Abstract
Early identification of patients at risk of hospital-acquired urinary tract infections (HA-UTI) enables the initiation of timely targeted preventive and therapeutic strategies. Machine learning (ML) models have shown great potential for this purpose. However, existing ML models in infection control have demonstrated poor ability to support explainability, which challenges the interpretation of the result in clinical practice, limiting the adaption of the ML models into a daily clinical routine. In this study, we developed Bayesian Network (BN) models to enable explainable assessment within 24 h of admission for risk of HA-UTI. Our dataset contained 138,250 unique hospital admissions. We included data on admission details, demographics, lifestyle factors, comorbidities, vital parameters, laboratory results, and urinary catheter. Models developed from a reduced set of five features were characterized by transparency compared to models developed from a full set of 50 features. The expert-based clinical BN model over the reduced feature space showed the highest performance (area under the curve = 0.746) compared to the naïve- and tree-augmented-naïve BN models. Moreover, models developed from expert-based knowledge were characterized by enhanced explainability.
Keywords
Introduction
Hospital-acquired urinary tract infection (HA-UTI) is the most common type of nosocomial infection,1,2 accounting for approximately 40%, 3 and is associated with increased morbidity, mortality, prolonged length of stay (LOS), as well as additional hospitalization expenditures.1,4 Management of HA-UTI relies on preventive hygienic measures 5 in the context of, e.g., urinary catheter care bundles, 3 which includes a careful clinical indication for use and instrumentation of the device,5,6 as well as rational use of antibiotics. 7
Recently, automated surveillance systems for HA-UTI, such as the Danish Hospital-Acquired Infections Database (HAIBA), 8 have been developed and implemented to monitor the incidence rates and enable benchmarking between Danish hospitals. 9 Additionally, studies mapping risk factors for HA-UTI have been widely conducted to create awareness of patterns leading up to HA-UTI.10–13 Notably, potential complex patterns of mutual risk factors may be used at admission to identify patients at risk of acquiring HA-UTI. 14 Machine learning (ML) may potentially be suitable for identifying such patterns.15–18 However, existing ML models in infection control – and healthcare in general –often demonstrate a lack of explainability and interpretability. 15 If the clinicians are challenged in interpreting the output of a given ML model, this may hamper the implementation of the ML model in daily clinical routines.15,16,18–25
Explainability of ML models for healthcare can be classified as being either model-specific or model-agnostic. 26 For instance, ML models, such as logistic regression or decision trees, are said to be explainable by nature as they allow direct insight into the inner mechanics of the model. We refer to such ML models as model-specific explainability. 26 On the other hand, ML models, such as neural networks or random forests, do not allow for immediate insights into the inner mechanics of the models, which is why they are often referred to as black-box models. For explainability, black-box models may benefit from a surrogate model, e.g., feature importance in the context of local prediction. We refer to such expression as model-agnostic explainability. 26 Examples of methods providing model-agnostic explanations of predictions include SHap Additive exPlanations (SHAP) 27 and Local Interpretable Model explanation (LIME). 21
A recent study by Rudin 28 advocates for ML models that enhance interpretability by promoting transparency and, e.g., using only the most meaningful features as predictors for the event of interest. A suitable ML model class that supports these design desiderata may be found within probabilistic graphical models, exemplified by Bayesian Network (BN) models.22,29–31 BN models support reasoning under uncertainty,22,32 relying on a transparent model specification that enables the clinician to inspect model assumptions in the form of probabilistic independencies and possibly causal statements, in addition to explanations of evidence.22,30 BN models also support the integration of expert knowledge in both data preprocessing and model development, which can be decisive for the successful adoption of ML models in daily clinical routines.23,24
In this study, the aim is to: (1) Capture probabilistic dependencies and independencies in BN models for stratification of patients within 24 h of admission for risk of HA-UTI. (2) Support BN model specification by combining clinical expert knowledge and clinical data to decide on relevant HA-UTI risk factors and guide model structure development. (3) Compare performance and discuss the explainability/interpretability of different types of BN models: 1. Models based on clinical expert-based knowledge, 2. naïve Bayes models, and 3. Tree-augmented-naïve (TAN) models.
Literature review
Studies on early identification of HA-UTI using explainable ML models are sparse.
Møller et al. 14 developed predictive models for HA-UTI based on data from admission time and within the first 48h of admission, respectively, reaching an Area Under the Curve (AUC) between 0.709 and 0.770. However, the study did not explore the meaning of explainability for their models. Zhu et al. 33 investigated ML models in the prediction of poststroke urinary tract infection (UTI) risk in immobilized patients, using SHAP for model-agnostic explainability, but without addressing model-specific explainability and only targeting a subgroup of HA-UTI. Jeng et al. 34 examined ML for predicting recurrent UTIs caused by Escherichia coli, demonstrating model-specific explainability through decision tree splits. However, the decision tree model achieved an AUC of 0.654 without the inclusion of, e.g., clinical expert knowledge in the model development. Taylor et al. 35 compared predictive performance between seven ML algorithms for community-acquired UTI, and Yelin et al. 36 used logistic regression and gradient boosting modeling to identify antibiotic resistance. However, they did not target HA-UTI nor discuss model-specific explainability of their models and predictions. Jakobsen et al. 37 explored model-agnostic explainability for early identification of HA-UTI but without delving into the model-specific explainability.
A few studies have applied BN models for infectious diseases.32,38,39 To our knowledge, no study has investigated early stratification for the risk of HA-UTI and addressed model-specific explainability in this context. Recently, Gupta et al. 32 applied a naïve BN and a TAN BN to assess the risk of sepsis by capturing dynamics between biomarkers, challenging the performance of existing scoring systems for sepsis. Ward et al. 38 developed a clinical decision support system (CDSS) named SepsisFinder, demonstrating how structures constructed by expert-based knowledge may effectively capture accurate dependencies; SepsisFinder is considered an extension of TREAT, a CDSS for guided antibiotic therapy in Denmark.32,40 Schrurink et al. 41 developed a BN to assist in diagnosing and treating nosocomial infectious diseases. They concluded that clinical experience with CDSS is still limited. Lastly, Visscher et al. 39 developed a BN model from expert-based knowledge for predicting pathogens causing ventilator-associated pneumonia, reaching an AUC between 0.51 and 0.77 for different pathogen groups when only relying on admission duration and ventilation, which significantly improved when including information on culture results. However, this study neither targeted HA-UTI nor addressed explainability. To our knowledge, BN models have never been applied for early risk-stratification of HA-UTI, nor have explainability been addressed in this context.
Materials and methods
Source of data and participants
Danish legislation requires no approval from an ethics committee or consent from participants for registry-based studies.
