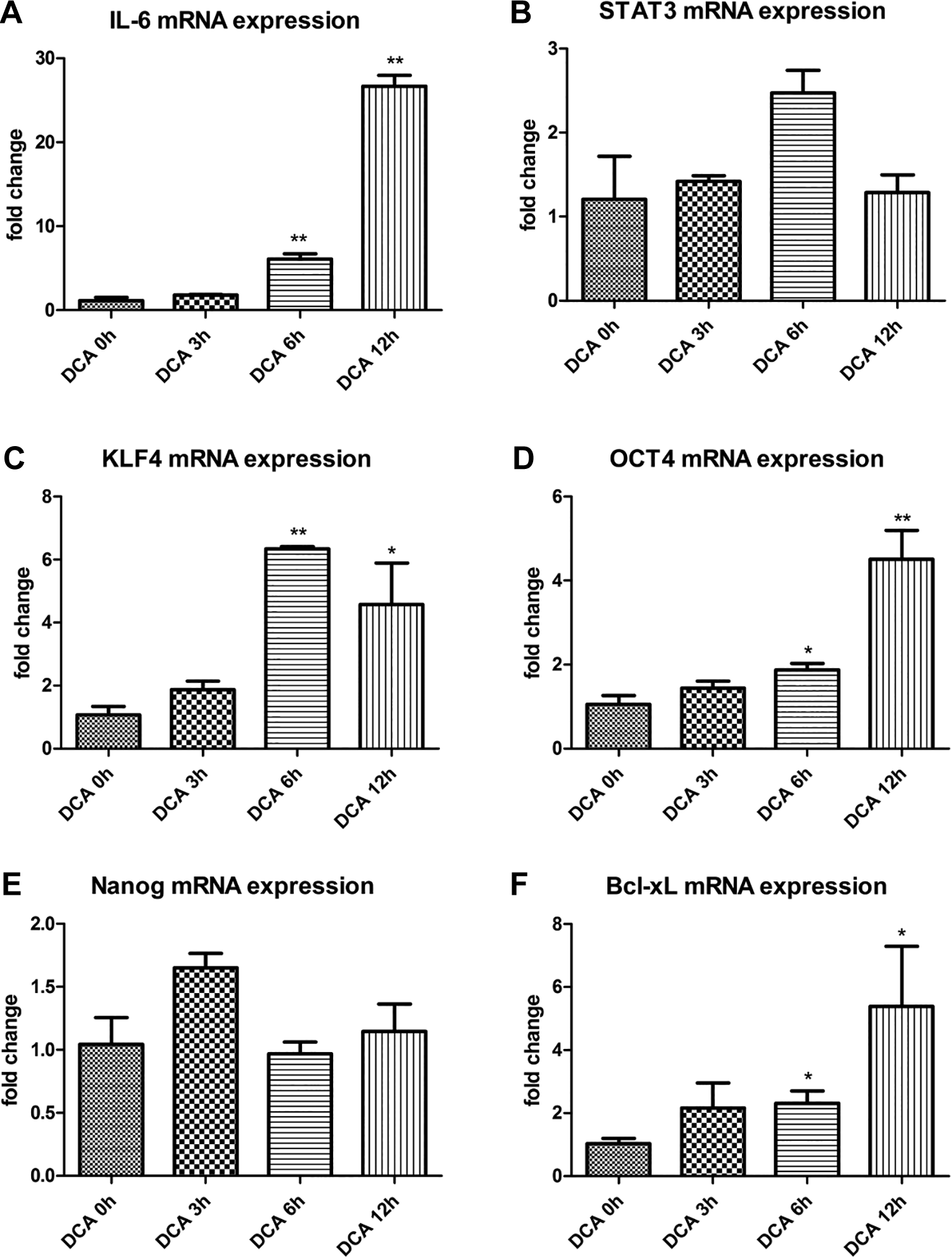
Fold change in IL-6 mRNA expression (A), STAT3 mRNA expression (B), KLF4 mRNA expression (C), OCT4 mRNA expression (D), Nanog mRNA expression (E), and Bcl-xL mRNA expression (F) after 0, 3, 6, and 12 hours of DCA (250 μM) treatment, *P < .05; **P < .01 compared with DCA 0 hour. DCA indicates deoxycholic acid; IL-6, interleukin-6; KLF4, Krüppel-like factor; OCT4, octamer-binding transcription factor 4; STAT3, signal transducer and activator of transcription 3.
