Abstract
Keywords
Introduction
Chronic Lymphocytic Leukaemia (CLL) represents a significant haematological malignancy, accounting for 25–30% of adult leukaemia cases and predominantly affecting elderly populations with a median age of 70. 1 This indolent disorder is characterised by the clonal proliferation of mature B-lymphocytes, manifesting through varied clinical presentations ranging from asymptomatic disease to systemic symptoms. Diagnosis employs a multimodal approach incorporating peripheral blood smear examination, flow cytometry immunophenotyping to identify prognostic markers that guides treatment. 2 The management landscape has evolved substantially, transitioning from traditional chemotherapy to targeted approaches including Bruton tyrosine kinase inhibitors (ibrutinib, acalabrutinib) and B-cell lymphoma-2 inhibitors (venetoclax), alongside established chemoimmunotherapy regimens for specific patient subgroups.3,4 Treatment strategies are individualised based on age, comorbidities, genetic profile, and disease stage, with asymptomatic patients often managed through watchful waiting. The disease course is complicated by significant immune dysfunction, leading to increased susceptibility to infections and various autoimmune manifestations including haemolytic anaemia and immune thrombocytopenia.5–8 Additional complications also include secondary malignancies and Richter’s Syndrome, a particularly aggressive transformation occurring in 2–10% of cases. This complex interplay of disease manifestations, treatment options, and complications, combined with its long natural history, presents an ideal scenario for ML applications to enhance classification, prognostication and treatment selection.
Given the challenges in CLL management, such as inefficiencies in flow cytometry gating, inconsistent genomic profiling, and difficulties in tailoring therapy for heterogeneous patient populations, ML models offer significant potential to address these gaps and enhance patient care across the complex CLL spectrum. In diagnosis, ML models could expedite identification of high-risk patients and optimise specialist referrals in resource-constrained settings. ML-based risk stratification may enable earlier treatment initiation for those at risk of rapid deterioration, while predictive models for life-threatening complications, particularly infections, could facilitate pre-emptive interventions to reduce mortality. In treatment selection, ML algorithms could personalise therapeutic choices by predicting individual responses and detecting early signs of resistance or transformation to Richter’s Syndrome. ML models could also be integrated into electronic patient records (EPRs) to provide real-time decision support and streamline data analysis for improved clinical workflows. However, challenges remain, including small, single-centre datasets limiting generalisability, a lack of prospective validation, and technical barriers in data integration.
