Abstract
Keywords
Introduction
Acute kidney failure (AKF) is characterized by a rapid decline in renal function and is frequently encountered among patients in intensive care units (ICUs) and other hospital setting. 1 The severity of AKF can range from mild impairment to life-threatening organ dysfunction. If not appropriately managed, AKF may lead to serious complications and increased mortality. Generally, patients with severe acute kidney failure may require Renal Replacement Therapy (RRT) to facilitate the removal of waste products from the bloodstream. 2
For patients admitted to the ICU, treatment for kidney failure, in addition to the RRT, may comprise pharmacologic interventions aimed at managing blood pressure, controlling inflammation, and addressing the underlying dysfunction. Continuous monitoring of fluid balance, electrolyte concentrations, and other vital parameters may also be required in these patients. 3 Typically, the prognosis of acute kidney failure in the intensive care setting can vary depending on the severity of the condition, the underlying cause, and the patient’s overall health. With timely and appropriate intervention, a substantial proportion of ICU patients with AKF can regain renal function and return to baseline health status. However, a subset may sustain chronic kidney impairment necessitating long-term dialysis or eventual renal transplantation.
The standard diagnostic evaluation for AKF includes laboratory tests, such as serum creatinine, blood urea nitrogen (BUN) levels, and urine analysis. Serum creatinine might be the most commonly used indicator of kidney function, and its level is a primary criterion for classifying the severity of AKF. Measurement of BUN provides additional assessment of renal excretory capacity, as it reflects the amount of nitrogen that is excreted by the kidneys. Urinalysis may provide critical insights into the underlying etiology of AKF by identifying abnormalities such as the presence of red blood cells, white blood cells, or urinary casts. In addition to the laboratory testing, clinical evaluation is a critical component of AKF diagnosis. This includes a thorough patient history, physical examination, assessment of fluid and electrolyte balance. Patients with AKF may present with symptoms such as decreasing of urine output, swelling, and shortness of breath. Hypertension, the swelling caused by excessive fluid trapped in the body’s tissues (or edema), and electrolyte imbalances are also commonly observed. 4 Based on these diagnostic tests, AKF can be classified into different stages of severity depending on the degree of kidney damage and the decreasing function of kidney. 5 The diagnosis of AKF helps healthcare providers determining an appropriate course of treatment, including the need for dialysis and other supportive interventions. In this context, the clinical guidelines outlined by KDIGO 6 or RIFLE 7 should be adhered to.
In this paper, an issue of data duplication in Electronic Health Record (EHR), which may comprise of medical history, diagnoses, medications, laboratory test results, treatment plans, and other relevant clinical information, is addressed for a kidney disease, Acute Kidney Failure (AKF), identification. Generally, data duplication in EHR refers to the presence of repeated patient and related information. It occurs when the same data elements are recorded multiple times within the EHR. Such duplication can manifest in various forms, e.g., having identical patient demographics excessively entered in multiple places, duplicate medication orders or prescriptions, redundant lab test results entered separately, or even the different presentation but same meaning as “HTN” and “Hypertension” or “DM” and “Diabetes”. These duplications may compromise data integrity and accuracy, thereby adversely affecting patient care, clinical decision-making, and research outcomes. 8
We propose to tackle the data duplication issue by applying feature engineering techniques to select, extract, and transform the most relevant features from the raw EHR data thereby improving data quality. In addition to structured data, note-event data comprising medical notes documented by physicians, nurses, and other healthcare professionals are incorporated to enhance the proposed methodology. Thus, an NLP technique, from a pre-trained model - scispaCy which has the biomedical terminology recognition capability, is applied to enhance the efficiency of eliminating duplicate data.9,10 Both feature engineering and NLP techniques are investigated in the experiments to reduce data duplication in the context of AKF. The proposed work is evaluated its outcome of duplication elimination as well as the application of a machine learning algorithm, KNN, on the de-duplicated well-known dataset, MIMIC-III database.
