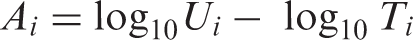Abstract
Healthcare-associated infections are of global concern, and textiles can contribute to the transmission of pathogens. In this study, we examined quantitatively the survival capability of 60 multidrug-resistant bacterial strains from four species (Klebsiella pneumoniae, Acinetobacter baumannii, Staphylococcus aureus and Enterococcus faecium) on untreated cotton textile in clinically relevant incubation periods. We determined the antibacterial efficiency of textiles treated either with quaternary ammonium compound (QAC)-containing Sanitized T99-19 liquid (50 m/m% Dimethyltetradecyl (3-(trimethoxysilyl)propyl) ammonium-chloride) or with silver salt-containing Sanitized T27-22 Silver liquid (2 m/m% AgCl and 8 m/m% TiO2) as well. Finally, we compared the results of the healthcare-associated, multidrug-resistant strains and antibiotic-sensitive, quality control standard strains (ATCC 25922, ATCC 11105 Escherichia coli, and ATCC 25923, ATCC 6538 Staphylococcus aureus) often used in antimicrobial efficiency tests. The results revealed that all investigated multidrug-resistant bacteria are able to survive on untreated cotton textile and pose health risk in hospitals. During one day the T27-22-Silver-treated textile was able to eliminate most of the Gram-positive pathogens, reducing the risk of cross-contamination, but none of the examined agents destroyed the multidrug-resistant, Gram-negative isolates. The antibiotic-susceptible and the multidrug-resistant Staphylococcus aureus strains had similar survival capability and biocide-tolerance, while the risk of infections caused by multidrug-resistant, Gram-negative pathogens could be extremely underestimated using only ATCC Escherichia coli standard strains. Our results also draw attention to the careful evaluation of antimicrobial efficiency tests and indicate that a significant reduction of bacterial count does not necessarily mean significant antibacterial efficiency that would be suitable to avoid infections.
Keywords
Introduction
Healthcare-associated infections (HAIs) are of global concern. By definition for a HAI, the infection must not be present before someone has been under medical care, the first symptoms must appear at least 48–72 hours after the admission into the hospital [1]. According to the data of European Centre for Disease prevention and Control (ECDC), more than 116,000 patients contact at least one HAI in European hospitals every day [2]. In Hungary 136 healthcare-associated outbreaks were reported in 2015 [3]. HAIs are associated with increased morbidity and mortality, longer length of hospital stay, and also represent a major financial burden on healthcare services. The lion’s share of HAIs are caused by a relatively small group of bacterial pathogens – the so-called ‘ESKAPE bugs’ – six key pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa and Enterobacter species) which compromise the healthcare provision [4]. These bacteria are highly virulent, and are able to survive on inanimate surfaces for a longer time and to develop antibiotic-resistance. Within the given species, the multidrug-resistant strains causing major outbreaks in hospitals all over the world represent a relatively few successful, ‘particularly fit to the task’ clones (i.e. genetically closely related strains).
In this study, the survival capability of four ‘ESKAPE bugs’ were investigated on untreated and on two kinds of antimicrobial cotton textiles treated with often used antimicrobial agents. The Gram-negative K. pneumoniae, A. baumannii and the Gram-positive S. aureus, E. faecium frequently occur in intensive care units (ICUs), and primarily attack immunocompromised patients who are subjected to invasive procedures and treated with broad-spectrum antibiotics.
These pathogens cause a variety of clinical infections ranging from minor local skin, soft tissue, surgical wound or urinary tract infections, to life-threatening bacteraemia, sepsis, pneumonia or meningitis [2,5–8]. They have intrinsic resistance mechanisms (chromosomally encoded innate ability) against several groups of antibiotics and they can possess acquired resistance mechanisms as well. Acquired resistance mechanism means that a microorganism obtains the ability to resist an antimicrobial agent to which it was previously susceptible [9–12]. These pathogens are usually resistant to a variety of classes of antibiotics causing serious therapeutic problems and public health concern worldwide.
These multidrug-resistant pathogens can have a significant capacity for long-term survival (even for months and years) on inanimate surfaces, with consequently enhanced opportunities for transmission between patients, either via human hosts or via inanimate materials [13]. Textiles in contact with the human body (bed clothes, pyjamas, towels, uniforms of healthcare workers, etc.) can be contaminated with resistant strains, contributing to the transmission of the HAIs and representing sources of outbreaks [14–19]. Antimicrobial finishing agents are able to eliminate or at least reduce the number of bacteria on abiotic surfaces, possibly preventing HAIs [20,21].
In our study we examined the antibacterial efficiency of two antimicrobial liquids from Sanitized AG, which contain antimicrobial agents often used against the aforementioned internationally important multidrug-resistant bacteria. Sanitized T27-22 Silver and T99-19 liquids contain 2 m/m% AgCl + 8 m/m% TiO2 and 50 m/m% Dimethyltetradecyl (3-(trimethoxysilyl)propyl) ammonium-chloride, respectively. Silver salts and Quaternary Ammonium Compounds (QACs) belong to the most common antimicrobials used for textile finishing [22]. Both agents proved to be microbicide at low concentrations against a wide range of microorganisms without adverse effects for humans. They have multiple target sites within the bacterial cell, for example they are able to destroy the cell membrane and inhibit the DNA replication [23–26]. Silver can inhibit the cell’s enzyme activities as well. Titanium dioxide is a photocatalytic molecule often used in combination with silver compounds. By creating free radicals, titanium dioxide can destroy the organic compounds and cause the death of the bacterial cells [23]. However, according to some studies titanium dioxide itself has no significant antibacterial effect [27,28].