9
Additionally, successful implementation will require addressing varying data standards across institutions, information governance and privacy concerns, and the need for clinician training to interpret and trust ML outputs. Despite these hurdles, ML holds promise for improving risk stratification and treatment optimisation, but success will depend on robust databases, standardised protocols, and careful clinical implementation, maintaining its complementary role with physician expertise.10–12 Given the complexity and heterogeneity of leukaemia and the rapid progress of ML techniques, it would be challenging to cover all leukaemia subtypes in a single literature review due to the significant amount of published literature and advances, as demonstrated in Figure 1. Number of studies exploring ML applications in haemato-oncology published on PubMed from 2001 to 2023.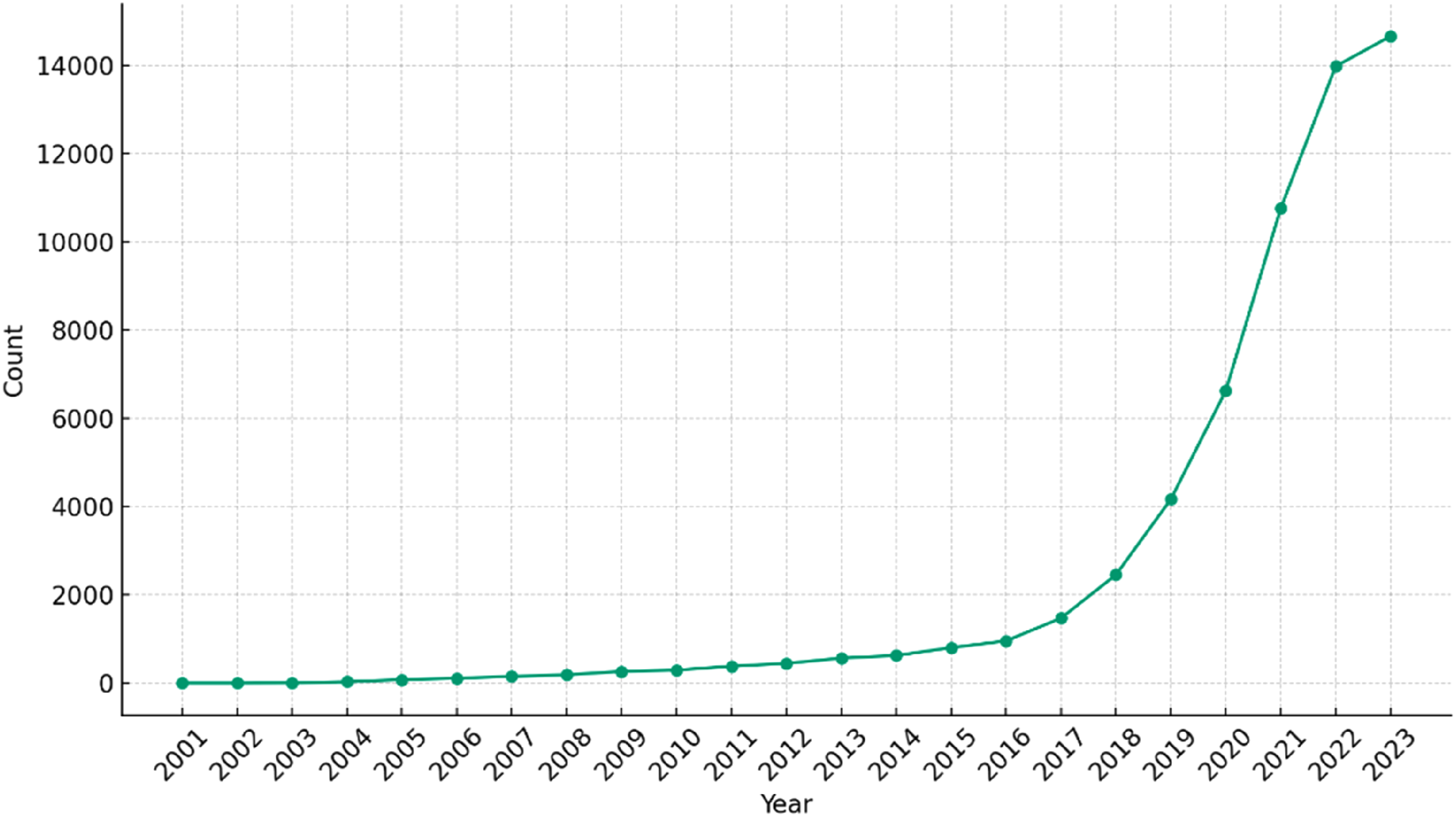
This review will focus specifically on CLL, as it is a particularly interesting subtype of leukaemia due to its highly variable clinical course and long natural history, often spanning years or even decades. This provides a unique opportunity to collect large longitudinal datasets that capture the complex dynamics of CLL, consisting of remissions, relapses, complications, and treatment strategies, such as watch-and-wait, chemotherapy, targeted therapies, and stem cell transplantation. The management of CLL is complex and requires a personalised approach based on patient characteristics, disease stage, molecular and genetic markers. ML techniques could play a crucial role in various aspects of CLL management, namely: • Improving accuracy and efficiency of CLL diagnosis and classification • Identifying novel prognostic markers and developing risk stratification models • Predicting treatment response and guiding the selection of optimal therapies • Predicting the likelihood of developing complications, such as infections or other malignancies and monitoring disease progression to detect early signs of relapse • Optimising supportive care and managing treatment-related complications
The primary objectives of this literature review are to provide an overview of the current state of ML applications in CLL, identify the key methodological approaches, data types used, and performance metrics used in existing studies, discuss the limitations and challenges of current research, and highlight potential areas for future research, development, and implementation.
