Abstract
Mycoplasma pneumonia may lead to hospitalizations and pose life-threatening risks in children. The automated identification of mycoplasma pneumonia from electronic medical records holds significant potential for improving the efficiency of hospital resource allocation. In this study, we proposed a novel method for identifying mycoplasma pneumonia by integrating multi-modal features derived from both free-text descriptions and structured test data in electronic medical records. Our approach begins with the extraction of free-text and structured data from clinical records through a systematic preprocessing pipeline. Subsequently, we employ a pre-trained transformer language model to extract features from the free-text, while multiple additive regression trees are used to transform features from the structured data. An attention-based fusion mechanism is then applied to integrate these multi-modal features for effective classification. We validated our method using clinic records of 7157 patients, retrospectively collected for training and testing purposes. The experimental results demonstrate that our proposed multi-modal fusion approach achieves significant improvements over other methods across four key performance metrics.
Keywords
Introduction
Mycoplasma pneumonia (MP), a type of community-acquired pneumonia (CAP), is responsible for 10 to 40% of CAP cases in hospitalized children.1,2 It can cause severe respiratory complications, leading to hospitalizations and potentially life-threatening situations in pediatric patients. MP spreads through droplets and direct contact with infected individuals, often displaying an epidemic infection pattern with seasonal peaks in autumn and winter.3,4 In clinical practice, pediatricians commonly treat children with MP using macrolide antibiotics, differing from treatment protocols for other pneumonias.5–7 Therefore, distinguishing MP from other pneumonia cases is crucial for ensuring child safety and minimizing inappropriate medication use.8–10
Recent advancements in artificial intelligence have greatly assisted clinicians in diagnosing and managing diseases using multi-modal data sources like electronic medical record (EMR) and medical imaging.11–13 EMRs provide comprehensive records of patient diagnosis and treatment, penned by hospital clinicians. Current research predominantly focuses on extracting and utilizing valuable Pulmonology information from EMRs for disease research and developing automated diagnostic systems, e.g., asthma,14,15 ICU mortality scoring 16 and infectious diseases identification. 17 Several machine learning approaches, e.g., XGBoost, 18 Bayesian network, 19 regression trees, 20 etc.,21,22 have also been developed for pneumonia diagnosis and management. Additionally, the integration of medical imaging and EMRs into disease diagnosis has also been explored, e.g., text-image semantic retrieval, 23 multi-modal pneumonia identification24–26 and COVID-19 diagnosis. 27
Despite the plethora of EMR-based respiratory disease diagnosis research, studies specifically focusing on MP diagnosis are lacking. In this study, we propose a diagnostic system for MP utilizing a combination of deep learning and multiple additive regression trees (MART). The methodology begins with extracting free-text and structured medical test data from EMRs via a preprocessing pipeline. A pre-trained BERT model
28
serves as the free-text feature extractor, while MART are employed to transform features from structured data, converting multi-source scalar numeric data into dimensionless multi-hot vectors. This facilitates the unification of diverse features into a single metric space. Subsequently, an attention-based fusion module merges free-text and structured data features, which are then inputted into a binary classifier for the final classification. Figure 1 provides an overview of the proposed system, which was trained and validated on 7157 retrospectively collected EMRs of CAP patients. Overview of the proposed system. (a) Overview of record preprocessing pipeline. Providing clinical text of EMR and extract both free-text and structured streams using pre-built regular expressions. (b) Overview of MP Identification. BERT extracts free-text features. Trained MART transforms structured streams. An attention fusion module is used to fuse features of free-text and structured data, and fed the fused features into the binary classifier for MP identification.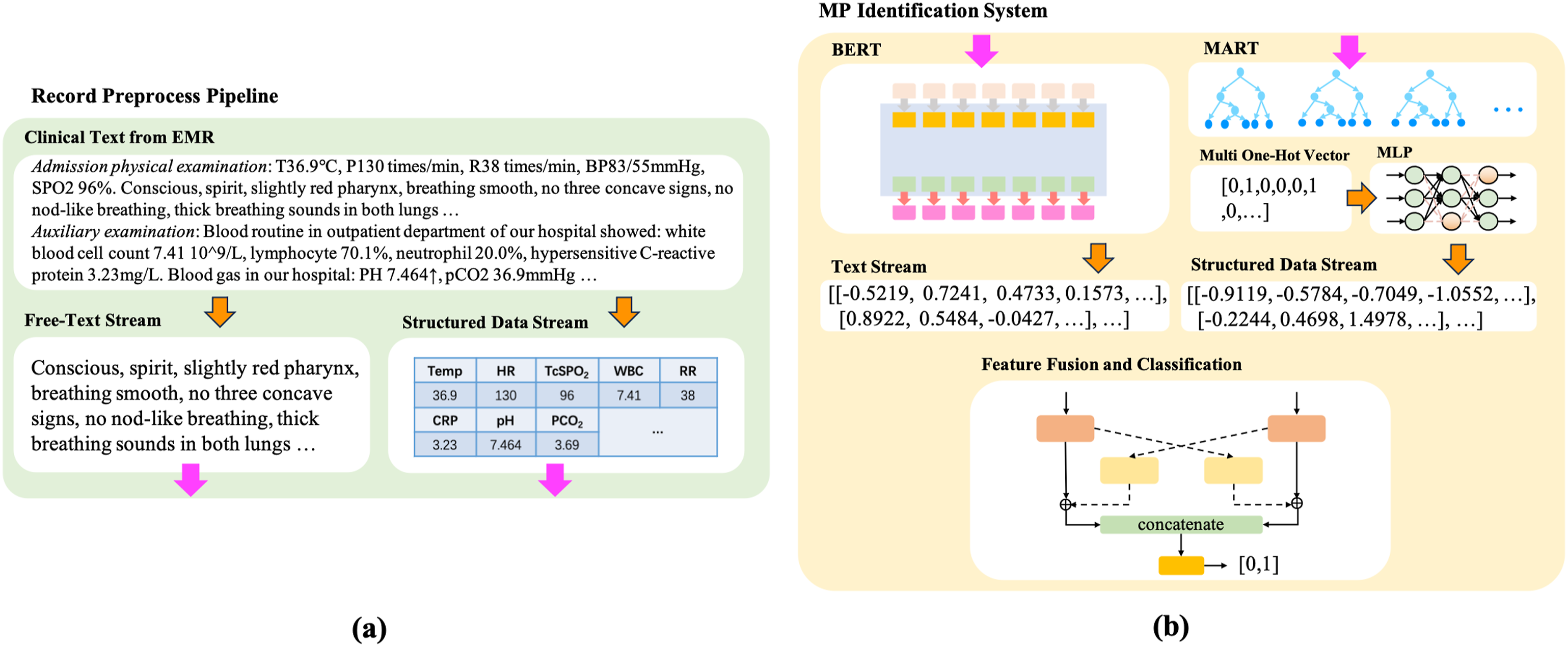
Materials & methods
Ethical approval
This research was approved by the Institutional Review Board (IRB) of the Medical Ethics Committee at Children’s Hospital, Zhejiang University School of Medicine, China (IRB Approval ID: 2020-IRB-058) and conducted in compliance with the Declaration of Helsinki. Given the retrospective nature of the data analysis involved in this study, the IRB granted a waiver for the requirement of informed consent.