Aims of the study
The aims of our study were i) to examine the survival capability of four multidrug-resistant healthcare-associated bacterial species on untreated cotton sheet, ii) to investigate the antibacterial efficiency of cotton textile treated with a silver-chloride or a QAC-containing agent, and finally iii) to compare the survival capability of the multidrug-resistant bacteria with antibiotic-sensitive quality control standard strains that are often used in efficiency tests of antimicrobial textiles.
Material and methods
A total of 60 healthcare-associated, multidrug-resistant isolates were examined for their survival capability on untreated and two kinds of antibacterial textiles. The Gram-positive bacteria we used were 15 vancomycin-resistant E. faecium (VRE) and 15 methicillin-resistant S. aureus (MRSA) strains. The Gram-negative isolates were 15 multidrug-resistant A. baumannii (MACI) and 15 multidrug-resistant K. pneumoniae (MRKP). The investigated collection consists of well characterized multidrug-resistant bacterial strains originated from different areas of Hungary between 1998 and 2014. Detailed information about the selected strains is shown in the Supplementary material – table 1.
To compare the survival capability of healthcare-associated pathogens and bacterial standards often used in antimicrobial efficiency tests [29], we selected four quality control strains from the American Type Culture Collection (ATCC). We examined two Gram-negative strains: ATCC 25922 and ATCC 11105 Escherichia coli and two Gram-positive ones: ATCC 25923 and ATCC 6538 Staphylococcus aureus.
Swatches of plain weaved, 100% cotton sheet (areal density: 104 g/m2) were treated either with QAC-containing Sanitized T99-19 (50 m/m% Dimethyltetradecyl (3-(trimethoxysilyl)propyl) ammonium-chloride) or with silver salt-containing Sanitized T27-22 Silver liquid (2 m/m% AgCl and 8 m/m% TiO2), according to the manufacturer’s (Sanitized AG) instructions. Hereinafter we refer to the Sanitized liquids as T99-19-QAC and T27-22-Silver. Homogeneous coating solutions of antibacterial agents were made in the concentration of 5.6 and 6.6 g/L of T99-19-QAC and T27-22-Silver liquid, respectively. The cotton swatches (10 × 20 cm) were submerged in either of the coating solutions for one minute. Then the excess was squeezed using a hand-powered laboratory padding machine (averaged percent expression: 106%). Finally, the coated swatches were dried at room temperature. According to the manufacturer’s instructions heat treatment of the textile was not necessary, as it only serves to increase the washing durability, which was not investigated in this study. Untreated cotton fabric was used as control. To examine the survival capability of bacteria, we cut the untreated and the antimicrobial textiles into square shaped swatches, with a side length of 2.5 cm (area: 6.25 cm2).
Bacterial strains were grown overnight on blood agar plates (Columbia agar with 5% sheep blood, manufactured in the National Public Health Institute, Hungary), and were examined one by one (Figure 1).
The examination of the survival capability of bacterial strains on untreated and two kinds of antibacterial textiles. McF: McFarland (optical density of the bacterial suspension, 0.5 McF = 1.5 × 108 CFU/ml); CFU: colony forming unit; T: temperature; Rh: relative humidity.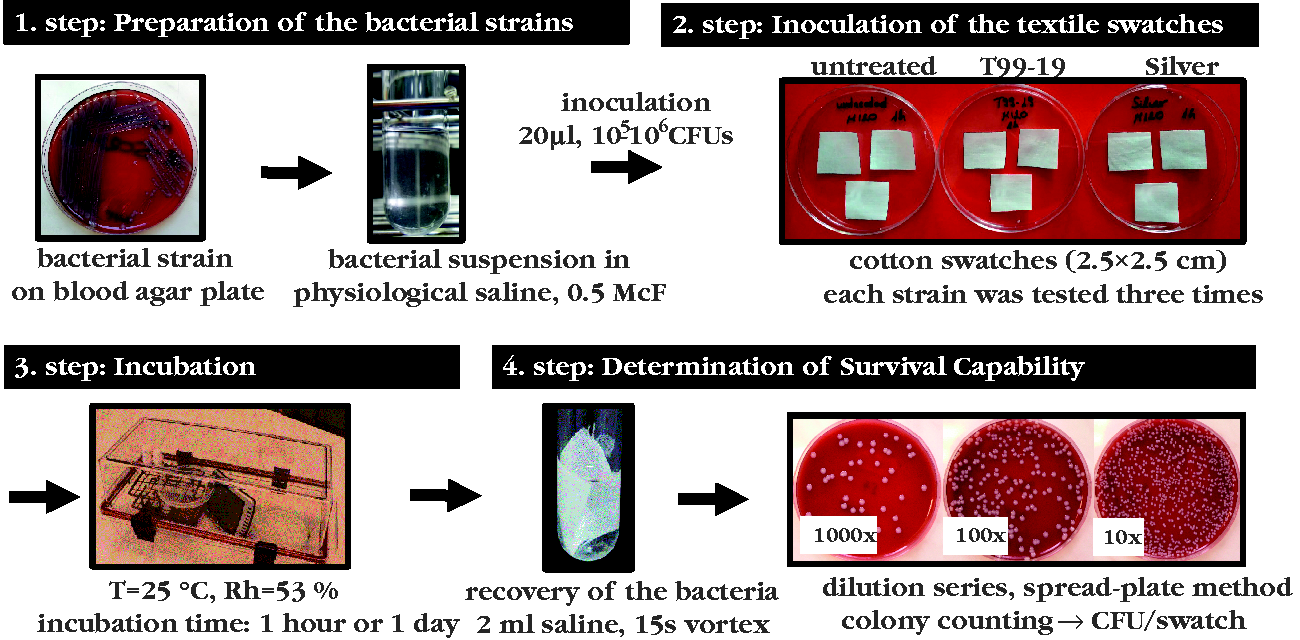
We adjusted the density of each bacterial suspension in 0.9 w/w% physiological saline solution to 0.5 McFarland using a McFarland turbidity meter. The working solutions were obtained after a tenfold dilution. 20 μl of working solution (containing approximately 105–106 Colony Forming Units (CFUs)) was inoculated onto the untreated and the two kinds of treated swatches. We made one working solution of each individual isolate and inoculated all three types of textile for each incubation period at the same time. We used the absorption method (ISO 20743-2013 [30]), that is, the working solution was inoculated at several points (four-six) of the swatches. The bacterial count in each working solution was determined by the plate count method. From each working solution, a dilution series was made (10,000×, 100,000× and 1,000,000×). Then 100 µl of each dilution was plated uniformly (spread-plate method) onto the surfaces of blood agar plates. After overnight incubation at 35℃, the bacterial colonies were counted manually. We tested each strain three times, the inoculated CFU of the strains was determined as the arithmetic mean of the three tests.