Methodology
For a study to be included in this review, the study must have been published between 2013 and 2023, be a full-access paper, and use data from at least 100 patients or samples to train and test classification, treatment recommendation or predictive models for CLL or its subtypes. The samples must be real-world human data, and not machine-generated synthetic data. The datasets used for training and testing must be cited or at least the data type(s) must be described. The diagnostic modalities that the study can include lab-based methods such as immunophenotyping, genomics, cellular morphology, and histology, as well as other sources such as demographics, blood tests, drug orders, or clinical notes. Predictions generated by models can include disease trajectory/prognosis, risk of developing complications or providing treatment recommendations. The primary search method was through Google Scholar and PubMed with combinations of search strings, booleans, and wildcards, such as: (“CLL” OR “chronic lymphoid leuk?emia” OR “chronic lymphocytic leuk?emia”) AND (“machine learning” OR “artificial intelligence” OR “ML” OR “AI” OR “deep learning” OR “neural network*” OR “support vector machine*” OR “SVM” OR “random forest*” OR “predictive model*” OR “NLP” OR “natural language processing”)
The results were supplemented with studies from the AIForHealth dashboard, which used a BERT model to identify studies on PubMED that utilised ML models, and categorised each study by subject matter, input data type, and algorithm type.
13
A Python script was used to automatically extract the latest cohort of studies and apply filters to rapidly identify CLL-specific studies (refer to Appendix 1). Despite the consistently increasing amount of literature in this field, a significant number of publications were excluded because they were editorials, letters, technical papers, and literature reviews (see Figure 2). A subgroup consisted of potentially qualifying publications studied other haematological diseases (classified as false positive studies), while many promising studies did not satisfy the search criteria because they had a small number of samples and/or patients. Consort diagram illustrating the literature review search process.
Results
Details of the qualifying studies published between 2014–23 that implemented ML models for CLL classification or prediction. Studies are grouped by the datatype(s) used.

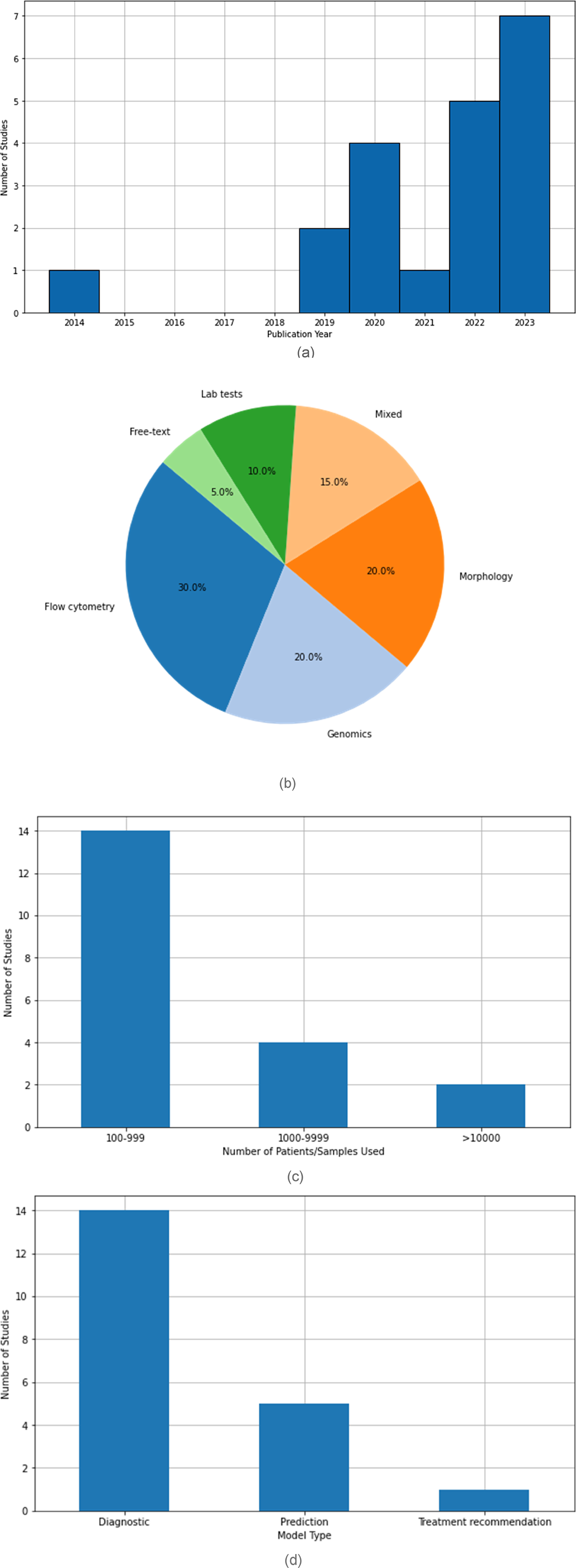
Visualisations illustrating characteristics of the qualifying studies. (a) Histogram of the qualifying studies by publication date. (b) Pie chart showing the breakdown of qualifying studies by data source. (c) Bar chart illustrating the qualifying studies by sample size used for ML model testing and training. (d) Bar chart illustrating the qualifying studies by the intended aim of the ML model output.
Discussion
Over the past decade, there has been a clear trend towards increasing complexity and diversity in the application of ML techniques in CLL. Early studies focused on using classical ML methods on single data types like many other models in other leukaemias, while more recent studies have explored the integration of multiple data types, the use of deep learning and unsupervised methods, and the application of techniques like transfer learning (TL) and convolutional neural networks (CNNs) for morphological analysis. The latest studies continue to refine these approaches, introducing novel XAI techniques and expanding ML applications, yet underscore a persisting gap in the use of NLP and large-scale, real-world datasets.