Data overview
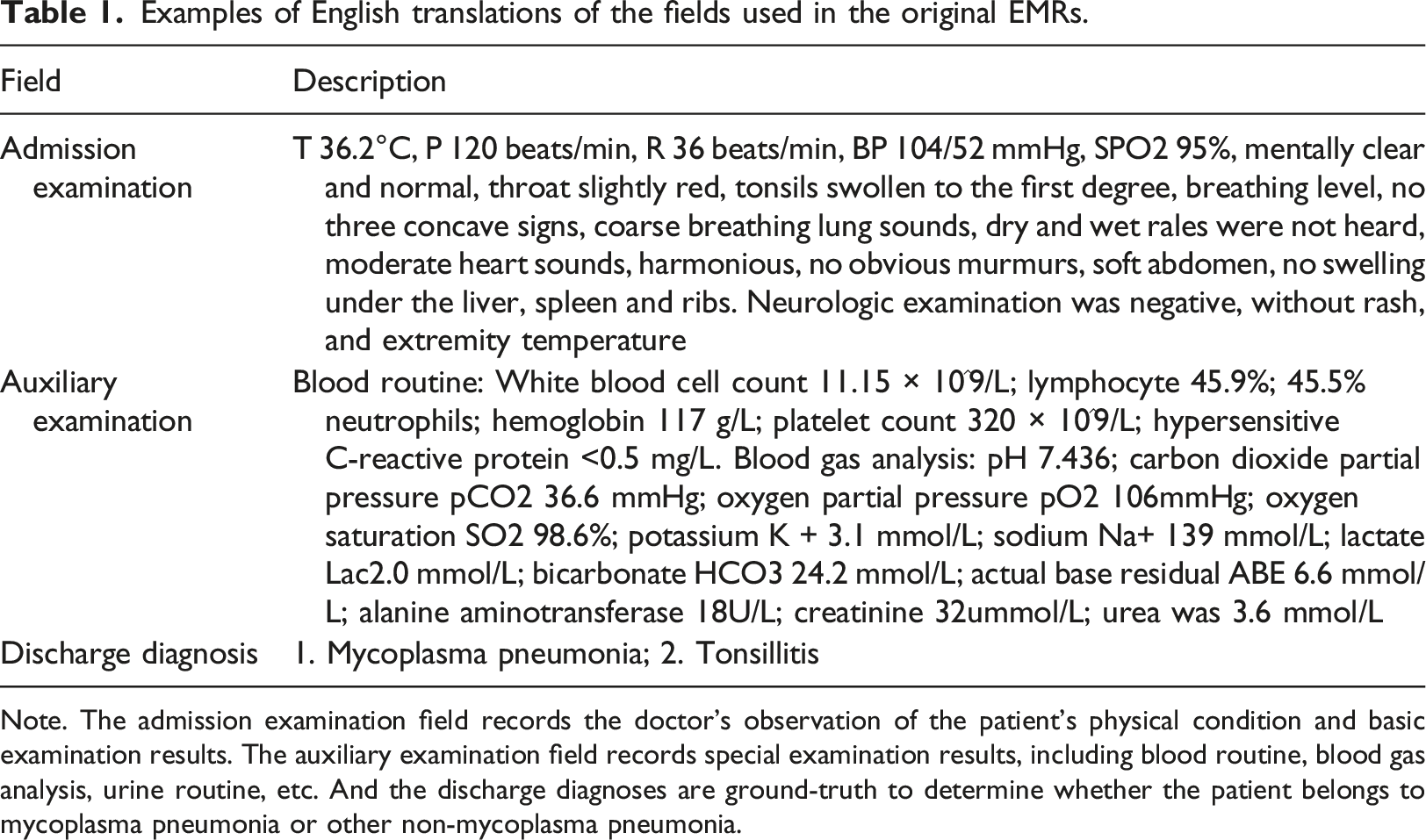
In this study, EMRs were retrospectively collected, consisting of a patient cohort of 7157 individuals diagnosed with CAP at the Department of Pulmonology, Children’s Hospital, Zhejiang University School of Medicine. This cohort was comprised of two subgroups: 3706 cases of MP and 3451 non-MP cases. The age distribution of these patients ranged from 29 days to 18 years, with a mean age of 3.57 years and a standard deviation (SD) of 3.05 years. Statistical insights derived from the patients’ EMRs are illustrated in Figure 2. The primary analytical focus of this study was on three critical aspects of the EMRs: admission examination, auxiliary examination, and discharge diagnoses. It is important to note that all free-text records within these domains were originally recorded in Chinese, as detailed in Table 1. Data overview. (a) Histogram of the age distribution of patients, ranging from 0 to 18 years old, with the majority of children under 4 years old. (b) Bar graph of the months of patient admission, where the peak seasons of MP and CAP are in autumn and winter. (c) Pie chart of gender distribution of patients, in which males account for 56.66% and females account for 43.34%. Examples of English translations of the fields used in the original EMRs. Note. The admission examination field records the doctor’s observation of the patient’s physical condition and basic examination results. The auxiliary examination field records special examination results, including blood routine, blood gas analysis, urine routine, etc. And the discharge diagnoses are ground-truth to determine whether the patient belongs to mycoplasma pneumonia or other non-mycoplasma pneumonia.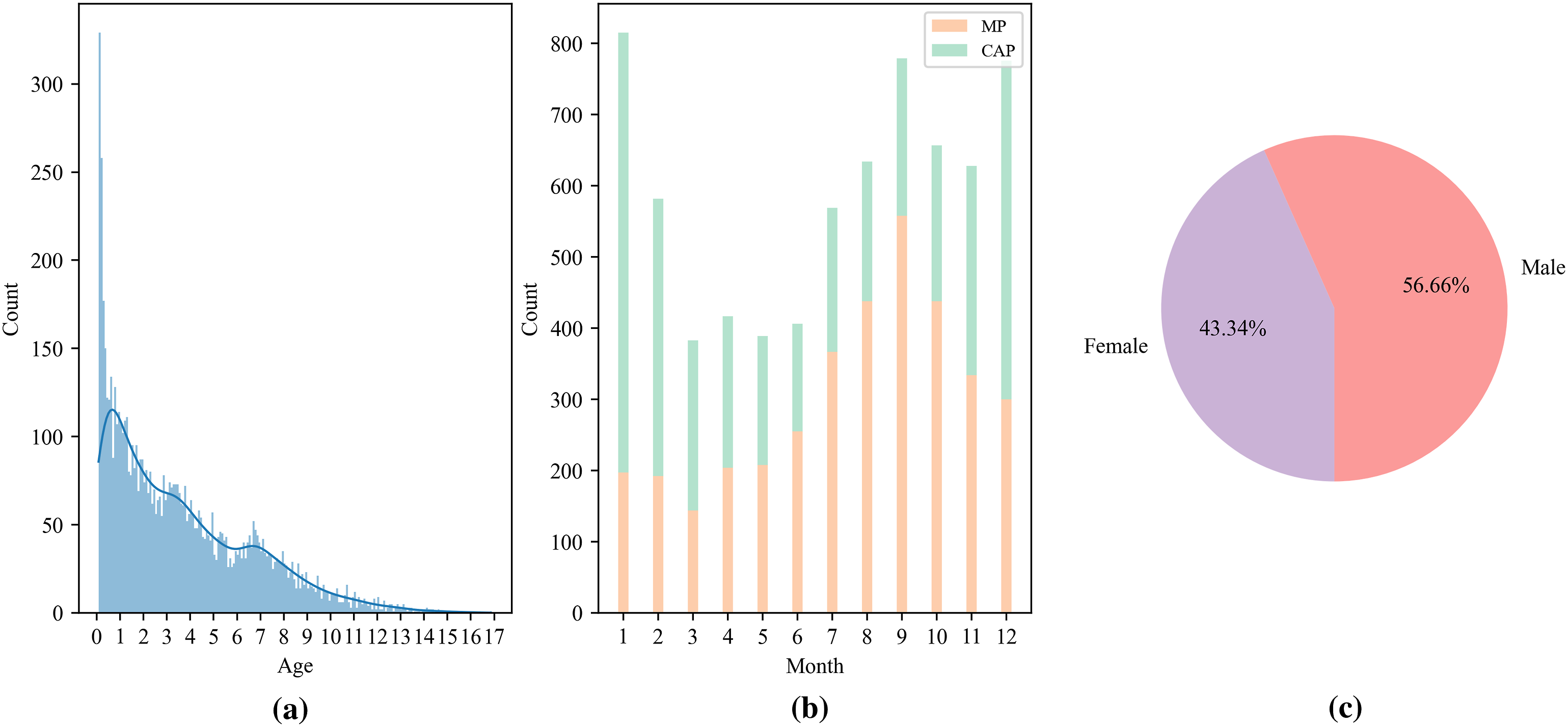

For this study, we randomly divided the EMR data into three distinct sets on a patient-by-patient basis: a training set (85% of the total data), a validation set, and a testing set (each constituting 7.5% of the total data). Specifically, the training set comprises 2966 MP cases and 3111 non-MP cases, totaling 6077 cases. The validation set, which we used for hyperparameter tuning, includes 370 MP cases and 170 non-MP cases, amounting to 540 cases in total. Similarly, the testing set, designated for model evaluation, mirrors the validation set in terms of case distribution, with 540 cases in total.
