Abstract
Objective:
To identify the effect of curcumin on tumor suppression and the possible molecular pathways involved.
Methods:
The expression of long noncoding RNA neighbor of BRCA1 lncRNA 2 (NBR2) was quantified using reverse transcription-polymerase chain reaction on cultured colorectal cancer cells. Next, we used Western blot to measure the activation of adenosine monophosphate-activated protein kinase and mechanistic target of rapamycin kinase (mTOR) signaling molecules. Both cell proliferation and viability were measured via MTT assay, and the cell ratio and S phase were detected by BrdU assay. Colorectal cancer cells were pretreated with curcumin or transfected with shNBR2 or adenosine monophosphate-activated protein kinase inhibitor Compound C to examine the molecular pathway involved.
Results:
Current data showed that glucose deficiency increased the expression of NBR2 in colorectal cancer cells, and NBR2 knockdown affected the progression of colorectal cancer cells under glucose starvation conditions. When NBR2 was silenced in the treated colorectal cancer cells, the proliferation, the clone formation, and the percentage of S-phase cells suppressed by glucose deprivation were compromised. Furthermore, NBR2 knockdown could suppress glucose deprivation-induced adenosine monophosphate-activated protein kinase activation plus mTOR inactivation. Similarly, when colorectal cancer cells were treated with curcumin, the expression of NBR2 was significantly increased. NBR2 knockdown reversed curcumin-suppressed proliferation, clone formation, and the percentage of S-phase colorectal cancer cells. Furthermore, NBR2 knockdown abolished curcumin-induced activation of adenosine monophosphate-activated protein kinase and inactivation of the mTOR signaling pathway.
Conclusion:
This study revealed a novel mechanism by which long noncoding RNA NBR2 mediates curcumin suppression of colorectal cancer proliferation by activating adenosine monophosphate-activated protein kinase and inactivating the mTOR signaling pathway.
Introduction
Colorectal cancer (CRC) has become the third most prevalent malignant tumor worldwide and one of the leading causes of cancer-induced mortality. 1 Despite recent progress in early-stage screening and the development of more effective treatments for CRC, the prognosis of advanced stage CRC is still unfavorable. 2 Therefore, predictive biomarkers are clearly and urgently needed to detect CRC and to identify the most effective treatments for specific patients with CRC.
Long noncoding RNAs (lncRNAs) are a group of endogenous RNA molecules that are more than 200 nucleotides in length but lack an open reading frame.
3
A previous study revealed that lncRNAs impact a variety of important biological processes.
4
Liang
NBR2 is a noncoding sequence that resides adjacent to the well-known tumor suppressor gene BRCA1. Its biological function has remained unknown for many years since its initial discovery. 7 A previous study found that NBR2 interacts with adenosine monophosphate-activated protein kinase (AMPK) and promotes the kinase activity of AMPK, and NBR2 deficiency dampens glucose starvation-induced AMPK activation and AMPK-mediated downstream biological processes. 8
Adenosine monophosphate-activated protein kinase is one intracellular energy sensor. Structural analysis reveals that it has a heterotrimeric form consisting of 1 catalytic subunit (α) plus 2 regulatory subunits (β and γ).
9
Once activated, AMPK can phosphorylate various downstream effector substrates, such as those that serve to shut down anabolic processes and facilitate adenosine triphosphate-generating catabolic metabolism, thus helping to restore energy homeostasis after stress challenge.
10
Although we have plenty of evidence showing that AMPK regulates downstream targets and their biological implications, the mechanisms of AMPK activation itself remain unclear. A previous study demonstrated the direct interaction between NBR2 and the kinase domain of the AMPK complex, and an
Curcumin is identified in the rhizomes of
Methods
Cell Culture
A total of 4 human CRC cell lines, including HCT116, HCT8, SW480, and SW620, were purchased from the Type Culture Collection, Chinese Academy of Sciences (Shanghai, China) and cultured in Dulbecco Modified Eagle Medium (DMEM) containing 10% fetal bovine serum (Gibco, Waltham, MA, USA). Cells were cultured in a humidified incubator with 5% CO2, and the temperature was 37°C. The concentration of curcumin used in the cell treatment was 10 μM based on a previous study. 13 In glucose starvation assays, cells were treated with different concentrations of glucose (0, 1, or 25 mM) within DMEM containing 10% fetal bovine serum.