Flow cytometry models
Zhao et al.’s deep learning approach, utilising Self-Organising Maps (SOMs) and CNNs, demonstrated great potential for automation in FC analysis. 14 Their impressive F1 score and visual explainability tools surpassed traditional manual gating strategies. Ng et al. further solidified the path towards automation with their groundbreaking application of Uniform Manifold Approximation and Projection (UMAP) for unsupervised feature extraction. 15 This method showcased not only high accuracy but also the potential for cost-effectiveness in clinical diagnostics. Salama et al. integrated deep neural network (DNN) models into clinical practice a step further, achieving promising accuracy and specificity. 16 However, their work also highlighted the ongoing challenge of detecting low-level minimal residual disease (MRD). Hoffman et al. introduced a novel approach with their Algorithmic population descriptions (ALPODS) algorithm, identifying cell populations predictive of outcomes and underlining the effectiveness of ML models in prognostic assessments. 17 Bazinet et al. built upon this by comparing their model within DeepFlow software against expert analysis, demonstrating strong correlation and accuracy. 18 Finally, Nguyen et al.’s work with Flow Self-Organising Map (FlowSOM) validated the capacity of ML models for rapid, accurate analysis and rare event detection, while emphasising the need for larger cohort validation for broader adoption. 19
While detecting low-level MRD is not currently an established aspect of CLL management, there is clear evidence of future utility to optimise patient management. 34 FC in CLL is a cornerstone in the initial CLL diagnostic workup, as well as MRD detection. Despite this, a recurring theme across the studies that developed and tested FC-specific models is the challenge of dataset limitations, either in size, diversity, or source. For instance, Zhao et al.’s model faced potential limitations due to FC data quality, and similar concerns were echoed in Bazinet et al.’s study, which relied on a small, single-centre dataset, which raises questions about the generalisability and robustness of these models across different patient populations and clinical settings.14,18 Ng et al. and Salama et al. both highlighted the cost-effectiveness and practical integration of AI models in clinical settings, yet they also acknowledged the need for further research to fully understand the feasibility and economic impact of these technologies.15,16 Additionally, Salama et al.’s model demonstrated a gap in detecting low-level MRD, which is a critical aspect in CLL management. Hoffman et al.’s study was limited by its retrospective design and relatively small sample size, restricting the extrapolation of its findings. 17 Moreover, a common shortfall across these studies is the lack of detailed analysis on the impact of demographic and clinical factors on model performance, as well as a general absence of detailed descriptions of the models’ architectures, which hinders model reproducibility, development, and prospective validation.