Additionally, this study also assembled an expansive dataset comprising 50,754 non-pneumonia-related EMRs denoted as ‘Extra-data’, which was sourced from three secondary-level children’s hospitals: Zhengzhou Children’s Hospital, Shangyu Maternity & Childcare Hospital, and Dongyang People’s Hospital. The collection spanned both pulmonary and non-pulmonary departments. The format and writing style of these records were analogous to the EMRs of patients with CAP in our study, encompassing fields such as admission examination and auxiliary examination. Notably, this dataset, which was not subjected to any preprocessing, was utilized as a domain-specific corpus for the purpose of unsupervised pre-training of a BERT model.
Data preprocessing pipeline
EMRs often contain semi-structured text that reflects both the hospital’s writing norms and individual clinicians’ habits. To address this, we have developed a three-step preprocessing pipeline, illustrated in Figure 1(a), for cleaning and extracting relevant information from free-text in EMRs: (1) Standardization of Test Item Expressions: Due to the varied representations of test items in EMRs, such as “temperature 37.1 degrees” and “T 37.1°C”, we applied 228 regular expressions to standardize these expressions. For example, “capillary refill time 3 s” is converted to “CRT 3s”. (2) Tabulation of Test Result Numerics: We utilized 74 regular expressions, e.g., “partial pressure of carbon dioxide ([0-9]+[.。]?[0-9]*)mmHg”, to extract numeric values from standardized test results and organize them into a tabular format. (3) Physical Descriptions Extraction and Refinement: The same 228 regular expressions from the first step are reused to extract physical descriptions, which involves removing text related to test results. Additionally, we employ a set of straightforward regular expressions to systematically eliminate redundant punctuations and speculative phrases that imply tentative diagnoses, like ‘suspected pneumonia’ or ‘suspected mycoplasma infection.’ This strategy is crucial for preventing label leakage, a common occurrence among clinicians during patient examinations.
Overview of structured data (75 items in total).
