Abstract
This study proposes a predictive model that uses structured data and unstructured narrative notes from Electronic Medical Records to accurately identify patients diagnosed with Post-Traumatic Stress Disorder (PTSD). We utilize data from primary care clinicians participating in the Manitoba Primary Care Research Network (MaPCReN) representing 154,118 patients. A reference sample of 195 patients that had their PTSD diagnosis confirmed using a manual chart review of structured data and narrative notes, and PTSD negative patients is used as the gold standard data for model training, validation and testing. We assess structured and unstructured data from eight tables in the MaPCReN namely, patient demographics, disease case, examinations, medication, billing records, health condition, risk factors, and encounter notes. Feature engineering is applied to convert data into proper representation for predictive modeling. We explore serial and parallel mixed data models that are trained on both structured and unstructured data to identify PTSD. Model performances were calculated based on a highly skewed hold-out test dataset. The serial model that uses both structured and text data as input, yielded the highest values in sensitivity (0.77), F-measure (0.76), and AUC (0.88) and the parallel model that uses both structured and text data as the input obtained the highest positive predicted value (PPV) (0.75). Diseases such as PTSD are difficult to diagnose. Information recorded in the chart note over multiple visits of the patients with the primary care physicians has higher predictive power than structured data and combining these two data types can increase the predictive capabilities of machine learning models in diagnosing PTSD. While the deep-learning model outperformed the traditional ensemble model in processing text data, the ensemble classifier obtained better results in ingesting a combination of features obtained from both data types in the serial mixed model. The study demonstrated that unstructured encounter notes enhance a model’s ability to identify patients diagnosed with PTSD. These findings can enhance quality improvement, research, and disease surveillance related to PTSD in primary care populations.
Keywords
Background and significance
Post-Traumatic Stress Disorder (PTSD) is a mental health disorder resulting from having experienced or witnessed a traumatic event such as an accident or war. 1 PTSD symptoms are manifested across several categories including intrusive thoughts, persistent avoidance, negative alterations in cognition and mood, and alterations in arousal and reactivity. PTSD can be difficult to diagnose. It requires that the symptoms persist for greater than 1 month; however, if a patient is reluctant to seek help, infrequent patient-clinician interactions can hinder diagnoses. Additionally, patients’ subjective and reporting biases, as well as variations in the symptoms of PTSD that can mimic other mental health conditions such as depression and anxiety, can prevent timely diagnoses. Although much work has been done on diagnosing PTSD,1–16 the majority of identified indicators represent group-level risk factors (e.g., injury) with less concern for personalized predictors of PTSD (e.g., patient demographics, symptom type, and severity). 1 Recent findings suggest that PTSD is associated with an array of multimodal risk indicators, which makes it unlikely that any single vulnerability factor will account for a large amount of variance in the prediction of this complex disorder. 2 To address multimodal risk factors, forecasting methods of PTSD using Electronic Medical Record (EMR) data exclusively must accommodate multiple combinations of risk indicators. Additionally, the forecasting method must account for missing risk indicators that might not be documented in some patients’ EMR and use prior knowledge to adjust the relative weights of putative predictors that are documented in the EMR.
The application of computational methods and machine learning techniques to health data is a promising field of research that aims to improve our understanding of health conditions, disease trajectories, and the quality of medical services. Machine learning techniques are well suited for knowledge discovery and outcome prediction for diseases that have complex etiology and multifaceted manifestations like PTSD. Machine learning methods, in particular, supervised learning methods, can discover structures and correlations within high dimensional multimodal data such as patients’ history, patient demographics, prescribed medications, laboratory results, health conditions and diseases, acute care presentations, and other biometric data that can inform prediction.14–16 Shickel et al. reported a large increase in deep-learning approaches to EMR datasets with the most common supervised models being the Recurrent Neural Networks (RNN), Long Short-Term Memory (LSTM), Gated Recurrent Units (GRU), and Convolutional Neural Network (CNN). The central idea behind computational methods in medical disease identification is to explore patients’ data, perform feature engineering, and then use statistical or machine learning algorithms to process the data with the goal of designing a model that is able to assist clinicians in large scale and accurate disease identification.1–16 Intelligent assistive systems can provide details that can increase diagnostic accuracy, reduce human errors, and allow efficient use of patients’ medical data. 17
Previous studies have explored different types of data sources such as EMR data4,5; qualitative data including self-reported data 7 ; telephone-based and face-to-face interviews1,8,9; surveys9,10; scales of psychiatric symptoms1,3,8,13; administrative data holdings8,11; event and emergency department (ED) features1,2,12; biochemical examination 8 ; injury etiology10,12; sleep quality experiment, 14 and data collected using wearable devices. 15 The complexity of PTSD requires data that can capture an array of multimodal risk indicators. 1 EMRs are a rich source of knowledge collected by primary care clinicians during every patient encounter that generally extends over multiple years. To develop methods and tools that could accurately predict or diagnose PTSD requires the capture of complex symptoms and variable interactions between presumed markers from a variety of fields within the EMR. 16
This study aims to develop a predictive model to accurately identify PTSD and associated symptoms using primary care community-based EMR data. We extend the existing prediction algorithms that use structured EMR data and propose a novel model that combines both structured and unstructured free-text encounter notes. Unstructured encounter notes may contribute to earlier diagnosis as well as reclassification of patients who may have been misdiagnosed with other mental health conditions that have similar symptoms as PTSD.
Materials and methods
The data used in this study was extracted from the EMRs of primary care clinicians participating in the Manitoba Primary Care Research Network (MaPCReN), a subnetwork of the Canadian Primary Care Sentinel Surveillance Network (CPCSSN). 18 The MaPCReN database contains information extracted from 266 primary care clinicians providing community-based health care in 48 clinics in Manitoba, Canada.
Based on the records, 154,118 patients were seen by participating MaPCReN primary care clinicians between 1 January 1995, and 31 December 2017. Structured data were available for all patients in the EMR dataset (n = 154,118). However, unstructured free text encounter note data were only available for 56,795 patients (who represented 2,125,961 encounter notes).
Using primary care EMR data in machine learning models presented many challenges due to the subjective note writing styles and diagnosis of the physicians, spelling mistakes, domain-specific terminology, and abbreviations, duplicate text from different patient encounters, and redundancy and ambiguity of information. To build a predictive model for PTSD diagnosis we (1) transformed and included various data types, (2) addressed imbalanced classes, and (3) applied filtering and other data pre-processing to control variations in the health information available for each patient.
Data aggregation
The dataset for this study was extracted and compiled from eight tables of the MaPCReN data repository including patient demographics, disease case, examinations, medication, billing records, health condition, risk factors, and encounter notes. The encounter notes table contained unstructured free text data entered by primary care clinicians during every primary care appointment with patients. We concatenated all encounter notes for each of the 56,795 patients to create a single note for every patient. The resulting note varied in length from a few words to 6,850 words. Additionally, we used the following database tables, which contained structured data recorded in the EMR: patient demographics, examinations, encounter diagnoses, billing records, health conditions/problem list, medication, and risk factors. These tables provided patients’ demographic information as different data types such as gender (binary); year and month of birth (numerical); examination results; for example, Body Mass Index (BMI) (numerical), systolic Blood Pressure (sBP) (numerical), and diastolic Blood Pressure (dBP) (numerical); prescribed medications (Anatomical Therapeutic and Chemical (ATC) code) (categorical); billing/health condition code (categorical), and risk factors (name of the risk factors of the patient) (categorical). The disease case table provided a categorical list of the chronic health conditions with validated case definition algorithms available in CPCSSN. 18 Case definitions captured chronic diseases using International Classification of Disease (ICD-9) codes from the billing, encounter diagnosis, and health condition tables, ATC codes for prescribed medication and laboratory results. 18
Variable creation
Summary of descriptive variables used in this research.