Cell Transfection
LncRNA NBR2 shRNA was purchased from GenePharma (Shanghai, China). HCT116 and SW480 CRC cells were seeded at a density of 1.8 × 105 cells per well in a 12-well plate. Transient transfection was conducted using a Lipofectamine 2000 kit (Invitrogen, Carlsbad, California, United States) following the instructions of the manufacturer.
MTT Assay
HCT116 and SW480 CRC cells were treated with 3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2-H-tetrazolium bromide (10% dilution in DMEM) at 37°C until the appearance of visible color. The cell proliferation rate was determined at 24, 48, and 72 hours posttransfection.
Colony Formation Assay
The colony formation assay was performed by seeding 200 cells into each well of the 6-well plate. After 14 days of incubation, cells were treated with 0.05% Crystal violet for 1 hour at room temperature. Under light field microscopy, the number of colonies in each well was enumerated. All groups were tested in triplicate.
BrdU Assay
HCT116 and SW480 CRC cells were seeded at 1.8 × 105 cells/well in DMEM containing 10 μM BrdU. After 3 hours incubation, cells were fixed for 15 minutes using 4% paraformaldehyde and were washed with phosphate-buffered saline (PBS) buffer followed by 0.5 mL Triton X-100. After further rinsing, the cells were treated with DNase (Tiangen, China) for 20 minutes at room temperature. Primary antibody against BrdU (Abcam, Cambridge, MA, USA) was then added following PBS washing for 12 hours incubation at 4°C. On the next day, cells were washed with PBS and incubated in secondary antibody for 60 minutes at room temperature. BrdU-positive cells were detected by flow cytometry.
RNA Isolation and Reverse Transcription-Polymerase Chain Reaction
Total cellular RNA was extracted using a Trizol kit (Takara, Kusatsu, Shiga, Japan) according to the manufacturer’s instructions. Cellular messenger RNA levels were quantified using SYBR PrimeScript RT-PCR Kit (Takara). The expression of NBR2 was measured using specific primers (Forward, 5′-GAGGTCTCCAGTTTCGGTAA-3′; Reverse, 5′-GAACCAAGGTGAAGGACCAA-3′). For the analysis of miRNA, we used stem-loop primers for reverse transcription and reverse transcription-polymerase chain reaction for miRNAs. These primers were purchased from RiboBio (Guangzhou, China). The relative expression of miRNAs was normalized to the internal control of the U6 RNA molecule (Forward, 5′-CGCTTCGGCAGCACATATAC-3′; Reverse, 5′-ATGGAACGCTTCACGAATTT-3′).
Western Blotting
Primary antibodies used in Western blotting, including anti-GAPDH (#5174), anti-p-AMPK (#4186), anti-AMPK (#2532), anti-p-ACC (#11818), anti-ACC (#3662), anti-p-S6 K (#2708), anti-S6 K (#9202), anti-p-S6 (#9209), and anti-S6 (#2317), were all purchased from Cell Signaling Technology (Danvers, MA, USA). In brief, cells were first lysed in Radio Immunoprecipitation Assay (RIPA) buffer. Proteins were quantified using the Bicinchoninic Acid (BCA) approach and were loaded onto 1× sodium dodecyl sulfate polyacrylamide gel electrophoresis for separation. Proteins were then transferred onto polyvinyl difluoride membranes (Millipore), which were incubated sequentially with primary and secondary antibodies (1:2000 dilutions for anti-GAPDH and 1:1000 for other antibodies).