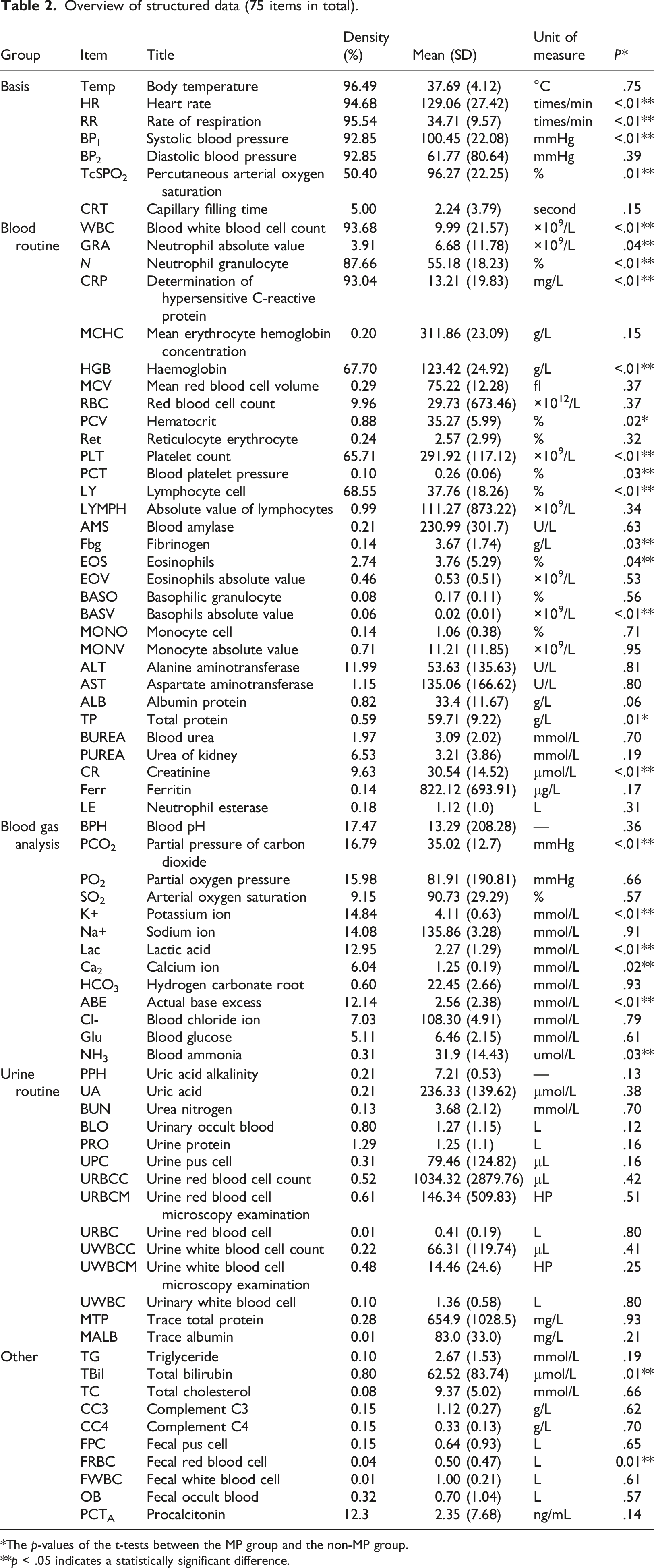
*The p-values of the t-tests between the MP group and the non-MP group.
**p < .05 indicates a statistically significant difference.
The proposed MP identification system
An illustrative overview of the proposed MP identification system is shown in Figure 1(b). In our study, we utilize both structured test data and free-text descriptions from the preprocessing pipeline to identify MP cases. Throughout this paper, we refer to these two types of data as ‘free-text stream’ and ‘structured data stream’, representing the dual input streams of our system.
Transformer encoder for free-text streams
For processing the free-text stream, we employ the well-known Transformer-based architecture BERT model 28 as our encoder. Specifically, we utilize the Chinese BERTBASE model from the Transformers package, 29 which comprises an initial embedding layer followed by a series of 12 Transformer blocks. 30 The free-text stream is first tokenized and then passed through the embedding layer. Subsequently, the output from the embedding layer undergoes feature interaction and extraction within the Transformer blocks.
Each Transformer block is composed of two primary sub-layers: a multi-head self-attention (MHA) layer and a feed-forward network (FFN) layer. The MHA layer, with 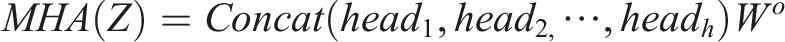
Here, 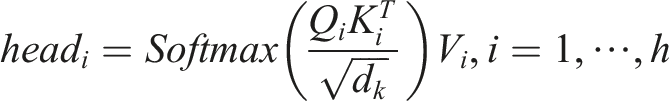
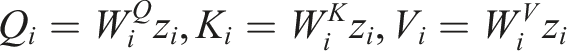
In this formulation, 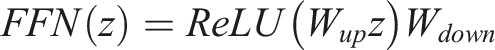
This layer consists of two linear transformations with a ReLU activation function in between. 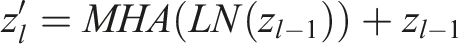
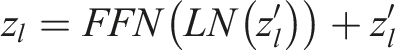
Here
In this study, the collected ‘Extra-data’ set serves as the training corpus for unsupervised pre-training of BERT, specifically tailored for the clinical text domain. Unsupervised pre-training in this context typically involves two tasks: the Masked Language Model (MLM) and Next Sentence Prediction (NSP). 28 In the MLM task, BERT is trained to predict randomly masked tokens within a sentence. In the NSP task, BERT assesses whether two sentences are contextually related. However, given that clinical text is often brief and lack clear contextual relationships, and considering that NSP has been found to be less effective than MLM in some scenarios,33,34 our study focuses solely on the MLM task for pre-training. We refer to the model trained through this pre-training process as ‘MLM BERT’.
