Abstract
We develop and validate a clinical guideline-integrated LLM for enhanced sepsis mortality prediction. Using MIMIC-IV data from 24,237 ICU sepsis patients, we fine-tuned a large language model with Low-Rank Adaptation, embedding clinical guidelines into the training process. The model’s predictive performance was evaluated using accuracy, F1-score, sensitivity, specificity, and area under the receiver operating characteristic curve (AUC). Ablation studies assessed the specific contributions of clinical guideline integration. The guideline-enhanced fine-tuned LLM demonstrated moderately higher performance across all evaluation metrics including predictive accuracy (0.819), F1-score (0.815), sensitivity (0.815), specificity (0.822), and AUC (0.852) in predicting mortality risk for septic patients compared to traditional machine learning (highest accuracy: 0.774, AUC: 0.850) and deep learning methods (highest accuracy: 0.762, AUC: 0.841). Ablation experiments demonstrated that explicit integration of clinical guideline knowledge substantially improved performance over both direct prompting (accuracy: 0.709, AUC: 0.706) and fine-tuning without clinical guidelines (accuracy: 0.786, AUC: 0.801). These findings demonstrate that incorporating clinical guidelines into the fine-tuning of large language models outperforms both traditional and deep learning baselines across multiple metrics in sepsis mortality prediction, highlighting the value of explicit domain knowledge integration for clinical AI’s robustness.
Introduction
Sepsis is an acute organ dysfunction syndrome that results from a dysregulated host response to bacterial, viral, fungal, or parasitic infections. 1 Approximately 48.9 million sepsis cases occur worldwide annually, causing around 11 million deaths and accounting for 19.7% of total global deaths. 2 In the United States alone, more than a third of hospital deaths are attributed to sepsis, with associated healthcare costs reaching approximately $38 billion in 2017, making it one of the most common and costly conditions leading to hospitalisation.3,4 Consequently, sepsis poses a severe threat to public health and imposes a substantial socioeconomic burden on healthcare systems.
To improve the quality of treatment and clinical outcomes of septic patients, the Surviving Sepsis Campaign (SSC) regularly updates clinical guidelines to standardise global clinical practice. 5 Despite significant advances in treatment and management strategies, sepsis mortality remains elevated and overall prognosis is frequently poor. 6 Accurate and effective prediction of epsis mortality risk could help clinicians identify high-risk patients quickly, allowing individualised treatment strategies, early initiation of palliative care discussions, and evaluating healthcare quality and effectiveness of treatment within clinical settings. 7
In recent years, traditional machine learning methods such as logistic regression (LR), random forest (RF), and gradient boosting decision trees (GBDT) have demonstrated efficacy in predicting mortality risk among septic patients in ICU settings.8–11 However, these algorithms exhibit intrinsic limitations in capturing complex interactions within high-dimensional, multimodal clinical datasets due to their linear assumptions and comparatively simple model structures.12,13 Deep learning approaches, including convolutional neural networks (CNN), recurrent neural networks (RNN), and Transformers, have shown promise in handling complex data structures, but typically require extensive annotated datasets and suffer from limited interpretability, hindering clinical implementation14–17
Existing predictive models for sepsis mortality often neglect the systematic integration of domain-specific clinical guidelines into large language models (LLMs) training processes, limiting their clinical applicability and robustness. 18 Recent advances in Transformer-based LLMs have demonstrated strong logical reasoning and contextual understanding capabilities, highlighting their substantial potential for medical diagnostic and prognostic tasks.19–21 However, current research predominantly relies on pretrained LLMs through simple prompting methods, lacking explicit incorporation of clinical expertise and guideline-based knowledge through supervised fine-tuning (SFT).
To address this critical gap, our study innovatively integrates explicit clinical guideline knowledge into supervised fine-tuning of the Qwen2.5-72B LLM using low-rank adaptation (LoRA) technology.22,23 We systematically evaluate our proposed guideline-enhanced LLM against traditional machine learning algorithms, classical deep learning models, and direct prompting methods, aiming to significantly improve both predictive accuracy and interpretability for sepsis mortality risk. Ultimately, this research seeks to provide ICU clinicians with a robust and reliable decision-support tool to enhance individualised patient care and clinical outcomes.