On the other hand, the results also showed that the relative mRNA expression levels of cytokine IL-6 at 3 hours, 6 hours, and 12 hours after DCA stimulation were 2.07 (P = .09), 6.67 (P < .01), and 22.95 (P < .01), respectively, compared to that at 0 hour after DCA stimulation (Figure 3). Since IL-6 is a secretory protein, it is secreted to the outside of cells after synthesis and there exerts its first signal effect. This experiment further detected the concentration of IL-6 in the cell culture medium of the different DCA-stimulated groups using ELISA. The concentrations of IL-6 in the supernatant in the DCA 0 hour, DCA 3 hours, DCA 6 hours, and DCA 12 hours treatment groups were 5.268, 8.029, 17.004, and 20.532 ng/L, respectively (Figure 5). Compared with DCA stimulation at 0 hour, there were statistically significant differences in cytokine IL-6 protein expression after 3 hours (P < .01), 6 hours (P < .01), and 12 hours (P < .01) after DCA stimulation (Figure 5). Moreover, pSTAT3 (Tyr705), a reactive protein downstream of cytokine IL-6, was significantly increased with the extension of DCA stimulation time (Figure 4A).

A, Protein expression of STAT3, pSTAT3, KLF4, OCT4, β-actin after 0, 3, 6, and 12 hours of DCA (250 μM) treatment. B, Relative protein expression of KLF4 after 0, 3, 6, and 12 hours of DCA (250 μM) treatment. C, Relative protein expression of OCT4 after 0, 3, 6, and 12 hours of DCA (250 μM) treatment. DCA indicates deoxycholic acid; KLF4, Krüppel-like factor; pSTAT3, phosphorylated-signal transducer and activator of transcription 3; STAT3, signal transducer and activator of transcription 3.

Interleukin-6 protein expression in media after 0, 3, 6, and 12 hours of DCA (250 μM) treatment. DCA indicates deoxycholic acid.
Signal Transducer and Activator of Transcription 3 Specifically Regulates the Expression of Stemness-Related Reprogramming Factors KLF4 and OCT4
In the first part, our experimental results showed that DCA stimulation can activate the IL-6/STAT3 inflammatory pathway. Prolonged DCA stimulation significantly increased the expression of active pSTAT3 protein and it also promoted the expression of reprogramming factors KLF4 and OCT4 (Figure 4B and C), Nanog, and antiapoptotic protein Bcl-xL. Considering that STAT3 is a transcription factor, its dimerized active form of pSTAT3 can enter the nucleus, target a specific gene, and initiate the transcription process of that gene. Therefore, whether STAT3 is involved in the upstream and downstream regulation of KLF4, OCT4, Nanog, and Bcl-xL genes requires further study. In this regard, the STAT3 gene was silenced by siRNA technology to observe the expression of KLF4, OCT4, Nanog, and Bcl-xL genes.
First, after siRNA silencing of STAT3 mRNA in OE33, its relative mRNA expression was reduced to 23% of that in the control group, which made the silencing efficiency of STAT3 in OE33 77%, which was statistically significant (P < .01; Figures 6A and 7A).Western blot analysis showed that STAT3 protein expression was significantly reduced after silencing (Figure 2). Therefore, the results of this siRNA experiment are suitable to study the expression of related genes after silencing STAT3 mRNA.

Fold changes in mRNA expression of STAT3 (A), KLF4 (B), OCT4 (C), and Bcl-xL (D) after STAT3 silencing, *P < .05; ** P < .01 compared with the control. STAT3 indicates signal transducer and activator of transcription 3.

A, Protein expression of STAT3, KLF4, and OCT4 after STAT3 silencing, (B) KLF4 protein relative expression after STAT3 silencing; (C) and OCT4 protein relative expression after STAT3 silencing. KLF4 indicates Krüppel-like factor; STAT3, signal transducer and activator of transcription 3.
Second, OE33 cells were transfected with siRNA, and the mRNA expression of STAT3 gene was silenced. The results showed that the mRNA expression of reprogramming factors KLF4 and OCT4 were downregulated by 12% (P < .01) and 31% (P < .05) of the mRNA expression before silencing, both of which are statistically significant (Figure 6A-C). Western blot analysis showed that KLF4, OCT4, and Bcl-xL protein expression were also significantly decreased after silencing STAT3 (Figure 7A-C).
