Abstract
Introduction
Early identification of high-risk pregnancies is crucial, yet conventional approaches often miss complex clinical and contextual factors. This study measured prevalence and determinants in Southern Ethiopia and applied explainable machine learning as a clinical decision-support tool, linking prediction to actionable digital health insights.
Methods
We conducted a retrospective cohort study of 3,954 mother–infant pairs using routine records from pregnancy to the postpartum. Five supervised machine learning algorithms - Logistic Regression, Random Forest, Support Vector Machine, an Artificial Neural Network and XGBoost were developed and validated. Performance was assessed using AUC, specificity, sensitivity, calibration, and F1-score. SHAP-based analysis enhanced interpretability, revealing the contribution of each predictor at both individual and cohort levels, supporting practical integration into digital maternal health systems.
Results
Obstetric complications occurred 16.3% of mothers, with higher incidence in rural settings. XGBoost achieved the highest predictive performance (AUC 0.86). Key predictors identified young maternal age, unplanned pregnancy, low education, previous complications, inadequate antenatal care, anemia, hypertension and long travel distance to health facilities.
Conclusions
Obstetric complications remain common in Southern Ethiopia. By applying explainable machine learning, this study not only predicts high-risk pregnancies with high accuracy but also provides actionable insights for clinical decision-making and digital health implementation.
Keywords
Introduction
Maternal health remains one of the most critical global priorities, specifically in low- and middle-income countries (LMICs), where obstetric complications persist to drive preventable morbidity and mortality. 1 According to the World Health Organization (WHO), nearly 295,000 mothers died from pregnancy-related causes in 2017, and 94% of these mortalities happened in LMICs; sub-Saharan Africa (SSA) alone reported almost two-thirds. 2 Ethiopia has made outstanding progress in maternal health improvement over the past two decades. However, complications such as sepsis, hemorrhage, obstructed labor, hypertensive disorders, and adverse perinatal outcomes continue as primary threats to neonatal survival and maternal well-being.3,4 These challenges are specifically critical in rural regions, where timely access to emergency newborn and obstetric care remains limited. 5 By leveraging digital health and machine learning tools, there is potential to enhance early risk identification and support timely intervention in these high-risk populations.
Earlier studies in Ethiopia and other LMICs have identified coherent determinants of obstetric complications, including inadequate antenatal care (ANC) attendance, maternal age extremes, unintended pregnancy, anemia, and rural residence.6–8 Nevertheless, most of this evidence is derived from case–control or cross-sectional designs, which limit the evaluation of temporal links and progressing risk patterns across pregnancy and childbirth.9,10 Moreover, conventional regression approaches may fail to capture complex, non-linear interactions among multiple predictors, underscoring the need for advanced computational methods in maternal health research. 11
Nowadays, attention has turned to leveraging digital innovations and data-driven approaches to enhance maternal care across the pregnancy, childbirth, and the postpartum period can be improved through digital innovations and data-driven tools. 12 Developments in machine learning (ML) and artificial intelligence (AI) give promising opportunities in this regard. ML methods such as extreme gradient boosting (XGBoost), random forests, artificial neural networks, and support vector machines can manage high-dimensional data and capture complex interactions that are difficult to model using traditional methods.13–17 Interpretability frameworks like SHapley Additive exPlanations (SHAP) further enhance transparency by quantifying the contribution of each determinant to both population and individual-level risk estimates. 18 The use of interpretable frameworks like SHAP translates these predictions into actionable insights for both population-level and individual-level risk management. 18 Additionally, adoption frameworks, including the Technology Acceptance Model and the Unified Theory of Acceptance and Use of Technology, emphasize that successful integration of digital tools depends on perceived utility, usability, and local contextual factors.19–21 By connecting predictive modeling to practical clinical decision-making, these approaches advance the field of health informatics beyond theoretical applications.
The growing utilization of AI in healthcare aligns with these theoretical and methodological developments. Current studies comprising systematic reviews of ML applications in maternal and perinatal health show that ML-based decision-support tools can improve early risk detection and enhance the allocation of limited medical resources.22,23 In Ethiopia, ML models applied to national survey data have been utilized to predict low birth weight, with algorithms like XGBoost and random forests demonstrating strong predictive performance. 24 Another study has employed ensemble ML techniques to predict perinatal mortality based on health-service and maternal health variables. 25 Worldwide, ML methods are progressively outperforming traditional statistical approaches in forecasting obstetric complications, comprising neonatal intensive care unit admission and postpartum hemorrhage, indicating their ability to capture complex, non-linear associations inherent in maternal health data.23,24 Nevertheless, there remains a gap: few studies in Ethiopia have analyzed a comprehensive set of maternal complications within a single regional cohort, and none have combined interpretable ML frameworks with actionable clinical insights, underscoring the originality of the current study.
To address this knowledge gap, we conducted a retrospective cohort study to assess temporal associations between maternal exposures and obstetric outcomes in the Sidama region of southern Ethiopia. By applying multiple ML algorithms alongside SHAP-based interpretability, we not only aimed to identify the strongest predictors of risk but also to provide implementation-ready insights for healthcare providers. The overarching goal was to develop validated, explainable ML models that can inform targeted interventions, support clinical decision-making, and guide policy in resource-limited environments, thereby contributing novel knowledge to the field of health informatics.
Novelty and contribution to health informatics
This study contributes to health informatics by applying explainable machine learning as a practical decision-support tool across the maternal care continuum in a low-resource setting. By integrating antenatal, delivery, and postpartum data, SHAP-based interpretability translates complex predictions into actionable insights, enabling providers to prioritize care while maintaining transparency and trust. Guided by this approach, the study addresses three research questions: (1) What is the prevalence of maternal obstetric complications in the Sidama region? (2) Which individual, clinical, and contextual factors most strongly predict complications? (3) How effectively can explainable machine learning models predict risk and support clinical decision-making in low-resource environments?
Methods
Stages of the study
This study used a structured approach to predict obstetric complications among 3,954 mother-infant pairs in the Sidama region of southern Ethiopia. Data were retrospectively collected from medical records and included demographic, obstetric, clinical, and contextual information. Data preprocessing comprised handling missing values through multiple imputation, standardizing continuous variables, encoding categorical variables, and addressing class imbalance using the Synthetic Minority Oversampling Technique (SMOTE). Five supervised ML models - Logistic Regression (LR), Random Forest (RF), Support Vector Machine (SVM), Artificial Neural Network (ANN), and Extreme Gradient Boosting (XGBoost) were developed and tuned using grid search.