Information on 50 features from 138,250 admissions is included in this study. We include data on adults between 18 and 100 years of age with a LOS between 0.1 and 365°days(s) at North Denmark Regional Hospital or Aalborg University Hospital in Denmark from 1 January 2017 to 31 December 2018. Data are linked using the unique civil personal registration (CPR) number issued to all Danish citizens together with the timestamps for the hospital admissions. A patient can appear more than once in the dataset if the patient has experienced multiple admissions at different time points during the 2-year study period. Statistical Analysis Software Enterprise Guide 8.1 is used for data management. Hugin Expert 8.9 is used for BN model development and model analysis.
Definition of Hospital-acquired urinary tract infection outcome
HA-UTI is included as a binary feature in the dataset. HAIBA registers new laboratory-diagnosed cases of HA-UTI if at least one urine culture is positive, including ≤2 microorganisms with at least 104 colony-forming units (CFU)/mL and at least one microorganism above 104 CFU/mL urine, measured between 48 h after hospital admission and 48 h after hospital discharge. Moreover, no case of UTI must be registered <14 days prior to a new case of HA-UTI.
Data preprocessing
Figure 1 Illustrates the data landscape. Illustration of the data landscape. Note: A combined code for patient ID, date, time, and the place was used as a unique primary key for each admission.HA-UTI (Hospital-acquired urinary tract infection).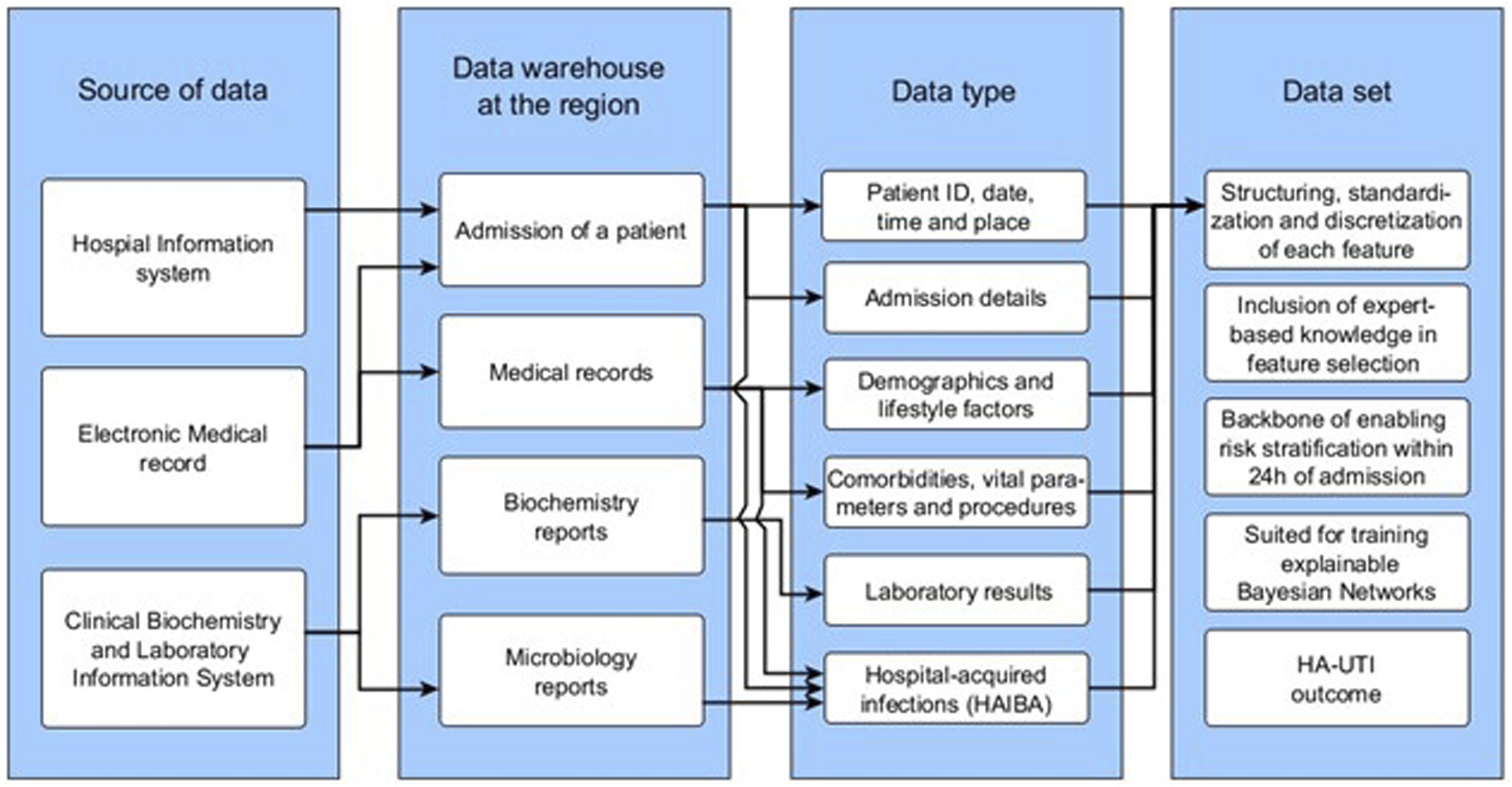
In Denmark, the medical condition leading to hospital admission is registered as an A-diagnosis, and for each additional condition, a B-diagnosis is added. Since 1994, diagnoses have been registered in accordance with the International Classification of Diseases, Tenth Revision (ICD-10). Diagnoses registered before 1994 are registered in accordance with the ICD-8. We choose a level of three digits in ICD-10 for A-diagnosis for the Admission cause-feature, resulting in a nominal feature of 19 categories. The ICD-10-codes used for admission cause are presented in Appendix A (supplementary).
The Charlson Comorbidity Index (CCI) 42 is used to assess comorbidities. Moreover, we add a feature of history of UTI because we consider patients to be more prone to acquiring a new incidence of HA-UTI if they have a past history of UTI. 35 The study does not include the CCI-score or the 10-years survival chance. Once again, we choose a level of three digits in the ICD-10- and ICD-8-structure (some comorbidities were also registered before 1994) for Comorbidities, as presented in Appendix B (supplementary).
Feature selection
The expert-based knowledge incorporated into this study is provided by a consultant in infectious diseases and a consultant in clinical microbiology, both of whom have vital insight into clinical routines. In addition, findings from previous studies on risk factors for HA-UTI10–14 were also taken into account.
Feature selection.