After STAT3 silencing, DCA was added to continue stimulating OE33 cells for 12 hours. The results showed that pSTAT3 protein expression in the siSTAT3 + DCA 12-hour group was significantly lower than that of the DCA 12-hour group. At the same time, there was a significant difference in the mRNA expression of KLF4 and OCT4 (P < .05; Figure 6A) and the expression of translated KLF4 and OCT4 proteins were decreased (Figure 6B). Furthermore, after STAT3 silencing, the mRNA expression of the antiapoptotic Bcl-xL gene was reduced to 29% of that of the negative control group, with a statistical difference (P < .05; Figure 6A), indicating that STAT3 was an upstream regulator of Bcl-xL and had an enhancing effect on the expression of Bcl-xL.
Activation of the IL-6/STAT3 Pathway Regulates the Expression of Stemness-Related Reprogramming Factors KLF4 and OCT4
Parts 2.1 and 2.2 demonstrated that the extracellular stimulation signal DCA can promote the expression of cytokine IL-6 and reprogramming factors KLF4 and OCT4. Additionally, KLF4 and OCT4 are downstream regulatory targets of STAT3. Therefore, recombinant human IL-6 was used to treat OE33 of EAC to further confirm that the IL-6/STAT3 signaling pathway can regulate the expression of vital reprogramming factors KLF4 and OCT4.
Recombinant human IL-6 with different concentrations were prepared to stimulate OE33 for 24 hours. The results showed that mRNA expression levels of KLF4 gene after 10 ng/mL and 100 ng/mL treatment of cells were 1.73 times (P = .097574) and 3.23 times (P < .05) that of the blank control group, respectively; while the relative mRNA expression of the OCT4 gene was 3.73 times (P < .05) and 4.43 times (P < .01), respectively (Figure 8). Western blot also showed that recombinant human IL-6 stimulation can promote the expression of pSTAT3, KLF4, and OCT4 proteins (Figure 9). Furthermore, 100 ng/mL recombinant human IL-6 can increase the mRNA expression of the Bcl-xL gene in OE33 by 8 times (P < .01; Figure 8).

Fold changes in mRNA expression of Bcl-xL, KLF4, and OCT4 gene after 10 ng/mL and 100 ng/mL of recombinant human IL-6 treatment, *P < .05, **P < .01, compared with control. IL-6 indicates interleukin-6; KLF4, Krüppel-like factor.

A, Protein expression of STAT3, pSTAT3, KLF4, OCT4, and β-actin after 10 ng/mL and 100 ng/mL of recombinant human IL-6 treatment; (B) KLF4 protein relative expression after 10 ng/mL and 100 ng/mL of recombinant human IL-6 treatment; (C) OCT4 protein relative expression after 10 ng/mL and 100 ng/mL of recombinant human IL-6 treatment. IL-6 indicates interleukin-6; KLF4, Krüppel-like factor; pSTAT3, phosphorylated-signal transducer and activator of transcription 3; STAT3, signal transducer and activator of transcription 3.
Discussion
Part 1. Roles of Stem Cell Associated Gene Expression in the Initiation and Progression of Esophageal Adenocarcinoma
Part 1.1. Signal transducer and activator of transcription 3 induced stem cell gene expression in HEEC and OE33
After the significant silencing of STAT3 mRNA expression in HEEC and OE33 cells (P < .01), the expression of 4 stem cell–related genes, OCT4, HIF-1α, Nanog, and CD44, were significantly downregulated or upregulated (P < .05 or P < .01), indicating that STAT3 could regulate the expression of these genes. Therefore, the expression of STAT3 is related to the expression of these stem cell–related genes, either upstream or downstream. The results support the targeting effect of STAT3 to the expression of stem cell–related genes such as OCT4, HIF-1α, and CD44. 30 The expression of stem cell–related genes is an important indicator associated with stemness acquisition. Tumor cells with high expression of stem cell-related genes could form spheres in vitro and tumors in vivo. 31 Exposure to DCA in normal esophageal epithelial cells would activate STAT3 inflammatory pathway and subsequently increase the expression of STAT3, 28 which may explain how DCA exposure would transform normal esophageal epithelial cells to BE cells. 7
In humans, embryonic stem cells are transformed into progenitor cells (also known as totipotent stem cells), which can be further transformed into pluripotent stem cells and then into various mature cell types. The upregulated gene expression of stem cells is an important sign of normal cells acquiring stemness and the ability to dedifferentiate. In the model of normal human esophageal epithelial cells developing into EAC cells, the upregulation of the gene expression of stem cells is an important sign of promoting normal esophageal epithelial cells to gain the ability to dedifferentiate and improve their reversal into pluripotent stem cells. Finally, this upregulated gene expression increases the possibility of normal esophageal epithelial cells developing in the direction of Barrett esophageal cells, EAC, or CSCs.