The inoculated textile samples were incubated at 25℃ and 52% relative humidity (maintained using saturated Mg(NO3)2 solution), which modelled the common environmental circumstances in a hospital ward. Based on our pilot study, we investigated the survival capability of the isolates after one hour and one day of incubation time. After the incubation, we added 2 ml of 0.9% w/v saline solution to each inoculated swatch, then vortexed them for 15 s. From the solution we got, a dilution series was made (10×, 100× and 1000×), and the viable CFU was determined using spread-plate method. Each strain was tested three times. The survival capability of a bacterial strain was determined as the arithmetic mean of the viable CFU/swatch obtained from the three parallel tests. Thus, we got one survival capability value for each textile type – incubation time – bacterial strain combination.
The Reduction Rate Value (R) of each individual strain on the untreated textile was calculated as the logarithmic reduction of the inoculated CFU/swatch value by different incubation times according to the next formula:
The Antibacterial Activity Value (A) of a treated textile was calculated according to ISO 20743-2013 [30] based on the paired survival capability values of each individual strain by different incubation times as
We calculated the mean (M) and the standard deviation (SD) of the individual Reduction Rate Values (R) and the individual Antibacterial Activity Values (A) of the isolates within each species. The standard deviation (SD) values were calculated according to the following formula
We used the IBM SPSS Statistics Data Editor to perform the statistical tests; P ≤ 0.05 was considered to be statistically significant. By means of one-way ANOVA model, we compared the Reduction Rate Values of the isolates on untreated textiles and the Antibacterial Activity Values of treated textiles according to Gram-classification, species and genetic diversity (Supplementary material – table 1). In case of all four species, we classified the bacteria by their clonal-types or resistance-types (MRKP: CG14/15 and CG258 international or sporadic clonal groups; MACI: CC1 and CC2 international clonal complexes; MRSA: hospital-acquired (HA-MRSA) and community-associated (CA-MRSA) strains or EMRSA-15 and New York/Japan international clones; VRE: strains carried vanA or vanB resistance genes). We used dependent t-test to compare the survival capability of the strains on the untreated and the two types of antimicrobial textiles.
Results
The survival capability of the bacteria after one hour incubation time
Survival capability on untreated textile after one hour incubation time
Mean Reduction Rate Values (RM) of the multidrug-resistant isolates and the ATCC standard strains by species on untreated cotton sheet.

RM: Mean Reduction Rate Values; SD: standard deviations; ATCC: American Type Culture Collection.
The swatches were inoculated with an average CFU values of 1.69 × 105, 8.16 × 105, 2.07 × 106 and 2.21 × 105 in case of MRKP, MACI, MRSA and VRE, respectively. All of the 60 multidrug-resistant bacterial strains retained their viability during one hour incubation period on the surface of dry cotton textile in nutrient-poor circumstances. Among the examined multidrug-resistant bacterial groups, MRKP and MRSA isolates had similarly high mean Reduction Rate Values (RM = 3.0 ± 0.6 and RM = 3.0 ± 0.4 in case of MRKPs and MRSAs, respectively) on the untreated textile after one hour incubation time (Table 1). The mean Reduction Rate Value of VRE and MACI isolates was 2.2 ± 0.3, 1.8 ± 0.4, respectively. MRKP and MRSA strains showed significantly higher Reduction Rate Values on untreated cotton surface than the MACI and VRE isolates (ANOVA, P < 0.01). Comparing the survival of MACI and VRE isolates, VREs showed significantly higher reduction during one hour incubation than MACIs (ANOVA, P < 0.01).
In the examination of ATCC standard strains, swatches were inoculated with an average CFU values of 4.91 × 105 E. coli and 2.09 × 106 S. aureus bacteria. The two ATCC E. coli strains showed a high mean Reduction Rate Value, (RM = 4.3 ± 1.2) even higher than MRKP and MRSA strains (Table 1). After one hour of incubation time only the ATCC 11105 strain was recoverable, ATCC 25922 perished. The mean Reduction Rate Value of ATCC S. aureus strains was RM = 3.1 ± 0.1, similar to MRSAs.
Survival capability on antibacterial textiles after one hour incubation time
The textile treated with T99-19-QAC antimicrobial agent reduced the survival capability of all examined Gram-positive and Gram-negative strains (Figure 2).