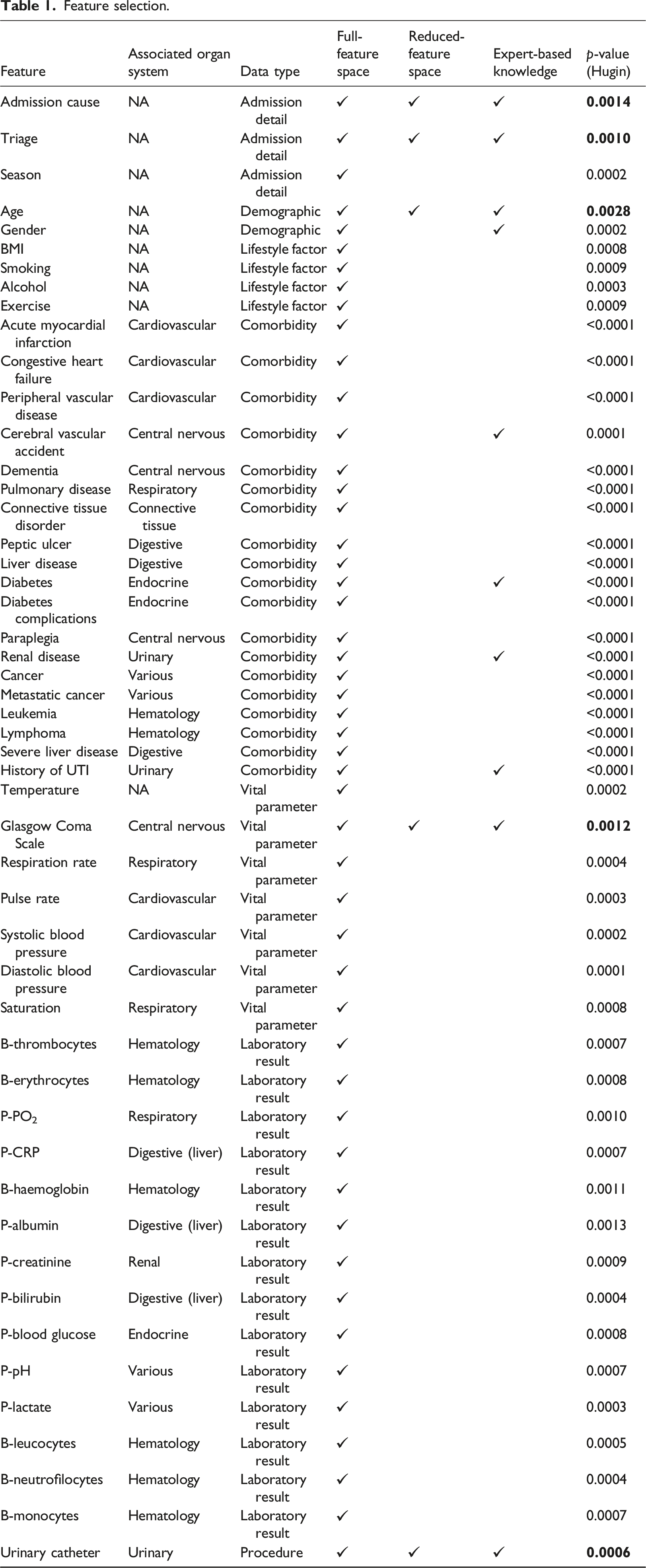
We randomly split the data into a training set containing 110,870 unique hospital admissions and a test set containing 27,380 unique hospital admissions.
Discretization
All features in the dataset are discretized from levels 0-6 according to the considered contribution for prediction of HA-UTI (Appendix C, Table C), based on clinical expert knowledge and previous literature, e.g., existing scoring systems used in the intensive care unit. 44 Feature values in intervals corresponding to the discrete level 6 are considered more associated with higher HA-UTI risk than the discrete level 0. If multiple values for the same feature are present within 24h of admission, only the value that is categorized in the highest discrete level is considered for our model. Values outside the discrete levels are deemed outliers and are omitted.
Bayesian network models
A BN model consists of a qualitative and a quantitative part and provides a compact representation of a joint probability distribution. The qualitative part of a BN model is represented by a directed acyclic graph (DAG), where the nodes correspond to random variables, and the directed edges specify dependence/independence relations between the variables. Note that we will use the terms variable and node interchangeably in the remainder of this paper. The quantitative part of the model captures the strengths of the dependence relations through a collection of conditional probability tables (CPT), one for each variable in the model. Together, these CPTs define a joint probability distribution over all the variables in the model. 45
We construct and evaluate eight BN models with different model structures. Specifically, we consider a model structure specified in collaboration with domain experts, referred to as a clinical model, as well as three model structures learned from data with varying levels of complexity expressed in terms of allowed conditional dependencies. The four modeling approaches are applied to both the full- and the reduced feature space, respectively, with all model parameters, i.e., conditional probabilities, being learned using the expectation-maximization (EM) algorithm. 46 The EM algorithm finds a local maximum likelihood estimate of the model parameters by iteratively alternating between an expectation step (involving inference in the underlying model), and a maximization step, where intermediate maximum likelihood parameter estimates are found based on fractional counts derived from the expectation step. 45
All BN models focus on performing stratification within 24 h of admission for the risk of experiencing HA-UTI, as illustrated in Figure 2. We will therefore refer to HA-UTI as the target variable in the following sections to distinguish it from the other model variables. Also, in the remainder of this paper, we will refer to a model defined over the reduced feature space as a reduced model and a model defined over the full features space as a full model. Illustration of how our Bayesian networks perform risk-stratification within 24h of admission for HA-UTI. Note: Within 