Abstract
Introduction
Within the healthcare industry, robust and generalisable real-world evidence (RWE) is increasingly required by decision-makers, such as regulators, payers and Health Technology Assessment (HTA) bodies.1,2 These studies often require the use of multiple databases to answer research questions that require large sample sizes and diverse study populations. 3 There are several networks that have been designed as platforms to carry out such multi-database studies. Examples include the Food and Drug Administration (FDA) Sentinel Initiative, the Patient-Centered Outcomes Research Network (PCORnet), the European Union Adverse Drugs Reactions (EU-ADR), Vaccine Safety Datalink and the Data Analysis and Real-World Interrogation Network (DARWIN EU).4–10
To homogenise analysis across databases, Common Data Models (CDMs) have been developed to standardize table structure, variable names and definition of key concepts. 11 The Observational Medical Outcomes Partnership (OMOP) CDM standardizes the structure and content of observational data to enable efficient analyses that can produce reliable evidence. 12 What makes this standardisation method different to other approaches (e.g. Sentinel/PCORnet) is the use of common vocabularies to which different coding systems, within the source databases, are mapped (e.g. for conditions, ICD-9, ICD-10 or Read codes are mapped to the common SNOMED-CT coding format). 13 Assuming that codes from different vocabularies, that represent common diseases, products, etc. are mapped to a common, standard vocabulary, then this would allow building one set of cohort definitions and analysis scripts that could be applied unchanged to every participating database.
Accurate transformation of source databases into CDM is essential for maintaining data integrity. Most peer-reviewed published studies assessing data transformation from the source data to a CDM have been conducted using Claims, Electronic Health Records (EHR) or Registry data in the US and the results have largely demonstrated a good mapping between the source and transformed data.14–18 However, relatively few studies have assessed European data sources.19–22 A study assessing the transformation of UK The Health Improvement Network data (THIN) to the OMOP CDM found that information loss occurred due to incomplete mapping of medical and drug codes as well as limitations in the data structure of the OMOP CDM. 19 In contrast, a study in UK Clinical Practice Research Datalink (CPRD) General Practitioner Online Database (GOLD) deemed all elements of the OMOP CDM transformation to be of high quality (99.9% of database condition records and 89.7% of database drug records were mapped and most unmapped drugs were devices or over-the-counter products). 21 Given these contrasting findings and the lack of formal evaluation of CPRD Aurum OMOP transformation, there is value in conducting an additional assessment of CPRD using a recent version of OMOP CDM. The aim of this study was to compare the source medical codes included in the source/non-transformed CPRD GOLD and CPRD Aurum databases to source medical codes within the OMOP-transformed data.
Methods
This is a methodological validation study designed to assess the transformation of CPRD GOLD and CPRD Aurum EHR databases into the OMOP CDM. The objective was to evaluate whether key clinical concepts (such as diagnoses, laboratory results, prescriptions and vaccinations) retained consistency in event counts after OMOP transformation. The study does not involve hypothesis testing, clinical outcome assessment, or inference about treatment effects.