Tree-based feature transformation for structured data streams
Drawing inspiration from the hybrid model structure presented in ref, 35 we trained a widely recognized MART model, namely LightGBM, 36 as a feature transformer for structured data streams. The primary objective of this training process is to execute binary classification between MP and non-MP cases, employing the binary cross-entropy function as the objective. However, this process exclusively considers structured tabular data as input within the dataset. Upon achieving training convergence, LightGBM transitions to serve as a feature transformer. This is achieved by converting multiple numeric features into a unified multi-hot categorical vector. Specifically, when a sample with various features is input into LightGBM, it ultimately reaches a leaf node within one of the model’s trees. This particular leaf node is designated as ‘1’, while all other nodes are labeled as ‘0’. As a result, the numerical feature of the sample, as it is distributed by the tree, is transformed into a one-hot vector. Given the presence of numerous sub-trees in LightGBM, all these one-hot vectors are concatenated to form a single multi-hot vector. This process effectively accomplishes the transformation of numerical features. Consider the scenario of an additive tree ensemble comprising two trees, where the first tree has three leaves and the second tree has two leaves. In this instance, if a sample is directed to the second leaf in the first tree and the first leaf in the second tree, it would be represented as [0,1,0,1,0]. This representation is an outcome of tree-based feature transformation. Essentially, this method acts as a supervised feature encoding technique, transforming continuous real-valued vectors into more concise binary-valued vectors. The basis for this transformation is a set of binary rules, which are established according to the training objective of the tree. In this context, the objective is to differentiate between MP and non-MP cases.
Attentive fusion of free-text streams and structured data streams
In our system, we employ a Multi-modal Attentive Fusion (MAF) module, as proposed in prior research.
37
This module is designed to effectively combine features from both free-text streams and structured data streams. The detailed architecture of the MAF module is illustrated in Figure 3 and defined as follows: The structure of the multi-modal attentive fusion module. (a) The structure of the fusion block. (b) The detailed structure of the adaptive feature infusion module.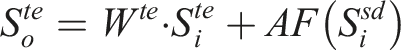
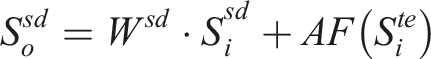
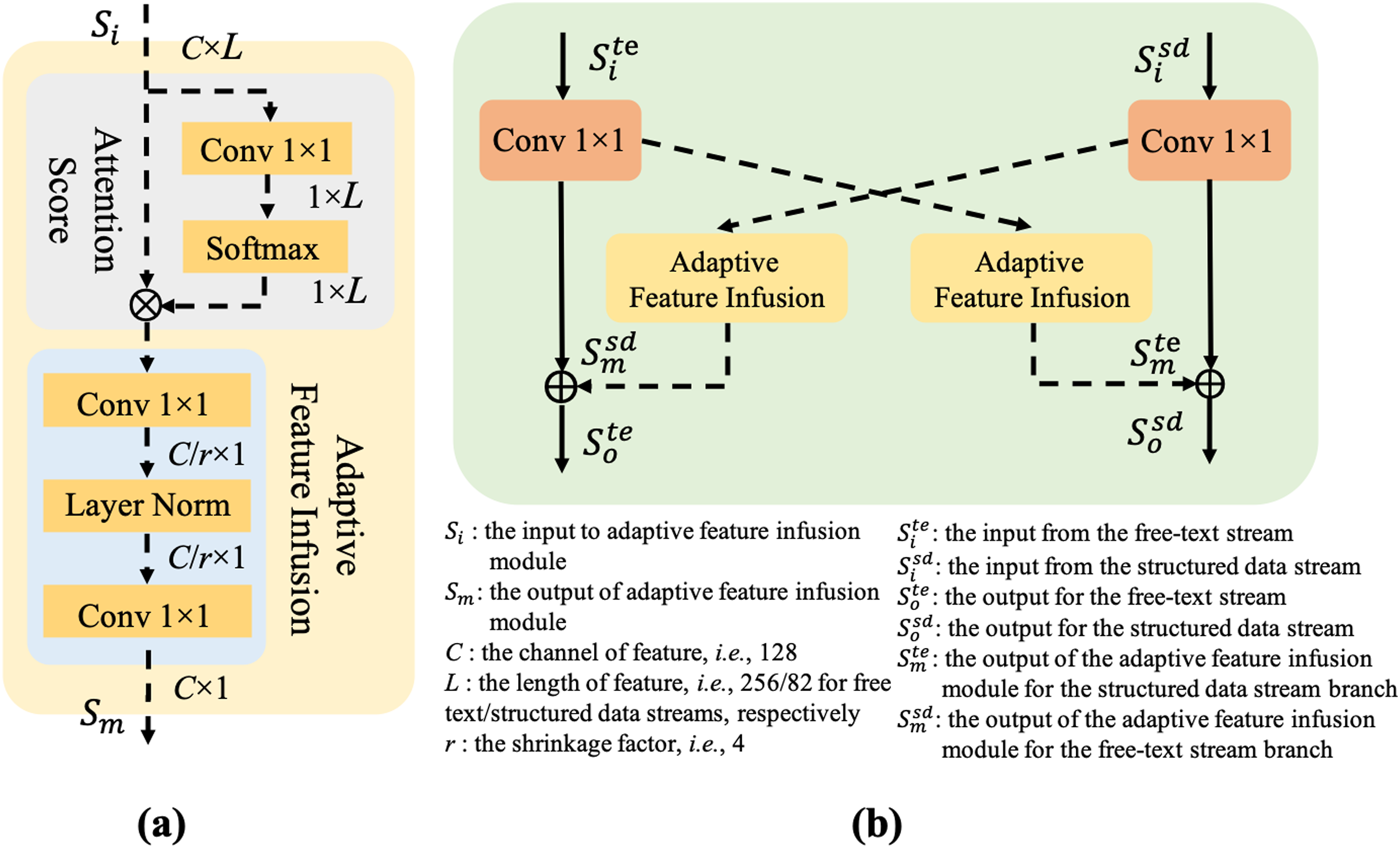
In this context, 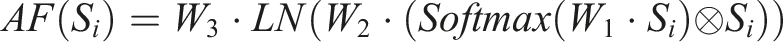
Here,
Results
In our study, we conduct a quantitative comparison of MP identification performance using uni-modal data streams, namely, free-text streams and structured data streams, and a multi-modal approach that fuses both types of data streams. To evaluate the effectiveness of these methods, we employ widely recognized evaluation metrics: accuracy (Acc), precision (P), recall (R), and F1 score (F1). Among these metrics, the F1 score is given the highest priority due to its balanced consideration of both precision and recall.