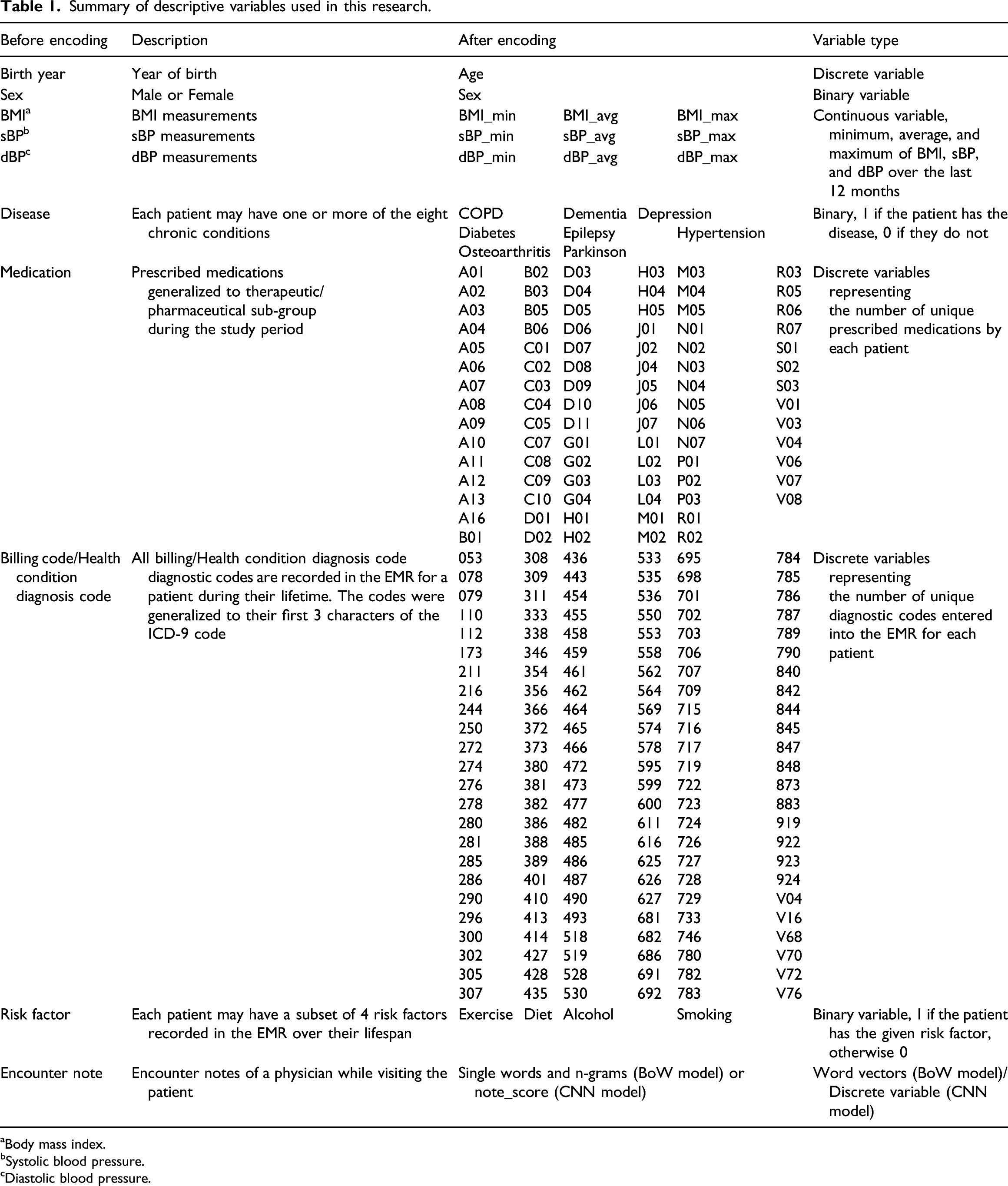
aBody mass index.
bSystolic blood pressure.
cDiastolic blood pressure.
Medications, billing codes, and health condition diagnostic codes are recorded as categorical features in the MaPCReN database. In our dataset, we had 1,522, 7,102, and 7,695 different values of medications, billing codes, and health condition diagnostic codes, respectively. This large number of unique values presented challenges when encoding with the one-hot encoding. To reduce the number of unique medication values present in the dataset and simplify the model, we grouped prescribed medications according to their therapeutic or pharmaceutical sub-group by taking the three left characters of the ATC codes. As a result, 1,522 different values were reduced to only 88 unique medication codes. A similar method was applied to the ICD-9 diagnosis codes found in the billing and health condition tables, which reduced the number of codes from over 7,000 in both cases to only 144 unique values.
After encoding, we excluded the ICD-9 code of 309.81 as it can denote the outcome variable that we are trying to predict. The variables used in the study are explained in Table 1.
We processed unstructured encounter notes using NLP and deep-learning algorithms. We defined a Bag of Words (BoW) model and a Convolutional Neural Network (CNN) model for feature extraction and transformation of the EMR text data. The CNN model converted the unstructured notes to a continuous value in the range of [0, 1]. The details of these models are presented in the following subsections.
Analysis preparation
A total of 154,118 patients were seen by the participating primary care providers of the MaPCReN between 1 January 1995 and 31 December 2017. The retrospective study was approved in 2018, at which time medical students spent 320 h reviewing medical charts to create a reference standard for this work. They reviewed the EMR charts of 1,137 patients to identify symptoms of PTSD or explicit mention of PTSD symptoms listed in the diagnostic service manual (e.g., distress, hyperarousal, functional impairment, and traumatic event). To build the gold standard from these 1,137 patients, 933 patients were selected as they had a diagnosis code in the EMR for an adjustment reaction (ICD-9 starting with 309). Previous validation studies comparing chart review to ICD-9 codes found that ICD-9 codes do not capture all patients with the condition. 18 We assessed the presence of an ICD-9 code of 309.81 (diagnoses code for PTSD) among our reference set. Not all of the patients in our reference group had a confirmed diagnosis of PTSD (ICD-9 code 309.81) or unstructured chart note data. As a result, the final reference set of 195 patients included patients with both an ICD-9 code for PTSD and structured and unstructured chart note data.