24h of hospital admission of a new patient (represented in the test set), the model uses all available patient data on relevant features to calculate the probability of acquiring HA-UTI. The red dashed timeline indicate the period outside the timeframes for HAIBA’s HA-UTI definitions, whereas the black timeline indicates the period inside. From a probabilistic decision boundary, a threshold, the BN classifies the patient as either ‘at risk patient’ or ‘not at risk patient’. The decision boundaries can be altered, e.g., due to preference. In all, this makes up our risk stratification. In the test of performance, we demonstrate how well the model classifies the patients. We also discuss the model-specific explainability related to our eight Bayesian Network models in our work. HA-UTI (hospital-acquired urinary tract infection), TP (true positive), TN (true negative), FP (false positive), and FN (false negative).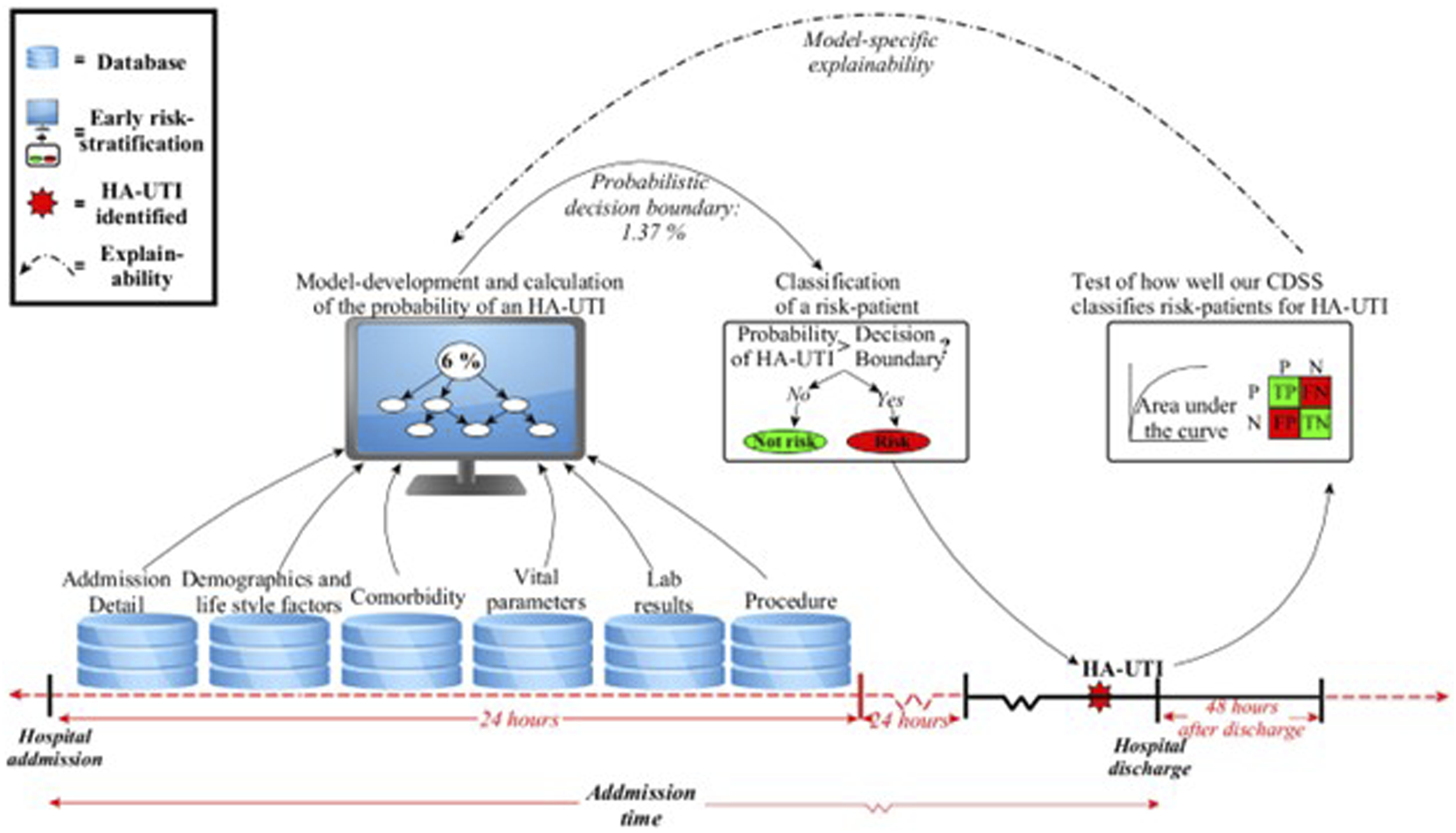
Expert-based clinical model structure
The expert-based clinical BN models were constructed using expert-based knowledge in a three-step approach. In the first step, we asked clinicians to decide on distinct dependencies between feature variables while excluding the HA-UTI target node and respecting the DAG properties. This resulted in, e.g., demographics and lifestyle factors being defined as parents to the comorbidities, which in turn is associated with parents to vital parameters and laboratory results. For instance, in the full feature space, we have body mass index (BMI) as parent of diabetes, alcohol status as parent of liver disease, diabetes as parent to level of blood glucose, and renal disease as parent to level of creatinine. In step two, we inspect the edges in the structure from step one and evaluate the suggested dependencies. In general, an edge between two nodes is altered if the implied dependency does not align with the clinical expectations. For example, the dependencies conveyed by edges between triage level and the use of urinary catheters to monitor urine output may be better described by an underlying disease or condition (meaning it is not a worsening in the triage that directly leads to the use of a urinary catheter). On the other hand, if an edge between two nodes reflects the expected clinical dependency, such as the Glasgow Coma Scale (GCS) affecting the triage level, the edge remains in the structure. We repeat this step for all edges in the model (in random order). In step three, we naively assume conditional independence between all the features and the HA-UTI target node by setting the HA-UTI node as the parent of all features. This ensures a tractable model structure, although we also recognize that the resulting edges incident to HA-UTI does not reflect causality and may violate expected clinical (in)dependencies. For the full feature space, we subsequently also remove the edges between comorbidities and HA-UTI, as preliminary experiments showed an improvement in AUC for this refined model structure. Figure 3 illustrates the process of developing the expert-based clinical model structures. Illustration of the three steps in developing our expert-based clinical Bayesian network model. Note: In 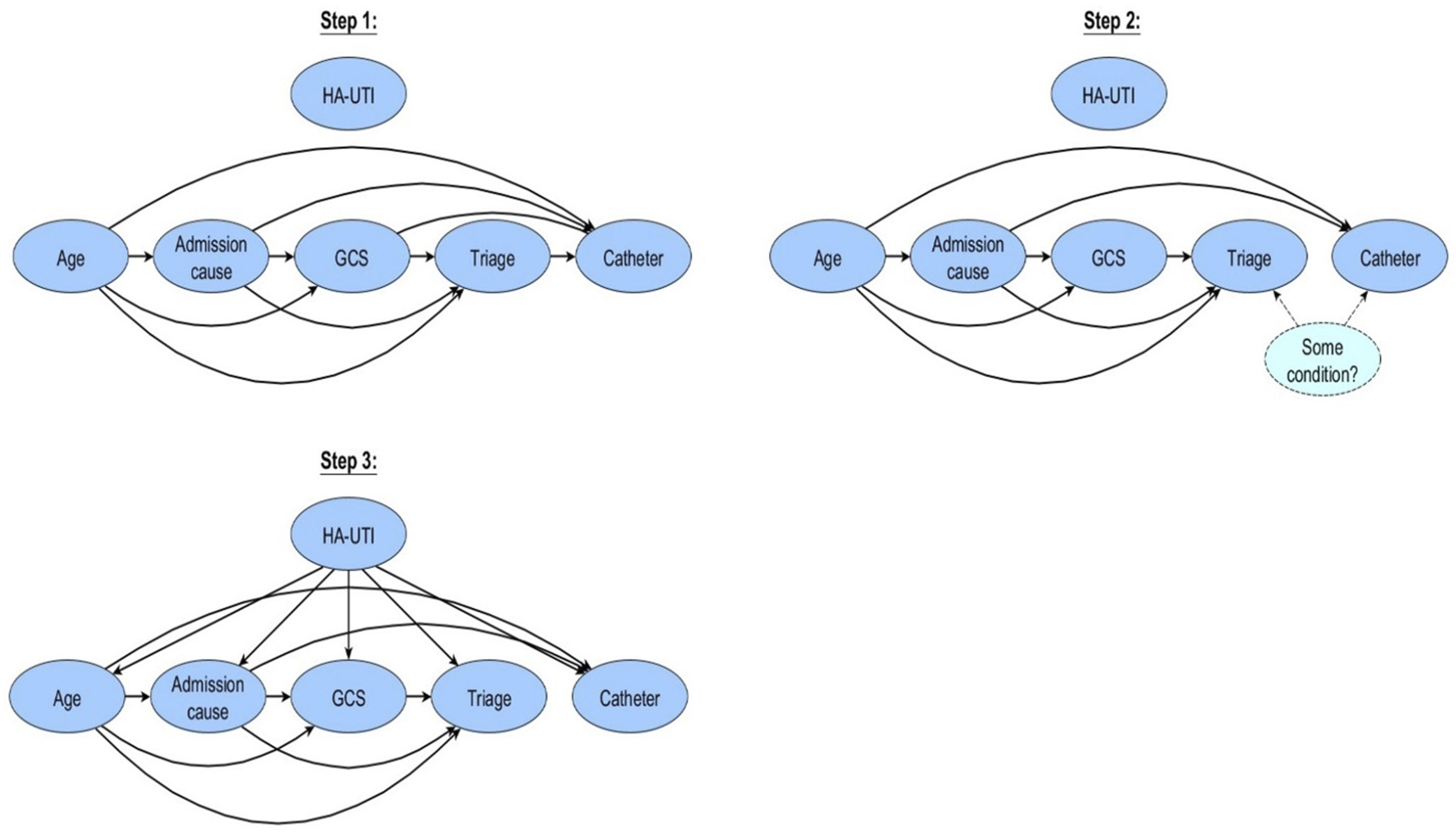