MP identification based on free-text streams
In our study, we emphasize the significance of text embedding/representation in analyzing free-text streams. We quantitatively compare three prevalent text-embedding strategies: Vector Space Model (VSM), Word2Vec, and Transformer-based method. For VSM and Word2Vec, the free-text is tokenized into word sequences using the Stanford Chinese Word Segmenter. 38 In contrast, Transformer-based methods tokenize free-text into character sequences. The VSM method utilizes Term Frequency–Inverse Document Frequency (TF-IDF) 39 for word representations, embedding text lines into VSM space. Word2Vec applies the Chinese word vector, from FastText,40,41 which has a vocabulary of 4,000,000 words, representing each word as a vector. In the Transformer-based method, BERT’s embedding layer provides word representations. For VSM-based classifiers, besides LightGBM, we also implemented SVM, LR and MLP via the Scikit Learn package. 42 The Word2Vec classifiers include FastText, 41 TextCNN, 43 and BLSTM. 44 The Transformer-based method employs a conv-tanh-linear composite layer, where BERT’s token representations are fed into a classifier, consisting of a 1-day conv layer, a tanh-activation layer, and a binary linear classification layer.
Performance comparison of various methods for different word embedding representation strategies based on free-text streams.
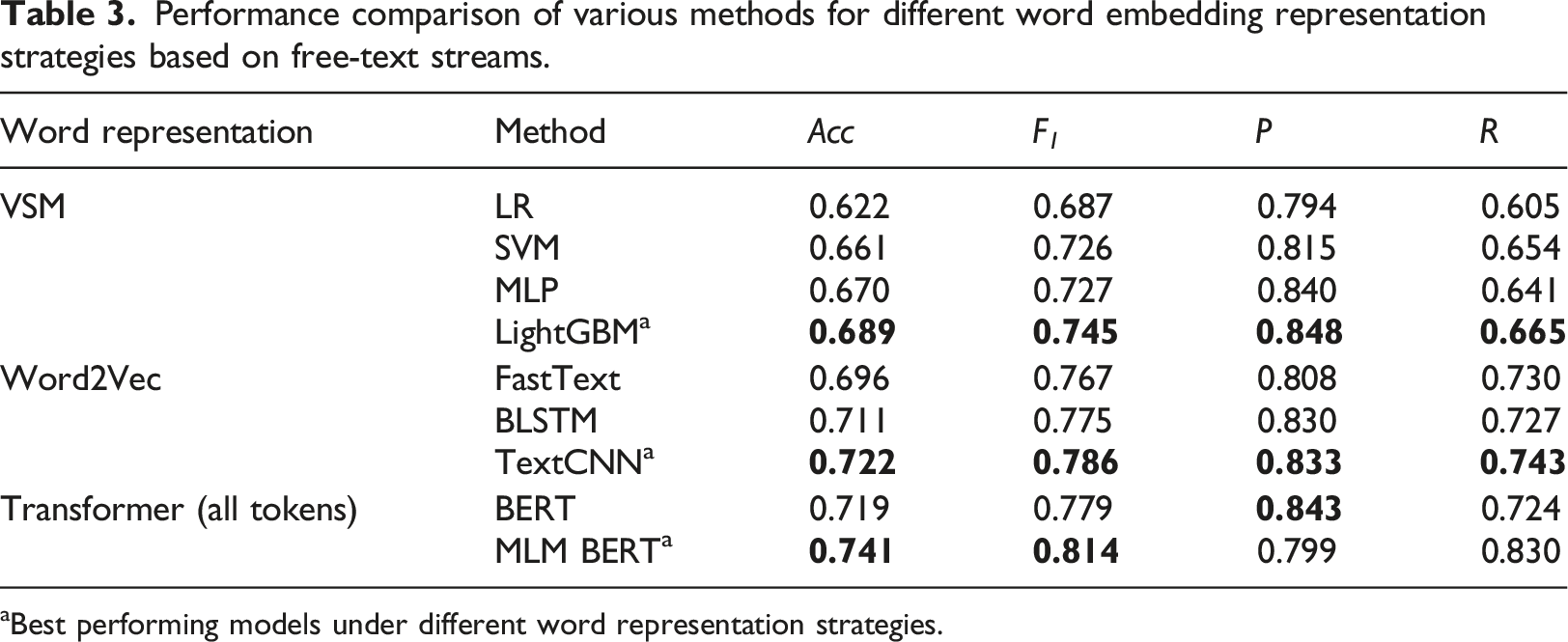
aBest performing models under different word representation strategies.
MP identification based on structured data streams
Performance comparison of different methods based on structured data streams.
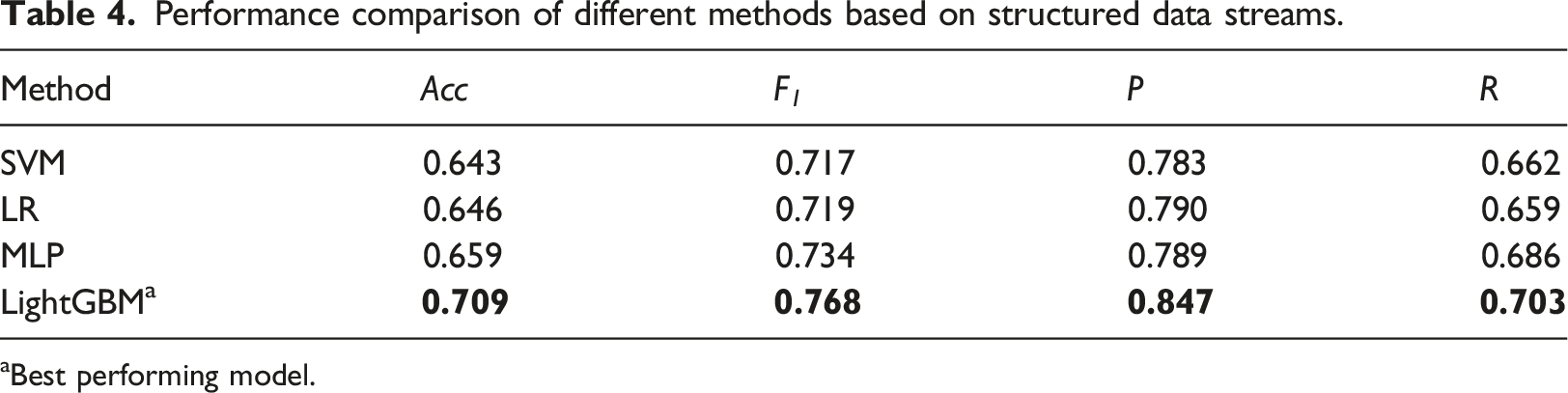
aBest performing model.