Materials and method
Data source and variable extraction
Data for this study were obtained from the Medical Information Mart for Intensive Care-IV database (MIMIC-IV, version 2.2), 24 provided by Beth Israel Deaconess Medical Center (BIDMC), Boston, USA. This publicly accessible database contains clinical data from over 500,000 patients admitted between 2008 and 2019, including detailed information on vital signs, nursing documentation, severity of disease scores, diagnoses, treatments, and laboratory results. Dr Ruiyi Zhu from our research team completed the required database training and was authorized to access the data (record ID: 59980404).
Patients met Sepsis-3 diagnostic criteria, defined as suspected infection with Sequential Organ Failure Assessment (SOFA) score of ≥2 points or septic shock, characterised by persistent hypotension requiring vasopressors to maintain mean arterial pressure (MAP) ≥65mmHg and serum lactate levels >2 mmol/L (18 mg/dL) despite adequate fluid resuscitation. Exclusion criteria were: (1) age under 18 years; (2) ICU stay under 24 h or exceeding 100 days; (3) multiple ICU admissions. The patient selection process is detailed in Figure 1: after screening 32,970 candidate admissions, 24,237 unique adult sepsis cases were retained and stratified 8:1:1 into training (19,389), validation (2,424) and test (2,424) cohorts. Flowchart of patients’ selection.
Data extraction was performed using PostgreSQL (Version 14.0; PostgreSQL Global Development Group). The extracted variables included demographic information, vital signs, laboratory measurements, and clinical interventions recorded within 24 h of ICU admission (Appendix A). The primary outcome was all-cause in-hospital mortality occurring after the first 24 h of ICU admission. Existing severity scoring systems, including the Simplified Acute Physiology Score-II (SAPS-II), Acute Physiology Score-III (APS-III), and Sequential Organ Failure Assessment (SOFA) scores, were utilized as comparative benchmarks for evaluating the predictive performance of the developed model.
Construction of guideline-enhanced fine-tuning datasets
We utilised the 2021 adult Surviving Sepsis Campaign guideline as the sole authoritative source, providing evidence-based standards for sepsis management. The computational pipeline for converting unstructured guidelines into executable knowledge representations comprised four sequential stages. Initially, guideline sections were segmented into individual recommendations. Subsequently, entities and relations were extracted using an instruction-tuned Qwen 2.5-72B model, identifying five core entity types (Indicator, Threshold, Action, TimeFrame, Outcome) and seven relation categories (including has_threshold and recommends_action). This was followed by an expert fusion phase where two board-certified intensivists reviewed and consolidated the outputs into a sepsis-specific knowledge base; representative outcomes included structured relationships such as (Lactate, has_threshold, ≥4 mmol L-1) and (MAP <65 mmHg, recommends_action, “vasopressor initiation”). Finally, the refined knowledge was serialized into a JSON-formatted knowledge graph (see Appendix B for complete explanation).
To explicitly evaluate the impact of integrating clinical guidelines into the fine-tuning of LLMs, we constructed two distinct datasets. The first dataset was purely data-driven, containing only raw clinical variables without explanatory context. The second dataset was enriched with explicit guideline-based interpretations for each clinical indicator, facilitating the model’s deeper understanding of clinical correlations and enhancing its interpretability and predictive capability. The data-driven dataset contains exclusively raw clinical variables including vital signs, laboratory values, and demographic features without any explanatory context or clinical interpretations. In contrast, the guideline-enhanced database enriched identical clinical variables through systematic injection of structured medical knowledge. Each data point was mapped to corresponding entities from the sepsis guideline knowledge graph (e.g., associating MAP <65mmHg thresholds with “vasopressor initiation” actions). The specific approaches to dataset construction are described in Figure 2. By constructing these two distinct datasets, we systematically assessed the value of explicitly integrating clinical guidelines into LLM fine-tuning, highlighting its potential to significantly enhance predictive accuracy and interpretability in clinical decision-making. For each patient, the worst 24-h value of every indicator is matched against SSC thresholds stored in knowledge graph. A guideline note—for example ‘Lactate 5.6 mmol L-1 — exceeds SSC threshold (≥4 mmol L-1), high risk’—is added only when the patient’s value crosses the relevant boundary; otherwise the phrase ‘within guideline range’ is used. These conditional comments are appended to the raw features to form the guideline-enhanced prompt. Construction of guideline-enhanced fine-tuning datasets.