Structure learning from data
We consider three approaches for learning BN models from data using both the reduced and full feature space. The underlying algorithms allow increasing levels of complexity in the model structures and cover naïve Bayes models,32,45,47 TAN models, 31 as well as models learned using the PC algorithm. 48
Given the target variable HA-UTI, the naïve BN model assumes conditional independence between all features. This assumption implies a model structure where all edges are directed from HA-UTI to the other variables in the model.32,45 The expert-based clinical-, naïve- and TAN model structure implies a naïve assumption to the HA-UTI target node. However, given the HA-UTI target node, they distinguish by the edges between the remaining features in the structure.
The TAN BN augments the naïve Bayes model by defining a tree structure over the feature variables (thus, not including the target HA-UTI), such that each node (except the root and target node) has two parents. 32 The first step of the TAN algorithm is to construct a weighted, fully connected, undirected graph over the features (excluding the target HA-UTI), where the edge weights correspond to the mutual information between the corresponding variables conditioned on the target variable HA-UTI. Next, a maximum weighted spanning tree is constructed, retaining the edges with the highest conditional mutual information between variables, such as the edge connecting the age node and admission cause node (Figure 5). We (somewhat arbitrarily) select the Age node as root node and direct all edges in the spanning tree away from age, thereby obtaining a directed tree structure. Finally, HA-UTI is included in the model as parent to all other nodes. 31
The PC algorithm
45
is a constraint-based algorithm relying on local conditional independence tests between variables.
48
The initial step of the PC algorithm is to construct a fully undirected network. An iterative second step performs conditional independence tests for all pairs of features, removing edges between nodes deemed (conditionally) independent. In step three, so-called v-structures in the model are identified based on the independence test performed during the previous step. Finally, in step four, any remaining undirected edges are oriented while respecting the DAG properties and avoiding new v-structures.45,48,49 Figure 4 llustrates the process of developing both the TAN- and the constraint-based PC model for early risk stratification of HA-UTI. Illustration of the four steps in the development of our tree-augmented-naïve- and PC Bayesian Network model. Note: In 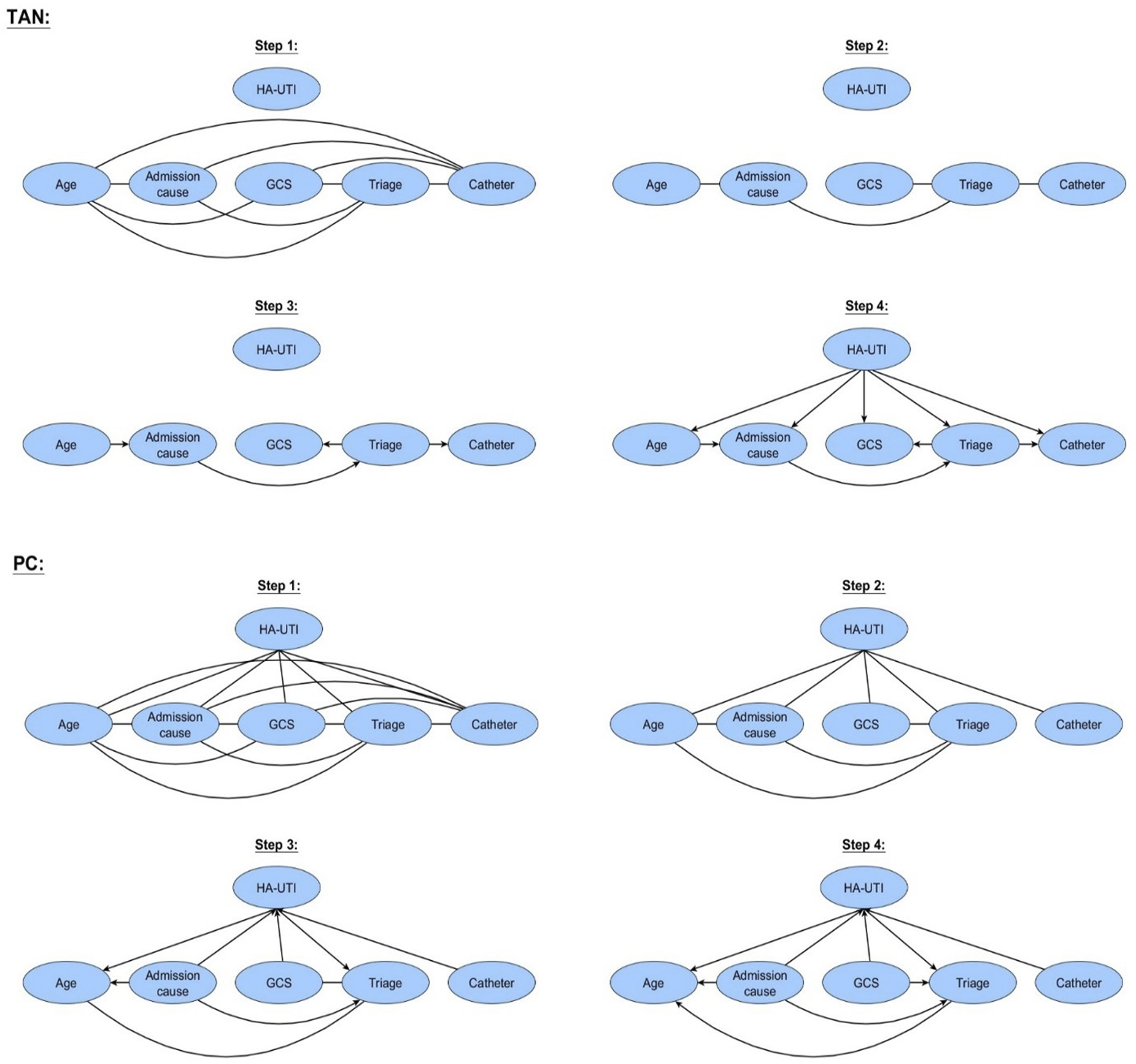
Evaluating the models
We report on model performance using the Reciever-operating-Curve (ROC) with an associated AUC and confusion matrix. We also discuss the model-specific explainability of the BN models concerning the degree to which the dependency structure of the models accurately represents the domain of HA-UTI.