MP identification based on fusion of both streams
Performance comparison of methods using different data modalities.
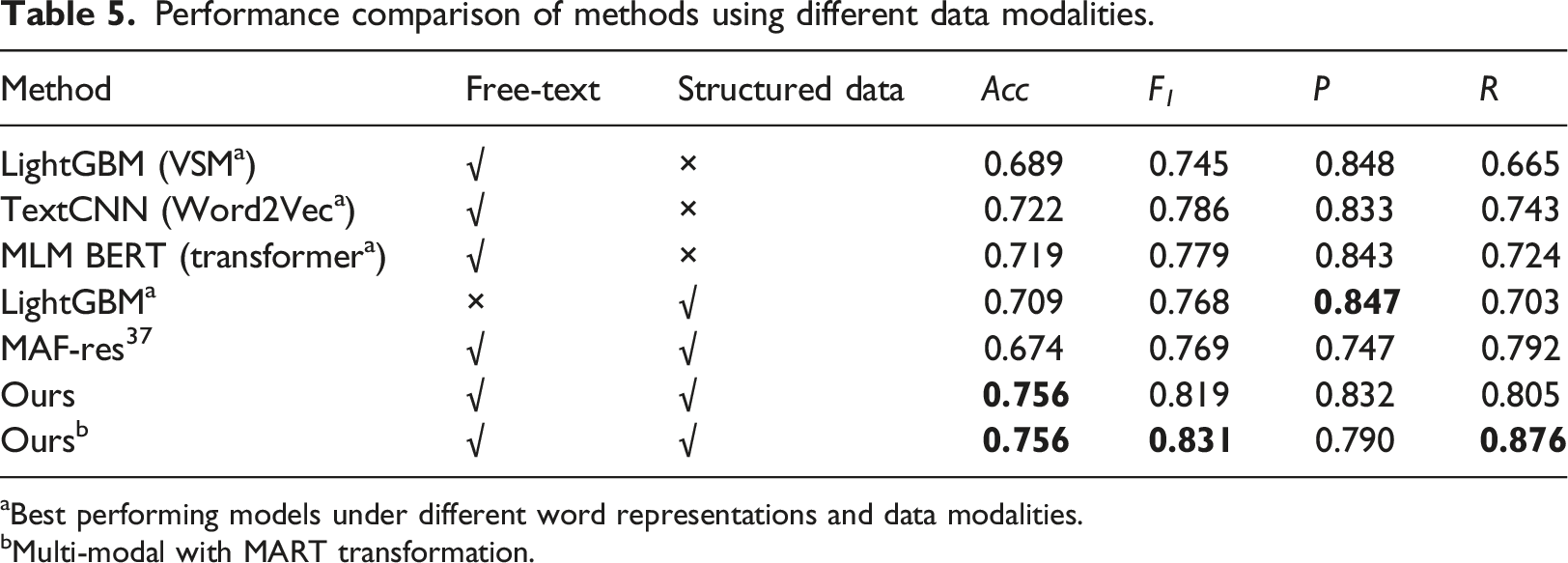
aBest performing models under different word representations and data modalities.
bMulti-modal with MART transformation.
Furthermore, when comparing our model with and without MART transformation (wherein the absence of MART, structured raw data is zero-filled for missing values), there are notable improvements in F 1 , and R (0.831 vs 0.819 in F 1 , and 0.876 vs 0.805 in R), while maintaining equivalent Acc (0.756 in both cases). This underscores the efficacy of tree-based transformation in enhancing model performance.
Discussions
Influence of word representation context on free-text stream analysis
In our study, we analyzed various word embedding representations and their corresponding classification methods, as shown in Table 3. When comparing VSM-based methods with Word2Vec-based methods, we equate MLP and FastText due to their similar complexities as shallow neural networks. FastText, however, demonstrates better performance than MLP (0.696 vs 0.670 in Acc and 0.767 vs 0.727 in F 1 ), indicating Word2Vec’s superior word representation capabilities. VSM’s corpus is restricted to the training set, while Word2Vec benefits from a much larger corpus. Despite VSM’s corpus being more relevant to specific tasks, word frequency alone proves insufficient for capturing the full importance of a word. This limitation includes the inability to reflect the positional information of words and their interrelationships. In contrast, Word2Vec, through unsupervised training on extensive corpora, learns more nuanced word representations, capturing implicit word relationships.
Interestingly, original BERT, despite its wide-ranging successes in natural language processing, does not show a marked advantage over Word2Vec in our analysis and is slightly outperformed by TextCNN. This observation is further evidenced by the improved performance of MLM BERT over the original BERT, suggesting that contextual word embeddings benefit from additional data and training. This leads us to conclude that for specialized domain language tasks like clinical text analysis, pre-training is essential to fully leverage BERT’s capabilities.
Impact of token representations on free-text stream analysis
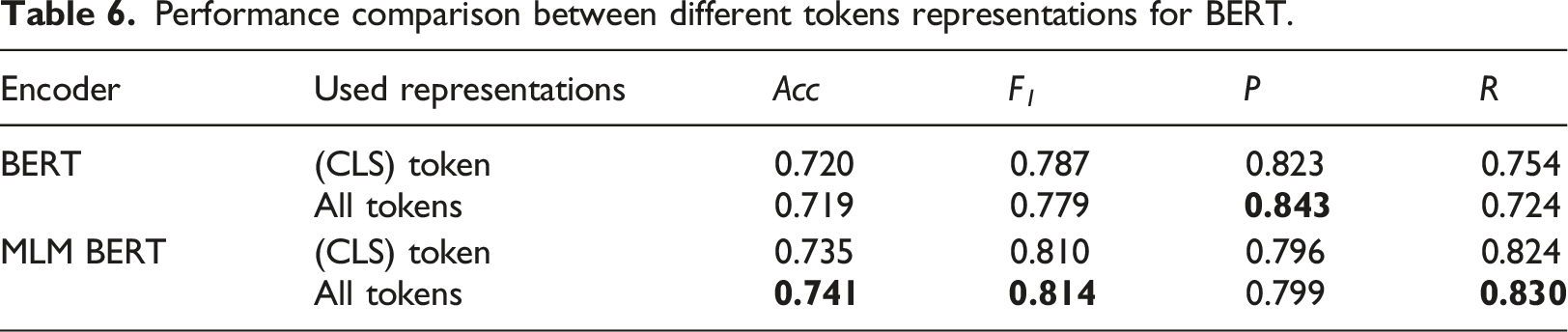
In the utilization of BERT as an encoder for downstream tasks, two primary approaches emerge for token representations. For tasks at the token-level, such as sequence tagging, representations of tokens other than [CLS] are utilized. Conversely, for document-level tasks like text classification, the [CLS] token’s representations are typically employed for classification, often viewed as encapsulating global textual information. However, our research findings indicate a more effective approach in leveraging all token representations for classification purposes.