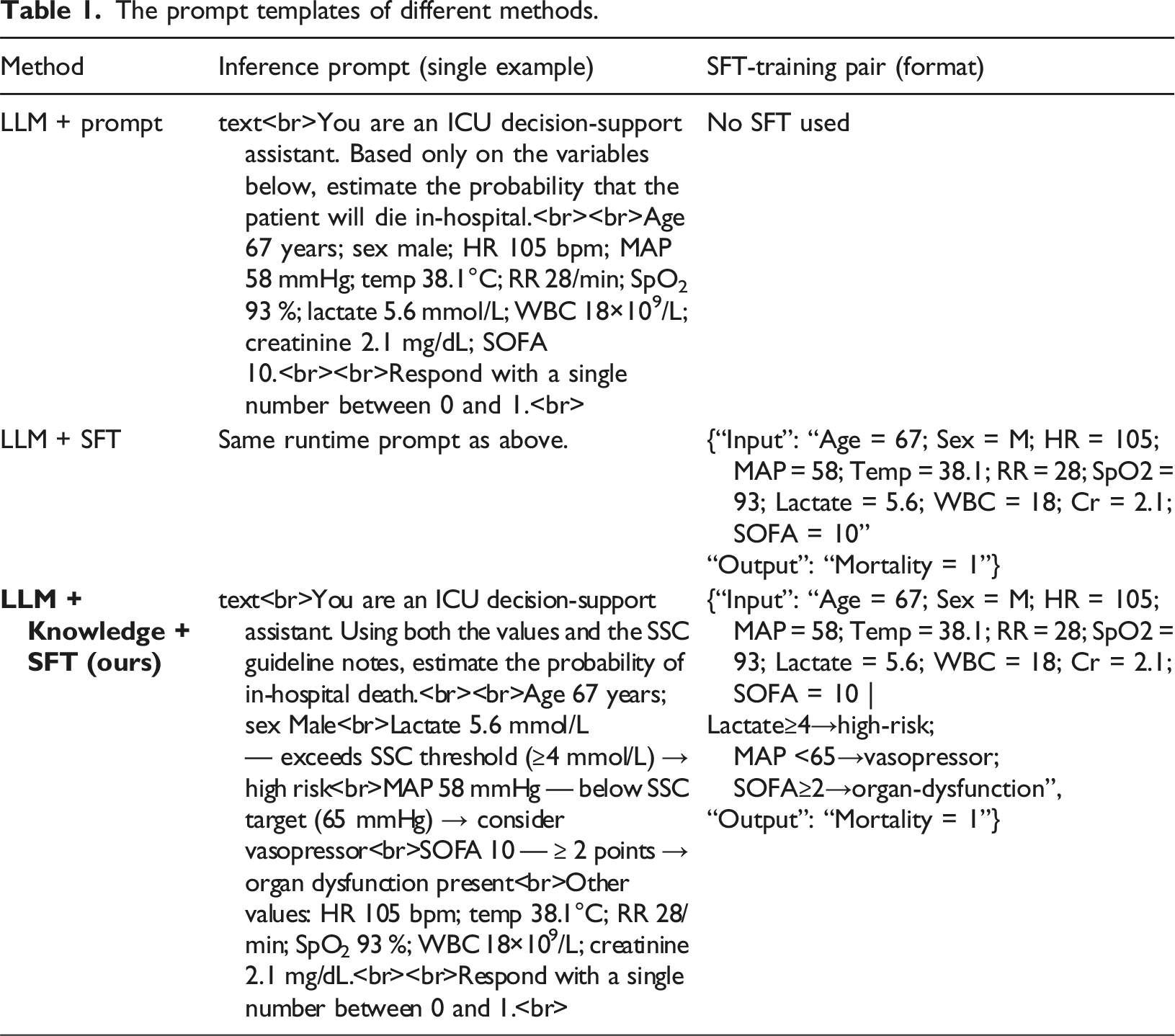
The prompt templates of different methods.