The selection of machine learning models was guided by their complementary strengths in handling structured health data, capturing complex non-linear relationships, and ensuring interpretability. LR provides a baseline for comparison; RF and XGBoost capture interactions and non-linear effects robustly; SVM is effective for high-dimensional classification; and ANN allow modeling of complex patterns. Together, these models enable comprehensive evaluation of predictive performance while balancing accuracy, interpretability, and applicability in low-resource clinical settings.
All analyses were performed in R software (v 4.3) using the caret, xgboost, randomForest, e1071, and SHAP for xgboost packages, Keras, and TensorFlow applied for neural network modeling. The study followed five main stages: data collection, data preprocessing and cleaning, feature preparation, model development, and model evaluation. The overall workflow is illustrated in Figure 1. Overview of the study process for predicting obstetric complications. Data were collected and cleaned, including handling missing values and balancing classes. Five machine learning models were developed and evaluated with 10-fold cross-validation, and SHAP values were used to identify the most important predictors.
In addition to developing predictive models, this study emphasizes the health informatics contribution by employing explainable machine learning as a clinical decision-support tool across the maternal care continuum. By integrating longitudinal data from antenatal, delivery, and postpartum periods, we translate risk prediction into actionable guidance for healthcare providers. The use of SHAP-based interpretability allows clinicians to understand why individual women are considered high-risk and prioritize interventions accordingly. Importantly, the modeling framework is designed for low-resource digital environments, facilitating future integration with electronic health records or mobile health platforms, and demonstrating a practical pathway for implementing machine learning in maternal healthcare.
Study design and setting
This retrospective cohort study was conducted in the Northern Zone of the Sidama region, Ethiopia, covering the period from January 2021 to January 2025. The study setting included four predominantly rural districts - Boricha, Hawela, Shebedino, and Bilate Zuriya, which collectively face considerable challenges in maternal and neonatal healthcare provision due to shortages of skilled personnel, inadequate infrastructure, and geographic barriers to referral facilities. Data were extracted from government hospitals and affiliated health centers, thereby ensuring broad representation of deliveries occurring within the public health system.
Reporting guidelines
This retrospective cohort study is reported in accordance with the STROBE (Strengthening the Reporting of Observational Studies in Epidemiology) guidelines for observational research and the TRIPOD-ML (Transparent Reporting of a multivariable prediction model for Individual Prognosis or Diagnosis - Machine Learning) guidelines for prediction model studies. Key items from each checklist were followed to ensure transparent reporting of participant selection, predictor definition, outcome measures, model development, validation, and clinical utility. The STROBE and TRIPOD-ML checklists are provided as Supporting Information 1 and 2, respectively.
Participants
The study population included all women who delivered live-born infants in the participating facilities during the study period - eligibility required complete maternal, obstetric, and neonatal records. Exclusion criteria were: multiple gestations beyond twins, congenital anomalies incompatible with life, and missing key clinical or demographic variables. After applying these criteria, a final analytic cohort of 3,954 mothers was retained. This sample size was considered sufficient for predictive modeling based on established guidelines indicating that several thousand observations are generally adequate to train and validate machine learning models of moderate complexity. To confirm this, we conducted preliminary analyses, including learning curve assessments, which showed that model performance stabilized at this sample size, demonstrating that the cohort provides reliable data for identifying associations between risk factors and obstetric complications.
Sampling procedure
A multistage sampling strategy was employed to ensure a representative sample of mothers across the Northern Zone of the Sidama region. In the first stage, one zone was randomly selected from the four administrative zones, resulting in the Northern Zone being included. In the second stage, four districts within this zone were chosen using simple random sampling. In the third stage, representative health facilities were selected from each of the four districts. Finally, study participants were systematically sampled from the registers and records of the selected facilities, using pre-determined intervals to capture deliveries proportionally over the study period. This approach minimized selection bias and ensured that participants reflected the geographic and facility-level diversity of the study area, thereby enhancing the generalizability of the findings.
Outcome variable
The primary outcome of interest was the occurrence of maternal obstetric complications during pregnancy, childbirth, and the postpartum period (up to 42 days after delivery). Obstetric complications were defined as the presence of one or more of the following: obstetric hemorrhage (antepartum or postpartum), hypertensive disorders of pregnancy (preeclampsia or eclampsia), obstructed labor, uterine rupture, sepsis or infection, maternal anemia requiring intervention, retained placenta, and other clinically significant complications documented in the maternal chart. Each complication was coded as a binary variable (1 = present, 0 = absent). Diagnoses were confirmed using standardized clinical and laboratory criteria documented in the medical records. Where applicable, ICD-10 codes were applied to classify diagnoses. Clinical definitions followed the World Health Organization (WHO) guidelines for maternal health26,27 and the Ethiopian Ministry of Health national obstetric guidelines. 28
Independent variables
Independent predictors encompassed maternal, obstetric, and contextual factors. Maternal factors included age, parity, educational status, antenatal care (ANC) attendance, pregnancy intention, history of prior obstetric complications, anemia, and hypertension. Obstetric factors comprised gestational age at delivery, mode of delivery, and any complications during labor. Contextual factors included place of residence (urban/rural) and distance to the nearest health facility. All variables were defined and coded prior to analysis. Maternal age was categorized as <20, 20–34, and ≥35 years. Residence was classified as urban or rural. Educational level included no formal schooling, primary, secondary, or higher, and was grouped as primary or less versus secondary or more for analysis. Parity was classified as 0–2 or ≥3 births. ANC attendance was coded as <4 versus ≥4 visits. Prior obstetric complications, maternal anemia (hemoglobin <11 g/dL), and maternal hypertension (systolic ≥140 mmHg or diastolic ≥90 mmHg) were coded as yes/no variables. Distance to the nearest health facility was grouped as ≤5 km or >5 km. Household wealth index included five national quintiles and was grouped as the lowest two versus the upper three quintiles. Marital status was categorized as married/cohabiting or not married. Maternal BMI was classified as underweight (<18.5 kg/m2), normal (18.5–24.9), overweight (25.0–29.9), or obese (≥30). Household food insecurity was classified as food-secure or food-insecure. Birth spacing was coded as <24 or ≥24 months, and iron–folate supplementation as <90 or ≥90 days. Facility type included health post, health center, or hospital, and skilled birth attendance was coded as yes, or no. Time of delivery was categorized as daytime (06:00–17:59) or nighttime (18:00–05:59).