NLP models
Loscertales et al. demonstrated the potential of NLP for real-world data extraction from unstructured EHR data by employing the proprietary EHRead model to generate clinical, treatment, and survival profiles of CLL patients in Spain. 20 While EHRead exhibited promising recall, precision, and F1 scores for CLL concept detection, certain limitations warrant consideration. The study’s reliance on a single-centre dataset up to 2018 raises concerns about the model’s generalisability and its applicability to more recent data. Moreover, the lack of transparency regarding EHRead’s architecture, training dataset, and performance across diverse sites hinders a robust assessment of its wider utility. Additionally, by incorporating only gender and date of birth alongside unstructured data, the study may have overlooked other influential factors such as ethnicity and mortality, potentially resulting in an incomplete patient profile. The exclusion of potential confounders like socioeconomic status and geographic location could further obscure variations in clinical characteristics and CLL management practices.
Genomics models
The use of a Support Vector Machine (SVM) model based on epigenetic biomarkers by Queirós et al. to identify CLL subgroups marked a significant advance in understanding the heterogeneity of CLLs and its implications for treatment and prognosis. 21 This paved the way for integrating epigenetic factors into CLL classification. Orgueira et al. further showcased the potential of transcriptomic patterns in predicting treatment timelines by applying the Gaussian Mixture Model-Expectation Maximisation (GMM-EM) algorithm for patient stratification based on gene expression, moving beyond traditional prognostic markers. 22 Their deep survival model achieved high precision and accuracy despite its small sample size, indicating the growing role of deep learning in CLL prognostics. Zhang et al.’s study using the Geometric Mean Naïve Bayesian Classifier (GMNB) algorithm for differential diagnosis marked a significant advancement in precision medicine by employing a targeted transcriptome approach to distinguish between various hematologic and solid tumours, including CLL. 23 Morabito et al. further expanded the scope by introducing the DeepSHAP Autoencoder Filter for Genes Selection (DSAF-GS), which combines deep learning with XAI to not only predict treatment outcomes but also to uncover new biological pathways and networks involved in CLL. 24
Several limitations were demonstrated in these studies, beginning with Queirós et al., a notable shortfall was the lack of external validation and reliance on a single DNA methylation analysis platform, raising questions about the model’s generalisability and robustness. 21 This issue of external validation was also evident in Morabito et al.'s study, where the model’s applicability beyond the initial Italian cohort remains untested. Furthermore, both studies, along with Orgueira et al.’s work, did not provide insights into the functional significance of the identified biomarkers or gene patterns, which potentially limits the understanding of their biological relevance in CLL. 22 Orgueira et al. and Morabito et al. also faced challenges due to small sample sizes and absence of comparative analysis with existing methods, in which the latter would provide a more nuanced understanding of their models’ efficacy.22,24 Zhang et al.’s study, while ambitious in scope, omitted comparisons between its algorithm and key diagnostic standards for CLL, specifically blood count, film examination, and FC, which are fundamental for initial diagnosis and confirmation. 23 The critique of not including immunohistochemistry is retracted, acknowledging its limited relevance to CLL diagnostics. This omission leaves a gap in understanding the relative strengths and weaknesses of their AI approach. Similarly to the FC models, there was a general lack of transparency regarding the models’ architectures.