The survival capability of the multidrug-resistant and ATCC standard strains on untreated (control) and two kinds of antibacterial textiles is shown by species after one hour incubation time. The mean inoculated CFU values were 1.69 × 105, 8.16 × 105, 2.07 × 106, 2.21 × 105, 4.91 × 105 and 2.09 × 106 in case of MRKP, MACI, MRSA, VRE, ATCC E. coli and ATCC S. aureus strains, respectively. Boxes show the median and interquartile range (IQR), whiskers indicate the values of the <75th percentile + 1.5 × IQR> and the <25th percentile – 1.5 × IQR>. The black diamonds show the outliers and the extreme values. At the top of the box-plots we indicated the significant efficiency of the treated textiles according to dependent t-test for paired samples (*(P < 0.05), and **(P < 0.01)), and the significant (A) antibacterial efficiency of the treated textiles based on Antibacterial Activity Values.
Based on dependent t-test for paired samples the survival capability of healthcare-associated MRKP (P = 0.02), MACI (P < 0.01), MRSA (P < 0.01) and VRE (P < 0.01) isolates were significantly lower on textile treated with T99-19-QAC than on untreated cotton. On the other hand only 3 MRKP isolates perished on the T99-19-QAC treated textile during one hour of incubation from the examined 60 multidrug-resistant isolates. That is, 12 MRKP (on average 1.39 × 102 CFU/swatch), all of the 15 VRE (4.92 × 102 CFU/swatch), all of the 15 MACI (2.11 × 103 CFU/swatch), and all of the 15 MRSA (5.11 × 102 CFU/swatch) strains retained their viability on the textile treated with T99-19-QAC.
On the textile treated with T99-19-QAC the average survival capability of the two ATCC E. coli standards was 1.00 × 101 CFU/swatch. The ATCC 25922 strain perished during one hour not only on the textile treated with T99-19-QAC, but also on untreated cotton. On average, 7.03 × 102 CFUs were recoverable in case of ATCC S. aureus standards from the swatches treated with T99-19-QAC.
Mean Antibacterial Activity Values (AM) of the treated textiles against the multidrug-resistant isolates and the ATCC standard strains by species.

*Indicates significant (2 ≤ A < 3) antibacterial activity.
AM: mean Antibacterial Activity Values; SD: standard deviation; T99-19-QAC: antibacterial textile treated with Sanitized T99-19-QAC liquid; T27-22-Silver: antibacterial textile treated with Sanitized T27-22-Silver liquid; ATCC: American Type Culture Collection.
The textile treated with T27-22-Silver liquid reduced the survival capability of all examined bacteria after one hour incubation time, but the efficiency was different against the Gram-negative and Gram-positive isolates (Figure 2).
The average survival capability of the Gram-negative MRKP isolates was 3.46 × 102 CFU/swatch, and only 3 strains perished from the examined 15 ones. According to dependent t-test and the Antibacterial Activity Values the textile treated with T27-22-Silver liquid did not have a significant effect on the survival capability of MRKP bacteria. The survival capability of the MACI isolates reduced significantly on the textile treated with T27-22-Silver liquid according to dependent t-test (P < 0.01), but at the same time none of the examined 15 strains perished. On average 3.59 × 103 CFUs of MACI were recoverable from the swatches. Based on Antibacterial Activity Values the textile treated with T27-22-Silver did not have significant antibacterial efficiency against MACI strains. Both examined ATCC E. coli strains perished on the T27-22-Silver antibacterial textile during one hour, but the ATCC 25922 isolate was not recoverable either from the untreated surface. According to the mean Antibacterial Activity Values (Table 2), the T27-22-Silver liquid did not have significant antibacterial efficiency against E. coli strains. The high standard deviation of the A-values indicated that the biocide-tolerance of the two ATCC E. coli isolates was very different (Table 2).
On T27-22-Silver-treated textile, the survival capability of Gram-positive isolates decreased to a greater extent than of Gram-negative ones (Figure 2). In case of the multidrug-resistant, Gram-positive strains the reduction of survival capability was significant compared to untreated textile (t-test, P < 0.01). 7 MRSA (1.16 × 101 CFU/swatch) and 7 VRE (on average 1.29 × 101 CFU/swatch) strains retained their viability, 8 MRSA and 8 VRE perished during one hour. In the Gram-positive group, the CFU values were significantly lower on the T27-22-Silver treated than on the T99-19-QAC treated fabric (t-test, P < 0.01). The ATCC 6538 S. aureus standard perished as early as 10 min on the fabric treated with T27-22-Silver liquid (data not shown). The Antibacterial Activity Values of T27-22-Silver treated textile showed significant antibacterial efficiency against VRE, MRSA and ATCC S. aureus isolates (Table 2).
The survival capability of the bacteria after one day incubation time
Survival capability on untreated textile after one day incubation time
The results are shown on Table 1. Swatches were inoculated with an average CFU values of 1.69 × 105 MRKP, 8.16 × 105 MACI, 2.07 × 106 MRSA and 2.21 × 105 VRE bacteria. From the examined 60 multidrug-resistant bacterial strains 55 retained their viability on untreated textile during the one day incubation period. All the 5 strains which perished belonged to MRKP. MRKP isolates had the highest Reduction Rate Values (RM = 4.3 ± 0.6) of the investigated four healthcare-associated bacterial species (Table 1). The mean Reduction Rate Value of MRSA, VRE and MACI isolates was 3.6 ± 0.3, 2.7 ± 0.4 and 2.3 ± 0.5, respectively. The Reduction Rate Values of MRKP and MRSA strains were significantly higher than those of MACI’s and VRE’s on untreated textile after one day incubation time (ANOVA, P < 0.01). Comparing the survival of MRKP and MRSA isolates, MRKPs showed significantly higher reduction during one day incubation than MRSAs (ANOVA, P < 0.01).
During the examination of ATCC standard strains we inoculated the textile swatches with on average 4.91 × 105 CFUs of E. coli and 2.09 × 106 CFUs of S. aureus. The ATCC E. coli strains showed higher Reduction Rate Values (RM = 5.1 ± 0.0) than the multidrug-resistant pathogens (Table 1). The mean Reduction Rate Value of the two ATCC S. aureus (RM = 3.5 ± 0.1) corresponded with the result of MRSA strains.