Of the total 154,118 patients, 56,795 had both structured data and narrative notes, and as outlined above, 195 patients were manually validated by chart note review to have a diagnosis of PTSD. With the 195 positive samples and plenty of negative instances, we created a training/validation and a hold-out test dataset. The training/validation dataset contains 85% of the total PTSD positive samples (n = 173) and an equal number of randomly selected non-PTSD cases for training, and an additional 1,700 randomly selected PTSD negative instances for validation purposes. A severely imbalanced dataset was created as the hold-out test dataset containing 15% of the total PTSD positive samples (n = 22) and 2,178 randomly selected PTSD negative instances. We created this dataset following the CCHS (Canadian Community Health Survey), which suggests that in Canada the prevalence of PTSD is ∼1%. 19
Predictive modeling
We investigated different models for assessing structured data, text data, and their combination. For assessing structured data, we developed a Random Forest (RF) and a Multi-Layered Neural Network (MLNN) model. For assessing free-text data, we developed two models, one based on the simple Bag of Words (BoW) model that serves as a baseline for text models, and a more sophisticated model that uses word embeddings and Convolutional Neural Networks (CNN). Comparing the results of the structured data models and the BoW baseline model with that of the CNN text model shows the value of text data in medical diagnoses as well as the effectiveness and applicability of deep learning in encoding medical text data. Finally, we explored two types of mixed data models, a serial model, and a parallel model, that were trained on both the structured and unstructured data for predicting PTSD.
Model implementation and validation
Structured data models
We developed two classification models based solely on the structured data: a Multi-Layered Neural Network (MLNN) model and a Random Forest (RF) model. The MLNN model accepted the 399 input variables in its first layer, and then processed them via four hidden layers, each having 50 nodes with a ReLU activation function. The binary cross-entropy was used as the loss function and the efficient Adam implementation of gradient descent was used to optimize the model. We trained the model for 20 epochs. Both RF and MLNN classifiers are implemented using Scikit-learn. 20 For the RF classifier, all the default settings were chosen including n_estimators = 100, splitting criterion = gini, max_depthint = none, and min_samples_split = 2. 21
Text data models
For assessing free-text data, we developed two models. The first model which serves as a baseline for text processing encodes the text using the simple BoW model and classifies them using a random forest (RF) classifier. The second model uses word embeddings to encode text documents and utilizes the CNN model for classification.
In the baseline text data model, the BoW and its extension n-gram 22 were used to convert the text fields into numerical feature vectors, which were then fed to the machine learning algorithms. During the pre-processing phase, the input text documents were tokenized and then non-alphabetic characters, stop words, emoticons, and special character strings were removed. We also lemmatized the words, using the spaCy lemmatizer function, to remove common morphological endings. All words were converted into lower case. As a measure of the presence of known words as well as assigning weight or importance to each word, we computed Term Frequency–Inverse Document Frequency (TF–IDF). Next, we used the TfidfVectorizer implemented in Scikit-learn to vectorize the text into n-gram integer vectors so that they could be passed as inputs to the classifiers. We empirically determined the best values of the parameters for the TfidfVectorizer function: ngram range of 1–3, and min_df of 0.005, which gave the best results on the validation dataset. We applied supervised learning to train the RF classifier using the training dataset to develop our baseline PTSD diagnosis model.
To create a more powerful model for processing free-text data, we used deep-learning algorithms. The non-linearity of these computational models, as well as the ability to apply more sophisticated word encoding based on word embedding methods, often lead to superior classification accuracy. 23 We used CNN for this purpose as they have proven to be successful at document classification problems. 24 CNNs are effective at document classification because they can extract salient features from documents represented using a word embedding in a way that is invariant to their position within the input sequences. 25 The architecture of our deep-learning model comprises three key pieces: word embedding, convolutional model, and fully connected model.
First, the narrative notes are preprocessed which involved converting all the words into lower case, removing all punctuation from words, and removing stop words. Next, a Keras Tokenizer was fitted on the training dataset to define the vocabulary for the embedding layer and transform the text notes to integers. This was followed by padding all document vectors, each document representing a collection of all notes of a patient, to the length of the longest note in the training dataset. Then, sentences were mapped to embedding vectors, and thus each document was transformed into an input matrix for the machine learning model. We used a 100-dimensional vector space for word embedding. The word embedding layers were initialized randomly to be trained along with the rest of the network. Then, a multi-channel convolutional neural network with 64 filters extracted and learned features from the input words using different sized kernels, when feeding 1, 2, and 3 words at a time. This allowed the document to be processed at different n-grams (1, 2, and 3-g). The resulting feature maps were then processed using a max-pooling layer to consolidate the output feature from the convolutional layer as depicted in Figure 1. Finally, these extracted features were interpreted by a fully connected model to a predictive output in the range of [0, 1]. The backend fully connected model had two layers, one with 10 nodes and a ReLU activation function and the other with one node and a sigmoid activation function. Finally, we fitted the network on the training data. We used a binary cross-entropy loss function and a batch_size of 16. The efficient Adam implementation of stochastic gradient descent was used and we kept track of accuracy in addition to loss during training. The model was trained for 10 epochs using the training data and validated via validation dataset in a 10-fold cross-validation fashion. Parallel architecture: Free text data and structured data are processed in two parallel branches of a multi-input model. Their results are concatenated and fed to a backend MLNN which makes the final prediction.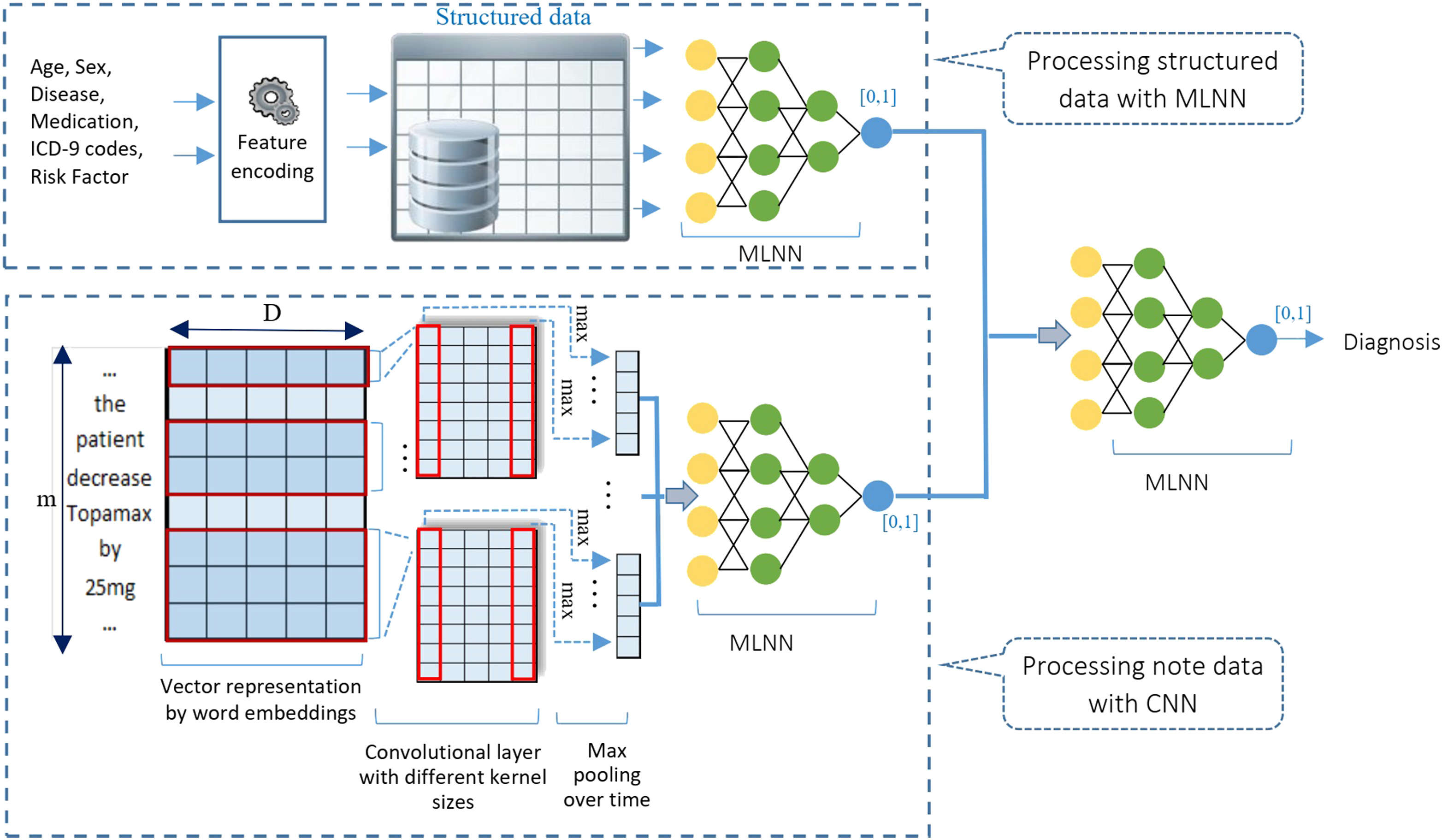
Mixed data models
We explored two types of mixed data models, a serial model, and a parallel model, to combine the model predictions from the structured and unstructured text data and to explore the combined predictive power of the two types of data in different orders in diagnosing PTSD.