Predictor variables with more than 20% missing observations were excluded from modeling to ensure data quality and validity. Continuous variables were standardized, and categorical variables were one-hot encoded before inclusion in machine learning models.
Data sources, quality assurance, and preprocessing
Data were abstracted from both electronic medical records and paper-based charts using a standardized data extraction tool developed for this study. To enhance accuracy, two independent data collectors cross-verified entries, with discrepancies resolved by referring to the original records. Missing data were addressed through multiple imputation with chained equations. Continuous variables were normalized, and categorical variables were encoded using one-hot encoding. To account for class imbalance in the outcome variable, oversampling of minority classes was conducted using SMOTE. The dataset was randomly partitioned into training (70%) and validation (30%) subsets.
Machine learning modeling
Five supervised machine learning algorithms were implemented in R (v4.3): logistic regression, random forest, XGBoost, support vector machine, and artificial neural network.13,15,16 Hyperparameters for each model were optimized through grid search with 10-fold cross-validation within the training set. 29 Predictive performance was evaluated using the area under the receiver operating characteristic curve (AUC), sensitivity, specificity, precision, recall, F1-score, and calibration slope. 30 The models were built using established R packages including caret, xgboost, random Forest, e1071, and nnet.31,32 Data preprocessing, including normalization, multiple imputation, and SMOTE oversampling, further enhanced model robustness and generalizability.33,34 To address the imbalance in our dataset, we employed the SMOTE. This method creates synthetic examples of the minority class (maternal obstetric complications) by interpolating between existing positive cases, rather than simply duplicating them. By doing so, SMOTE enables the models to better learn patterns associated with complications, reducing the risk of overfitting and improving predictive performance across both classes. 34
Model interpretability
To enhance clinical interpretability, SHAP values were computed using the iml and shapper packages in R.18,35 SHAP analysis quantified the contribution of each predictor to model predictions and identified interaction effects between maternal and contextual factors. 18 This approach ensured that the models generated not only accurate predictions but also clinically actionable insights. 17
Hyperparameter optimization
For each machine learning model, including the ensemble XGBoost model, hyperparameters were carefully tuned using a grid search in combination with 10-fold cross-validation. Key parameters such as nrounds, max_depth, learning rate (eta), subsample, and colsample_bytree were systematically varied to identify the configuration that provided the best predictive performance while minimizing overfitting. The same approach was applied to the support vector machine, artificial neural network, and random forest models to ensure consistent, robust, and reproducible evaluation across all algorithms.
Data preprocessing and missing data handling
Variables with more than 20% missing values (household income and body mass index) were excluded from analysis. 36 For variables with ≤20% missingness, including facility type (5%), iron–folate supplementation (8%), marital status (7%), and nighttime delivery (3%), we applied multiple imputation by chained equations (MICE) within each 10-fold cross-validation to prevent data leakage.29,33 Imputation was performed using m = 5 datasets over 10 iterations, with all remaining predictors included in the imputation models and a fixed random seed for reproducibility. Diagnostic checks confirmed convergence and plausible distributions of imputed values.
Model transparency and reproducibility
To enhance transparency and reproducibility, all models were developed in R version 4.3 using a fully documented preprocessing pipeline. 31 Continuous variables were standardized within each resampling fold, and categorical variables were encoded using one-hot encoding to avoid leakage. 31 For each algorithm, we reported all key hyperparameters and the ranges explored during tuning. The SVM used an RBF kernel with a grid search over C = 0.1–10 and γ = 0.001–0.1. 16 The ANN consisted of three hidden layers (32, 16, and 8 units) with ReLU activation, the Adam optimizer, a learning rate of 0.001, and early stopping after 10 epochs without improvement. 15 The XGBoost model explored nrounds = 200–600, max_depth = 2–6, eta = 0.01–0.3, subsample = 0.6–1.0, and colsample bytree = 0.6–1.0. 13 We also generated SHAP dependence plots, including interaction profiles such as anemia × ANC less than four, to complement the global ranking. 18 To support reproducibility, we have provided the complete analysis code, a de-identified metadata dictionary, and a Model Card describing intended use, limitations, and subgroup behaviour as supplementary material (Supplemental Files 3).
Statistical analysis
Descriptive analyses summarized baseline maternal and contextual characteristics. Categorical variables were reported as frequencies and percentages, while continuous variables were expressed as means with standard deviations (SD) after checking for normal distribution. 9 Group differences were assessed using chi-square tests for categorical data and independent t-tests for continuous variables. 9 All analyses were conducted in R, 31 with statistical significance defined as a two-sided p-value <0.05. To strengthen model validation, we applied nested cross-validation 29 and bootstrapping techniques. 29 These approaches provided robust estimates of predictive performance and reduced the risk of overfitting, 11 supporting the reliability of our findings. To assess the relative performance of different approaches, we benchmarked XGBoost 13 against multiple machine learning algorithms, including Logistic Regression, Random Forest, 32 Support Vector Machine, 16 and Artificial Neural Networks. 15 All candidate predictors were included in the models. XGBoost and Random Forest inherently manage irrelevant features via tree-based selection,13,32 and SHAP analysis was applied post-hoc to quantify feature contributions. 18
Results
Study population and baseline characteristics
Baseline obstetric, maternal, and other contextual characteristics of study subjects by maternal obstetric complication status (n = 3,954).