Data Analysis
All statistical analyses were performed using the SPSS software package (version 17.0, IBM, Armonk, NY, USA). All data are shown as the mean ± standard error. Student
Results
Energy Starvation Could Increase the Expression of NBR2
Previous knowledge has revealed the activation of the AMPK pathway by starvation of cells. To verify the connection between glucose starvation-induced AMPK activation and NBR2 in CRC cells, we tested whether glucose-free medium would induce the change of NBR2 in CRC cell lines. The results revealed that glucose deficiency induced the increased expression of NBR2 in HCT116, HCT8, SW480, and SW620 cells compared to 25 mM glucose-treated cells (Figure 1A and 1B,

Glucose starvation-induced NBR2 upregulation in cultured colorectal cancer (CRC) cells. A and B, Reverse transcription-PCR showed the expression change of NBR2 in HCT116, HCT8, SW480, and SW620 cells treated with 0 or 25 mM glucose for 24 hours using 2 primer sets. C and D, Reverse transcription-PCR showed the expression of NBR2 in HCT116 and SW480 cells treated with 0 or 25 mM glucose plus Compound C treatment for 24 hours. E to G, Total, nuclear (Nul), and cytosolic (Cyt) expression of NBR2 plus normalization genes GAPDH1 and U1. Three independent experiments were performed, and the results are presented as the mean ± standard deviation. *
NBR2 Knockdown Suppressed the Glucose Deprivation-Induced AMPK/mTOR Signaling Pathway
To investigate the downstream signaling pathway of NBR2, we suppressed NBR2 expression in both HCT116 and SW480 cells. First, the transfection of shNBR2 effectively suppressed NBR2 transcript levels in both HCT116 (Figure 2A,

Knockdown NBR2 in HCT116 and SW480 cells and the detected expression of glucose starvation-induced changes in protein phosphorylation in AMPK/mTOR signaling. A and B, Reverse transcription-PCR showed the level of NBR2 in both HCT116 and SW480 cells after transfection with shNBR2. C-F, Western blotting protein bands and quantitative analysis showed changes in the phosphorylation levels of AMPK, ACC, S6K, and S6 proteins in both HCT116 and SW480 cells treated with 0 or 25 mM glucose for 24 hours. Three independent experiments were performed, and the results are presented as the mean ± standard deviation. *
NBR2 Knockdown Affects the Progression of CRC Cells Under Glucose Starvation Conditions
Then, we detected whether NBR2 affects the progression of CRC cells under glucose starvation conditions. First, MTT assay results revealed that shNBR2 suppresses the proliferation of either HCT116 or SW480 cells when compared to the control group under glucose deprivation conditions (Figure 3A and B,