Survival capability on antibacterial textiles after one day incubation time
The textile treated with T99-19-QAC agent reduced the survival capability of all examined Gram-positive and Gram-negative bacteria (Figure 3).
The survival capability of the multidrug-resistant and ATCC standard strains on untreated (control) and two kinds of antibacterial textiles is shown by species after one day incubation time. The mean inoculated CFU values were 1.69 × 105, 8.16 × 105, 2.07 × 106, 2.21 × 105, 4.91 × 105 and 2.09 × 106 in case of MRKP, MACI, MRSA, VRE, ATCC E. coli and ATCC S. aureus strains, respectively. Boxes show the median and the interquartile range (IQR), whiskers indicate the values of the <75th percentile + 1.5 × IQR> and the <25th percentile − 1.5 × IQR>. The black diamonds show the outliers and the extreme values. At the top of the box-plots we indicated the significant efficiency of the treated textiles according to dependent t-test for paired samples (* (P<0.05), and ** (P<0.01)), and the significant (A) antibacterial efficiency of the treated textiles based on Antibacterial Activity Values.
According to the dependent t-test for paired samples MRKP (P = 0.02), MACI, MRSA and VRE (P < 0.01) strains showed significantly lower survival capability on T99-19-QAC treated textile compared to untreated textile. Nevertheless 4 MRKP (on average 5.44 × 100 CFU/swatch), 12 MRSA (on average 3.20 × 101 CFU/swatch), 14 VRE (on average 2.03 × 102 CFU/swatch) and all of the 15 MACI strains (on average 6.60 × 102 CFU/swatch) retained their viability on the textile treated with T99-19-QAC agent. The Antibacterial Activity Values of T99-19-QAC did not show significant antibacterial efficiency against any of the healthcare-associated species we examined (Table 2). In case of MRSA isolates the close to significant A-values and the high standard deviation indicated the different biocid-tolerance of the strains.
The ATCC 25922 E. coli isolate perished even on the untreated textile during one hour incubation time, thus the efficiency of the antibacterial agents was not measurable in that case. The ATCC 11105 E. coli isolate was recoverable from the untreated textile, and perished on the T99-19-QAC treated fabric during one day of incubation (Figure 3), but based on the Antibacterial Activity Value the antibacterial effect was not significant. On the textile treated with T99-19-QAC, the ATCC S. aureus strains had higher survival capability (on average 1.70 × 102 CFU/swatch) than MRSA strains. Significant antibacterial efficiency was not found in this case either (Table 2).
The T27-22-Silver liquid showed low long-term antibacterial efficiency against the multidrug-resistant Gram-negative strains (Figure 3). Although it reduced the survival capability of MACI strains significantly (t-test, P < 0.01), 14 isolates from the examined 15 retained their viability (on average 1.42 × 103 CFU/swatch). Even 5 MRKP isolates retained their viability (on average 1.42 × 101 CFU/swatch) on T27-22-Silver treated fabric, although only 10 MRKP strains were recoverable from the untreated textile. The Antibacterial Activity Values confirmed the low antibacterial efficiency of T27-22-Silver-treated cotton against Gram-negative species (Table 2).
The T27-22-Silver treated textile eliminated 24 isolates from the 30 examined healthcare-associated, multidrug-resistant Gram-positive bacteria. After one day incubation 2 MRSA (on average 1.33 × 100 CFU/swatch) and 4 VRE (on average 3.11 × 100 CFU/swatch) isolates retained their viability on T27-22-Silver-treated textile. The significant antibacterial efficiency was confirmed by dependent t-test for paired samples (P < 0.01) and by Antibacterial Activity Values (Table 2) as well. The ATCC 6538 S. aureus standard strain perished already during one hour of incubation, but the ATCC 25923 S. aureus strain was recoverable (2.00 × 101 CFU/swatch) after one day as well. Based on Antibacterial Activity Values, the effect of T27-22-Silver treated textile was significant against ATCC S. aureus stantard strains (Table 2).
The comparison of survival capability within species
We grouped the isolates within species based on their clonal-types and resistance-types (Supplementary material – table 1). Several significant associations were detected.
The MACI strains belonged to the CC1 international clone showed significantly higher survival capability on untreated textile than CC2 MACI strains (ANOVA, P < 0.01, one hour and one day incubation time as well). At the same time, the T27-22-Silver treated antibacterial textile had significantly higher antibacterial efficiency against CC1 than CC2 MACI isolates (ANOVA, P < 0.01, one hour and one day incubation times).
The CA-MRSA strains showed significantly higher survival capability on untreated textile than HA-MRSA isolates (ANOVA, P = 0.02, one day incubation time).
The textile treated with T99-19-QAC agent had significantly higher antibacterial efficiency (ANOVA, P < 0.01) against vanB VRE than vanA VRE isolates.
Discussion
Survival capability of bacteria on untreated textile
We examined the survival capability of 60 healthcare-associated, multidrug-resistant and 4 ATCC standard strains on untreated cotton sheet in a dry, nutrient-poor environment. In the study of Messaud et al., both the E. coli and Listeria innocua strains increased their CFUs by two orders of magnitude on untreated cotton-polyester textile after one day of incubation [27]. However, in our investigation all the 64 investigated strains were unable to proliferate in nutrient-poor environment, incubated at room temperature and normal relative humidity. Moreover, the survival capability decreased in a species-dependent quantity over the incubation time. This indicates that textiles themselves do not provide a suitable environment to the proliferation of pathogens, even those made of cotton-based natural fibers. The role of organic and inorganic contaminants, humidity and temperature in the survival of bacteria is still controversial [13].