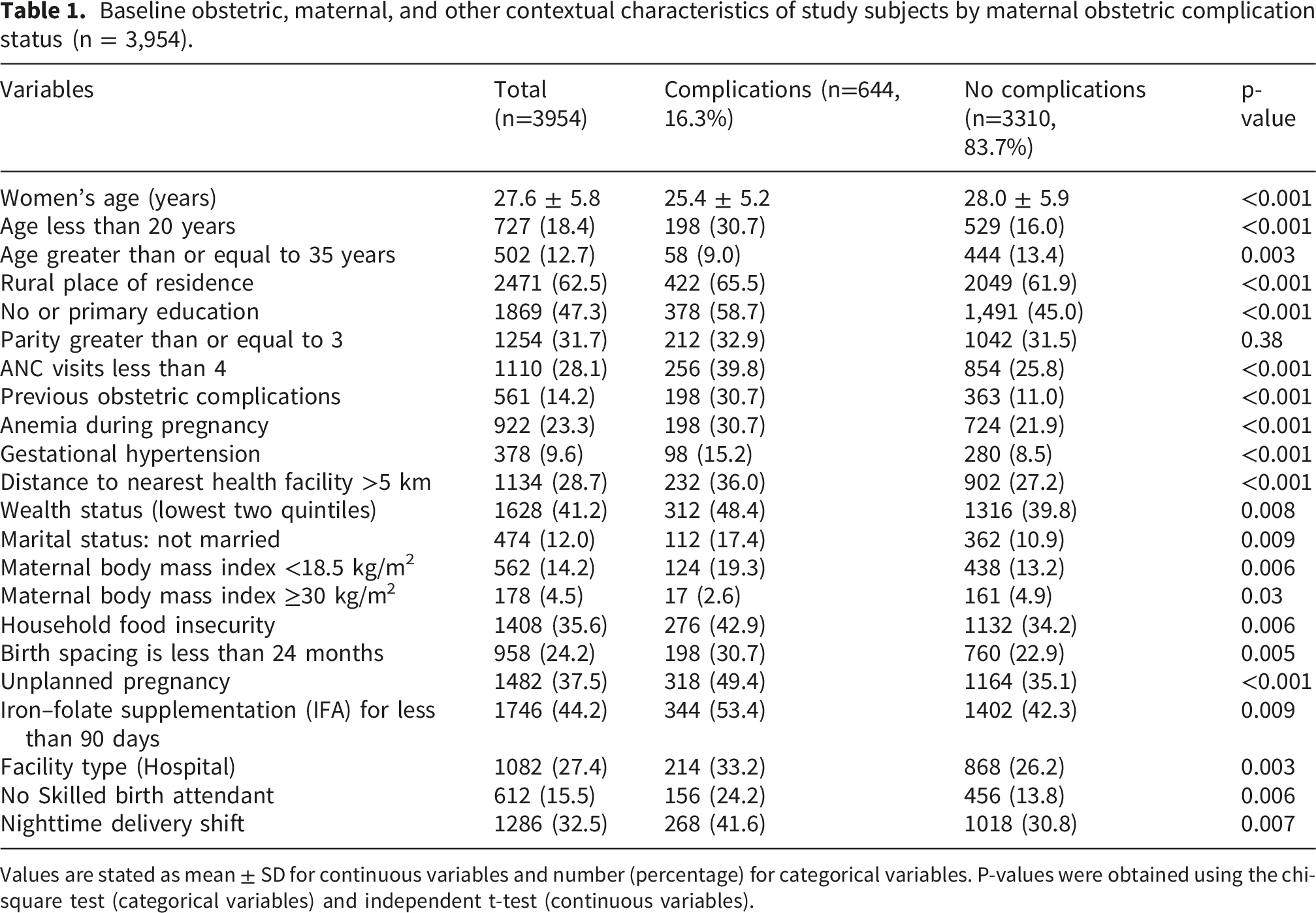
Values are stated as mean ± SD for continuous variables and number (percentage) for categorical variables. P-values were obtained using the chi-square test (categorical variables) and independent t-test (continuous variables).
Missing data and cohort characteristics
After preprocessing, 3,954 mother–infant dyads were retained for analysis. A supplemental File 4 summarizes the proportion of missing values per variable. The excluded variables had >20% missingness, while all other predictors were fully imputed, ensuring completeness for machine learning model development and evaluation.
Incidence of maternal obstetric complications
Overall, 16.3% (95% CI: 15.0–17.7) of women experienced at least one maternal obstetric complication during pregnancy, delivery, or the postpartum period. The most frequent complications were obstetric hemorrhage (6.8%), hypertensive disorders of pregnancy (5.4%), obstructed labor (3.9%), and postpartum sepsis (2.1%). The incidence of complications was higher in rural districts (18.7%) compared with urban catchment areas (12.4%, p < 0.01), highlighting geographic disparities in obstetric outcomes (Figure 2). (a) Overall incidence of maternal obstetric complications among women in the study cohort (n = 3,954). (b) Percentage distribution of major obstetric complications among women (n = 3,954). (c) Proportion of maternal obstetric complications by place of residence. (d) Proportion of maternal obstetric complications across districts.
Machine learning model performance
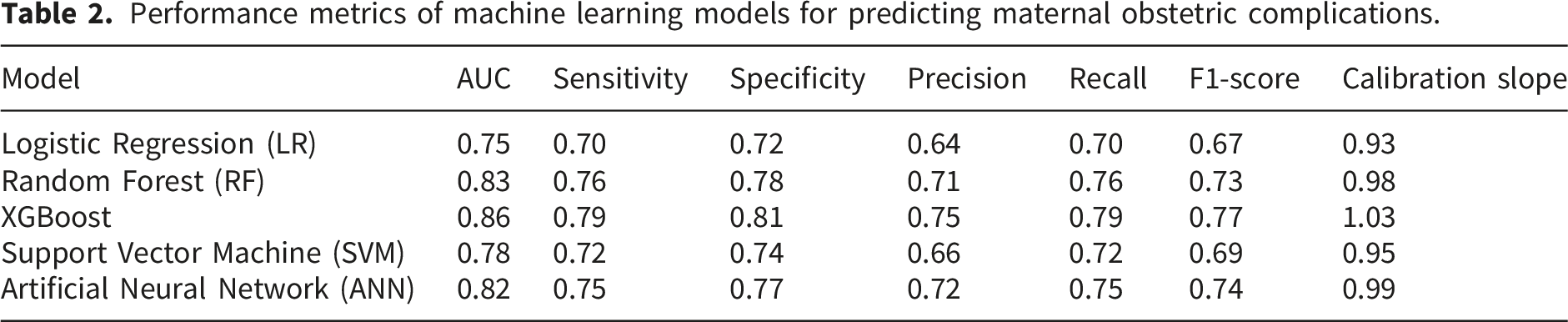
Performance metrics of machine learning models for predicting maternal obstetric complications.

Presents the receiver operating characteristic (ROC) curves for the five machine learning models, generated using 10-fold cross-validation. In these plots, the horizontal axis represents the False Positive Rate (1 – specificity), and the vertical axis represents the True Positive Rate (sensitivity), following standard ROC conventions.
Among these, XGBoost demonstrated the highest predictive performance, achieving an area under the receiver operating characteristic curve (AUC) of 0.86, with sensitivity 79%, specificity 81%, and F1-score 0.77. Random forest and artificial neural network (ANN) models also performed robustly, with AUC values of 0.83 and 0.82, respectively. Logistic regression and support vector machine (SVM) models exhibited moderate performance (AUC = 0.75 and 0.78, respectively). Calibration analysis indicated good agreement between predicted probabilities and observed outcomes, with XGBoost showing the closest fit (calibration slope = 1.03). Model performance metrics, including AUC, sensitivity, specificity, and F1-score, were averaged across 10-fold cross-validation. Fold-level accuracy and F1-score for all models are provided in Supplemental File 5, demonstrating stable predictive performance across folds.