NBR2 knockdown affects the progression of colorectal cancer (CRC) cells under glucose starvation conditions. A and B, MTT assay results revealed the proliferation of HCT116 and SW480 cells after transfection with shNBR2 and treatment with 0 or 25 mM glucose for 24 hours. C and D, Clonal formation assay showed the proliferation of HCT116 and SW480 cells under high-glucose or low-glucose conditions with shNBR2 transfection and treated with 1 or 25 mM glucose for 24 hours. E and F, BrdU assay showed the percentage of S-phase cells under high-glucose or low-glucose conditions in HCT116 and SW480 cells treated with 0 or 25 mM glucose for 24 hours. Three independent experiments were performed, and the results are presented as the mean ± standard deviation. *
NBR2 Knockdown Abolished the Inhibitory Effect of Curcumin in CRC Cells
Due to the known AMPK activation by curcumin, we further detected the role that curcumin played in HCT116 and SW480 cells, as curcumin significantly upregulated NBR2 in HCT116, HCT8, SW480, and SW620 cells (Figure 4A,

Curcumin elevated the level of NBR2 in CRC cells. A, RT-PCR showed the expression level of NBR2 in HCT116, HCT8, SW480, and SW620 cells after treatment with 0 or 10 μM curcumin for 24 hours. B, C, Reverse transcription-PCR showed NBR2 levels in curcumin-treated cells after shNBR2 transfection treated with 0 or 10 μM curcumin for 24 hours. Three independent experiments were performed, and the results are presented as the mean ± standard deviation. *
We further examined whether the suppression role of curcumin was dependent on NBR2. MTT results revealed a curcumin-induced suppression of proliferation of HCT116 and SW480 cells (Figure 5A and B,
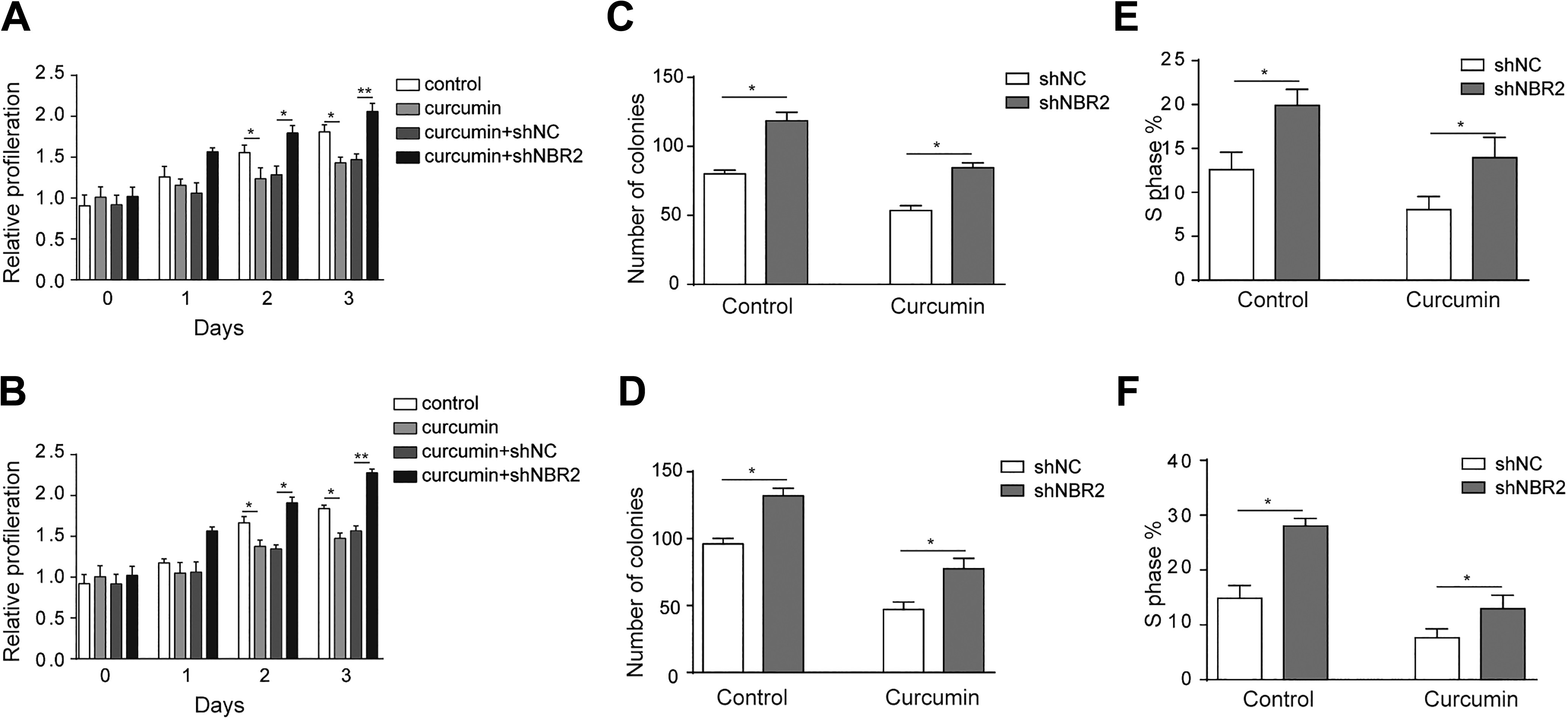
Inhibition effect of curcumin in CRC is mediated by NBR2. A and B, MTT assay revealed the proliferation of HCT116 and SW480 cells after treatment with 0 or 10 μM curcumin for 24 hours. C and D, Clone formation in curcumin-treated HCT116 and SW480 cells after shNBR2 transfection and treated with 0 or 10 μM curcumin for 24 hours. E and F, BrdU assay showed the percentage of S-phase cells induced by curcumin in HCT116 and SW480 cells after shNBR2 transfection and treated with 0 or 10 μM curcumin for 24 hours. Three independent experiments were performed, and the results are presented as the mean ± standard deviation. *
NBR2 Knockdown Eliminated the Effect of Curcumin on AMPK/mTOR Signaling
Finally, we investigated p-AMPK, p-ACC, p-S6K, and p-S6 in HCT116 and SW480 cells treated with curcumin. Similar to the glucose starvation treatment, curcumin significantly decreased the phosphorylation of p-AMPK and p-ACC (Figure 6A and C) and increased the level of p-S6K against total S6K (Figure 6B and D). With NBR2 knockdown in the curcumin-treated cells, the suppression of p-AMPK and p-ACC induced by curcumin was notably abolished, while the levels of p-S6K and p-S6 were unchanged upon curcumin treatment (Figure 6A-D). These results demonstrated that NBR2 is mediated by curcumin regulating AMPK/mTOR signaling.