Laboratory models
Haider et al.’s development of a predictive model using morphological and immature fraction-related parameters from full blood count (FBC) results is a notable advancement. 25 The Radial Basis Function Network (RBFN)-based model was demonstrated to be effective in early differentiation among various types of leukaemia. However, the study’s lack of external validation, particularly with datasets from multiple centres, limits the generalisability of its findings. Similarly, Padmanabhan et al.’s application of Extreme Gradient Boosting (XGB) for CLL diagnosis and screening based on routine FBC results, represented a significant advancement. 26 However, there was a reliance on a small dataset from a single centre and both contextual clinical data and unstructured data were not integrated.
Multimodal models
Chen D et al. broke new ground with their use of unsupervised clustering alongside demographics and laboratory test results to predict time to first treatment for CLL patients. 27 This work not only improved prognostic prediction but also demonstrated the performance of RSF and GBM in patient risk stratification. Building on this, Agius et al. developed a multimodal ensemble model utilising a genetic algorithm (GA) to build an ensemble model and data from the Danish CLL registry. 28 Their model proved effective in combining diverse data types, offering resilience to incomplete data, and improving interpretability through SHAP analysis. Finally, Meiseles et al.’s gradient boosting machine (GBM) model effectively predicted treatment necessity while incorporating feature importance analysis. 29
A notable shortcoming in Chen et al.’s study lies in its lack of exploration into the specific features learned by the models, obscuring potential details regarding the underlying biological mechanisms. 27 Both Chen et al. and Meiseles et al. relied on single-centre datasets, which raises concerns about the generalisability of their findings.27,29 Agius et al.’s bag-of-words approach for extracting features from clinical data may not fully utilise the richness of such data; implementing more advanced NLP techniques on clinical free-text could potentially add depth to the analysis. 28 While their model proved robust with missing data, further validation, and exploration of its generalisability across diverse populations are crucial for wider applicability.
Morphology models
Zhang et al.’s application of TL and Principal Component Analysis (PCA) achieved a perfect cross-validation accuracy for CLL, setting a new standard in classification accuracy and demonstrating the potential of automated subtype differentiation. 30 Steinbuss et al.’s use of Efficient Neural Networks (EfficientNet) achieved high accuracy in classifying lymph node subtypes based on cellular morphology, suggesting the potential to improve accuracy in the diagnosis and treatment of lymph node cancers. 31 Chen et al.’s unsupervised clustering-based model successfully identified distinct morphological cellular phenotypes associated with CLL progression stages, offering insights into CLL progression and potential new treatment targets. 32 Finally, Wang et al.’s deep CNN algorithm demonstrated high accuracy in lymphocyte identification, showcasing its potential to develop new diagnostic tests for CLL and other blood cancers. 33
Despite the advances, certain limitations should be noted. The practical applicability of the TL/PCA-based model in Zhang et al.’s study is questionable due to the narrow dataset, lack of interpretability and clinical validation. 30 This indicates potential issues with generalising the model to other datasets and limits its immediate use in clinical practice. Steinbuss et al.s use of EfficientNet faced challenges due to a relatively small sample size considering the diversity of subtypes analysed, and the potential skill bias introduced by single annotator involvement. 31 These limitations suggest the model may not be accurate for all subtypes of lymph node cancer and could be influenced by the annotator’s specific skills. Chen et al.’s unsupervised clustering-based model was constrained by a small, single-centre cohort and the absence of detailed model architecture, potentially limiting its generalisability, and hindering understanding of how the model functions. 32 Lastly, Wang et al.’s deep CNN algorithm was limited by its retrospective nature and its inability to distinguish between disease progression and infection. 33 This highlights that the model cannot predict future disease development and might misinterpret CLL symptoms as those of other infections.
These findings underscore the potential for morphology models to automate histopathological analysis and refine CLL classification as a result. However, for effective clinical integration, these models require further validation, clarity, and adjustments to address dataset diversity and interpretability challenges.