Cancer might be a kind of disease derived from the stem cell dysregulation. Previous studies have observed their strong association. Pathways associated in stem cell–related genes, for example, Wnt, Hedgehog, and Notch signaling pathways, are involved in tumor cells. Although CSCs are suspected to be involved in tumor formation, the source of CSCs is still to be a question required further clarification. The source of CSCs might derive from the transformation of adult stem cells, mutations of adult cells, the activation of proto-oncogenes, or from the dedifferentiation of tumor cells expressing stem cell–related genes. Whenever the tumor cells could express stem cell-related genes, which indicates the tumor cells have already in the process of acquiring stem cell properties, that is, self-renewal abilities and multiple developing capabilities. Similarly, normal tissue cells induced with the expression of stem cells markers would dedifferentiate into pluripotent stem cells under certain condition and finally promote the transformation to tumor cells. Therefore, understanding the biology in relation with the microenvironment might help in developing the therapeutic strategies for tumor stem cells. 3 Current research has observed the possible relationship between STAT3 and stem cell–related genes, but further studies are required to clarify their involvement in the initiation and progression of esophageal squamous cancer.
In this experiment, after significant silencing of STAT3 mRNA expression in HEEC, the expression of 4 stem cell–related genes, OCT4, HIF-1α, Nanog, and CD44 was significantly inhibited (P < .01). It showed that, upon stimulation with DCA, STAT3 mRNA expression was significantly silenced in HEEC, and the expression of these 4 stem cell–related genes was significantly inhibited, that is, the downregulation of STAT3 inhibits the expression of these genes. This indicated that the expression of STAT3 could be controlled to regulate the expression of these stem cell–related genes, thus regulating the tendency of HEEC to become cancerous under the stimulation of DCA. Therefore, STAT3 might be an important factor influencing the development of HEEC toward BE and EAC.
On the other hand, a high expression of stem cell–related genes is one of the key markers of stemness acquisition in tumor cells. The results from this experiment support the notion that cancer stem cells are converted from cancer cells, thereby increasing the probability of recurrence, metastasis, and chemotherapy resistance. In this experiment, after the significant silencing of STAT3 mRNA expression in OE33, the expression of stem cell–related genes OCT4, HIF-1α, and Nanog were significantly downregulated (P < .01). This suggested that under DCA stimulation, STAT3 mRNA expression was significantly silenced in OE33, and the expression of the stem cell–related genes OCT4, HIF-1α, and Nanog was also inhibited, that is, STAT3 downregulation inhibits the expression of these genes.
Similarly, activation of the JAK/STAT3 signaling pathway can induce CD44 expression in breast cancer, while STAT3 inhibitors could reduce CD44 expression. 32 Signal transducer and activator of transcription 3 can be recruited into the promoter region of OCT4 and Nanog in the process of acquiring stemness in breast cancer, suggesting that STAT3 could regulate the expression of OCT4 and Nanog, promoting the formation of breast cancer stem cells. 33 It was also found that STAT3 could target HIF-1α and lead to chemotherapy resistance in breast cancer cells. 34 Therefore, silencing STAT3 could inhibit the expression of stem cell–related genes, and that silencing STAT3 might be one of the important factors for inhibiting the acquisition of stemness in EAC cell lines and reducing its malignant progression, proliferation, recurrence, metastasis, and radiotherapy/chemotherapy resistance. In summary, upon DCA stimulation, the inhibition of STAT3 expression was an important factor in controlling the carcinogenesis of normal esophageal epithelial cell lines and the progression of EAC cell lines.