Parallel model
The first mixed data model handles the structured and unstructured data in two parallel data modeling pipelines as shown in Figure 1.
The parallel mixed data model was implemented using Keras APIs 26 which allows the different components of the mixed data model to be trained at the same time. It accepts structured data as the input into one processing pipeline (structured data branch) and free text encounter notes as input in the other parallel data processing pipeline (text data branch). Outputs from the two parallel pipelines are passed into a final decision model that predicts the binary outcome of PTSD or non-PTSD (mixed data branch). Specifically, in the structured data branch, a MLNN converts the structured data into a real number. It includes four hidden layers following the input layer that accepts the 399 input variables. The hidden layers have 50, 50, 50, and 20 nodes with a ReLU activation function. The final layer contains one node with a sigmoid activation function to create a value in the range [0,1] indicating the probability of a patient being PTSD positive or negative based on structured data. In the text data branch, a CNN model is responsible for processing free-text data which is the same as the one explained above under the Text Data Model. Outputs from the two parallel branches are then concatenated using Keras functional API 26 and fed to a backend MLNN decision model. The final MLNN decision model includes two layers with 20 and 1 nodes, respectively. It interprets the extracted features and makes the final prediction (Figure 1).
Performance metrics for different proposed models on 10-fold cross-validation on the validation dataset.
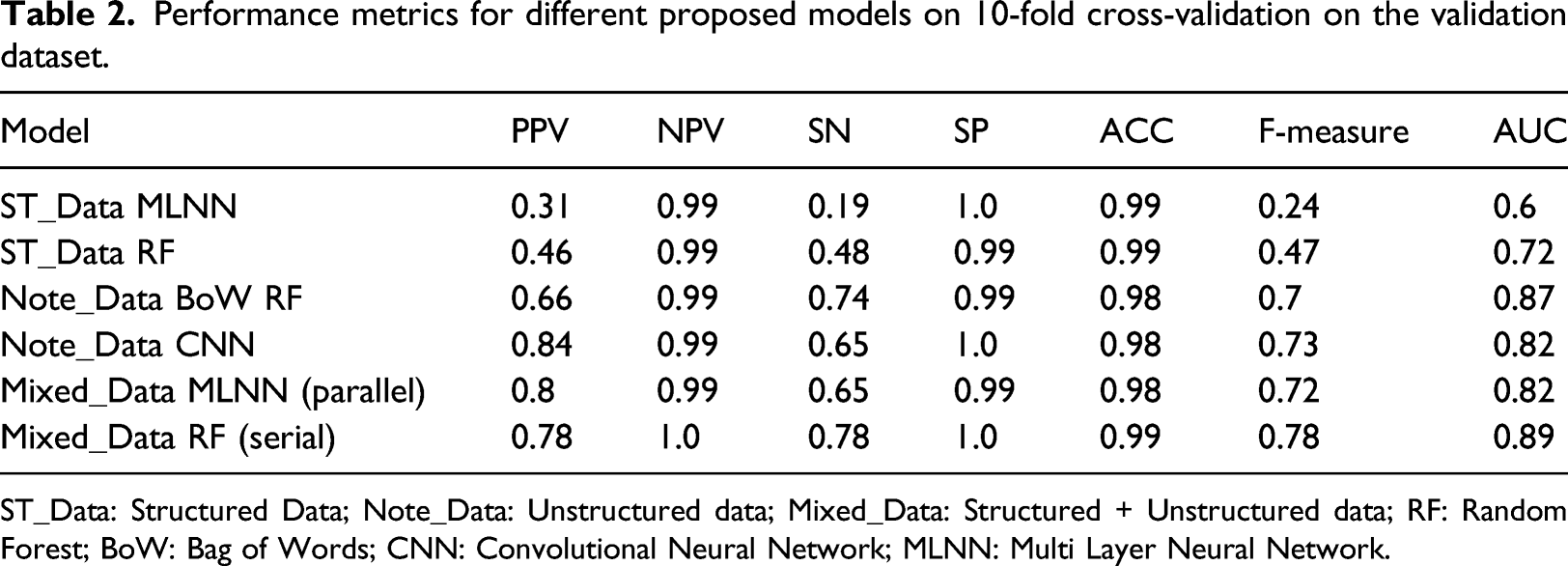
ST_Data: Structured Data; Note_Data: Unstructured data; Mixed_Data: Structured + Unstructured data; RF: Random Forest; BoW: Bag of Words; CNN: Convolutional Neural Network; MLNN: Multi Layer Neural Network.