Method
In this section, the proposed framework to address data duplication in EHRs is proposed. Generally, it incorporates seven sequential processes, outlined as shown in Figure 1. Proposed data management for de-duplication.
Initially, data joining is generally conducted to integrate data from related sources, ensuring comprehensive and interconnected information. This step could be omitted, if the data integration is in-place before the de-duplication process is applied. However, it is generally necessary due to the inherent structure of electronic medical records (EMRs) which are collected and stored in healthcare information systems.

In Step 2, basic de-duplication techniques are employed to identify and remove duplicate records for redundant reducing in the joined dataset based on metadata. 11 This is proceeded by automated removal of any dirty text elements and replacing with empty string then storing it back to the dataset. Subsequently, in Step 3, patient medical history (PMH) is extracted from the note-event sources by constructing algorithm as shown in Algorithm 1. The algorithm extracts patient’s morbidity from the note-event source using a function to recursively split the keywords until the patient’s morbidity is found. For example, suppose that the note-event text is “HPI: PMHx: PMH: Prostate CA w/spinal mets, Gastric volvulus, Constipation, Depression, Lacunar infarct PSH: Gastropexy, Hiatal Hernia Repair... Current medications: ...PMH: (copied from prior clinic note) HTN; DM2; CKD stage 3; dyslipidemia.”, the first extract text will be “HPI:” as underlined, as well as all the second part after PMHx:, then the algorithm will extract the second part text into: “PMH: Prostate CA w/spinal mets, Gastric volvulus, Constipation, Depression, Lacunar infarct PSH: Gastropexy, Hiatal Hernia Repair... Current medications: ... ” and “(copied from prior clinic note) HTN; DM2; CKD stage 3; dyslipidemia.” recursively. Note that medical history keyword can be varied, the algorithm can be adapted and applied to extract the information.
To further improve the de-duplication process, a pre-trained model - scispaCy 12 (en_core_sci_lg mode) is applied in Step 4. This process helps identifying and eliminating any remaining duplicated data. The identification is performed by the scientific or medical term tokenization such that the terms, e.g. mg/dL are not parsed separately. Also, such duplicate data, e.g. “HTN” and “Hypertension”, “DM” and “Diabetes”, “PVD” and “Peripheral Vascular Disease” or “CAD” and “Coronary Artery Disease”, can be eliminated by the Unified Medical Language System (UMLS) in which the terms are compared with string similarity with the context data. It is trained on a vast amount of biomedical and scientific text data contained approximately 785,000 terms, and 600,000 word vectors which were trained with GloVe algorithm. 13 Such algorithm captures semantic relationships between words through the analysis, maximizing the likelihood of observed word co-occurrences within the given text. Additionally, an AbbreviationDetector, a Spacy component that executes the abbreviation detection algorithm, is also applied to verify similarity between its abbreviated and full-term words. 9 Eventually, PMH data from the previous step is mapped back into the dataset to further seamless data integrating in Step 5.
Subsequently, to facilitate the effective application of the designated analytical tasks, two additional steps are performed. First, irrelevant data and outliers potentially present in the dataset are identified and removed to minimize bias and improve the validity of the results in Step 6.
Secondly, in Step 7, data imputation and normalization procedures are performed. Note here regarding to the data imputation, missing data in EHRs can be classified into several types, namely missing completely at random (MCAR), missing at random (MAR), and missing not at random (MNAR). 14 This study focuses primarily on MCAR, as it frequently occurs in medical datasets. MCAR arises when certain healthcare measurements cannot be completed uniformly in practice. For instance, patients admitted to the ICU who remain bedridden may be unable to have their height accurately measured, resulting in missing values for height in the clinical notes. Such instances represent MCAR, where data absence is unrelated to observed or unobserved patient characteristics.