Performance comparison between different tokens representations for BERT.
Advantages of tree-based feature transformation
The efficacy of tree-based feature transformation is clearly exhibited in the results presented in Table 5. Additionally, as shown in Table 4, MART demonstrate superior performance over neural networks like MLP, particularly when handling structured data, which often exhibits significant diversity in range, variance, and measurement units, and accompanied by numerous missing values as indicated in Table 2. We attribute this advantage to two key factors: a ranking-based splitting principle and robustness to missing values.
Neural networks update parameters based on the gradient derived from the difference between the output and target, thereby, their effectiveness can diminish when dealing with large input feature distributions. Although normalization techniques (such as feature norm and norm layer) are commonly used in neural networks to mitigate issues arising from large magnitude differences in input features, MART’s ranking-based approach is more adept in this context as the depth-based splitting inherent in trees facilitates implicit feature interaction, enhancing the model’s understanding of complex data relationships. Moreover, by transforming continuous values into multiple sets of categorical one-hot features, MART aligns well with neural networks’ strengths, thereby optimizing the subsequent feature fusion process in MAF modules.
The other advantage of MART lies in its handling of missing values. Neural networks typically require missing values to be replaced with fixed values (like zero or the mean), which can inadvertently affect parameter updates through gradient changes. In contrast, MART assesses the impact of missing values by measuring their gain in different branches of the tree. This approach not only maintains sensitivity to the true data distribution but also circumvents the distortions that can arise from arbitrary value imputations.
Comparison with other related work
Previous EMR diagnostic systems, as referenced in,37,46 often rely on word-wise features like word2vec. Such features, while useful, overlook the sequential nature of words, thereby failing to capture the full semantics of sentences. Notably, the superior performance of word-wise features over the BERT model, as reported in ref, 37 may be attributed to the absence of fine-tuning BERT with a specialized medical field corpus. In contrast to these studies, our approach leverages sentence-level embeddings, utilizing the richness of unlabeled corpora to enhance the accuracy of EMR diagnosis. Our methodology, though inspired by the feature fusion module in ref, 37 addresses a critical shortcoming in their numerical feature input: the lack of normalization. Even with normalization, the differing ranges of medical test features could disproportionately influence the model, potentially leading to biased predictions. Our implementation of MART mitigates this bias through a ranking-based split principle for feature transformation.
Limitations
While our method demonstrates high accuracy, its effectiveness might vary in other clinical centers due to differences in clinical text writing conventions. Despite this, we posit that the semantic interpretation of EMRs based on BERT offers better generalizability compared to word-based text understanding. Furthermore, the data preprocessing pipeline in our study includes an intensive manual template design for structured data extraction. This aspect could potentially restrict the practical deployment of our model. Nonetheless, as mentioned earlier, this preprocessing pipeline is not mandatory in situations where structured test data is readily available, particularly via a lab module, and can be directly employed. Finally, the successful application of our model is contingent upon the integration with a hospital’s information system infrastructure, including ETL (Extract, Transform, Load) tools for the automated extraction, preprocessing, and storage of necessary test numerical values.
Conclusions
In our study, we developed and evaluated a hybrid model that integrates clinical free-text descriptions with structured numerical medical test data for identifying MP in CAP patients. The model was trained and validated on a dataset of 7157 EMRs. Our findings show that the proposed model achieves an accuracy of 0.756, an F1 score of 0.831, a precision of 0.790, and a recall of 0.876 on a test set comprising 370 MP cases and 170 non-MP cases. It significantly surpasses the performance of the state-of-the-art MAF-Res method and all leading uni-modal methods across key metrics. This highlights the effectiveness of our approach. Furthermore, our experiments demonstrate the critical role of pre-training and the efficiency of tree-based feature transformation. Looking ahead, potential improvements include expanding the model’s capabilities to identify more CAP sub-types and enhancing its interpretability.
Footnotes
Acknowledgements
We gratefully acknowledge the funding support received from the Key R&D Program of Zhejiang, the National Key Research & Development Program of China, the National Natural Science Foundation of China, and the Hong Kong Research Grants Council through the General Research Fund.
Author contributions
(1) Conceptualization: Qiuyang Sheng, Xiaoqing Liu, Jingna Xie, Yingshuo Wang; (2) Methodology: Qiuyang Sheng, Xiaoqing Liu, Yizhou Yu; (3) Formal analysis and investigation: Yingshuo Wang, Jing Li, Jingna Xie, Gang Yu; (4) Writing - original draft preparation: All authors; (5) Writing-review and editing: All authors; (6) Funding acquisition: Xiaoqing Liu, Yiming Li, Yizhou Yu, Gang Yu; (7) Resources: Jing Li, Fenglei Sun, Yuqi Wang, Shuxian Li; (8) Supervision: Yizhou Yu, Gang Yu.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by grants from the Key R&D Program of Zhejiang (Grant No. 2023C03101), the National Key R & D Program of China (Grant No. 2023YFC2706400 and 2019YFE0126200), the National Natural Science Foundation of China (Grant No. 62076218 and 82171934), Zhejiang Province Research Project of Public Welfare Technology Application (Grant No. LGF22H180004), and the Hong Kong Research Grants Council Through General Research Fund (Grant No. 17207722).