Even if the bacteria cannot proliferate on dry textiles, they are able to keep their survival capability and virulence for some time, even for years in extreme cases [13,15,31,32]. The duration of survival can vary largely within one bacterial species. For example, while one K. pneumoniae strain was able to survive for more than 30 months, another one perished after 2 h [13]. From the perspective of HAIs, the most relevant question is the average survival capability of the most important multidrug-resistant strains for a few hours or maybe a day. In our study, 60 and 55 multidrug-resistant strains retained their survivability after one hour and one day of incubation, respectively. MRKP strains were the least viable, the highest CFU/swatch values were measured in case of MACI isolates. Our results were supported by the data in the literature [31,33–35]. Based on our investigation, all examined multidrug-resistant pathogens were able to survive on untreated cotton fabric for enough time and in adequate numbers to cause cross-contaminations. According to the review of Dancer, an MRSA inoculum containing ten to several million CFUs could potentially cause infections [36]. Experimental epidermis lesions, which modelled the insertion sites of medical devices (e.g. vascular catheter) can be infected by as few as 15 CFUs of S. aureus [37]. Only 10 CFUs of S. aureus introduced on suture material or cotton dust can produce skin abscesses in mice [38]. Each species we investigated showed several orders of magnitude higher survival capability on untreated cotton textile after one hour and – except for MRKP – after one day incubation time as well.
There is no need for long-term bacterial survival to cause cross-contaminations. According to Bhalla et al., only a few touches of environmental surfaces near hospitalized patients could be enough to contaminate the healthcare workers’ hands with multidrug-resistant pathogens [39]. Based on the investigation of Hayden et al., 52% of healthcare workers contaminated their hands or gloves with VRE bacteria after touching only the environment of a patient [40]. In other studies, indistinguishable MRSA strains were isolated from the patients and their environments [41,42], and in the study of Dancer et al. indistinguishable coagulase-negative staphylococci were detected not only from the patient’s blood and environment but also from the hands of staff [43]. According to Riley et al. cross-contamination could happen during laundering at 40℃ [44]. These findings support the presumption that a pathogen transmission can occur among the patients, the environment and the healthcare workers. All the species we investigated were able to survive at least one day on untreated cotton textile, which is more than enough to multiply the risk of cross-contamination.
According to our results there were differences in the survival capability of strains within the species, but the differences were greater among the species. Thus, the survival capability seems to be species-dependent in the examined circumstances.
Because of the differences in viability, it is important to be careful in the selection of bacterial species and strains for the survival tests. Our findings suggest that the widely used ATCC 6538 and ATCC 25923 S. aureus isolates can be appropriate to investigate the survivability of Gram-positive pathogens on untreated and on antimicrobial textiles. However, the ATCC 11105 and ATCC 25922 E. coli strains proved to be significantly less viable than most of the investigated Gram-negative, multidrug-resistant isolates. With the aforementioned ATCC E. coli standards, it is possible to underestimate the risk of infection caused by Gram-negative pathogens.
The efficiency of antimicrobial textiles
We examined the antibacterial efficiency of cotton sheet treated with two kinds of antibacterial agents, often used in textile industry. The incubation periods of one hour and one day were used to study the short- and long-term effects of the agents. According to Mitchell et al., not only the hospital textiles, but also the clothing and hands of staff and visitors could be contaminated with pathogens [16], thus quick-acting antibacterial agents are needed to reduce the chance of cross-contaminations. Therefore, the results of our one hour efficiency tests are particularly important.
It is difficult to compare the efficiency of antimicrobial textiles described in different investigations, because a lot of factors can influence the results. Such influencing factors are the properties of the examined bacterial strains, parameters of the antimicrobial coating (agent, concentration, method), the size of the inoculum (CFUs) and the circumstances of the incubation. The careful evaluation of the results is equally important. We analysed our data in several ways. Our results showed that a significant decrease in the bacterial CFU does not necessarily mean significant antibacterial efficiency.
The exact concentration of the agents is not described in every scientific article, which makes the comparison of the efficiency more complicated. The agent content of the antibacterial textiles we used was ∼3000 mg QAC/kg dry textile and 140 mg AgCl/kg dry textile in case of T99-19-QAC and T27-22-Silver treated sheets, respectively (based on calculation). This is in accordance with the recommendations and practices described in publications [22,45,46]. We used the bath-method for the treatment of cotton fabric, which is more suitable for the application of T99-19-QAC than the spraying method according to Gutarowska et al. [47].
Antibacterial agents of Sanitized were investigated previously, but according to our knowledge the antibacterial efficiency of these agents has not been examined using such numerous, multidrug-resistant, healthcare-associated pathogens so far [47–54].
The antibacterial efficiency of textiles treated with QAC agents
Budimir et al. examined the efficiency of 100% cotton fabric treated with T99-19-QAC. After one day incubation time, the Antibacterial Activity Values against ATCC 6538 S. aureus (examined by us as well) and ATCC 10536 E. coli were 4.71 and 3.9, respectively [50]. The significant effect during one day incubation time is inconsistent with our results. According to our experiments, the cotton treated with T99-19-QAC did not show significant efficiency neither against ATCC standard strains nor against healthcare-associated isolates. The two ATCC E. coli strains we examined showed low survival capability even on untreated textile, therefore significant A-values were not expected. It is possible that the E. coli isolate examined by Budimir et al. is a more desiccation-tolerant and QAC-sensitive strain. The experienced inconsistency in the behaviour of ATCC 6538 S. aureus strain can be explained by the different inoculating method, as Budimir et al. used broth, we used physiological saline. The presence of nutrients or the different agent content in the antibacterial textiles might be the cause of the discrepancy (Budimir et al. did not publish the applied concentration of T99-19-QAC).