Results and discussion
Of the 138,250 admissions included in this study, 1,889 (1.37 %) admissions include at least one HA-UTI. Percentage frequency distributions and missingness for gender, smoking-, alcohol-, and exercise status, comorbidities, and presence of urinary catheter, are presented for cases with and without HA-UTI for both the training- and the test datasets. In addition, the median and IQR for age, BMI, vital parameters, and laboratory test results in training- and test set, respectively (Appendix D, supplemental).
Model performance
Figure 5 demonstrates the overall performance of the models using ROC curves. The figure and the subsequent discussions do not include the results of the model learned using the PC algorithm over the full feature space, as the learned model exhibits complex dependency structures for which parameter learning is not feasible with the available data. We expect that this is due to the dataset not covering all relevant variables, including a representation of the underlying admission cause, but a more rigorous analysis of this issue is outside the scope of the present paper. Illustration of the ROC for our models. Note: The left ROC curves represent the performance of the Bayesian network models developed over the reduced feature space. On the contrary, the right ROC curves represent the performance of the BN developed over the full feature space. The blue curves are the expert-based models, the yellow curves are the naïve models, the black curves are the TAN models, and the red curves are the PC model.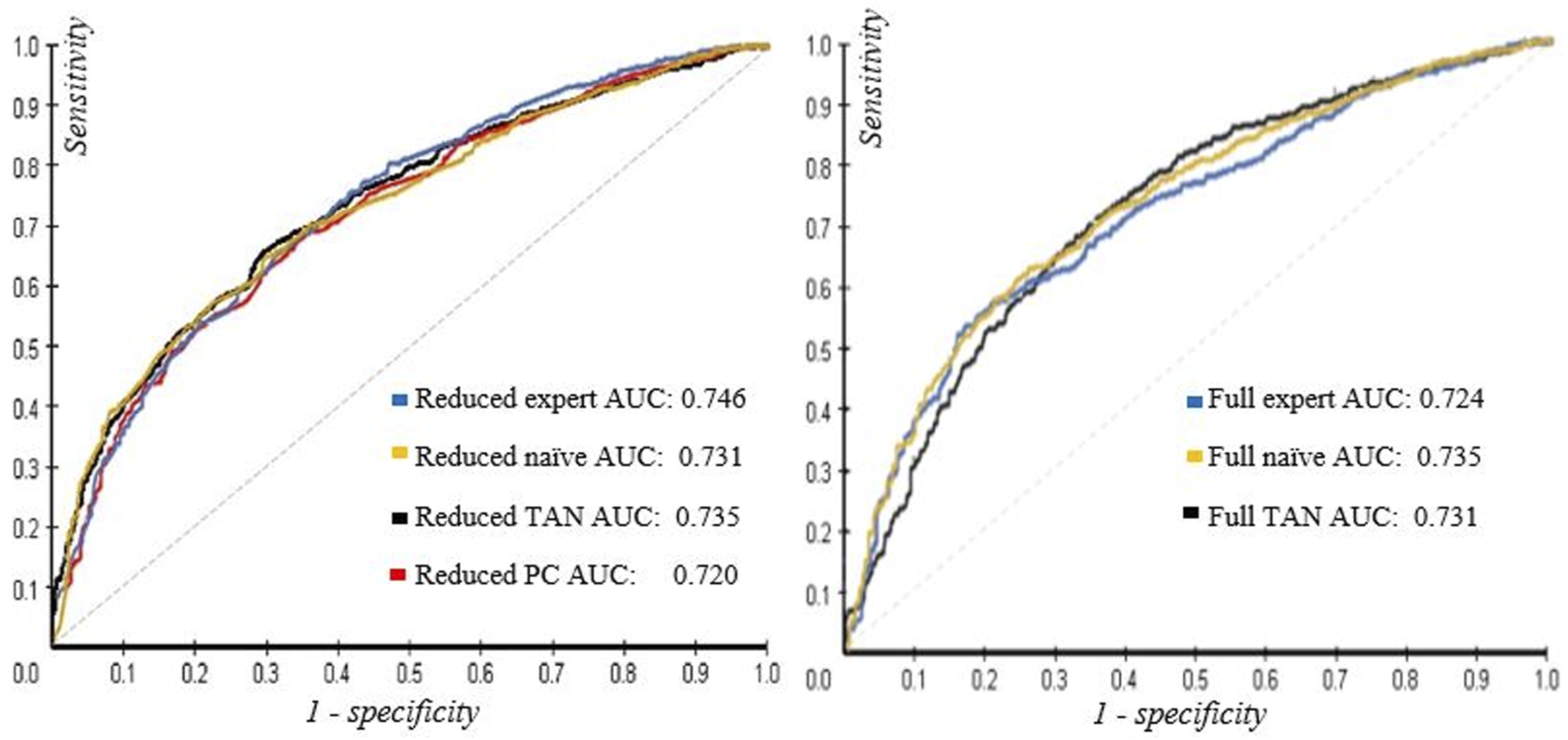
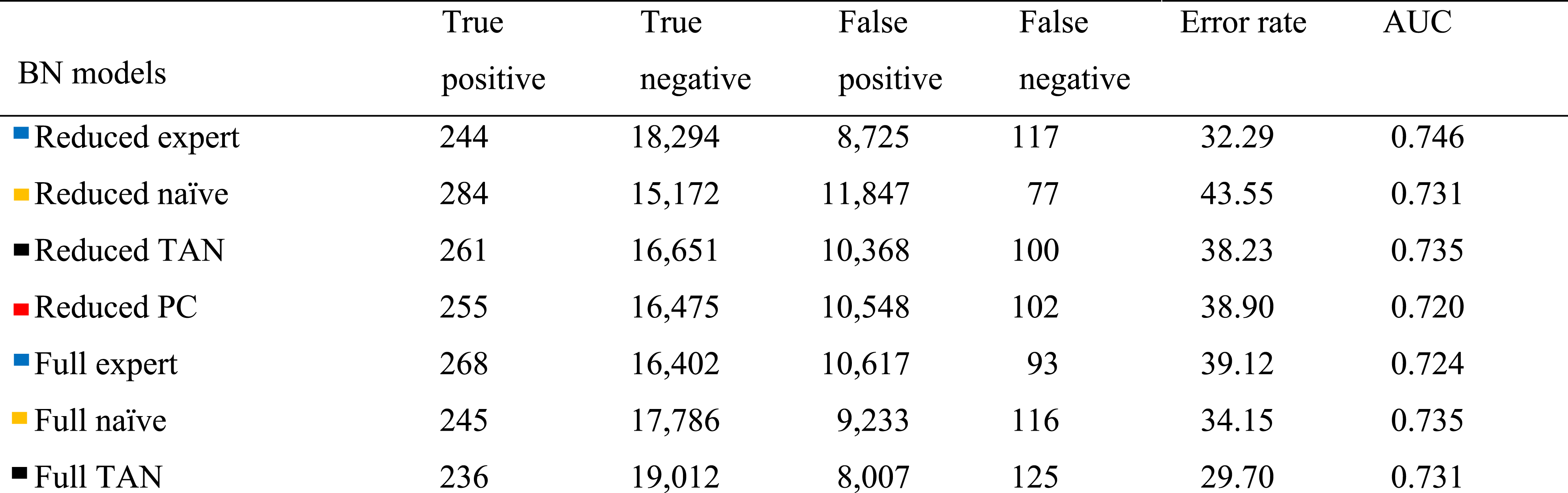
Confusion matrix.
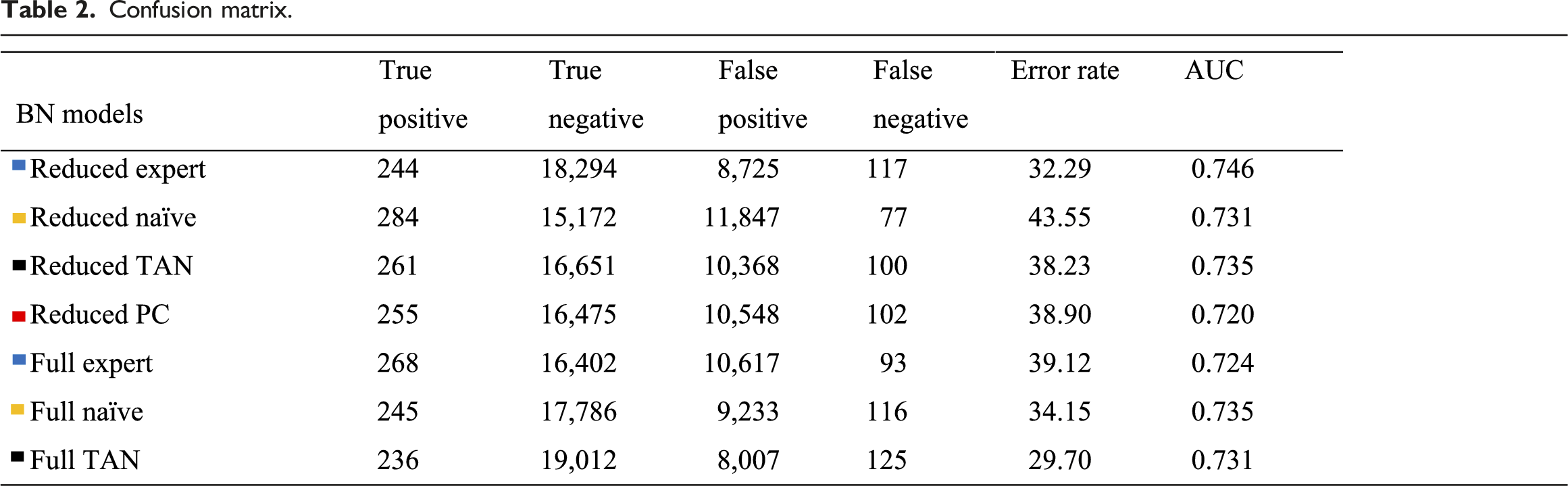