Thus, in Step 7, KNN imputer 15 is applied to impute missing data within the dataset. The KNN imputer is particularly effective in this context as it considers the characteristics of related observations, clusters, or nearest neighbor data, the approach is well-suited to medical data where features often exhibit strong interdependencies. This can result in accurate and reliable imputation of missing values, especially for data characterized as MCAR. The optimal value of k for the KNN imputer is determined via GridSearchCV, 16 with uniform weighting and the Euclidean distance metric applied. Furthermore, data normalization is conducted to standardize the scale and range of features, thus facilitating unbiased comparison and more robust analysis across variables.
To evaluate our proposed work, two experiments are to be conducted. Firstly, the confusion matrix from the manual-labeled duplication identified dataset compared to the duplication identified by our proposed work is presented. The criteria for the manually duplication identification are that two records are entirely identical, and the timestamp is in the same day. This evaluation serves as a direct evaluation whether our proposed work can eliminate the duplication. Such experiment will be conducted to the result dataset up to Step 5, before any specified data analytic task, of our proposed framework. Secondly, a machine learning algorithm, KNN classification between the AKF and Unspecified kidney condition (UKC) classes, is applied to the de-duplicated result datasets. This can evaluate how the result of our proposed work can be utilized in the further analytical tasks. The comparing methods are the baseline de-duplication 11 under our framework without NLP process (referred as basic de-duplication), and result of the classification on the original dataset without any modification (referred as Org). Note here that the GridSearchCV 16 is applied for fine-tuning the parameters in training process to automate the process of searching for the optimal combination of hyper-parameters within a predefined range in all the three comparing methods.
Results
Dataset exploration
Our proposed work is evaluated on the public available medical dataset, MIMIC-III database version 1.4 (MIMIC-III v1.4), the details of the dataset as well as some key findings from applying our framework to the dataset are as follows.
First, the MIMIC-III database consists of 26 tables, which is classified into four types: identifier, events, dictionary, and patient care. The identifier tables, i.e. PATIENTS, ADMISSIONS, and ICUSTAYS, employ distinct primary keys with the ‘ID’ text suffix to uniquely identify patients, hospital admissions, and intensive care unit stays, respectively. The events tables, i.e. OUTPUTEVENTS, NOTEEVENTS, CHARTEVENTS, and PROCEDUREEVENTS, have comprehensive records of distinct events and measurements pertaining to patients’ data, e.g. lab results or medical notes. The dictionary tables play a crucial role of facilitating cross-referencing between tables through ID of the defined terms, e.g. D_ICD DIAGNOSES, D_ICD_PROCEDURES, D_ITEMS, and D_LABITEMS for diagnoses ICD, medical procedure ICD, and lab item ID respectively. Last, the patient care table contains important information, e.g. physiological measurements, caregiver observations, or comprehensive billing details.
The results of applying our 7-steps proposed framework as shown in Figure 1 on MIMIC-III database are as follows. First, the related tables from the database are retrieved and joined hierarchically as shown in Figure 2. For example, patient’s information is to be joined with hospital admission, i.e. admission number, preliminary diagnosis, and also laboratory results, i.e. BUN, sodium, potassium, or creatinine. The tables are to be joined to the top of the hierarchy. The size of each joining result is shown in Table 1. Hierarchy of data joining. MIMIC-III joining result.

Then, in the basic de-duplication step, some duplications existing in dataset are removed initially for minimizing duplicate data based on the metadata. We construct an automated algorithm to remove such the dirty text elements, e.g. parentheses “()”, brackets “{}”, square brackets “[]”, chevrons “⟨⟩”, numbers, symbols, etc., in note-event column. After this step, the number of records is reduced to 735,609 records from originally 812,995 records.
Subsequently, the PMH extraction algorithm is applied for reducing excessive PMH records in note-event column. Typically, PMHs information in the MIMIC-III database are written on paper-based systems and stored to EHR system. This results in long texts and redundant data storing. Furthermore, the keywords of PMHs in the note-even column can be labeled variably, i.e. “PMH:”, “PMHx”, and “PMH”. With Algorithm 1, these keywords are split from the note-event column for finding patient’s morbidity. After the splitting, it is recursively separated into two chunks, until the patient’s morbidity is found in the second one. Then, the second chunk will be stored back into the dataset. In some cases, there are redundant PMH keyword found in the second chunk caused by excessive recording of practitioners. Therefore, the second chunk is re-split to exactly meet patient’s morbidity and store it back to the dataset.