Part 1.2. Relationship between STAT3 and Bcl-xL in EAC
The data from part 1.1 of the Results indicated that STAT3 is an upstream regulator of Bcl-xL, promoting its expression. This suggested that STAT3 activation could upregulate the expression of the antiapoptotic protein Bcl-xL, improve the survival of EAC cells, inhibit the programed death of tumor cells, and increase the antiapoptotic ability of EAC. 4-parvifuran (PVN) was found to upregulate the expression of antiapoptotic proteins Bcl-xL and Bcl-2 by inhibiting JAK2/STAT3 activation. 35 Liu et al 36 reported that the activation of STAT3 and NF-κB by chronic lymphoma cells could upregulate the levels of antiapoptotic proteins Bcl-xL, Mcl, and IL-6, leading to chemotherapy tolerance. Zaanan et al 37 showed that STAT3 promoted the apoptotic resistance induced by KRAS mutation by upregulating Bcl-xL expression. Matsumoto et al 38 found that the vacuolar formation toxin of Helicobacter pylori could reduce STAT3, thereby downregulating Bcl-xL and Bcl-2 to induce apoptosis of gastric adenocarcinoma cells. These studies are similar to the results of this experimental group, which indicated that STAT3 activation could target the upregulation of the antiapoptotic protein Bcl-xL and increase the antiapoptotic ability of tumor cells. This suggested that clinical treatment could limit tumor cell survival and promote apoptosis of tumor cells by targeting STAT3 inhibition.
Esophageal adenocarcinoma is known to be an inflammatory tumor. Signal transducer and activator of transcription 3 is a functional protein involved in the signal transduction pathway of tyrosine phosphorylation. Its phosphorylation forms pSTAT3, which, after dimerization, enters the nucleus. It can bind with other transcription factors to target gene expression, mediate inflammation, and tumor formation. Signal transducer and activator of transcription 3 is a key molecule in inflammation-related tumors. In EAC, the signal of the STAT3 inflammatory signaling pathway is activated, which plays a central role in enhancing its malignant potential, inducing the transformation of tumor cells into CSCs. In recent years, the academic community has proposed the theory of CSCs, which was considered to have a close relationship with the existence of CSCs. Cancer stem cells are a heterogeneous subgroup of tumor cells that have the characteristics of self-renewal, polygenesis, and asymmetric division. Thus far, the source of CSCs in vivo is not clear, which may be derived from stem cells, progenitor cells, or it could be driven by the dedifferentiation of tumor cells. In EAC, this experimental group found that STAT3 could target the regulation of stem cell–related genes OCT4, HIF-1α, Nanog, and CD44, suggesting that activation of the STAT3 pathway might induce EAC to acquire stemness and transform into CSCs.
The activation of the STAT3 pathway in EAC laid a foundation for the subsequent development of EAC cells: the enhancement of malignancy, proliferation, invasion and metastasis, antiapoptosis behavior, or the transformation of EAC cells and CSCs. Both tumor progression and transformation to CSCs undoubtedly shorten the survival of patients with EAC. Therefore, the development of targeted molecular treatments for STAT3 has promising prospects for the treatment of EAC.
Deoxycholic Acid Regulates the Expression of Reprogramming Factors KFL4 and OCT4 in Esophageal Adenocarcinoma Cells Through the IL-6/STAT3 Pathway
The current theory proposed that heterogeneous subsets of tumor cells that acquire self-renewal abilities, multidirectional development, and asymmetric division, 39 called CSCs, are closely related to radiation resistance, chemotherapy resistance, recurrence, invasion, and tumor metastasis. Tumor stem cells can be isolated from many solid tumor tissues. The most commonly used markers for CSC identification include the transmembrane glycoproteins CD24, CD34, CD38, CD44, CD90, CD133, acetaldehyde dehydrogenase, and some other molecular phenotypes specifically expressed only in CSC. Separation methods for CSCs include flow cytometry fluorescence separation, magnetic bead separation, side group cell separation, serum-free liquid medium method, and the acetaldehyde dehydrogenase luminescence method. Cancer stem cells can overcome the programmed cell death induced by breaking from the surrounding extracellular matrix, forming colonies and suspension cultures into pellets in soft agar, which is a feature of cell stemness. Thus, the isolated cells for further identification would conduct tests of tumorigenesis in vivo or pelletization in vitro.
Cancer stem cells are usually inactive and are less responsive to radiation and chemotherapeutic treatment, where they can escape the killing effect of current cancer treatments by ATP-binding cassette transporters, DNA damage repair, cell cycle arrest, attenuated substances in cells, apoptosis inhibition, and through the epithelial mesenchymal transformation mechanism. As long as they survive, they would differentiate and proliferate again under suitable conditions, causing tumor recurrence, invasion, and distant metastasis. To date, the source of CSCs in vivo has not been determined, which may be transformed from the activation of stem cells or progenitor proto-oncogene mutations, or the dedifferentiation of tumor cells to drive the formation of CSCs. 40 Therefore, there is a mutual transformation relationship between differentiation and dedifferentiation between CSCs and tumor cells; thus, the cell fate has a considerable degree of plasticity.