Key predictors of maternal obstetric complications
SHAP analysis identified the most important predictors of maternal obstetric complications, with clear quantitative and interaction-based support. Overall, 16.3% of women experienced at least one complication. Complications were more frequent among mothers <20 years (30.7% vs 16.0%), anemic mothers (30.7% vs 21.9%), those with fewer than four ANC visits (39.8% vs 25.8%), and rural residents (18.7% vs 12.4%). Feature importance analysis confirmed that maternal age <20 years, prior obstetric complications, low educational attainment, unplanned pregnancy, fewer than four ANC visits, maternal anemia, hypertension, rural residence, and greater distance to the nearest health facility were the strongest predictors (Figure 4). Interaction analyses revealed compounded risk: for example, anemic women with inadequate ANC attendance and a history of prior complications had substantially higher predicted probabilities than women with a single risk factor. Similarly, rural residence combined with a long travel distance (>5 km) further increased predicted risk. These results provide quantitative and interpretable evidence that multiple individual, clinical, and contextual factors jointly contribute to maternal obstetric complications. Representative SHAP dependence and interaction plots are presented in Figure 5. SHAP (SHapley Additive exPlanations) summary plot showing the top nine predictors of maternal obstetric complications. Features are ranked by their overall contribution to model predictions, with color indicating feature value (high to low). Subgroup and sensitivity analyses of maternal obstetric complications. The plots display predicted complication rates across key subgroups, highlighting variation by maternal age, anemia status, ANC attendance, and rural residence. These analyses provide quantitative insights into how risk differs across clinically and contextually relevant subpopulations.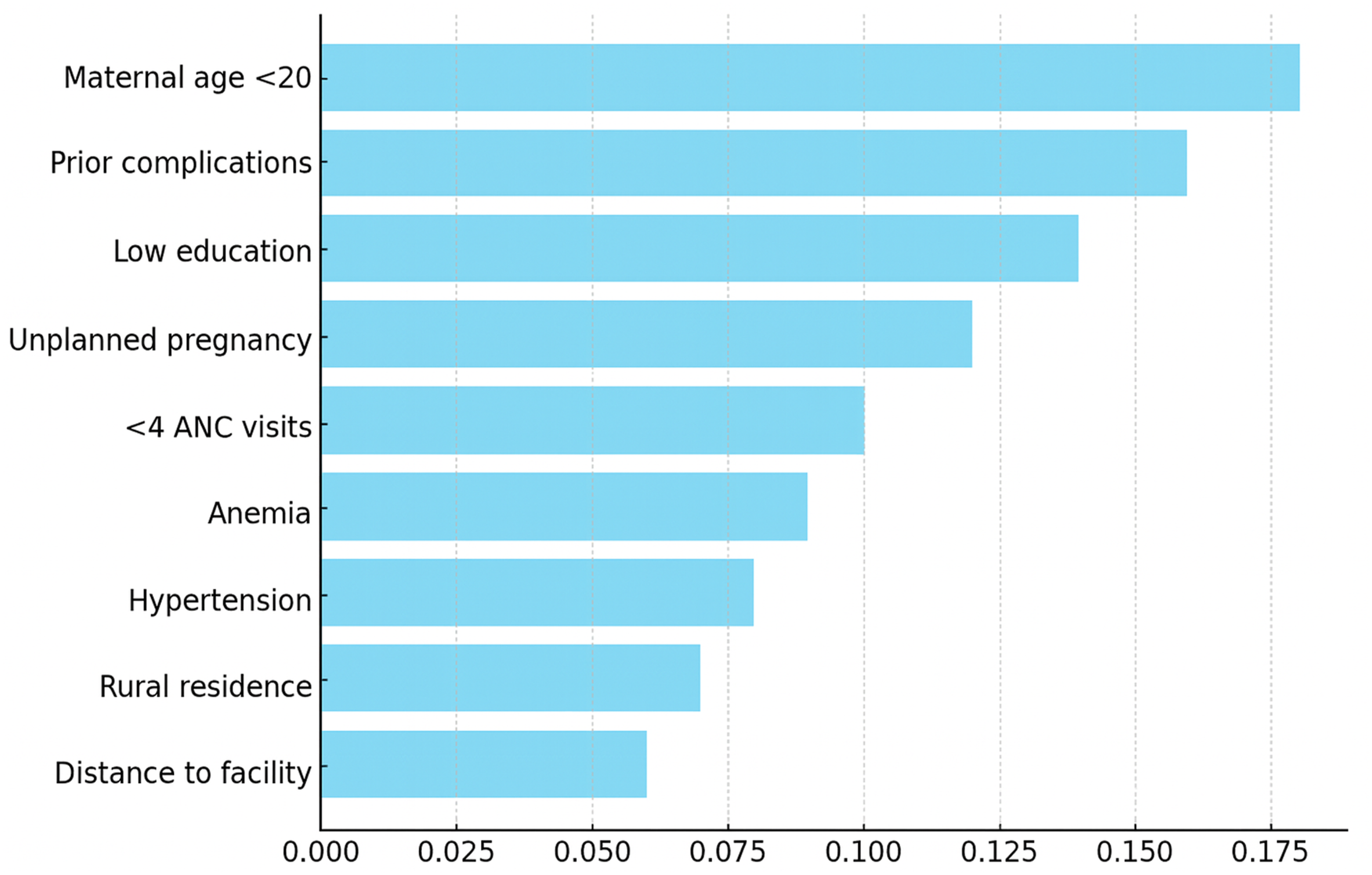

The SHAP analyses provide actionable insights for maternal care, showing how individual predictors drive risk throughout antenatal, delivery, and postpartum periods. This interpretability enables clinicians to prioritize interventions and allocate resources effectively. Designed for low-resource settings, the approach can integrate into electronic health records or mobile health platforms, translating machine learning from theory into practical health informatics tools.
Comparison of machine-learning studies on maternal and obstetric outcomes in Ethiopia, emphasizing outcome scope, sample size, feature composition, modeling techniques, and interpretability.