NBR2 knockdown compromised curcumin-induced changes in protein phosphorylation in AMPK/mTOR signaling. A and B, Western blotting bands and quantitative analysis showed phosphorylation of AMPK, ACC, S6K, and S6 proteins in curcumin-treated HCT116 cells with shNBR2 transfection and treated with 0 or 10 μM curcumin for 24 hours. C and D, Phosphorylation of AMPK, ACC, S6K, and S6 proteins in curcumin-treated SW480 cells with shNBR2 transfection and treated with 0 or 10 μM curcumin for 24 hours. Three independent experiments were performed, and the results are presented as the mean ± standard deviation. AMPK indicates adenosine monophosphate-activated protein kinase.
Effects of Curcumin on AMPK/mTOR Signaling Under Glucose Deprivation
To further elucidate the mechanism of curcumin, we treated cells under glucose-deprivation conditions. Under glucose-supplemented conditions, curcumin treatment significantly activated p-AMPK and p-ACC (Figure 7A and C) and suppressed the p-S6K/p-S6 pathway (Figure 7B and D). Furthermore, glucose-deficient conditions facilitated the curcumin-activated AMPK-ACC pathway and inactivated the S6K-S6 pathway (Figure 7). These results demonstrated that curcumin and glucose deficiency have a synergetic effect in regulating the AMPK/mTOR pathway under glucose-deficient conditions.
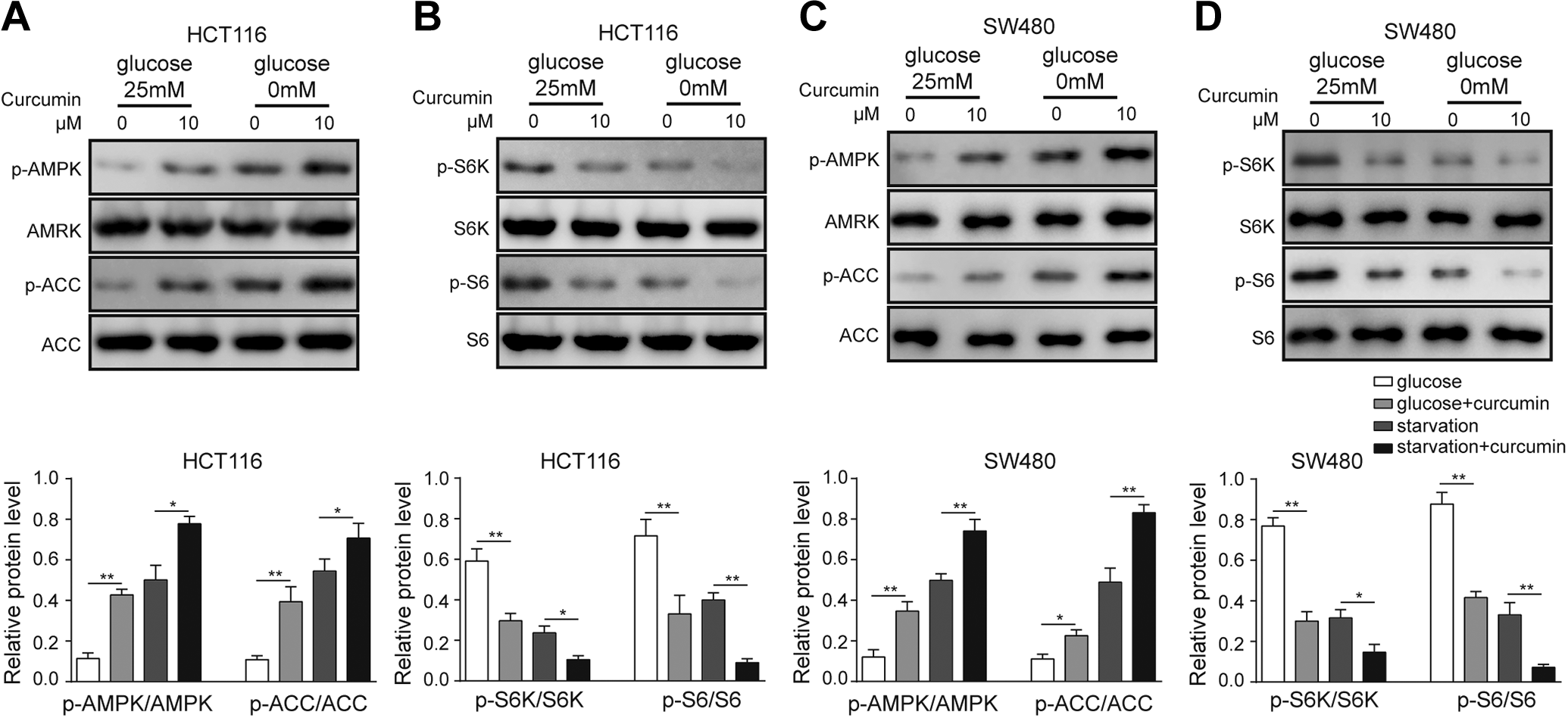
Regulation of the AMPK/mTOR pathway by curcumin under glucose-replenishment and glucose-deficient conditions. A, Western blotting and quantitative analysis of p-AMPK and p-ACC in HCT116 cells by curcumin treatment. B, Western blotting for the relative expression of the p-S6K/p-S6 pathway in HCT116 cells. C, Western blotting and quantitative analysis of p-AMPK and p-ACC in SW480 cells by curcumin treatment. D, Western blotting for the relative expression of the p-S6K/p-S6 pathway in SW480 cells. Three independent experiments were performed, and the results are presented as the mean ± standard deviation. *