Cell reprogramming is the process in which mature, terminally differentiated cells are reversed under certain conditions to return to totipotency or pluripotency or to form pluripotent stem cells. In short, it is the reverse differentiation of mature cells. Cell reprogramming can be involved in the occurrence and development of diseases. Induced pluripotent stem cells are adult cells directly transformed into embryonic stem cell–like pluripotent cells through gene reprogramming, with which tumor cells would alter their malignancy. In the early stages of reprogramming, tumor cells become more stem-like, which could translate into more malignant CSCs. In the late stage of reprogramming, the diverse potential of tumor cells (pluripotency) increases with the decline in the degree of malignancy; they are then transformed into tumor iPSCs. Current views suggest that the reprogramming process is accompanied by the overexpression of 4 Yamanaka transcription factors, including KLF4, OCT4, SOX2, and MYC, collectively known as OSKM. In 2006, Yamanaka, a Japanese scientist, first introduced OCT4, SOX2, KLF4, and c-MYC into mouse fibroblasts via retroviral transduction and produced iPSCs. 41 Later, Thomson further proposed Nanog combined with the abovementioned Yamanaka factors also has the potential to induce multidirectional network effects. 42
Our experimental results indicate that the bile portion of the gastroesophageal reflux substance, DCA, promoted the expression of cell reprogramming factors KLF4, OCT4, and Nanog in EACs, which would induce the reprogramming of EACs into CSCs with self-renewal ability and multi-differentiation potential. Nanog is believed to play a role only in the early stages of this transformation process, after which KLF4 and OCT4 mediate the formation of CSCs in EAC.
OCT4 is thought to be necessary in the cell reprogramming process, which shows various functions according to its expression level and is called the “variant” gene. There is an interaction between STAT3 and OCT4. In mouse embryonic stem cells, leukemia inhibitory factor (LIF) activates important pluripotency-associated gene expression, such as OCT4, preventing embryonic stem cells from differentiating and maintaining their pluripotency by activating the JAK1/STAT3 pathway. Moreover, the target gene of STAT3 is also regulated by OCT4. Krüppel-like factor is directly downstream of OCT4 and STAT3, which is necessary to maintain the self-renewal and multi-directional differentiation of embryonic stem cells. Furthermore, STAT3 upregulates the expression of stem cell factor Nanog by binding to OCT4 and further maintain the self-renewal ability of stem cells. 43 Therefore, STAT3 and OCT4 are closely related. Studies have shown that utilization of the STAT3 inhibitor WP1066 can inhibit the expression of OCT4 in iPSCs. 44 In cervical cancer, overexpression of STAT3 significantly increased the mRNA and protein expression of OCT4; OCT4 expression was decreased when STAT3 expression was inhibited by siRNA. After transfection with an STAT3 active mutant, pancreatic cancer cells expressed the stem cell marker OCT4. 45 Subsequently, studies have shown that STAT3 can be recruited into the OCT4 promoter region in breast cancer cells. 33 Similarly, our experimental results also show that OCT4 is regulated by STAT3, and they are positively correlated.
Krüppel-like factor plays an important regulatory role in a variety of cellular self-renewal processes, including proliferation, differentiation, apoptosis, cell transformation, and embryonic development. 46 Similar to transforming growth factor β1, KLF4 exhibits both anticancer and procancer effects, depending on cellular function and cancer type. 47 In colorectal cancer, 48 gastric cancer, 49 esophageal squamous cell carcinoma, 50 lung cancer, 51 prostate cancer, 52 and bladder cancer, low expression of KLF4 can promote excessive cell proliferation and malignant transformation, supporting the role of KLF4 as a tumor suppressor gene. In addition, KLF4 is upregulated in primary ductal carcinoma of the breast 53 and oral squamous epithelial carcinoma, which promotes the survival and progression of tumors, reflecting the function of KLF4 as a proto-oncogene. Previous studies on KLF4 have focused on the promotion and inhibition of cancer, until 2006, when