The description of each column in the dataset.
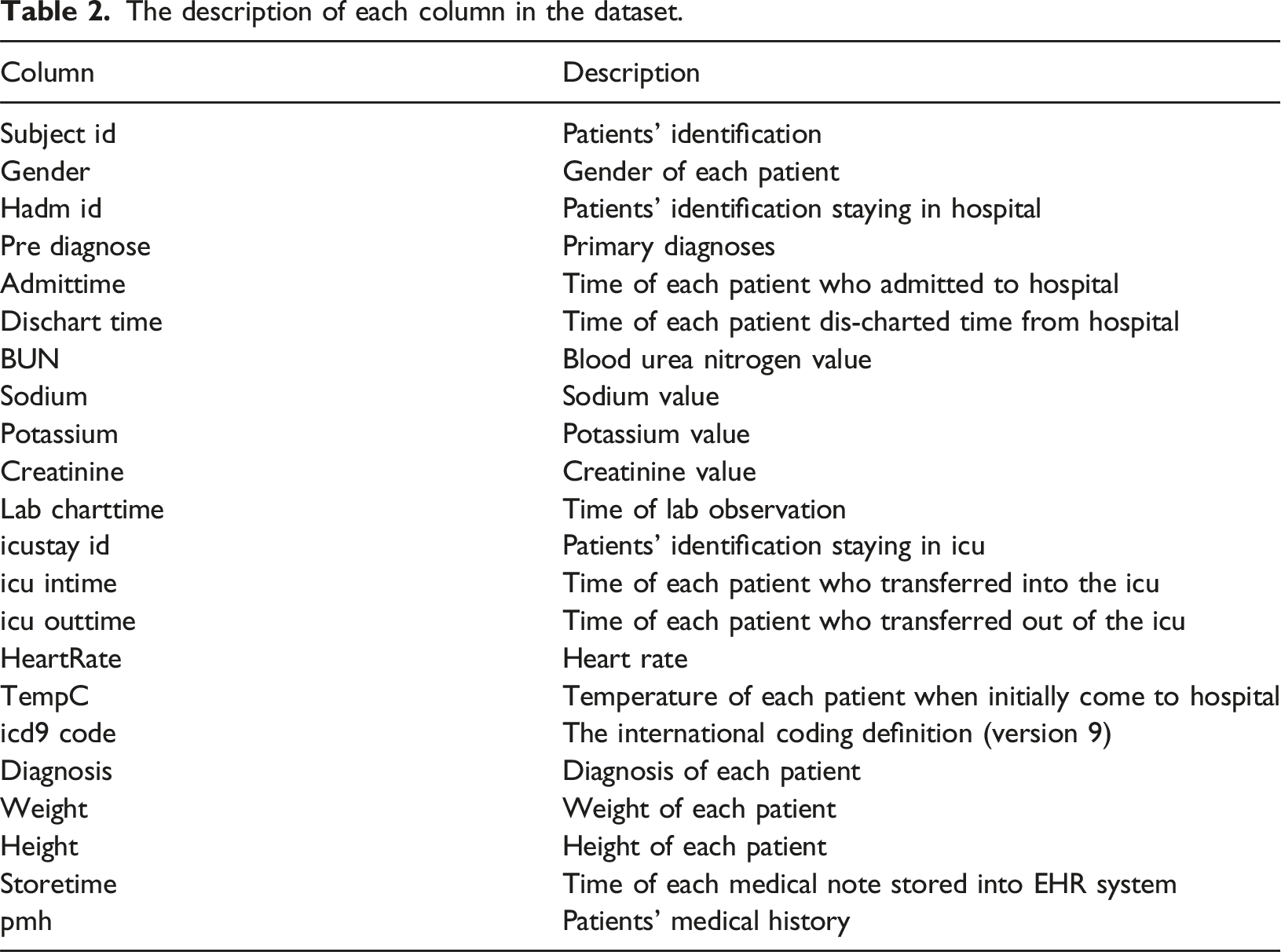

For the outlier elimination, we employ two techniques, i.e. percentile capping as suggested by the domain experts
17
as the following steps: (1) First, all instances of Acute kidney failure (AKF) such as Acute kidney failure with lesion of tubular necrosis and Acute kidney failure, unspecified, are combined under the category of “Acute kidney failure.” (2) Secondly, the other cases, which are not belonging in condition that result in kidney injury, are discarded. In addition, any records in the diagnosis column, which are less than 10% of the total dataset, as well as those with the highest occurrence, are excluded to mitigate potential biases during the training process. As the result of this elimination process, the final dataset consists of 39,264 records.