Data source
The CPRD primary care electronic records have been used for research for over 30 years. CPRD databases are among the most thoroughly described and validated primary care databases in the world with over 3,500 peer-reviewed publications (https://www.cprd.com/bibliography). The UK National Health Service (NHS) provides universal health coverage. From the CPRD databases patients are excluded if they choose to opt out or if their General Practitioner (GP) practice does not give its consent to be included. 23 The study was conducted using CPRD GOLD and CPRD Aurum databases. The two databases include GP practices that use different patient management software (Vision and EMIS for CPRD GOLD and CPRD Aurum, respectively). 24 The CPRD GOLD database was established in 1987 and includes over 21 million historical patients in England, Wales, Scotland and Northern Ireland. The geographical distribution of the ∼3 million active patients has changed resulting in very few practices in England. 25 Within the Vision system, diagnoses and other non-prescription data are recorded using the Read coding system and prescriptions are coded using Gemscript. The CPRD Aurum was launched in 2017 and includes ∼45 million patients from 1987 onwards of which >15 million are active patients. 26 It includes GP practices in England and Northern Ireland. The EMIS system uses a combination of SNOMED CT (UK edition), Read Version 2 and local EMIS Web® software-specific codes that have been cross-mapped to a single code dictionary by National Health Service Digital for diagnoses and other non-prescription data. Prescriptions are coded using the Dictionary of Medicines and Devices (dm + d) codes which are a subset of the SNOMED CT terminology. Some of the characteristics of the GP systems and the CPRD databases have been described in the past.24,27–29
Observational Medical Outcomes Partnership (OMOP) Common Data Model (CDM)
The OMOP CDM is managed by the Observational Health Data Sciences and Informatics (OHDSI) community. As of October 2024, 544 data sources across 54 countries have been converted into OMOP CDM.12,30 Within the OMOP CDM, clinical events are expressed as concepts which represents the semantic notion of each event. The concepts cover any event related to patient experience (e.g. conditions, procedures, drug exposures etc) as well as administrative information (e.g. visits, care sites etc). Each standard concept has a concept id (concept_id) and is assigned to a domain (e.g. “Condition”, “Drug”, “Procedure”, “Visit”, “Device”, “Specimen” etc) which direct to which CDM table and field a clinical event or event attribute is recorded. 31 Records from the source database are mapped from the original table to the domain table in the OMOP version of the database in which the standard vocabulary belongs. This mapping is described in the Vocabulary tables that are an integral part of the OMOP CDM. A detailed online browser of the OMOP Vocabularies is also available.32,33
The transformation of the CPRD databases into OMOP CPD was performed using an Extract, Transform, Load (ETL) process, which describes how the data can be systematically converted into the standardised OMOP structure. For this study, the CPRD databases were transformed into version 5.3.1 of the OMOP CDM by Odysseus Data Services. The databases containing data up to 31st December 2019 were used.
The COde list DEvelopment and eXploration (CODEX) approach for generating medical code lists
A novel methodology is introduced here, termed CODEX, designed to systematically generate comprehensive medical code lists. This approach was applied to produce code lists for 12 diseases, 9 medications, 10 laboratory measures, 2 lifestyle measures, 3 vaccinations and 3 procedures. For specific measures such as Body Mass index (BMI), smoking status and laboratory values the presence of recorded values (and not the actual measurement) was used. The CODEX approach can be summarised in the following three steps. • Step 1: Relevant medical terms were searched for (using combinations of “OR” and “AND” logical operators) within the database dictionaries, producing a “broad code list”. Only medical codes that appeared in the database at least once were included. • Step 2: Each medical code description within the “broad code list” was reviewed by a researcher line-by-line to determine inclusion in the final list based on clinical relevance. • Step 3: A second researcher independently repeated Step 2 to ensure consistency. Any discrepancies were discussed before a final decision on the classification was made.
A complete record of the audit trail was maintained to make the process fully transparent and reproducible. This process was carried out separately to generate medical code lists for CPRD GOLD and CPRD Aurum. A comprehensive description of the CODEX process will be provided in a forthcoming publication.
Medcodes selected for peripheral arterial disease.

Analytical methods
Prevalent and incident event counts for the selected diseases, medications, laboratory records, lifestyle measures, vaccinations and procedure/tests were compared between the source and OMOP versions of CPRD databases. For example, for the “diagnoses” domain we compared counts of incident acute myocardial infarctions (identified via Read code sets). For “vaccinations” we evaluated influenza and Measles, Mumps, Rubella (MMR) vaccine administrations. The year 2019 was selected as the reference year to calculate incident and prevalent cases. To identify incident events in 2019, we screened all records prior to 1st January 2019, to confirm that no earlier occurrence of each event (diagnosis, medication, etc.) existed. For prevalence on 1st January 2019, we reviewed every record before that date to determine whether the event had already occurred and should therefore be counted as prevalent. For each of the 38 clinical concepts, the number of prevalent and incident events was calculated in the source CPRD and in the OMOP-formatted data. The absolute difference in counts was then derived between the two datasets and expressed as a percentage of source CPRD count. The formula for the difference can be written as:
This procedure was performed separately for prevalent and incident measures.
Datasets/domains were used to search codes by database.

aSame domains used for CPRD GOLD and CPRD Aurum.