Takahashi and Yamanaka first identified KLF4 as one of the 4 reprogramming factors. It is necessary to induce mouse embryonic development and transformation from mature fibroblasts into multipotent stem cells. Krüppel-like factor has been explored as a stem cell gene. Yu et al 54 found that KLF4 is necessary to maintain the stemness of breast cancer stem cells, and the clearance of KLF4 reduces the ability of tumor formation. Chang et al 55 also proposed the necessity of KLF4 in prostate cancer CSCs. Qi et al 56 found that KLF4 can activate the p38 MAPK signaling pathway to enhance the stemness of osteosarcoma cells. In contrast, Shum et al 57 believed that KLF4 inhibited the stemness of neuroblastoma. Yan et al 58 also showed that KLF4 had an adverse effect on the maintenance of stemness in pancreatic ductal adenocarcinoma. In human glioma, the paracrine PGI signal can promote the self-renewal and tumorigenic potential of glioma stem cells by inducing KLF4 expression. 59 These contradictory results of about KLF4 on the maintenance of stemness in tumor cells may be related to tissue specificity. Regarding the relationship between the STAT3 pathway and KLF4, in embryonic stem cells, the LIF/STAT3 pathway can selectively promote KLF4 expression and maintain the pluripotency and self-renewal of embryonic stem cells. 60 Research has shown that the use of the STAT3 inhibitor WP1066 can inhibit KLF4 expression in iPSCs, suggesting that STAT3 is crucial in maintaining the pluripotency of iPSCs. 44 In breast cancer cell lines, the small molecule inhibitor of STAT3, Stattic, significantly reduced the mRNA expression level of KLF4. 40 These experiments are consistent with our results and also indicate that KLF4 is a downstream regulator of STAT3, which can promote KLF4 expression. In contrast, after treatment with the STAT3 inhibitor stx-0119, KLF4 was unexpectedly upregulated and other Yamanaka factors were downregulated in glioblastoma strains. 61
The self-renewal process of CSCs involves several important cellular signaling pathways, such as the Wnt/β-catenin and Notch signaling pathways. In colon CSCs, the Wnt/β-catenin signaling pathway improves their ability to self-renew. Mesenchymal cells promote the self-renewal of CSCs by activating the IL-4-Notch signaling pathway of CSCs. Hepatocyte growth factor also plays a role in maintaining the stemness of colon CSCs, preventing their differentiation. 62 Various stemness-related transcription factors, such as OCT4, Nanog, SOX2, and B-cell-specific Moloney leukemia virus insert site 1 are involved in regulating the self-renewal and stemness maintenance of CSCs. In the small intestine, expression of OCT4 causes dysplasia and expansion of progenitor cells. 63 Knocking out the OCT4 gene in human and mouse cell lines can reduce CSC subsets. 64 Studies have reported that SOX2 can maintain the key signaling cascade of tumorigenesis and that tumor cells silenced by SOX2 have a decreased expression of c-MYC, Wnt1, and Notch1, which reduces the ability of immune-deficient nude mice to be transplanted with tumors. 65 B-cell-specific Moloney leukemia virus insert site 1 plays a crucial role in the self-renewal of normal stem cells. Knocking out bm-1 in CD133+ laryngeal cancer cells resulted in decreased cell growth, clonal formation, cell invasion ability, and inhibition of tumorigenesis ability in vivo. 66 In pancreatic cancer, the KRAS-cMYC signaling pathway promotes the formation of stem cell characteristics. 67 In animal models, the NF-κB signaling pathway can stimulate Wnt, induce intestinal epithelial cells to differentiate, as well as acquire tumorigenic ability. 68 In addition, incomplete reprogramming can trigger tumor cells to acquire stemness features. Knappe et al 69 used doxycycline to induce melanoma cells to express OSKM, which caused their partial reprogramming, increased invasion potential in vitro, and pulmonary metastasis in vivo.
The analysis above indicates that there is plasticity between tumor cells and CSCs, which would change their fate. Tumor cells reprogram their fate through extracellular and intracellular signaling pathways in which they can obtain stem cell characteristics or transform into tumor-iPSCs, change the malignancy degree of tumors, and promote the survival and progression of tumors.