XAI and NLP applications
Only a quarter of the reviewed studies explored XAI, highlighting an area for expanded focus. Agius et al., Meiseles et al., and Morabito et al. applied SHAP to enhance interpretability.24,28,29 Zhao et al. used saliency maps and density plots to pinpoint important features, potentially aiding human experts. 14 However, such visualisations might oversimplify model decision-making and can be computationally demanding in high-dimensional settings. Further development is needed to optimise their robustness, accuracy, and practicality. Hoffman et al.’s ALPODS offers sample-based explanations of CLL-specific immune cell populations relevant to outcomes. 17 While valuable, the study lacks clarity on how these explanations were produced and their ease of use by healthcare professionals. Future studies that utilise ALPODS should prioritise transparency to build trust and assess its impact on clinical workflows. Morabito et al. employed DeepSHAP with a neural network (NN) to pinpoint genes predictive of treatment outcomes. 24 Yet, the benefits of validating their findings with external datasets, compare its performance against other XAI methodologies, and enhancing the computational efficiency of SHAP would help to ascertain the utility of this XAI approach. Agius et al. and Meiseles et al. both used SHAP to rank feature importance and reveal risk factors, although potential variability of SHAP values calls for a more consistent metric within these approaches.28,29 Larger and more diverse datasets could further boost robustness. Notably, only Loscertales et al. focused exclusively on free-text data, likely due to the scarcity of accessible real-world clinical datasets and inherent complexities in integrating textual and non-textual data. 20 Developing strategies to address these challenges would greatly expand the utility of free-text in future research.
The case for multimodal ML models
While Table 1 illustrates the performance of various ML models using reported metrics, it is crucial to consider their practical applicability in diverse clinical environments encountered in CLL management. Monomodal models often trained on homogenous datasets may not adequately reflect the complexities of real-world data, which are characterised by variability and incomplete data. In contrast, multimodal models that integrate heterogeneous data types are typically more robust, as exemplified by CLL-TIM’s performance. These models leverage multiple data sources to effectively compensate for missing features in any single data type. Such models are more suited to address challenges such as interoperability and data standardisation prevalent in healthcare settings. Therefore, while the metrics in controlled settings are informative, the real value of ML models lies in their ability to operate effectively within the intricate ecosystem of EPRs, thereby offering a more comprehensive approach to patient management in CLL. This integration is pivotal in harnessing the full potential of ML to enhance disease characterisation, diagnostic precision, and the personalisation of therapeutic strategies. However, interoperability with existing EHR systems remains a major challenge, as data formats and standards vary widely across healthcare providers, making the integration of ML tools cumbersome. Additionally, the legal and regulatory landscape, including compliance with TRIPOD guidelines and distinguishing the definition of the ML application as either research and development or a medical device, adds layers of complexity. 35
To address these challenges, future research should also focus on the development of ML models that can operate effectively under these constraints. This involves designing models that are adaptable to different data standards and can provide reliable outputs even with incomplete data. Furthermore, ensuring that these models adhere to stringent regulatory standards will be crucial for their acceptance and integration into clinical practice. These steps are necessary to move from the experimental application of ML in CLL management to their routine use in patient care, providing a more holistic approach to disease management that leverages the full potential of ML technologies
Limitations of the review
Several limitations of this review warrant consideration. The substantial heterogeneity across the reviewed studies encompassed diverse ML architectures, varied dataset characteristics and inconsistent outcome measures across diverse modalities including flow cytometry, genomics and morphological data. This variability precluded meaningful quantitative synthesis and meta-analysis. The heterogeneity coupled with insufficient dataset descriptions in some studies, such as fundamental details like indicating the number of samples per patient (e.g., whether there was one or more than one sample per patient) complicated the assessment of data quality and limited the generalisability of findings across different clinical settings. Furthermore, the reliance on a single reviewer for study selection and data extraction introduces potential selection bias, although this was partially mitigated through structured inclusion criteria.