The data analysis for this paper was generated using SAS software, Version 9.4 of the SAS System for LIN X64. Copyright © SAS Institute Inc. SAS and all other SAS Institute Inc. product or service names are registered trademarks or trademarks of SAS Institute Inc., Cary, NC, USA.
Results
The incident and prevalent event counts before and after OMOP transformation for OMOP CPRD GOLD and CPRD Aurum databases.

The most prominent percentage difference in incident event counts between source and OMOP CPRD GOLD was observed for PAD (3.0%) and Bacille Calmette-Guérin (BCG) and Pneumococcal vaccines (1.6% and 1.0%, respectively). For CPRD Aurum, the greatest incident event count difference was for Albumin blood test result (1.6%) (Table 3). The largest prevalent event count differences between source and OMOP CPRD GOLD were for MMR vaccine (17.5%), PAD (2.9%) and BCG vaccine (1.9%). For CPRD Aurum, the largest prevalent event count differences were observed for the Albumin blood test result (1.2%) (Table 3).
The mapping of events from the clinical tables in source CPRD GOLD database to their respective domains in the OMOP-transformed version is shown in Figure 1. For CPRD GOLD, events from the source “Clinical” table were split between “Observation”, “Condition Occurrence” and “Procedure Occurrence” tables following the OMOP transformation. Events from the “Additional” table were split between “Observation” and “Measurement” domains. Events from the tables “Test” and “Therapy” were mainly moved into tables “Measurement” and “Drug Exposure”, respectively. Events from the “Immunisation” table were also moved into the “Drug Exposure” domain. Records (rows of data) for CPRD GOLD before and after OMOP transformation. Note: This figure could not be replicated for CPRD Aurum as there was no direct access to the full database.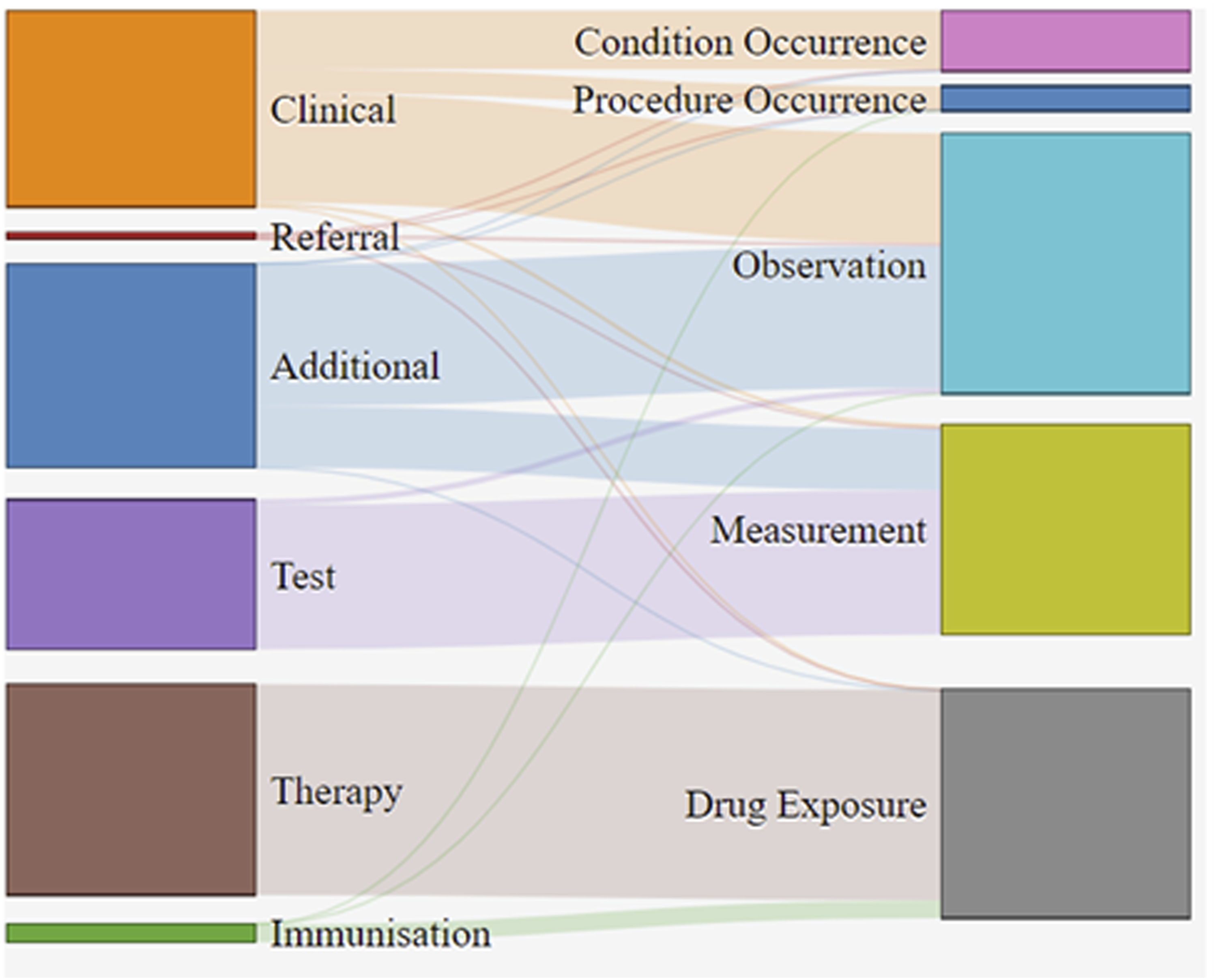
Discussion
In this study, CPRD GOLD and CPRD Aurum databases were transformed (separately) into the OMOP CDM (version 5.3.1). A total of 38 diseases, medications, laboratory records, lifestyle measures, vaccinations and procedures/tests were used for comparing the source and OMOP-transformed CPRD databases. The results showed that for CPRD GOLD database, 89.5% of events had no or very small differences in prevalent and incident event counts between the source and OMOP versions. CPRD Aurum showed even better alignment, with 97.4% of events showing no or very small differences in counts between the source and OMOP data. When looking at individual domains, diagnoses and prescriptions showed nearly perfect alignment (differences ≤ 0.1%), whereas vaccinations had the largest mapping discrepancies. Laboratory measurements exhibited marginally higher variability compared with other domains. These differences were due to how the