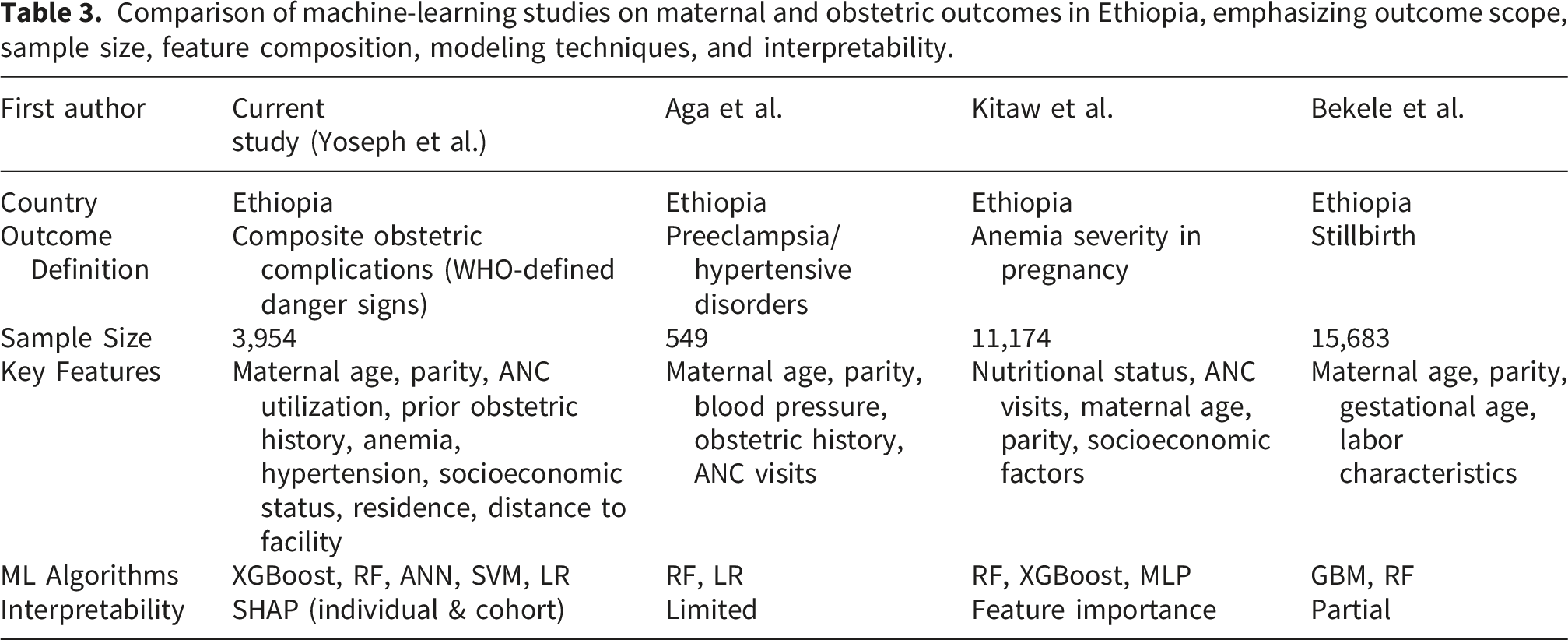
Individual- and cohort-level SHAP analysis
To enhance interpretability, we expanded SHAP analysis to both cohort-level and individual-level insights. At the cohort level, feature importance from XGBoost confirmed the strongest predictors described above. Individual-level SHAP values illustrated how these features contributed to risk predictions for specific mothers. Some patients exhibited high predicted probabilities primarily driven by anemia and limited ANC visits, whereas others had lower predicted risks despite elevated values in some variables. Comparing individual-level contributions with cohort-level rankings revealed alignment in overall trends but also highlighted patient-specific deviations, emphasizing the value of SHAP for personalized risk assessment. Representative SHAP plots are presented in Figure 6. SHAP analysis of key predictors for maternal obstetric complications.
Model transparency, tuning performance, and explainability
Across all models, hyperparameter tuning improved discrimination and calibration compared with default settings. For XGBoost, the optimal configuration included nrounds = 400, max_depth = 4, eta = 0.05, subsample = 0.8, and colsample_bytree = 0.8, yielding the highest AUC (0.86). The tuned SVM with an RBF kernel (C = 1, γ = 0.01) achieved an AUC of 0.78, while the ANN architecture with three hidden layers (32–16–8 units, ReLU activation, Adam optimizer, early stopping at epoch 28) produced an AUC of 0.82. Tuning results for all algorithms are provided in Supplemental File 6.
Dataset class distribution and model performance
The dataset was imbalanced, with 16.3% of women experiencing maternal obstetric complications and 83.7% without complications. To clearly illustrate model performance across both classes, we included a confusion matrix for the XGBoost model (Figure 7), showing the number of true positives (508), false negatives (136), false positives (631), and true negatives (2,679). This visual representation complements the metrics reported in Table 2 and confirms that predictive performance is robust despite the class imbalance. Confusion matrix for the XGBoost model predicting maternal obstetric complications. The matrix shows the number of true positives, false negatives, false positives, and true negatives, illustrating model performance across both classes in the imbalanced dataset (16.3% complications, 83.7% no complications).
The synthetic minority oversampling technique
To mitigate the class imbalance, SMOTE was applied during model training. This allowed the XGBoost and other models to better learn patterns associated with maternal obstetric complications, resulting in stable and robust predictive performance. The metrics reported in Table 2 and the confusion matrix (Figure 7) confirm that the models achieved high discrimination and calibration across both classes, indicating that oversampling effectively improved model learning without overfitting.
Subgroup and sensitivity analyses
SHAP analyses provided clear evidence of both main effects and meaningful interactions among key predictors. The dependence plots showed that anemia, fewer than four ANC visits, and a history of prior obstetric complications exerted strong and often non-linear effects on the likelihood of adverse outcomes. The joint effect of anemia and inadequate ANC attendance was especially pronounced, with the combined profile producing a much higher predicted risk than either factor alone. A similar pattern emerged for rural residence paired with long travel distance (>5 km) to a health facility, highlighting the compounded disadvantage faced by women in remote settings. Representative interaction plots illustrating these relationships are provided in Figure 5.
Discussion
Principal findings
This cohort study demonstrates that 16.3% of women experienced at least one maternal obstetric complication during pregnancy, delivery, or postpartum, with rural women disproportionately affected (18.7% vs. 12.4% in urban areas). The most incident complications were obstetric hemorrhage (6.8%), hypertensive disorders of pregnancy (5.4%), obstructed labor (3.9%), and postpartum sepsis (2.1%). Machine learning modeling, particularly using XGBoost, effectively predicted maternal complications with high discrimination (AUC = 0.86) and reliable calibration (slope = 1.03). SHAP analysis revealed multifactorial determinants, including maternal age <20 years, prior obstetric complications, low educational attainment, unplanned pregnancy, fewer than four ANC visits, maternal anemia, hypertension, rural residence, and greater distance to health facilities.