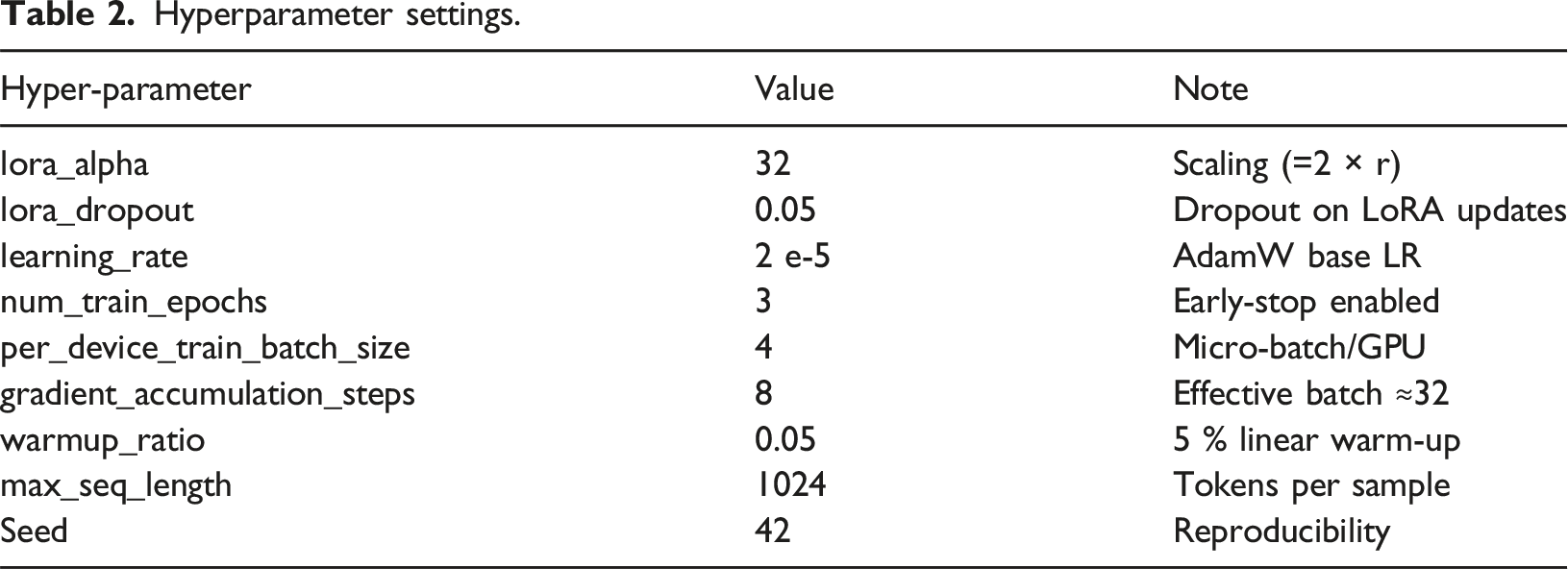
Hyperparameter settings.
Model training
In this study, we used a pre-trained LLM (Qwen2.5-72B) to predict the risk of mortality for sepsis patients. We implemented two distinct model-inference approaches to systematically evaluate the effectiveness of fine-tuning with clinical domain knowledge. First, we applied a direct prompt-based strategy, utilizing clinical indicators from the patient as input without modifying the pre-trained parameters. Second, we employed supervised fine-tuning (SFT) enhanced by Low-Rank Adaptation (LoRA) to efficiently optimize the selected layers of the pre-trained Qwen2.5-72B model. LoRA updates it according to:
Fine-tuning data sets were constructed from the MIMIC-IV database in two forms: a purely feature-based data set and a guideline-enhanced dataset explicitly integrating clinical guideline interpretations. Model optimization was performed using the AdamW optimizer (learning rate 2 × 10−5) with a linear scheduler and accumulation of gradients. Early stopping was implemented based on validation loss to prevent overfitting. The final optimized LoRA parameters were subsequently merged with the original model weights, resulting in a highly efficient and clinically interpretable predictive model. The entire model training framework is illustrated in Figure 3, outlining the end-to-end workflow comprising two distinct approaches: a purely data-driven version and a knowledge-enhanced version enriched with guideline annotations. The model-training phase proceeds through five sequential steps: loading the pre-trained LLM while configuring LoRA adapters; preparing and preprocessing the input data; selecting training strategies while optimizing hyper-parameters; defining the loss function and evaluation metrics; and finally training and validating the model. After validation, the LoRA-optimised parameters are merged with the base weights to produce a compact, clinically interpretable predictor. The framework of guideline-enhanced fine-tuning of LLM.
Baselines
To comprehensively evaluate the performance of our proposed LLM-driven method, we compared it with three categories of baseline methods.