Serial model
The second mixed data model applies an innovative serial data processing approach and models the structured and unstructured data based on deep learning and ensemble learning methods (Figure 2). Serial architecture: There are two components in this method, the first for processing free text data and producing the input for the second component to be processed with the structured data.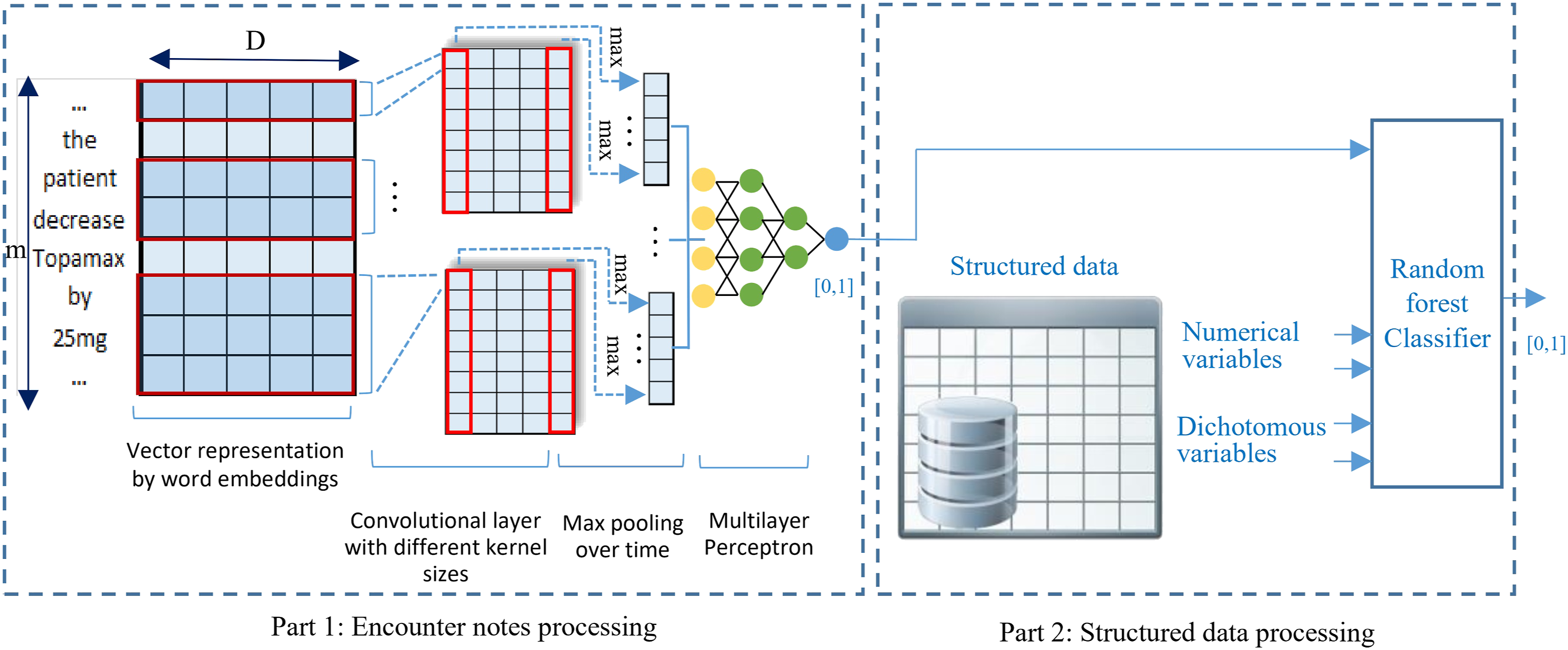
This model consists of two components. The first component processes unstructured text data (text data component) using a CNN deep learning-based model very similar to that explained under the text data model and outputs a numeric value in the range of [0, 1] indicating the probability of a patient being PTSD positive or negative based only on the notes. This component is trained first to generate the note score values in a cross-validation fashion. Next, the second component accepts the output of the text data component and combines it with the other structured features as listed in Table 1, to generate the final prediction. Unlike the parallel model, whose components were trained at the same time, we had to train the components of the serial model separately. By stacking the two sub-models, the combined model is able to ingest mixed data and make a binary prediction of PTSD and non-PTSD. The RF classifier is implemented using Scikit-learn 20 with all the default settings chosen as explained in Structured Data Models.
Validation
We validated our models on the validation data set with 10-fold cross-validation to obtain the optimal values of the hyper-parameters for each model. Considering the skewed nature of our dataset, we aimed for a threshold that maximized the F-measure value. We evaluated our models with several metrics including Positive Predictive Value (PPV), Negative Predictive Value (NPV), specificity (SP), sensitivity (SE), F-Measure (F1), overall accuracy (ACC), and the Area Under the curve (AUC) of a Receiver Operating Characteristic (ROC) curve. Significance was assessed at 0.05. The equations used for calculating these metrics are presented below

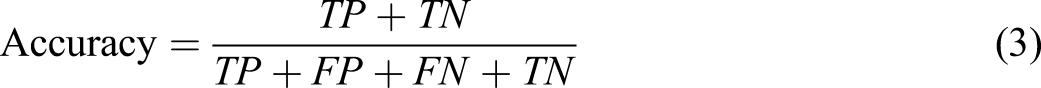


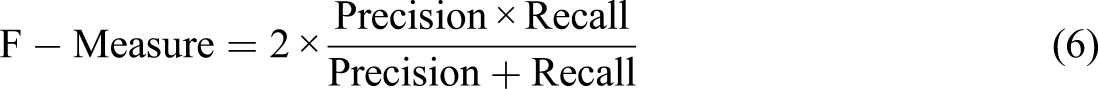
Analyses
To study the contribution of each predictor in the serial model applied to the PTSD hybrid dataset, we performed a feature importance ranking. Feature importance was assessed for the features having importance ≥0.005 in the RF model. This is calculated based on Gini impurity score or Mean Decrease Impurity (MDI) which is the impurity reduction achieved by splitting the features. Therefore, the sum of the impurity reductions in all the trees for a variable is calculated as the importance of the variable. For impurity reduction, classification trees use Gini coefficient index or information gain of variables. The equation for calculating the importance of variable x
j
is as follows
27
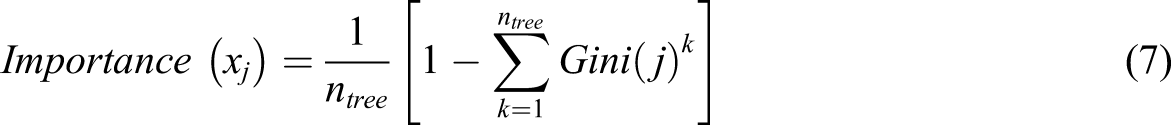
As the impurity importance is known to be biased in favor of variables with many possible split points28,29 (e.g., continuous variables), we also investigated feature importance based on Pearson correlation coefficient score 30 between feature and class label.
This study was approved by the Health Research Ethics Board at The University of Manitoba with the research Ethics number of HS21053(H2017:257).
Results
Performance metrics for different proposed models on the hold-out test dataset.