Comparison with previous findings
Global context
Maternal complications remain a significant cause of morbidity and mortality worldwide, with low- and middle-income countries disproportionately affected. 37 Hemorrhage, hypertensive disorders, and sepsis are consistently reported as leading contributors to adverse maternal outcomes. 38 In our study, 16.3% of women experienced at least one maternal obstetric complication, which aligns with global estimates, although slightly lower than the upper range reported in some LMICs. This consistency suggests that the underlying biological and social determinants of maternal complications are broadly similar across diverse settings.39,40
Ethiopian context
Previous studies in Ethiopia reported maternal obstetric complication prevalence ranging from 20% to 33.9% during delivery and the postpartum period.3,4 Our slightly lower prevalence may reflect improvements in antenatal care coverage, differences in urban-rural distribution, or variation in outcome definitions. Consistent with prior Ethiopian studies, we identified young maternal age, prior obstetric complications, anemia, low educational attainment, and rural residence as key risk factors.5–8 This agreement reinforces the validity of our findings and underscores the ongoing influence of structural determinants on maternal health disparities in Ethiopia.
Our study extends previous research by applying interpretable machine learning models to predict a broad set of maternal obstetric complications. For instance, Bogale et al. applied gradient boosting to predict perinatal mortality in Ethiopia, identifying anemia, maternal education, rural residence, and birth interval as top predictors, with reported accuracy around 90%. 25 Similarly, our XGBoost model achieved an AUC of 0.86, highlighting anemia and rural residence among the strongest predictors. Differences in predictive performance likely reflect variations in outcome definition, sample size, and selected features. Other ML-based studies, such as Bekele’s prediction of low birth weight using random forest, 24 have focused on single outcomes, whereas our study captures a comprehensive set of complications, providing a more holistic risk assessment for LMIC settings.
Explaining similarities and discrepancies
The similarity between our findings and prior reports underlines the universality of key risk factors, including maternal age, parity, and ANC utilization.37–40 Conversely, the lower complication prevalence in urban areas compared to national reports, where urbanization is sometimes associated with lower maternal risk, may be explained by better healthcare infrastructure, shorter distances to health facilities, and higher health literacy in our study urban catchment areas.41–45 Additionally, differences in study design, data collection methods, and definitions of complications likely contributed to variability across studies.
Interpretation of machine learning findings
The application of XGBoost, a robust ensemble learning method, 13 revealed not only individual risk factors but also their interactions. For instance, anemia compounded with inadequate ANC attendance and prior obstetric complications substantially elevated predicted risk, highlighting the importance of multidimensional risk assessment.24,25 The strong performance of XGBoost relative to Random Forest, ANN, SVM, and logistic regression models emphasizes the potential utility of advanced predictive analytics in maternal health surveillance and risk stratification.14–16,22 Such models can identify high-risk women earlier, enabling targeted interventions and resource allocation. 17
A key contribution of this study is the translation of machine learning predictions into actionable health informatics tools. By using SHAP-based interpretability, clinicians can understand the factors driving risk and prioritize interventions throughout antenatal, delivery, and postpartum care. Importantly, the approach is designed for low-resource settings, allowing potential integration into electronic health records or mobile health platforms. This demonstrates a step beyond conventional predictive modeling, offering a practical framework for digital maternal healthcare that supports informed decision-making and equitable resource allocation.
The predictive models demonstrate strong performance; however, false positives may lead to unnecessary interventions, while false negatives could delay critical care. 30 By balancing sensitivity and specificity, our models aim to optimize early identification of high-risk pregnancies while minimizing potential harm.31,32 These considerations are vital for translating machine learning predictions into safe clinical practice.
The deployment of machine learning models in maternal healthcare carries important ethical, practical, and policy considerations. Algorithmic bias may arise if training data do not adequately represent all subpopulations, potentially leading to inequitable risk predictions. 18 Ensuring model transparency and interpretability is crucial for clinician trust and informed decision-making, particularly in high-stakes obstetric care. 35 Data privacy and security must be rigorously maintained, especially when integrating predictive tools with electronic health records.17,46,47
Practical implementation also requires alignment with existing healthcare infrastructure. Integration with clinical workflows, staff training, and resource allocation is necessary to maximize the utility of ML-based risk prediction.12,26,28 From a policy perspective, establishing guidelines for responsible AI deployment can help prevent misuse and ensure equitable access, particularly in low-resource and rural settings where maternal mortality is highest.1,2
Overall, addressing these ethical, practical, and policy considerations is essential for translating machine learning models into safe, effective, and equitable maternal health interventions.
Clinical utility and decision-making considerations
The predictive models developed in this study have the potential to support real-time decision-making in antenatal clinics and community health programs by identifying women at high risk of obstetric complications.13,14,22 Operational thresholds, such as acceptable false-positive and false-negative rates, must be carefully considered to balance sensitivity and specificity.30–32 High sensitivity ensures that most high-risk pregnancies are identified, reducing missed complications, while controlling false positives is important to avoid unnecessary interventions and strain on limited healthcare resources.31,32
By providing early risk stratification, these models could inform targeted monitoring, timely referrals, and allocation of preventive resources, enhancing both maternal and neonatal outcomes.12,17,26,28 Implementing such predictive tools alongside clinician judgment allows for proactive, data-informed interventions while maintaining safety and feasibility in low-resource and rural settings.1,2,26,48
Strengths and limitations of the study
The study possesses several notable strengths that enhance the validity and applicability of its findings. The longitudinal cohort design enabled accurate temporal assessment of maternal obstetric complications, aligning with STROBE reporting standards. Inclusion of a large and diverse sample, encompassing both rural and urban women, improves the external validity of the results and ensures that findings are broadly representative of the population. Furthermore, the application of advanced machine learning analytics, particularly XGBoost, allowed precise prediction of complications and identification of interacting risk factors, offering actionable insights for both clinical management and public health interventions. Subgroup and sensitivity analyses further confirmed the robustness of the results, demonstrating consistent patterns across high-risk populations and excluding women with incomplete antenatal care records. We minimized overfitting through careful model selection, cross-validation, and comprehensive preprocessing. These steps strengthen confidence in the model’s predictive performance, while future validation on independent datasets will help confirm broader applicability. Future research could explore hybrid ensemble methods, such as stacking or blending, which may further enhance predictive accuracy by combining the strengths of individual models. Comparative benchmarking provides insight into which algorithms perform best under specific data conditions and helps guide selection for clinical deployment.