Traditional Machine Learning Methods: These included Support Vector Machine (SVM), Naive Bayes, K-Nearest Neighbor (KNN), Logistic Regression (LR), Gradient Boosting Decision Tree (GBDT), Decision Tree (DT), and Random Forest (RF). These methods classify data based on linear separability (SVM), posterior probabilities (Naive Bayes), proximity of data points (KNN), logistic transformations (LR), ensemble strategies to sequentially correct prediction errors (GBDT), entropy-based partitioning (DT), and ensemble averaging to minimize overfitting (RF).
Deep Learning Methods: Deep learning approaches included Long Short-Term Memory (LSTM) networks, which capture temporal dependencies, Convolutional Neural Networks (CNNs) designed to identify local feature patterns, and Transformer models leveraging self-attention mechanisms for modeling complex, long-range dependencies in sequential data.
Large Language Model without Fine-Tuning (Prompt-based): As an additional baseline, we employed the pretrained LLM (Qwen2.5-72B) without fine-tuning, making predictions based solely on crafted input prompts.
Statistical analysis
Normally distributed measurement data are typically expressed as mean ± standard deviation (X ± S) and compared between groups using the independent samples t-test. Data that do not follow a normal distribution are represented as median (P25, P75) and compared using the Mann-Whitney U test. Categorical characteristics are expressed as frequencies (percentages) and compared via the χ2 test. A two-sided p-value of less than 0.05 is considered statistically significant. Statistical analyzes were performed using SPSS software (Version 29.0).
To quantitatively evaluate the performance of the model in predicting mortality risk in patients with sepsis, we used the following metrics 1 : Area Under the Receiver Operating Characteristic Curve (AUROC), indicating overall predictive capability 2 ; Sensitivity (Recall), the proportion of correctly identified death cases 3 ; Specificity, the proportion of correctly identified survival cases 4 ; Accuracy, reflecting overall correct predictions; and 5 F1-score, the harmonic mean of precision and recall, particularly suitable for imbalanced classification tasks.
All model training and evaluations were performed on the following hardware platform. GPUs using NVIDIA 8*A100 (80 GB memory), CPUs using Intel Xeon Gold 6248 (64 cores, 2.5 GHz frequency), 512 GB RAM, and a 4 TB NVMe SSD storage device. The software environment comprised Python 3.10, PyTorch 2.1, and CUDA 12.6 for GPU acceleration. Mini-batches use stratified 1:5 death–survival sampling.
Results
Baseline characteristics of patients
Baseline features of 24,237 sepsis patients categorized by in-hospital mortality.

SOFA sequential organ failure assessment, SAPS II simplified acute physiology score II, GCS Glasgow coma scale, CHD coronary heart disease, CKD chronic kidney disease.
Comparative evaluation of model performance
To systematically evaluate the performance of the proposed guideline-enhanced fine-tuned LLM (Qwen2.5-72B + LoRA) for predicting mortality risk in patients with sepsis, we conducted comprehensive comparative experiments. Traditional machine learning algorithms including Support Vector Machine (SVM), Naive Bayes (Bayes), K-Nearest Neighbor (KNN), Logistic Regression (LR), Gradient Boosting Decision Tree (GBDT), Decision Tree (DT), and Random Forest (RF) were selected as baseline models. Furthermore, classic deep learning models such as Long Short-Term Memory (LSTM), Convolutional Neural Network (CNN), and Transformer were included to enrich the comparative analysis. The proposed method is denoted as LLM+Knowledge+SFT. All baseline hyper-parameters were optimised by 8-fold cross-validated grid/random search. To ensure internal robustness, the 80 % training set was further subjected to stratified eight-fold cross-validation. Model hyper-parameters were tuned on the fold-specific development subset, and the optimised model was retrained on the entire 80 % before final evaluation on the independent 10 % test cohort.
Experimental results comparing various models on the sepsis mortality prediction task.

SVM, Support Vector Machine; KNN, K-Nearest Neighbor; LR, Logistic Regression; GBDT, Gradient Boosting Decision Tree; DT, Decision Tree; RF, Random Forest; LSTM, Long Short-Term Memory; LLM, Large Language Model.
Superior performance across multiple metrics compared with traditional machine learning methods (e.g., SVM, Bayes, KNN, LR, GBDT, DT, and RF) and classic deep learning approaches (LSTM, CNN, Transformer), achieving the highest Accuracy (0.819), F1 Score (0.815), and Area Under the ROC Curve (AUC = 0.852). Among traditional machine learning algorithms, ensemble methods such as Gradient Boosting Decision Tree (GBDT) and Random Forest (RF) exhibited relatively strong predictive capability, whereas deep learning models like Long Short-Term Memory (LSTM) and Transformer models showed moderate performance, likely limited by their dependency on larger annotated datasets and the inherent complexity of clinical data features.