Discussion
In this study, we showed that NBR2 expression was strongly induced by glucose deprivation in cultured CRC cells. Activated NBR2 may further activate the AMPK signaling pathway and thus suppress the proliferation of tumor cells. These results are consistent with previous reports showing the tumor suppressor function of NBR2. 8 Both RNA sequencing and analysis of noncoding RNA profiles have identified NBR2 as a candidate lncRNA with a response to glucose starvation. 14 In the present study, we found that glucose deficiency induces the expression of NBR2 in CRC cells via the energy stress sensor AMPK. This NBR2 upregulation sequentially inactivated the mTOR signaling pathway and activated AMPK, thus forming a positive feedback loop to further potentiate AMPK under energy stress. These cellular events further led to cell proliferation suppression, whereas NBR2 knockdown abolished this process.
NBR2 has been demonstrated as one tumor suppression factor via transregulation with the BRCA1 promoter using a breast cancer cell model.
15
Our study is the first to report NBR2 upregulation induced by glucose deprivation in CRC cells
In conclusion, this study revealed the inhibitory effect of curcumin on the progression of CRC via modulating the NBR2/AMPK/mTOR pathway. Further studies are thus required to illustrate the
Footnotes
Authors’ Note
Our study did not require an ethical board approval because it did not contain human or animal trials.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The project was supported by the Ningbo Natural Science Foundation (Grant Nos: 2018277842, 2018262634, 2017A610145, 2017A610158, 2016A610135) and the Zhejiang Medical and Health Bureau (2017KY593, 2017KY594).