Nevertheless, the study has certain limitations that warrant consideration. Incomplete data, particularly regarding ANC visits, prior obstetric history, and some contextual factors, may have introduced bias. The generalizability of findings is also potentially limited, as results may not fully apply to regions with substantially different healthcare systems or demographic profiles. Additionally, while predictive modeling effectively identifies associations and interactions between risk factors, causal inferences cannot be definitively established. Variations in clinical documentation and reporting of obstetric complications may also have influenced prevalence estimates, potentially affecting the accuracy of outcome measurement. Because our dataset is drawn from a single region in southern Ethiopia, the generalizability of our findings may be constrained, underscoring the need for future validation using multi-center or national datasets that encompass more diverse populations and health-system contexts.
Hyperparameter tuning was a critical step to balance predictive performance and generalizability. While grid search efficiently identified near-optimal values, exploring automated optimization methods such as Bayesian optimization could further improve model performance and reduce computational cost. This represents a potential area for future research, particularly for scaling predictive models in low-resource clinical settings.
Practical and policy implications
The practical implications of these findings are substantial. Clinically, integrating XGBoost-based risk prediction into routine antenatal care workflows can enable the early identification of high-risk women, allowing timely referral and management to prevent adverse outcomes. Community-level interventions, including educational programs promoting ANC attendance, recognition of warning signs, and facility-based deliveries, are crucial, particularly for adolescent and anemic women who are at heightened risk. Moreover, predictive models can support efficient allocation of limited healthcare resources by prioritizing women at the most significant risk, ensuring that interventions are both targeted and effective.
From a policy perspective, the study highlights the need for continued investment in healthcare infrastructure, particularly in rural areas. Strengthening health facilities and improving transport networks can mitigate geographic disparities in maternal outcomes. Incorporating machine learning-based risk assessments into national maternal health policies may facilitate early identification of complications and more proactive management strategies. Finally, community-based health education campaigns and the engagement of local health workers are essential to improve maternal health literacy, promote timely care-seeking behavior, and support the overall goal of reducing maternal morbidity and mortality.
Data diversity and future research directions
Future studies should aim to enhance data diversity by incorporating additional variables such as environmental exposures, healthcare utilization indices, and socio-economic markers to improve model comprehensiveness. Collaborations across multiple regions or national datasets would create more representative cohorts, allowing for external validation and strengthening generalizability. Expanding both feature sets and geographical coverage will enhance the predictive power and clinical relevance of machine learning models in maternal healthcare.
Conclusion
This cohort study underlines a substantial burden of maternal obstetric complications in Ethiopia, particularly among rural women. Machine learning modeling, especially XGBoost, reliably identified women at elevated risk and elucidated multifactorial determinants. Addressing these risk factors through targeted interventions, strengthened health systems, and predictive analytics has the potential to reduce maternal morbidity and mortality. Continued research should evaluate the integration of machine learning tools into routine maternal healthcare and monitor their impact on health outcomes. Moreover, this study demonstrates a clear contribution to health informatics by translating machine learning predictions into actionable, interpretable tools that can be integrated into electronic health records or mobile health platforms, particularly in low-resource settings. This practical focus advances digital maternal healthcare and supports equitable, data-informed decision-making.
Supplemental material
Supplemental material - Explainable machine learning algorithms for predicting maternal obstetric complications in Ethiopia: Evidence from a retrospective cohort in southern Ethiopia
Supplemental material for Explainable machine learning algorithms for predicting maternal obstetric complications in Ethiopia: Evidence from a retrospective cohort in southern Ethiopia by Amanuel Yoseph, Yohannes Seifu Berego, Mehretu Belayneh, Francisco Guillen-Grima in Health Informatics Journal
Supplemental material
Supplemental material - Explainable machine learning algorithms for predicting maternal obstetric complications in Ethiopia: Evidence from a retrospective cohort in southern Ethiopia
Supplemental material for Explainable machine learning algorithms for predicting maternal obstetric complications in Ethiopia: Evidence from a retrospective cohort in southern Ethiopia by Amanuel Yoseph, Yohannes Seifu Berego, Mehretu Belayneh, Francisco Guillen-Grima in Health Informatics Journal
Supplemental material
Supplemental material - Explainable machine learning algorithms for predicting maternal obstetric complications in Ethiopia: Evidence from a retrospective cohort in southern Ethiopia
Supplemental material for Explainable machine learning algorithms for predicting maternal obstetric complications in Ethiopia: Evidence from a retrospective cohort in southern Ethiopia by Amanuel Yoseph, Yohannes Seifu Berego, Mehretu Belayneh, Francisco Guillen-Grima in Health Informatics Journal
Supplemental material
Supplemental material - Explainable machine learning algorithms for predicting maternal obstetric complications in Ethiopia: Evidence from a retrospective cohort in southern Ethiopia
Supplemental material for Explainable machine learning algorithms for predicting maternal obstetric complications in Ethiopia: Evidence from a retrospective cohort in southern Ethiopia by Amanuel Yoseph, Yohannes Seifu Berego, Mehretu Belayneh, Francisco Guillen-Grima in Health Informatics Journal
Footnotes
Acknowledgements
We sincerely thank the Sidama Region President Office and Hawassa university for their generous financial support, which was instrumental in enabling the successful completion of this study. We are also grateful to the Sidama Regional Health Bureau and the Districts Health Office for their invaluable collaboration. Our deep appreciation goes to the data collectors, field assistants, and supervisors, whose dedication and effort were essential to this research. Finally, we acknowledge the School of Public Health at University Saskatchewan for their expert technical guidance throughout the study design and data analysis.
Ethical considerations
The Institutional Review Board of Hawassa University College of Medicine and Health Sciences gave its approval to the study protocol (Approval No. IRB/027/16). All study participants provided the required information and were informed of their right to withdraw from the study at any time without any penalty.
Consent to participate
Written informed consent was obtained from all participants prior to data collection. De-identified data were used for analysis, and patient confidentiality was rigorously upheld.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Sidama President Office and Hawassa university. The funders were not involved in the study design, data collection, management, analysis, or interpretation; in the preparation, review, or approval of the manuscript; or in the decision to submit the manuscript for publication.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