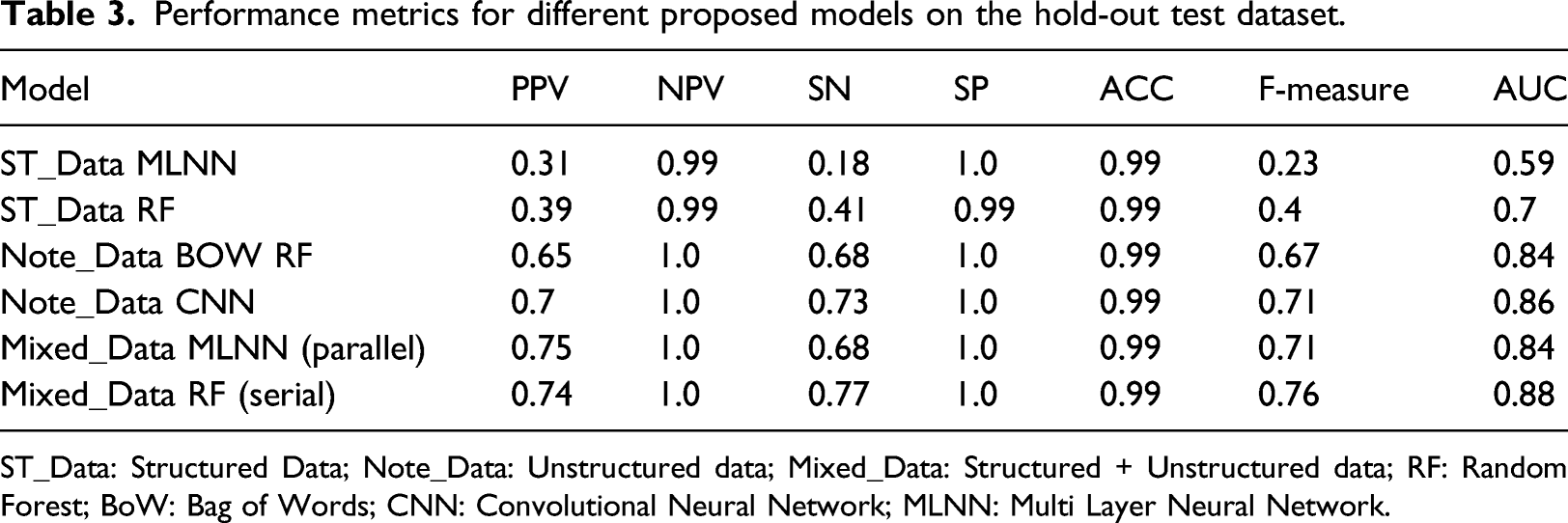
ST_Data: Structured Data; Note_Data: Unstructured data; Mixed_Data: Structured + Unstructured data; RF: Random Forest; BoW: Bag of Words; CNN: Convolutional Neural Network; MLNN: Multi Layer Neural Network.
The serial model represented in Table 3 as Mixed_Data_RF yielded the highest values in sensitivity (0.77), F-measure (0.76), and AUC (0.88). However, the parallel model represented as Mixed_Data MLNN, obtained the highest PPV (0.75).
Feature importance
Our first feature importance model is based on the impurity reduction of splits in the RF classifier and can highlight which variables are contributing more to the prediction. Figure 3 shows the relative importance of the features with the importance ≥0.005 in the RF model applied on the mixed data represented as Mixed_Data RF (Serial). The most important feature is the value assigned to the encounter note (note_score), followed by the health condition category of 309 (HC_309) which denotes adjustment disorder, depressed mood, and anxiety disorder. Other interesting features on this list are nervous system medications, and in particular, antidepressant medications (ATC N06*** and N02***). HC_305 which denotes tobacco, alcohol, drug, and opioid abuse, is also on the list. Depression, a comorbid disease, was also an important feature. The rest of the list is continuous variables like age, blood pressure, and BMI as well as discrete variables of ATC codes (Figure 3). Average importance from the cross-validation tests of the RF model indicates the proportional value contributed by a feature towards the final model prediction of PTSD. Shading indicates the score of the features with importance ≥0.005 based on their importance.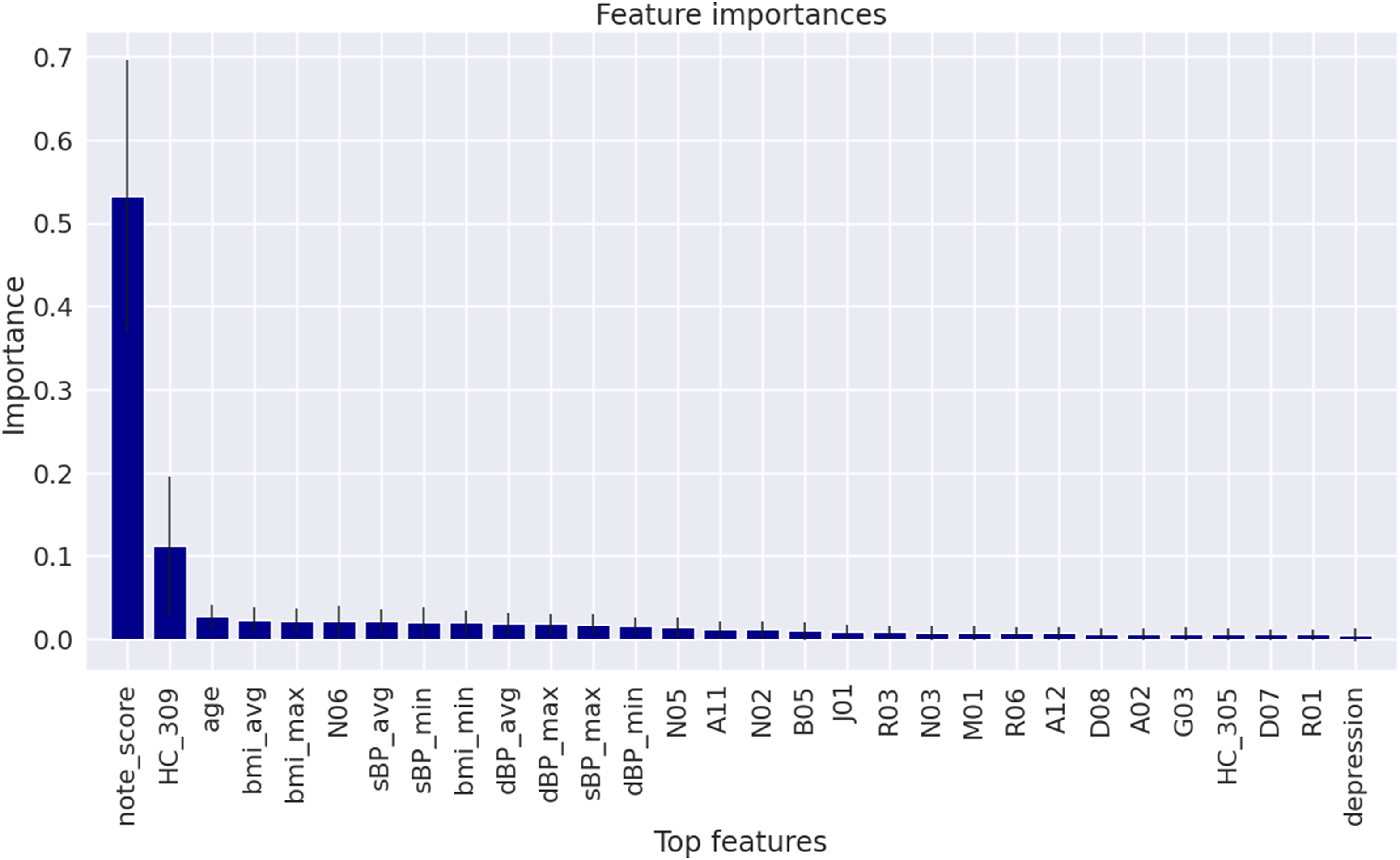
Features with a correlation ≥0.05 with the outcome.