Lastly, the data imputation and normalization are proceeded. First the dataset is examined to assess the extent of missing data, as illustrated in Figure 3. The analysis reveals considerable variation in the proportion of missing values among variables, with a standard deviation of 0.34. Then, the KNN imputer is applied because its MCAR nature as mentioned before. Missing values in each of variables.
Distribution of kidney condition within the dataset.

De-duplication results
Performance of the proposed method for the de-duplication.

Classification results
To evaluate the classification performance between the AKF and UKC classes, first, the dataset was divided into 80:20 for the training and testing purpose respectively. Note that the number of records in each method was different due to its process, i.e., the number of records for the proposed method, the basic de-duplication, and the Org is 39,264, 43,258, and 43,258 respectively. The testing dataset from the 80:20 splitting for the proposed method, the size of basic de-duplication, and the Org result was 7,853, 8,652, and 8652 respectively. As mentioned before, GridSearchCV 16 was applied for fine-tuning the parameters in training process for hyper-parameters optimizing for all the three comparing methods.
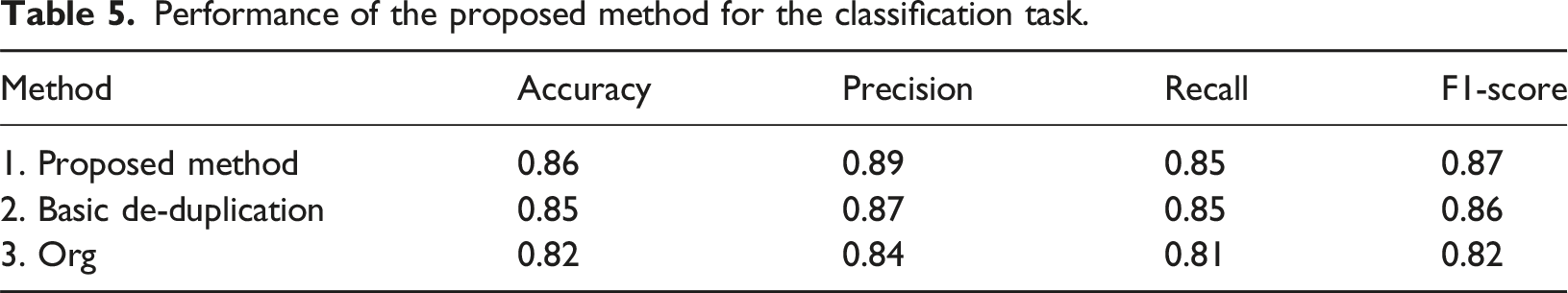
Performance of the proposed method for the classification task.
Confusion matrix of the comparing methods.