The best-performing model in terms of AUC is the reduced expert-based clinical BN model with an AUC of 0.746. The full naïve BN- and the reduced TAN BN models performed second-best with AUCs of 0.735. The reduced expert-based clinical- and the full TAN BN models also demonstrate the lowest error rates of 32.29% and 29.70% for the decision threshold, respectively (Table 2). Note that a model that never classifies a patient at risk of HA-UTI will have a low error rate of only 1.37%, as it will correctly identify all non-HA-UTI cases. However, this model will also be of no clinical value.
For the given decision threshold, the reduced naïve BN model reaches the highest number of true positives (TP) with 284 patients correctly identified as developing HA-UTI (of 402 HA-UTI cases in the test set). However, this model also has the second-highest number of false positives (FP), meaning the model tends to be more aggressive in classifying a patient as being at risk of HA-UTI, which also describes the high error rate of 43.55% (second highest number of FP of all BN models). The full expert-based clinical model reaches the second-highest number of TP but without the high error rate caused by a high number of FP (Table 2). The reduced expert-based clinical- and full TAN BN models reach the highest number of true negatives (TN) of 18,294 and 19,012 and the lowest FP of 8,694 and 8,007. The reduced naïve BN model demonstrates the lowest number of false positives (FP) of 77, followed by the full expert-based clinical BN model with a number of FP of 93. Note that an FN (i.e., not initiating preventive strategies for a patient at risk of HA-UTI) is considered more critical than an FP, where an unnecessary preventive strategy is initiated. The reduced PC BN model has a performance with an AUC of 0.720 (Table 2), which is the second-worst AUC of the BN models.