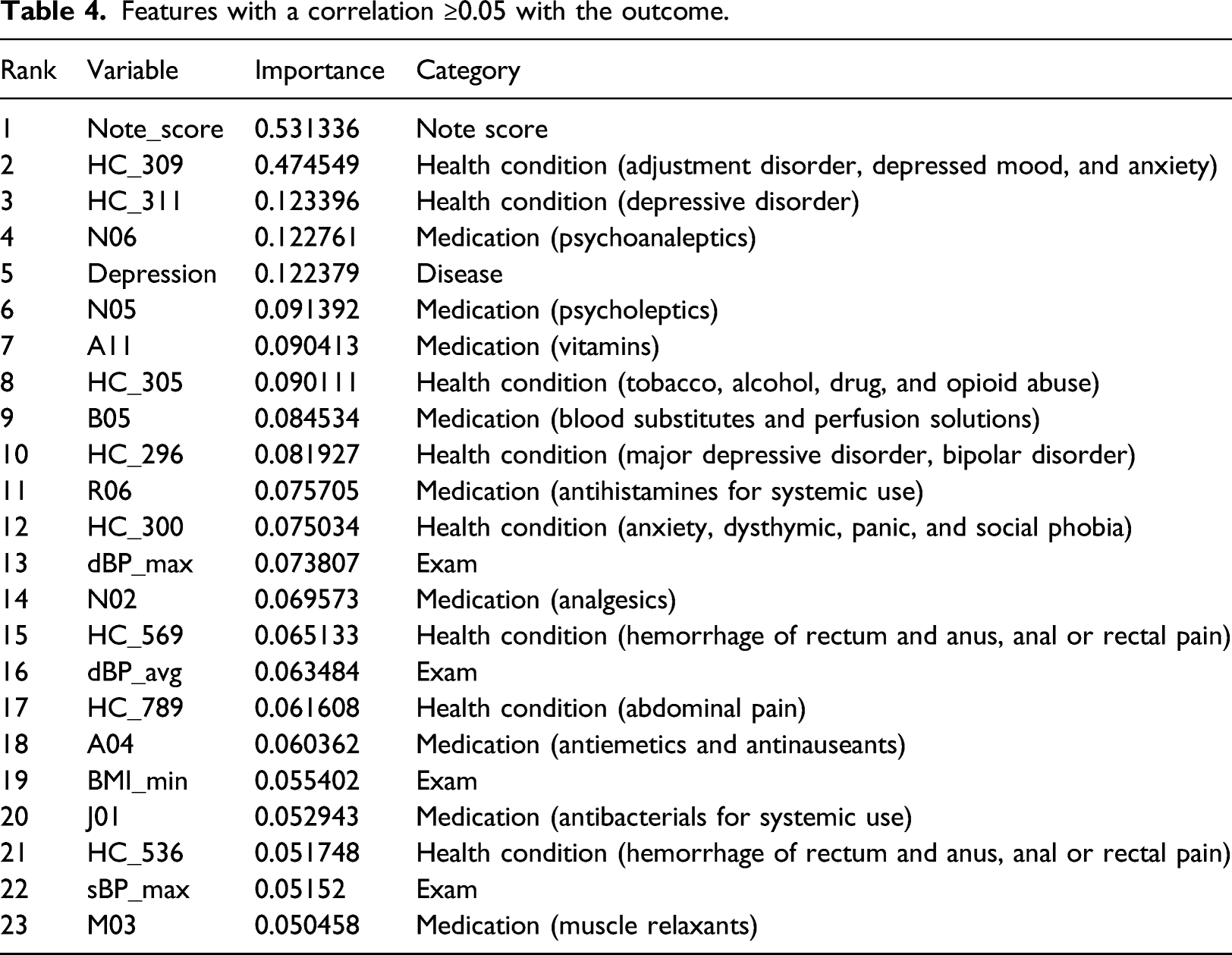
As shown in Table 4, from the 23 features with a correlation >0.05 with the outcome, again the most important feature was found to be note_score, followed by HC_309. While the RF model associated depression and PTSD via a comorbid disease variable (depression), the correlation found this association through the health condition codes of 311 and 296 (the third and 10th rows in Table 4). The nervous system medications of N06*** and N02*** were also identified by this method. The remaining features are from other health conditions, medications, and exams.
Discussion
In this study, we applied machine learning techniques to develop models for diagnosing PTSD based on both structured and unstructured EMR data. One distinctive characteristic of our work is the utilization of mixed data modeling that is able to ingest a mixture of structured and free-text data from community-based EMRs to identify PTSD. We discuss (a) the contributions of this study by comparing our work with the state-of-the-art medical research, (b) possible reasons behind the observed results achieved by the various computational models to serve as a guide for future research studies, and finally, (c) challenges with the data and the limitations of this work.
Research contributions
Previous studies have mainly focused on structured data or exploration of data obtained using qualitative data collection techniques, either in specialized clinics or other PTSD-specific studies.1–16 Our work, on the other hand, used noisy EMR data from primary care which broadly describes the health conditions of each patient instead of in-depth rich data focus on selected health conditions as obtained from the specialized clinics. While specialized data can result in models with high accuracy, this kind of data is expensive, time-consuming to acquire, and is unable to comprehensively explore the patients' health outcomes. The readily available nature of EMR data, on the other hand, makes it easy to access for diagnostic and prognostic purposes.
While EMR data systems contain both structured and free text data, most studies focused either on structured fields or text notes.4,5,31 However, increasingly more studies are using such diverse sets of data. 32 Diao et al. 33 developed five machine learning prediction models of common etiologies in patients with suspected secondary hypertension. Both structured data and CT text reports were assessed in this study; however, they leveraged regular expressions to extract structured data out of text and fed them along with other features to an XGBoost model. Liu et al. 34 proposed a general framework for disease onset prediction that combined both free-text medical notes and structured information using deep-learning architectures including CNN, LSTM, and hierarchical models. Baxter et al. 35 generated predictive models to predict the need for glaucoma surgical intervention using EMR data. By excluding free-text notes, they only utilized structured data in their system. Harrington et al. 4 utilized rich EMR data to identify PTSD in the U.S. veteran population. Our dataset did not have the in-depth details available in some specific PTSD datasets limiting our model in terms of data features and therefore preventing diagnosis of PTSD with high accuracy. However, our models informed primary care providers to make preliminary diagnosis and as necessary, refer patients to a specialist.
Researchers such as Ma et al. 16 suggest the need for more advanced models to identify and predict the severity of PTSD. Harrington et al. 4 found that using Lasso algorithm showed modestly higher agreement with a chart review as compared to an ICD rule-based algorithm. Ma et al. 16 developed a clinical decision support pipeline using patient data from telephone interviews having a sensitivity of 0.62–0.67 and a specificity of 0.69–0.73. Shickel et al. 32 suggest that code-based representations of clinical concepts and patient encounters are only the first step towards working with heterogeneous EMR data. Judd et al. 36 developed a generic decision support system to diagnose chronic low back pain by utilizing machine learning algorithms on unstructured notes in EMR. LaFreniere et al. 37 developed an artificial neural network to predict hypertension from clinical data with an accuracy of 82%. Although some community-based EMRs such as ours do not include imaging data, we were able to include measurement data including laboratory results, vital signs, etc. The structured EMR data fields ensure that key diagnoses, medications, and information about the patients are entered in a systematic way. Clinicians may elaborate on their observations, assessment, and treatment plan within the unstructured free text data fields, and as shown in the result, our mixed data models that used both structured and unstructured free-text data had the highest predictive power for identifying PTSD.
Previous application of traditional NLP techniques has explored free text data collected during PTSD-focused interviews with patients or clinicians. 7 To the best of our knowledge, this is the first model to apply state-of-the-art text processing techniques on Canadian community-based free text EMR data to identify PTSD. The study also incorporates a combination of the structured EMR and text data and compares the performances of a variety of models trained and validated using both data types.