QAC agents similar to T99-19-QAC show greater antibacterial activity against Gram-positive bacteria than Gram-negative ones [55–57]. In our study the T99-19-QAC agent was indeed more efficient against the Gram-positive, multidrug-resistant MRSA than against the other species. Nonetheless only three MRSA isolates perished completely from the treated textile during one day incubation time. It is important to note that none of the aforementioned publications examined the antibacterial efficiency against VRE isolates, which had great survival capability on T99-19-QAC treated textile in our study. Thus, the application of VRE isolates in efficiency tests could be recommended.
The efficiency of silver-containing antimicrobial textiles
Kuthong et al. used T27-22-Silver liquid in five different concentrations to treat cotton fabric, and examined their antimicrobial efficiency against ATCC 6538 S. aureus and ATCC 25922 E. coli strains [53]. Their results are in line with ours, and the T27-22-Silver liquid proved to be more effective against Gram-positive than Gram-negative pathogens.
Lorenz et al. compared eight commercially available, silver-containing antibacterial textiles [54]. Three of them did not show any antibacterial effect against the ATCC 4352 K. pneumoniae strain. Other three products caused a five orders of magnitude decrease in bacterial CFU. In the study of Messaud et al., the fabrics treated with Ag-TiO2 nanoparticles proved to be efficient against a Gram-positive and a Gram-negative strain [27]. They found that the bacteria were able to proliferate on textile areas poorly impregnated with silver. In the study of Gerba et al. the silver-impregnated textile showed bacterial reduction between ≥2.7 log10 and 8 log10 in case of five ATCC strains after 24 h, although the initial inoculum of the test strains contained only ∼104–105 CFUs in 0.1 ml solution [58]. According to the in vitro study of Hanh et al., the silver nanoparticles showed >99.9% reduction of bacterial CFU against ATCC 6538 S. aureus and ATCC 4352 K. pneumoniae compared to untreated textile [59]. Their silver nanoparticles proved to be effective against several clinically isolated bacteria as well, but the size and the composition of the investigated strain collection were not described precisely. Silver proved to be effective in all of the aforementioned studies, but none or only a few multidrug-resistant Gram-negative isolates were investigated. Our results indicate that the selection of the test strains is one of the critical steps of efficiency tests. Although the T27-22-Silver-treated textile had significant antibacterial efficiency against all of the Gram-positive isolates and showed >90% reduction of bacterial CFU in case of some Gram-negative isolates (ATCC 11150 E. coli, 3 MRKP and 5 MACI) as well, but overall according to the averaged Antibacterial Activity Values the T27-22-Silver liquid had no significant antibacterial efficiency against the Gram-negative isolates. These results indicate that silver agents should be tested either with numerous or very carefully selected isolates.
According to the literature and to our results, silver agents proved to be effective against Gram-positive bacteria, but more investigations are necessary to improve the effectiveness of antimicrobial coatings against the Gram-negative pathogens. One possible solution could be the combination of antibacterial agents with different mechanisms of action. For example, according to Kang et al. cotton textile treated with both a quaternary ammonium compound and silver nanoparticles showed synergistic antibacterial properties against the Gram-negative ATCC 15692 P. aeruginosa [60]. In another study, the combined use of silver nanoparticles and antibiotics showed synergistic antibacterial activity [61].
In our experiments, the survival capability of ATCC S. aureus and MRSA strains were similar on both kinds of antimicrobial textiles. However, we can recommend the use of other Gram-positive species in efficiency tests, because VRE isolates are more viable on inanimate surfaces. The ATCC 11105 and ATCC 25922 E. coli standards we used are less appropriate for efficiency tests because of their low desiccation tolerance and their higher sensitivity against antibacterial agents.
The comparison of survival capability within species
According to literature the survivability of bacterial species varies within broad limits, from a few hours even to 46 months [13,15]. We quantitatively examined the survival capability of healthcare-associated bacteria during health-relevant incubation times. The results showed a standard deviation of one to four orders of magnitude among strains that belong to the same species. To examine this phenomenon, we grouped the bacterial strains based on their genetic characteristics and resistance patterns to compare their survival capability on untreated and on two antimicrobial textiles.
The difference in the survival capability of sporadic (only independent, isolated occurrence) and epidemic (outbreak-causing, multiple occurrence) strains on untreated surfaces is still controversial [62–64]. In a healthcare environment, those bacterial strains become the most successful which have several resistance mechanisms against antibiotics and biocides, are highly virulent, are able to multiply quickly and to survive on inanimate surfaces. Such bacteria are able to spread successfully all over the world and become prevalent, pandemic clones. Thus, it is not surprising that pandemic strains proved to be more viable on inanimate surfaces than sporadic ones. Nevertheless the accumulation of resistance genes can lead to the reduction of bacterial fitness, especially among Gram-positive bacteria. In an antibiotic- or biocide-free environment, the sensitive or moderately resistant strains can prove more viable than the multidrug-resistant ones [65]. In our study, the CA-MRSA (community-associated MRSA) strains showed significantly higher survival capability on untreated textile than the HA-MRSA (hospital-acquired MRSA) isolates. Unlike to HA-MRSA isolates, the CA-MRSA strains are usually resistant against less antibiotics, nonetheless 5–10% of CA-MRSA infections are invasive and potentially life threatening [6].