While the absolute AUC improvement over the strongest baseline (GBDT) is modest (+0.002), the consistent gains across all evaluation metrics, particularly specificity (+8.2%) and accuracy (+5.8%), highlight the clinical utility of guideline-enhanced fine-tuning.
Furthermore, the ROC curves presented in Figure 4 clearly illustrate the superior predictive performance of the guideline-enhanced fine-tuned LLM compared with other baseline methods, highlighting its improved stability and robustness. Overall, our novel integration of clinical guidelines into supervised fine-tuning significantly enhances the accuracy, interpretability, and clinical applicability of the LLM-driven mortality risk prediction, demonstrating considerable potential for informing clinical decision-making and patient management in ICU settings. ROC curves presenting the performance of machine learning methods in the MIMIC cohort (N = 24237). ROC,receiver operating characteristics; SVM, Support Vector Machine; KNN, K-Nearest Neighbor; LR, Logistic Regression; GBDT, Gradient Boosting Decision Tree; DT, Decision Tree; RF, Random Forest; LSTM, Long Short-Term Memory; LLM, Large Language Model.
Ablation study
To deeply investigate the specific contributions of model fine-tuning and domain knowledge enhancement, we designed detailed ablation experiments, comparing three setups: LLM + Prompt (direct prompting without fine-tuning), LLM + SFT (fine-tuning without domain knowledge), and LLM+Knowledge+SFT (complete method).
Ablation study comparing different training strategies.

Compared with the GBDT baseline, the LLM + Knowledge + SFT model correctly re-classified 22 additional deaths and 252 additional survivors in the 4,848-case test set. A paired AUROC test yielded p = 0.02, indicating that the performance difference between the two models is statistically significant at the 95 % confidence level.
Discussion
Sepsis remains a leading cause of in-hospital mortality particularly in immunocompromised populations, 25 necessitating accurate and timely risk prediction for early intervention. Traditional methods for predicting sepsis mortality have primarily relied on structured clinical data and statistical models, such as logistic regression and the Sequential Organ Failure Assessment (SOFA) score. 26 Although these models based on structured clinical data have provided useful insights into sepsis mortality risk, their predictive performance remains limited due to inadequate incorporation of comprehensive multidimensional and multi-omics information, underscoring the need for predictive tools in infection management. 27 Recently, machine learning (ML) techniques have shown superior performance in mortality prediction by leveraging high-dimensional data, enabling capture of intricate patterns overlooked by traditional models.
Multiple investigations have leveraged ML models to forecast mortality from sepsis, utilizing structured data from extensive clinical databases such as MIMIC-III, MIMIC-IV, and eICU Collaborative Research Database. 28 Techniques including Random Forest (RF), 29 XGBoost, 30 and LightGBM have consistently outperformed traditional regression-based approaches. For instance, researchers 31 developed an RF model in the MIMIC-IV database, achieving significantly superior results compared to conventional SOFA-based models. Notably, prior studies utilizing traditional ML approaches have achieved comparable or marginally superior discriminative ability. Some reaseachers 30 reported an AUC of 0.857 using XGBoost on MIMIC-III data, while some one 32 attained an AUC of 0.888 with gradient-boosted methods in a large administrative cohort. However, Our guideline-enhanced LLM demonstrates robust sepsis mortality prediction (AUC: 0.852). Though these slight differences in absolute performance metrics warrant acknowledgment, our approach confers distinct advantages that extend beyond discriminative power alone. The distinct advantage lies in generating clinically interpretable guideline-anchored rationales that enhance trust and actionability beyond “black-box” predictions. While computational demands and single-center validation require targeted improvement, this approach prioritizes evidence-based clinical utility over marginal metric gains, bridging a critical gap in AI-assisted critical care decision-making. Deep learning approaches, such as Long Short-Term Memory (LSTM) networks 33 and Transformer-based architectures, 34 have also been explored. These methods effectively capture temporal dependencies within the ICU time series data. Although deep learning methods theoretically possess superior feature extraction abilities, their performances in our study did not surpass that of traditional methods. This outcome may arise from the complexity and heterogeneity of clinical data, where deep learning approaches require substantial amounts of annotated data to achieve effective generalization. Given the relatively limited dataset used in our study, the full potential of deep learning models might have been restricted. However, their clinical applicability is impeded by poor interpretability. 35 To mitigate this, some studies have incorporated Shapley Additive Explanations (SHAP) to enhance model transparency and perform more detailed feature importance analyses. 32
However, our guideline-enhanced LLM synchronously generates evidence-grounded rationales during clinical predictions through real-time knowledge graph queries to the guideline ontology. This self-contained interpretability mechanism produces clinically actionable narratives that directly support clinical decision-making, rendering supplementary validation via SHAP or attention weight analyses superfluous.