corresponding codes were mapped to tables during the OMOP transformation. The results suggest that vaccine codes (and to a lesser extent laboratory codes) require extra attention when building OMOP cohorts. Moreover, to mitigate information loss and ensure no medical codes are missed, it is important that researchers search for relevant codes across all OMOP tables and domains, rather than relying on a single domain search.
The code lists were generated for the purpose of validating the OMOP transformations for this paper. They were developed using a novel methodology that systematically searches within the description associated with each medical code. Using this approach, it is crucial to include all potentially valid medical terms associated with the disease, medication, etc. Missing a relevant search term could lead to incomplete code list and hence, patient cohort. The advantage with this approach is that it is fully transparent and reproducible. It requires suitable search terms and criteria for deciding whether to include each code within the code list. The limitation of this approach is that it is resource intensive as it requires manual review of large numbers of codes. However, the latter issue can be mitigated by applying exclusion criteria.
OMOP CDM standardizes different structures across disparate data sources into common tables with a single structure, field datatypes and conventions. This re-formatting of the data is designed to avoid information loss. However, the standardisation process may in some cases inadvertently result in data loss. For example, past OMOP transformations excluded patient information prior to the dates GP surgeries started to provide information according to a set of data quality standards that was introduced at the time. Although it is likely that this information may not be completed as consistently or as fully, it is still believed to be accurate and valuable and therefore, should be used.
The 38 cohorts were chosen to cover a broad range of clinical events. We believe these events provide a sufficient test for validating the measures derived using the source against the OMOP-transformed CPRD versions. However, these may not be representative of all possible use cases in CPRD data, particularly for conditions captured infrequently (rare events).
Converting CPRD to the OMOP CDM can be challenging because of the unusual data structure some of the source tables have. Our evaluation relied on the quality and transparency of the ETL process provided by the vendor. Moreover, vocabularies such as Read codes are very comprehensive and granular which means that some source code types may not be mapped to the OMOP Vocabularies. 34 However, all Read codes were retained (whether mapped to a concept id or not) and placed in the Observation table.