Bayesian networks and explainable network structures
In this study, we suggest risk-stratifying patients within 24h of admission (Figure 2) using a strong case definition of the target HA-UTI that combines multiple high-quality administrative and clinical data sources.8,9 The nodes in the reduced feature space are identified using expert-based knowledge in addition to feature selection based on tests of marginal independence. Using expert-based knowledge in both feature selection and model development may be decisive for applications of ML models in a daily clinical routine.23,24 We use BN models in this study as these types of models naturally support certain types of explanations/interpretability through analysis of the assumed dependencies within the models as well as principled reasoning mechanisms .22,50 In what follows, we discuss potential insights and explanations derived from the learned model structures, particularly to what extent the models reflect the HA-UTI domain in terms of the existence and orientation of the directed arcs that define the model structures. Appendix E (supplementary) illustrates the four BN models defined over the reduced feature space.
For the expert-based clinical BN, a trade-off is made between capturing correct conditional dependencies (i.e., supporting model-specific explainability) and potentially pursuing higher performance in terms of AUC. For instance, we expect the age-, triage-, and urinary catheter nodes to be parents of the HA-UTI target node because higher age can be associated with a weakening of the immune system at both cellular and humoral levels, and adverse triage reveals an ailing severity of the acute condition of the patient. In addition, the use of a urinary catheter is associated with intra-luminal transmission of bacteria as well as forming of biofilm, which together cause a higher risk of acquiring HA-UTI. However, by reversing the edge between the features and HA-UTI in the expert-based clinical model, which conforms to the relationships expected in the underlying domain, the AUC in the performance test reduces to 0.697 from 0.746. Note that the reduced PC BN models have a higher number of conditional dependencies similar to those from the reduced clinical expert BN model (Appendix E, supplemental). Appendix F (supplementary) illustrates the four BN models developed over the full feature space.
The naïve BN models do not capture the correct clinical conditional dependencies, which is an immediate consequence of the conditional independence assumption between the feature variables given the target HA-UTI variable. Despite the TAN BN model capturing edges from the age node to the admission cause node, and from the admission cause node to the triage node, these edges’ orientation results from orientating all edges away from the root node in the tree structure (Figure 4). The PC algorithm over the reduced feature space (Appendix E, supplemental) captures the admission cause-, GCS- and urinary catheter node as parents to the HA-UTI target node, which may be explained by admission causes such as neurological- or renal diseases can be associated with decreased bladder function (increase in the risk of recurrent urine), that may increase the risk of HA-UTI. Thus, severe admissions may be associated with adverse GCS and an increased likelihood of using a urinary catheter to monitor the urine output, which may also cause a higher risk of acquiring an HA-UTI.
Limitations and strengths
Predicting events occurring >48 h after admission within 24 h of admission may be challenging because the outcome is likely to be affected by many confounders, such as an underlying disease or given intervention, in a potentially large timeframe between prediction time and the adverse event (Figure 2). However, we see significant clinical potential in the ability to identify risk patients at early onset, because this may allow for initiation of preventive hygienic measures, such as decreasing the duration of potential catheterization, or for supporting suspicion of HA-UTI during the course of admission, e.g., if the patient experiences a fever at a later onset.
Unfortunately, we did not have access to information on antibiotic therapy, a well-established risk factor for HA-UTI. 11 This knowledge may significantly impact the HA-UTI risk, leading to better conditions for identifying risk patients for the BN models, potentially resulting in improved performance in terms of higher AUC. Future BN models might be extended with information about antibiotic therapy, motivated by Leibovici et al., 51 who included information on antibiotic therapy as a parent node to a node representing a positive bacterial culture result in a BN model. Such an extension could also easily be accommodated by the models in the present study.
Conclusion
This study evaluates the use of BN models for risk stratification within 24 h of admission for HA-UTI to enable timely targeted preventive and therapeutic strategies. We present how risk stratification within 24 h of admission may be performed by BN-based ML models having transparent model structures that can provide a degree of model-specific explainability for the clinical personnel. We considered two feature spaces as the basis for model construction: 1. A full feature space including data on admission detail, demographics, lifestyle factors, comorbidities, vital parameters, laboratory results, and procedures. 2. A reduced feature space (selected based on expert-based knowledge and the tests of marginal independence), which indicates that admission cause, age, GCS, triage level, and urinary catheter are the most essential features of model construction for risk stratifying HA-UTI. In all, seven out of eight BN models with model structures either fixed a priori (a naïve Bayes model), learned from data (using the TAN and PC algorithms), or derived from expert knowledge, reached AUC scores between 0.746 and 0.720. This work’s reduced clinical BN model reached the highest AUC of 0.746. Notable, the reduced feature space only has five predictive features. For comparison, Møller et al. 14 performed risk stratification after 48h of admission for HA-UTI using >20 variables, reaching an AUC of 0.735 and 0.701 for their logistic regression and decision tree, which are model classes also considered explainable. 26 Their black-box neural network reached an AUC of 0.770, but this model class does not directly support model-specific explainability.
The findings are in line with studies suggesting a trade-off between performance and explainability52,53 in contrast to studies suggesting that a trade-off is not necessary at all22,54,55 – at least for model-specific explainability for risk stratification within 24 h of admission for acquiring HA-UTI. Moreover, the study may serve as proof of concept for how BN models might constitute a promising choice for improving both performance and explainability in future ML applications for infection control.
Supplemental Material
Supplemental Material - A study on the risk stratification for patients within 24 hours of admission for risk of hospital-acquired urinary tract infection using Bayesian network models
Supplemental Material for A study on the risk stratification for patients within 24 hours of admission for risk of hospital-acquired urinary tract infection using Bayesian network models by Rune Sejer Jakobsen, Thomas Dyhre Nielsen, Peter Leutscher and Kristoffer Koch1 in Health Informatics Journal.
Footnotes
Acknowledgment
The authors are grateful for the academic feedback and inspiring inputs from professor emeritus Steen Andreassen and the technical assistance in data management from consultant Andreas Egmose at the Business Intelligence and Analysis Unit in the Region North Denmark.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Region North Denmark Health Innovation Foundation. This study was conducted as a retrospective cohort data analysis in a collaboration between Centre for Clinical Research at the North Denmark Regional Hospital, Department of Computer Science at the University of Aalborg in Denmark, and Business Intelligence and Analysis Unit in the North Denmark Region, providing data from the electronic health records. Legal representatives approved the project of the North Denmark Region (ID-number 2019-46). Collaboration was also established with Statens Serums Institut in Denmark, providing data on HA-UTI in HAIBA.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.