A recent advancement in sepsis mortality prediction involves the integration of large language models (LLMs) with clinical datasets. Unlike traditional machine learning (ML) approaches, LLMs can effectively process unstructured clinical notes, capturing valuable qualitative insights about patient conditions. Studies have demonstrated that combining structured clinical data with summaries derived from LLMs significantly enhances predictive accuracy. 36 For instance, integrating ChatGPT-generated clinical summaries with ICU data notably improved the area under the receiver operating characteristic curve (AUC), underscoring the benefits of multi-representational learning. 37 In contrast, we embed the 2021 Surviving Sepsis Campaign as a machine-readable knowledge graph and inject its rule-based annotations into the prompt during LoRA fine-tuning. Notably, our approach fundamentally differs from prior LLM-based frameworks like the Sepsis Early Risk Assessment (SERA)algorithm, which utilize natural language processing (NLP) to extract clinically relevant information from unstructured notes, further demonstrating the potential of leveraging LLMs alongside domain-specific knowledge. While SERA demonstrates the value of unstructured data, our model explicitly embeds guideline-based diagnostic criteria and therapeutic pathways through real-time knowledge graph traversal, ensuring alignment with evidence-based protocols38,39 Given the promising performance of guideline-enhanced LLM,it could be leveraged within clinical workflows to facilitate early triage decisions, prioritizing high-risk septic patients for intensive care unit admission and aggressive resuscitation, or prompting earlier reassessment and treatment escalation for deteriorating patients in general wards. However, successful integration necessitates overcoming barriers such as seamless interoperability with diverse Electronic Health Record (EHR) systems and fostering clinician trust through robust validation and the provision of interpretable, explainable AI techniques.
However, most current LLM-based studies rely heavily on simple prompting strategies and have not systematically integrated domain-specific clinical guidelines into their training process, potentially limiting their predictive capabilities and clinical applicability.40–42 To address this gap, our study proposes an innovative approach that explicitly integrates clinical guideline knowledge with supervised fine-tuning (SFT) of an LLM (Qwen2.5-72B) using low-rank adaptation (LoRA) techniques. Our results show that this guideline-enhanced fine-tuned LLM (LLM+Knowledge+SFT) significantly outperforms traditional machine learning models such as SVM, GBDT, and RF, as well as classical deep learning approaches including LSTM and Transformer, in terms of accuracy, F1 score, and AUC. Ablation studies further confirmed that supervised fine-tuning notably improved predictive performance (accuracy increased from 0.709 to 0.786), and explicit incorporation of guideline-based knowledge provided additional significant performance gains (accuracy improved to 0.819, AUC to 0.852). These findings clearly highlight the critical value of embedding expert clinical guideline knowledge into LLM fine-tuning processes to enhance predictive robustness and interpretability, representing a significant advancement in clinical decision support.
Despite these promising results, our study has limitations. First, the model was developed and validated using data from a single-center database (MIMIC-IV), potentially limiting the generalizability of findings. This single-center database, derived from an urban tertiary hospital, was characterized by a predominantly Caucasian population (median age ≈65 years) with a mean SOFA score of 5. This limits generalizability to other demographic groups and healthcare settings. Second, The substantial computational burden associated with fine-tuning the Qwen2.5-72B architecture presents significant deployment barriers in clinical settings. Even with parameter-efficient LoRA techniques, this process requires approximately 9 GPU-hours across eight A100-80 GB cards. Such resource-intensive procedures may prove impractical in resource-constrained clinical settings. 43 More critically, the requisite hardware configurations including multiple high-end GPUs and supporting cooling systems entail substantial initial capital investment that could impose prohibitive financial burdens on healthcare institutions, creating particularly acute challenges for intensive care units in community hospitals or developing regions. 44 Third, guideline dependency introduces potential biases. Our knowledge graph exclusively encodes the 2021 adult Surviving Sepsis Campaign guidelines, which predominantly reflect Western medical practices. This framework currently lacks adaptation for paediatric or obstetric populations, excludes considerations for traditional medicine protocols in low-resource settings, and may not accommodate rapid evidence updates, potentially diminishing performance in these contexts. Fourth, real-world integration barriers remain unaddressed. The model was evaluated on retrospectively batched data without direct interfaces to bedside monitoring systems. Future implementations must integrate real-time ICU data streaming to process live vitals and lab results, while rigorously evaluating latency, alert fatigue, and adoption thresholds to ensure timely identification of high-risk patients.