Ethical considerations
As ML models become increasingly integrated into the CLL clinical care pathways, it is crucial to address ethical concerns with inclusive development approaches such as improved transparency, patient and public involvement (PPI) events and strong adherence to information governance (IG) legislation. Holding regular PPI events would serve as a forum for patients to share their thoughts regarding the models’ intended use, potential benefits, and risks, which would also help inform the direction of research and development. Transparency in the provision of clear and accessible documentation about the models’ data sources, architecture, performance metrics and XAI implementation is critical for building trust with healthcare professionals as end-users and patients.36–38 In addition to these considerations, data privacy and security remain paramount, with robust IG frameworks, anonymisation techniques and sharing protocols must be enforced to protect patient confidentiality. All these principles align with National Institute for Health and Care Excellence (NICE)’s commitment to involving patients and the public in healthcare decision-making, ensuring that ML models serve the best interests of all users. 39 An additional critical ethical consideration in the future, when models become more sophisticated and autonomous, is accountability and liability in the event of clinical errors. To alleviate this, clear guidelines and policies must be established to determine roles and responsibilities of healthcare providers, researchers, and model developers. Ongoing education and training for healthcare professionals on the appropriate use and interpretation of model outputs are also necessary to ensure the safe model integration into clinical practice.
Conclusion
This review has highlighted the advances in ML applications in CLL management, particularly in diagnosis and classification. Several studies demonstrated that ML models can accurately diagnose CLL, especially using FC and morphological data. However, this progress also underscores critical gaps and limitations in the current research landscape, which align with broader trends in ML applications in haematology and oncology.40,41
Current ML research in CLL management has largely produced narrow, single-purpose models using limited datasets or cohorts and often utilise single-modal data. While effective within their limited scope, these models do not address the complex, multifaceted nature of CLL management. Even the more advanced multimodal approaches fall short of a truly comprehensive solution, notably lacking integration of NLP to leverage the vast number of insights contained in unstructured clinical data. This fragmented approach has resulted in a collection of specialised tools rather than a cohesive system capable of supporting the full spectrum of CLL management.
Future research should prioritise the development and validation of NLP models capable of extracting clinically relevant information from unstructured data sources in CLL. These models could significantly enhance ML-assisted CLL management by incorporating previously underutilised data from clinical notes, pathology reports, and patient-reported outcomes. The output of such models would serve dual purposes: providing actionable information for clinical decision-making or research and generating additional input features for more complex predictive models. This could pave the way for novel methods to integrate diverse data types as additional inputs while addressing challenges of data harmonisation, quality, and IG. To facilitate multi-centre collaborations while maintaining patient confidentiality, future studies should explore advanced federated learning approaches and innovative data anonymisation techniques. Access to large longitudinal, real-world datasets would enable the development of ML models capable of capturing the temporal aspects of CLL progression and treatment response. XAI techniques are instrumental for effective prospective evaluation of the impact of these models on patient outcomes, clinical decision-making, and healthcare resource utilisation.
Equally important is the development of robust frameworks for continuous monitoring and mitigation of potential biases in ML models for CLL. This includes ensuring equitable performance across diverse patient populations and addressing potential disparities in model outcomes.
By pursuing these directions, the role of ML models is likely to change from isolated tools and become part of a more comprehensive, ML-assisted CLL management system. Such a system would not only improve diagnostics, but also enhance treatment planning, predict outcomes, and ultimately improve patient care across the entire CLL journey. This holistic approach to ML in CLL aligns with the broader roadmap for AI in haematology and oncology. 41 Realising this potential will require collaborative efforts between clinicians, data scientists, and healthcare systems.
Supplemental Material
Supplemental Material - Systematic review of machine learning applications in early prediction and management of chronic lymphocytic leukaemia
Supplemental Material for Systematic review of machine learning applications in early prediction and management of chronic lymphocytic leukaemia by Mohammad Al-Agil, Piers Patten EM and Anwar Alhaq in Health Informatics Journal
Footnotes
Author contributions
MA: Conceptualisation, Methodology, Investigation, Writing – Original Draft Preparation and Writing – Review & Editing. PEMP and AA: Conceptualisation, Supervision and Writing – Review & Editing.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: PEMP was supported by MRC grant MR/T005106/1. The other authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