Two other controversial topics are the differences in the biocide-sensitivity between the sporadic and epidemic strains [66,67] and the cross-resistance between the antibiotics and biocides [26,66,68–71]. Probably these findings have not yet been answered in a satisfying manner, because there are lots of biocides and antibiotics with different mechanisms of action and against them bacteria can have lots of different resistance mechanisms. So it is essential to take into consideration as many background data of biocides and bacterial strains as possible. In our study, we compared two groups of VRE strains with different resistance genes against glycopeptides (vanA and vanB) [12]. VanA gene cluster causes combined vancomycin and teicoplanin resistance, vanB gene cluster usually causes only vancomycin resistance. In our study, the vanA VRE isolates proved to be significantly more viable on antibacterial surfaces than vanB VRE strains. According to literature a transferable, plasmid-located copper resistance gene (tcrB) was identified, which is genetically linked to vanA and other (ermB) antibiotic-resistance genes [72]. It is possible that a similar phenomenon could explain the higher QAC-tolerance of vanA VRE isolates.
Within the given species, the survival capability of prevalent, international bacterial clones can also diverge. Based on our results, the MACI strains which belong to the CC1 international clonal complex showed significantly higher viability on untreated textile than CC2 MACI isolates. At the same time, the antibacterial textile treated with T27-22-Silver agent showed significantly higher antibacterial efficiency against CC1 MACI isolates. Since 2010, the rate of CC2 MACI strains has increased worldwide. Between 2005 and 2010, most Hungarian MACI isolates belonged to the CC1 lineage, but between 2010 and 2015 an increased rate of the CC2 clone was determined [3]. This might be explained by the higher biocide-tolerance of CC2 MACI isolates and the use of silver in healthcare products [73], or the intensive use of QAC disinfectants [74].
As our results indicate, a plethora of factors and parameters can influence the survival capability of bacteria even within one species. The number of the strains we examined was not enough to reveal the contexts, and this topic requires further, comprehensive investigations.
One thing is for sure, bacteria have incredibly great evolutionary potential, accordingly many biocide-resistant strains were described [75–79], and the resistance genes were often found on conjugative plasmids [71,72,80]. Thus, one successful clone could be enough to the world-wide emergence of biocide-resistant bacteria [81,82]. The prudent use of antibiotics and biocides, and the further investigation of the resistance mechanisms are indispensable to avoid the emergence of an antibiotic-like, extensive biocide-resistance.
Conclusion
We examined multidrug-resistant bacterial pathogens belonging to four species (MRKP, MACI, MRSA, VRE) which cause healthcare-associated infections worldwide. In the first step, we determined the survival capability of 15 well-characterized, Hungarian strains of each species on untreated cotton. We performed this investigation with two clinically relevant incubation periods (one hour, one day). According to our results, all examined bacterial strains were able to survive on textiles in an extent to pose health risk in the hospitals.
We selected two antibacterial liquids of Sanitized AG (T27-22-Silver and T99-19-QAC) which contain antibacterial agents widely used in textile industry. We performed quantitative method to examine their antibacterial efficiency against the 60 multidrug-resistant bacterial strains. Our research showed that during a longer period (one day) the T27-22-Silver liquid was able to eliminate most of the Gram-positive pathogens (MRSA, VRE), reducing the risk of cross-contamination. At the same time, none of the examined agents were able to destroy the multidrug-resistant, Gram-negative isolates (MRKP, MACI) on the textiles. Their high desiccation-tolerance and low biocide-sensitivity partly could explain the increasing occurrence of MACI isolates in the HAIs.
We found significant differences in the survival capability of isolates within species according to their clonal-types or resistance-types (MACI: CC1 vs. CC2 international clone; HA-MRSA vs. CA-MRSA; VRE with vanA vs. vanB resistance-gene).
The comparison of multidrug-resistant isolates and ATCC standard strains revealed that the antibiotic-sensitive S. aureus and the multidrug-resistant MRSA strains had similar survival capability and biocide-tolerance. Thus, the ATCC 6538 and ATCC 25923 S. aureus standards could be appropriate to investigate the survivability of Gram-positive pathogens on untreated and on antimicrobial textiles; however, we consider the examination of other, well-surviving Gram-positive bacterial groups (e.g. VRE strains) to be important as well. The ATCC 11105 and ATCC 25922 E. coli strains proved to be more sensitive against the effects of antibacterial agents and desiccation than the multidrug-resistant MRKP and MACI isolates. According to our results, the examined ATCC E. coli strains are less suitable for survival and antibacterial efficiency tests because their use can underestimate the risk of cross-contamination caused by multidrug-resistant Gram-negative pathogens.
Our results draw attention to the necessity of the use of healthcare-associated, multidrug-resistant bacterial strains and to the careful evaluation of the results in antibacterial efficiency tests. The evaluation of our data with several methods indicated that a significant decrease in the bacterial CFU does not automatically mean significant antibacterial efficiency that would be enough to reduce the risk of cross-contamination and infection effectively.
Footnotes
Acknowledgements
The authors wish to express their sincere thanks to Árpád Tóth from Clariant Hungaria Trading Company for making available the Sanitized T27-22 and T99-19 liquids, and also to INNOVATEXT Textile Engineering and Testing Institute Co. (Hungary) for the cotton sheet, we used in our investigation. We would like to thank Zsuzsanna Frank (Department of Organic Chemistry and Technology, BUTE, Budapest, Hungary) for her assistance in the antibacterial finishing process and Judit Borsa (Doctoral School on Materials Sciences and Technologies, Óbuda University, Budapest, Hungary), Hosam E.A.F. Bayoumi Hamuda (Rejtő Sándor Faculty of Light Industry and Environmental Engineering, Óbuda University, Budapest, Hungary) for their permanent support and advices. The authors are grateful to Gyula Vörös for his assistance in the statistical evaluation.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported through the New National Excellence Program [code number: ÚNKP-16-3/IV.] of the Ministry of Human Capacities.
Supplementary Material
Supplementary material is available for this article online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.