Furthermore, our experimental results showed DCA stimulation could activate the IL-6/STAT3 signaling pathway of EAC cells. There is an upstream and downstream regulatory relationship between STAT3 and the reprogramming factors KLF4 and OCT4, in which STAT3 is the upstream target promoting the expression of KLF4 and OCT4. Next, DCA activates the STAT3 pathway, where pSTAT3 regulation promotes the expression of downstream reprogramming factors KLF4, OCT4, and Nanog, which induces stemness-acquiring characteristics and transformation of EAC cells to CSCs. The reduction of STAT3 phosphorylation is correlated with the therapeutic effect of curcumin on esophageal cancer. 70 In addition, the activation of STAT3 could upregulate the expression of the antiapoptosis protein Bcl-xL, improving the survival of EAC cells, inhibiting programed death of tumor cells, and increasing the antiapoptotic ability of EAC. Bcl-xL is an antiapoptotic protein of the Bcl-2 family. The inflammation caused by deoxycholic acid increases cellular reactive oxygen species that are associated with the increase of circccRNA of miR-21 and miR-21, which are directly regulated by the transcription factor (NF-κB), 71 and the activated NF-κB pathway could also produce antiapoptotic protein Bcl-2, making DNA-damaged cells resistant to apoptosis. 8,70 Evaluation of circRNA expression in esophageal squamous cell carcinoma has revealed differentially expressed circRNAs in cancer-related cellular and molecular pathways. 72 Our results suggested that DCA stimulation promoted the expression of Bcl-xL, improved the antiapoptosis ability of EAC cells, and inhibited the programed death of tumor cells. It further increased the malignancy and improved the survival rate of EAC cells.
In the case of the relationship between STAT3 and antiapoptotic protein Bcl-xL, studies have found that PVN can upregulate the expression of antiapoptotic proteins Bcl-xL and Bcl-2 by inhibiting JAK2/STAT3 activation. 35 Liu et al 36 reported that the activation of STAT3 and NF-κB in chronic lymphoma cells could upregulate the levels of antiapoptotic proteins Bcl-xL, Mcl, and IL-6, leading to chemotherapy tolerance. Zaanan et al 37 showed that STAT3 promoted KRAS mutagen-mediated apoptosis resistance by upregulating Bcl-xL expression. Matsumoto et al 38 found that the vacuolating cytotoxin of Helicobacter pylori can induce apoptosis of gastric adenocarcinoma cells by lowering STAT3, thereby downregulating Bcl-xL and Bcl-2. The above studies support our findings that STAT3 activation could upregulate the antiapoptotic protein Bcl-xL and increase the antiapoptotic ability of tumor cells.
These results indicate that the IL-6/STAT3 signaling pathway mediates the expression of reprogramming factors KLF4 and OCT4 to induce the transformation of EAC cells into CSCs. The IL-6/STAT3 signaling pathway can also promote the expression of the antiapoptotic protein Bcl-xL, increase the antiapoptotic ability of EAC cells, and reduce their malignancy. This experiment revealed that the gastroesophageal reflux constituent DCA has toxic effects that can activate the IL-6/STAT3 inflammatory signaling pathway. On one hand, expression of the reprogramming factor Nanog was upregulated in the early stage, followed by the expression of reprogramming factors KLF4 and OCT4 and were upregulated in the later stages. They promote the stemness of EAC cells and possibly induce the formation of EAC CSCs. On the other hand, DCA activates the IL-6/STAT3 pathway to upregulate the antiapoptosis protein Bcl-xL, increasing the antiapoptosis ability of EAC cells, which inhibits programed cell death, improves their survival rate, and worsens the malignancy of EAC. Thus, targeting STAT3 inhibition may improve the clinical treatment of EAC to limit the survival of tumor cells and promote tumor cell death.
Conclusion
There are upstream and downstream relationships among STAT3, KLF4, OCT4, and the antiapoptotic protein Bcl-xL. Deoxycholic acid was shown to have a malignant induction effect, activating the IL-6/STAT3 pathway, and promoting the upregulation of KLF4, OCT4, Nanog, Bcl-xL, and other genes. It also has the potential to induce the transformation of EAC cells into CSCs, improving the antiapoptosis ability of tumors, and further increasing the malignancy of EAC. Since the formation of CSCs is closely related to chemo- and radiotherapy resistance, recurrence, invasion, and metastasis, small-molecule inhibitors targeting the IL-6/STAT3 signaling pathway and DCA neutralization drugs may be considered to improve their clinical efficacy and inhibit the formation of CSCs in EAC.
Footnotes
Authors’ Note
Mei Chen and Xiaojun Ye are equal weighting first author, sharing the equal amount of effort in performing experiments, and writing of the papers. Guarantor of article: Karen Poon and Ruihua Wang are acting as the submission’s guarantors. Mei Chen and Xiaojun Ye designed, performed the experiments, collected and analyzed the data, and wrote the paper. Jingxi Wei performed some of the experiments. Karen Poon and Ruihua Wang corrected the paper and design. All authors approved the final version of the article, including the authorship list.
Acknowledgments
The authors thank Liu Chang for conducting some of the laboratory works.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Beijing Normal University – Hong Kong Baptist University United International College research grant R201801.