Although our analysis focused on UK primary-care data, the same principles can be applied to any databases transformed to OMOP CDM. These can include specialty medicine, hospital or claims data. Researchers working with these data types can follow a similar approach (generating code lists, manually reviewing mappings, and comparing event counts) to evaluate potential information loss in these data. Moreover, because OMOP Vocabularies are country-agnostic, this approach could be extended to healthcare data outside the UK, enabling consistent validation across international datasets. To replicate and validate CPRD OMOP transformations without having direct access to patient-level data, researchers can use high-fidelity synthetic datasets that have been generated by CPRD (available at https://www.cprd.com/synthetic-data).
Studies carried out on multiple databases have become more common due to the increased accessibility of data.35,36 Data sources transformed into CDMs are often used as they provide analytic efficiencies.9,37 The aim of this study was to evaluate the success of the OMOP transformation for CPRD, a major EHR database, by comparing the prevalent and incident event/code counts between the source and OMOP-transformed data. The event count estimates were not produced with the same rigour as when investigating diseases/episodes and should not be used for that purpose. Some differences were due to how codes were mapped to different tables, highlighting the importance for researchers to search across all datasets and domains when defining cohorts. Overall, the findings confirm the consistency of output following the OMOP transformation of both CPRD GOLD and CPRD Aurum and provide confidence in analyses conducted using OMOP-transformed data. The availability of CPRD datasets in OMOP format promotes methodological consistency and streamlines multi-database analyses. Citing OMOP transformation evaluations (for CPRD and other databases) is essential for the scientific community to ensure high-quality research. Future research could include validation of other aspects of OMOP transformation and specifically the mapping of source codes to concepts.
Conclusion
This study evaluated the transformation of CPRD GOLD and CPRD Aurum databases into the OMOP CDM. A comparison of prevalent and incident event counts across 38 clinical concepts between the source and OMOP-transformed datasets, showed a high degree of consistency, particularly for diagnoses and prescriptions. Small discrepancies were observed in certain domains such as vaccinations and laboratory results, largely due to variation in code-to-domain mappings.
These findings support the reliability of CPRD OMOP-transformed data for real-world evidence generation. However, it is important to note that this study assessed only selected aspects of the OMOP transformation process, focusing on event count consistency, and did not evaluate the accuracy of medical code mappings. Comprehensive validation of OMOP-transformed data requires further examination of these additional components. To enhance confidence in the use of OMOP CDM across diverse data sources, it is critical that researchers working with other OMOP-transformed datasets conduct similar validation exercises tailored to their data context. Consistent validation practices across OMOP data sources are essential not only to improve efficiency for multi-database studies, but also to ensure transparency, reproducibility and credibility of real-world evidence.
Footnotes
Ethical consideration
This study used data from the Clinical Practice Research Datalink (CPRD). The CPRD data are collected in compliance with relevant legal and regulatory requirements, and studies conducted using CPRD data are approved by the Independent Scientific Advisory Committee (ISAC) for Medicines and Healthcare products Regulatory Agency (MHRA) Database Research (protocol number: 19_044). This study did not require additional ethical approval as it relied solely on anonymized, de-identified data provided by CPRD.
Author contributions
George Kafatos: Conceptualisation, Methodology, Analysis review, Writing. Joe Maskell: Conceptualisation, Methodology, Analysis, Writing review. Olia Archangelidi: Conceptualisation, Methodology, Analysis review, Writing review. David Neasham: Conceptualisation, Methodology, Analysis review, Writing review.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: GK, JM, OA and DN are Amgen Ltd employees and own Amgen Inc shares.
Data Availability Statement
The data used in this study are derived from the Clinical Practice Research Datalink (CPRD) and were transformed into the OMOP Common Data Model. Access to individual patient-level data is not permitted due to contractual agreements with the data provider. Researchers interested in accessing CPRD data should contact CPRD directly (![]() ) to obtain the necessary permissions and licenses.
) to obtain the necessary permissions and licenses.