Discussion
First, with regard to de-duplication performance, our proposed method demonstrated an effective reduction in both false positive and false negative rates compared to the basic de-duplication. The false positive caused by the same notations in the EHR were written in the difference context in which the basic de-duplication cannot detect effectively. Meanwhile, the false negative caused by the small variation when recording the data in the EHR by different personal in a single tuple, thus the NLP applying can eliminate more effectively. From such processes, our proposed method can outperform the basic de-duplication in a large margin, the accuracy at 99.59% comparing with the basic de-duplication at 80.59 accuracy. In addition, when the discordant incorrect outcomes between the two methods, i.e. when any method detects the duplicates incorrectly while the other incorrectly detects, are to be considered, the McNemar’s test was statistically significant (χ2 at 79,800.00, p << 0.001), which can present the difference between the methods clearly.
For the classification performance, our proposed method aided the KNN classifier to achieve the highest accuracy result at 0.86 compared with the other two comparing methods. Also, the McNemar’s test between the proposed method and the basic de-duplication was also statistically significant (χ2 at 199.24, p << 0.001), which can present the difference between the two methods clearly. It can also be seen that the recall of our proposed method was lower than the recall. In general, the higher recall could be more challenging because the automatic process cannot justify the duplication exactly in the same way as manual process. Based on our experiment results, there were different interpretation of creatinine levels which may be ambiguous by each practitioner. There was variation in measurement units (e.g., mg/dL or µmol/L), or the normal range differs across the dataset. Moreover, there also were some incorrect laboratory test results remains in the dataset. For example, a patient had negative creatinine level which was not valid. This issue could be prevented by either a more effective data entry processes, or a semi-automated de-duplication process with human interaction.
Furthermore, the presence of diagnostically similar conditions may contribute to misclassification. Although elevated BUN and creatinine levels are commonly indicative of AKF, overlapping value ranges can occur across different renal conditions. For example, a patient presenting with a BUN level of 30 mg/dL and a creatinine level of 1.5 mg/dL was diagnosed with AKF, whereas another patient with a BUN level of 29 mg/dL and a creatinine level of 1.6 mg/dL was classified as having an UKC. Such similarities contributed to an increase in false negative rates, as evidenced by the 8.16% rate reported in Table 6, which subsequently affected the recall performance demonstrated in Table 5.
However, when considering F1 score, our proposed method still achieved the highest effective result at 0.87. In contrast, the baseline basic de-duplication method exhibited a slightly lower score of 0.85, as shown in Table 5. The difference was primarily contributed to a decline in precision. Such method cannot adequately handle the ambiguous cases existed in the dataset, i.e., semantically equivalent terms, but different representation. For example, “hypertension” and “HTN”. Furthermore, in certain instances, misclassification occurred due to the highly distinction feature, e.g., “Chest pain” which often occurred in pre-diagnostic records and was associated with AKF and some UKC.
Discrimination performance comparisons.

Conclusion
This paper presents a comprehensive data duplication framework comprising of data acquisition, basic de-duplication by the baseline algorithm, medical note-event extraction, NLP methods application, data mapping and unrelated and outlier data elimination, eventually missing data imputation. The proposed de-duplication method was evaluated against the baseline algorithm on MIMIC-III database in both de-duplication and the classification task based on AKF. The experiment results showed that the proposed work achieved up to 99.59% accuracy in de-duplication task, achieved a classification accuracy at 86 % and the F1-score at 0.87, which outperforming the comparing methods. Future work will focus on improving the recall metric, particularly addressing duplicates recorded at different timestamp through more semantic-aware approaches. Such distinguishing between variations in timestamps that represent genuine clinical updates versus redundant duplicates remains a significant challenge. Additionally, further investigations will explore other data analytical tasks within EHRs beyond AKF classification.
Footnotes
Ethical considerations
This research, utilizing solely open and publicly anonymous available data with no reasonable expectation of privacy issue, thus, does not necessitate ethical board review, as such data falls outside the purview of ethical oversight.
Author contributions
Chomchanok Yawana is responsible for initial research design, method implementation, experiment conduct, and manuscript drafting. Wachiranun Sirikul is responsible for method validation and data analysis. Juggapong Natwichai is responsible for the research design and manuscript review and editing.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Chiang Mai University.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