In conclusion, this study demonstrates that incorporating clinical guidelines into the fine-tuning process of large language models significantly enhances the prediction accuracy, stability, and interpretability of sepsis mortality risk assessment. Building on these results, we have begun coordinating multicentre experiments with three hospitals in our regional medical consortium to evaluate the model’s performance across different case-mixes and practice patterns. In parallel, we are collaborating with the ICU information-system vendor to create a lightweight middleware that streams real-time vital signs and laboratory data to the model and feeds the resulting risk score plus guideline-based rationale back to the bedside dashboard. These efforts will enable prospective validation and seamless deployment in routine critical-care workflows.
Conclusion
Our guideline-enhanced LLM outperformed clinical scoring systems and computational baselines across key metrics, validating that explicit domain knowledge integration is essential for clinically viable AI. This approach enhances model robustness and generalisability, providing a replicable framework for reliable mortality prediction in critical care. The methodology enables trustworthy clinical decision-support for early risk stratification and personalised interventions.
Footnotes
Author note
Anonymized information revealing author’s identity in the main manuscript: Dr Ruiyi Zhu from our research team completed the required database training and was authorized to access the database of MIMIC (record ID: 59980404).
Acknowledgement
The authors would like to acknowledge the support provided by various individuals and organizations during the course of this research. We thank Dr Miaorong Xie for his valuable insights and technical assistance. We also extend our gratitude to the National Natural Science Foundation of China for their financial support through the project (No:82374069).
Ethics approval
The Ethics Committee of Beijing Friendship Hospital Affiliated to Capital Medical University waived the need for ethics approval and patient consent for the collection, analysis and publication of the retrospectively obtained and anonymised data for this non-interventional study.
Authors Contribution
All authors contributed to the conceptualization and design of the study. Zhen Zhao and Bo An are co-first authors. They conceived the study, designed the experimental protocols, conducted the literature review, undertook the data analysis, drafted the initial manuscript and contributed to the subsequent drafts. Tianpeng Zhang performed statistical analyses, validated the data, and created visualizations. Ruiyi Zhu and Zihao Fan interpreted the data, summarized the patient of MIMIC features. Guoxing Wang provided conceptual guidance, revised the manuscript, approved the final version, secured funding and oversaw the research integrity as corresponding author. All authors read and approved the final version.
Funding
This research was supported by the National Natural Science Foundation of China (grant number 82374069).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.

Catagory
Content
Demographic Information
Gender, Age, Body Mass Index (BMI)
Vital Signs (first 24 hours)
Heart Rate, Mean Arterial Pressure (MAP), Respiratory Rate, Peripheral Oxygen Saturation (SpO2), Arterial Oxygen Partial Pressure to Fractional Inspired Oxygen Ratio (PaO2/FiO2 Ratio), Central Venous Pressure (CVP)
Laboratory Results (first 24 hours)
White Blood Cell Count (WBC), Red Blood Cell Count (RBC), Hemoglobin, Hematocrit (HCT), Platelet Count (PLT), Prothrombin Time (PT), Activated Partial Thromboplastin Time (PTT), Lactate Levels, Arterial Blood pH, Arterial Oxygen Partial Pressure (PaO 2), Alanine Aminotransferase (ALT), Aspartate Aminotransferase (AST), Albumin, Total Bilirubin, Blood Urea Nitrogen (BUN), Creatinine, Total Calcium, Chloride, Potassium, Sodium, Urine Output, Fluid Balance
Treatments (first 24 hours)
Use of Vasopressors, Mechanical Ventilation, Renal Replacement Therapy (RRT)
Baseline Comorbidities
Hypertension, Diabetes Mellitus, Acute Myocardial Infarction (AMI), Congestive Heart Failure, Chronic Kidney Disease (CKD)