Analysis of Computational Models
We explored six models to assess their ability to screen PTSD using structured data, unstructured notes, or both types of data. We tested our models on a highly skewed hold-out test dataset which has a PTSD positive ratio close to the prevalence of this disorder in the Canadian population. Two models, a RF and a MLNN model, were developed using the structured EMR data including ICD-9 codes for PTSD identification. The RF obtained a F-measure of 0.4, which is higher than the F-measure of 0.23 acquired by the MLNN model, however, none of them were informative enough on the highly skewed test dataset compared to the other four models that utilized the chart note data. This shows that structured data models provide a moderate prediction of PTSD. Similar to Harrington et al., 4 we found that using ICD-9 codes alone did not produce the highest accuracy in identifying PTSD. This study, therefore, highlights the value of note data in medical data analytics. We also developed two models based only on note data, a RF classifier in combination with the BoW encoding model and a CNN model. Comparing the results of these two models, the CNN model obtained about 5 points higher values in PPV, SN, and F-measure. This indicates that the CNN model outperforms the traditional BoW model in extracting relevant features from the clinical text data. Other researchers have found the CNN-based model is effective in using real clinical EMR data to predict congestive heart failure or chronic obstructive pulmonary disease. 38
We also investigated the impact of incorporating a mixture of both structured and text data with a parallel and a serial mixed data model. With the highly skewed data used, we achieved a high value of almost 1.0 for all of our models for the validation metrics of NPV, SP, and ACC. The validation metrics used the True Negative (TN) value in their calculation, suggesting that we have to rely on the other metrics for performance comparison. The serial mixed data model that combined the data using a RF classifier outperformed all other models by obtaining the highest values in SN, F-measure, and AUC metrics while the parallel mixed data model acquired the highest PPV value. This suggests that the RF classifier had a much higher degree of separability when distinguishing between classes. RF models are known to be unhindered by adding weakly predictive attributes, 8 which is the case in our dataset. Another advantage of the serial model is that it has better interpretability compared to the deep learning-based parallel model. Comparing the CNN note data model that works solely based on text data (Note_Data CNN) and the parallel mixed model, they both have the same F-measure; however, their performance is different in terms of PPV and SN. While the Note_Data CNN model performed better in terms of SN (0.73 vs 0.68), the parallel mixed data model obtained a higher PPV (0.75 vs 0.7). This suggests that structured data informed the parallel model to have fewer False Positive (FP) cases. On the other hand, the note data used by Note_Data CNN model helped it to have higher True Positive (TP) and lower False-Negative (FN) rates. In terms of AUC, again text-based models outperformed structured data models, both in the validation and test datasets, with the serial mixed data model being the best performing model followed by the CNN text model.
Tree-based methods and deep learning-based models are both powerful machine learning algorithms. However, each of these methods has its strengths and weaknesses. While neural network approaches are necessary for image analysis or NLP-related tasks, they are often overkill for tabular data and prone to overfitting. 39 Neural networks require explicit handling of missing values prior to modeling, but gradient boosted trees handle them automatically. 40 Ensemble methods like RF reduce bias and variance by incorporating different estimators with different patterns of error, to diminish the impact of a single source of error. 40 As we can see in Tables 2 and 3, the CNN model outperformed the RF in assessing note data. In the case of tabular or structured data, on the other hand, tree-based models are advantageous. 40 Structured data is natural for a decision tree while a neural network is overkill for tabular data prediction. 40 That is why our serial model outperformed the parallel model in both validation and hold-out datasets. However, the parallel model is easier to implement as we trained it at the same time with the help of Keras functional APIs.
In designing the parallel mixed data model, we experimented with several ways to combine the structured and the text data using the Keras functional APIs. In particular, we considered concatenating different formats of feature vectors obtained from the structured and the text data processed by MLNN and CNN sub-networks, respectively. These include applying different activation functions, applying a flatten layer on the feature vectors, using different vector sizes, and converting the feature values to a probability value.
Shickel et al. 32 reported that the human interpretability of models is important for clinical application. Assessing important features in a model is a valuable way to validate a model for clinical relevance.32,42 Many features that were found to be important in our study are consistent with factors associated with PTSD in other research.42–46 However, some features associated with PTSD in the literature42–46 were not important features in our model. The data recorded in a community-based EMR is different compared to the data found in the PTSD/psychiatric specific datasets. PTSD symptoms identified in primary care by a family physician or nurse practitioner would likely be referred to a psychiatrist for formal assessment. Primary care clinicians would assist with the management of comorbid medical conditions. However, treatment for PTSD may be sought at an operational stress injury clinic or co-managed with a psychiatrist. Therefore, not all the details specific to PTSD would be available in the primary care EMR. 47 The developed models need to consider the population under investigation and use the available features in the dataset. When we expanded our analysis to assess the correlation of variables with the outcome, we found many features were similar to those from the impurity reduction of splits in the RF. Further work is required to improve machine learning outcomes using a larger cohort that contains more positive PTSD patients. As well as investigate an alternative solution for the skewed class problem by applying a cost function with different miss-classification weights for the majority and minority classes. 48
Limitations
Positive cases of PTSD were identified using a structured data algorithm that identified patients with an ICD-9 code for PTSD (309.81) 18 when available. We confirmed the diagnosis using a manual chart review. However, encounter notes are not available for all patients in the MaPCReN data repository due to variations in data-sharing agreements with the data custodians. Patients without encounter notes in the data repository could not be included in our manual chart review and therefore were not used as a reference standard in our models. Future work will aim to validate the structured data case definition for PTSD (ICD-9) with a chart review using EMR data from other provinces to obtain a larger sample of patients with PTSD.
PTSD is a rare chronic condition producing a highly skewed dataset due to the substantially lower number of patients with PTSD compared to the total number of patients in the data repository. To mitigate this problem, we applied down-sampling techniques to create a training dataset. The down-sampling may have introduced bias in the dataset and had some negative effects on the performance of our models. An alternative solution to deal with the class imbalance problem, as future work, would be to impose an additional cost on the model for making classification mistakes on the minority class during training.
Despite these limitations, the key contribution of this study is the demonstration that a mixture of structured and text data can be used to create a promising PTSD diagnosis model based on EMR data of patients. Future work can use encounter dates to explore the progression of PTSD among identified patients and the diagnosis of PTSD at each visit level. Unstructured encounter notes, medications, and risk factors should be explored to refine the details into more specific groups and categories. Future work will continue to expand on the features that our models found to be important.
Conclusions
Unstructured encounter notes from primary care community-based EMRs can strengthen model prediction of PTSD, particularly when using a mixed data model that combines both structured and unstructured data.34,49 We found a serial model that assessed both structured and unstructured data to identify PTSD, had the highest sensitivity, F-measure, and AUC. Many of the features significant in the model have been found to be associated with PTSD42–46 suggesting accuracy in our machine learning models. However, there were features not significant in this community-based population that may be found in a PTSD/psychiatric dataset. Detection of PTSD based on existing primary care EMR data is feasible and can inform primary care quality improvement, research, and disease surveillance. Future studies can extend this approach by applying it to larger cohorts with more positive cases and more unstructured free text. Our findings are also promising foundational work for developing models that can trend disease trajectory for PTSD and other diagnoses.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This article is supported by Advanced Analytics Grant from IBM and Canadian Institute for Military and Veteran Health Research.
