Abstract
Colorectal cancer incidence has continually fallen among those 50 years old and over. However, the incidence has increased in those under 50. Even with the recent screening guidelines recommending that screening begins at age 45, nearly half of all early-onset colorectal cancer will be missed. Methods are needed to identify high-risk individuals in this age group for targeted screening. Colorectal cancer studies, as with other clinical studies, have required labor intensive chart review for the identification of those affected and risk factors. Natural language processing and machine learning can be used to automate the process and enable the screening of large numbers of patients. This study developed and compared four machine learning and statistical models: logistic regression, support vector machine, random forest, and deep neural network, in their performance in classifying colorectal cancer patients. Excellent classification performance is achieved with AUCs over 97%.
Introduction
The incidence of colorectal cancer (CRC) increases beginning at the age of 50. 1 Screening for CRC is widely recommended for those of average risk in this age group.2-4 As a result, incidence has continually fallen among those 50 and older. 5 CRC incidence increased by 50% in the age group 40–45 over the years 1987 to 2006. 6 Patients younger than 50 years tend to have a worse prognosis, presenting with more advanced disease.7-10 For these reasons, several guideline organizations now recommend age 45 for screening.11,12 Even if fully implemented and followed, half of all cases of CRC in persons younger than age 50 would be missed because of CRC instances in those below the age of 45. A method is needed to identify other high-risk individuals.
Traditionally, domain experts have identified risk factors by review of all available data on a subject. In medical settings, this is generally accomplished through a process of chart review;13,14 chart review is labor-intensive, requiring large time commitments from skilled researchers and clinicians. Automating some of that effort can make the process more efficient, and more information can potentially be analyzed.
There are several prior studies related to the identification of CRC cases using EHR data.15–18 The most pertinent to our study is a 2011 study by Xu et al. 19 It describes an algorithm combining machine learning and natural language processing to detect patients with colorectal cancer (CRC) from entire EHRs at Vanderbilt University Hospital. The algorithm achieved an excellent F-measure of 0.93. This study, however only focused on identifying CRC cases that had CRC ICD codes or CRC diagnosis in free text notes. It also did not take any risk factors into count. In contrast, the algorithm we report in this paper does not depend on the CRC diagnostic codes because our ultimate goal is to identify high-risk individuals for targeted screening who would not have CRC diagnosis codes present.
Known risk factors for early onset CRC include age, male gender, body mass index (especially in late adolescence and early adulthood), cigarette smoking, and alcohol consumption.5–10 Other candidate risk factors include metabolic syndrome, low physical activity, processed meat consumption, and oral antibiotic use in childhood. Some of these variables (e.g., prior diagnoses) can be easily obtained from structured data tables in the electronic health records (EHR). Other variables including social history and lifestyle factors are often documented more in the free text notes. We thus developed and applied natural language processing and machine learning techniques, which have shown impressive results in many cases.5–7
To extract both known and unknown CRC risk factors, we used an unsupervised method of topic modeling, the Latent Dirichlet allocation (LDA) algorithm. 5 Topic modeling is typically used to discovering common themes, called “topics,” that are shared by documents in a large text corpus. These topics are technically represented as a series of words that are thought to be semantically related. LDA is one of the most widely used topic modeling approaches and makes assumptions that topics are probabilistic distributions over words and that documents are mixtures of topics. It has found applications in many areas, including biomedicine.7,20–22
There are many machine learning methods. We trained and evaluated the performance of logistic regression (LR), support vector machine (SVM), random forest (RF), and deep neural network (DNN) models in the identification of CRC in US military Veterans aged 35–49 from medical records.
Methods
Data set
Structured and unstructured data from the Veterans Administration’s Corporate Data Warehouse (CDW) was used. 23 Statistical and ML models were trained and tested using a subset of CDW data that had undergone a chart review for a case-control study on risk factors for early-onset CRC in Veterans. In this dataset, study patients were initially classified into three groups: (1) Cases diagnosed with CRC during 2008–2015; (2) Colonoscopy controls, who underwent colonoscopy for diagnostic purposes (e.g., rectal bleeding) but were not diagnosed with CRC; and (3) Clinic controls, who did not undergo colonoscopy, were not diagnosed with CRC and were seen in a primary care clinic at least once per year for each of the 2 years immediately preceding the index date of a matched case. Inclusion and exclusion criteria are shown in Appendix 1. Colonoscopy controls and clinic controls were matched to cases on healthcare facility (based on the location of where the majority of primary care encounters occur) and year of index date in a 2:1 ratio (4 total controls per case), and then merged into a single control group. CRC diagnosis was determined by CRC oncology reports in CDW and through the VA’s cancer registry. In total, there are 722 cases and 3617 controls.
Feature extraction
Features from structured data
Summary of variables.
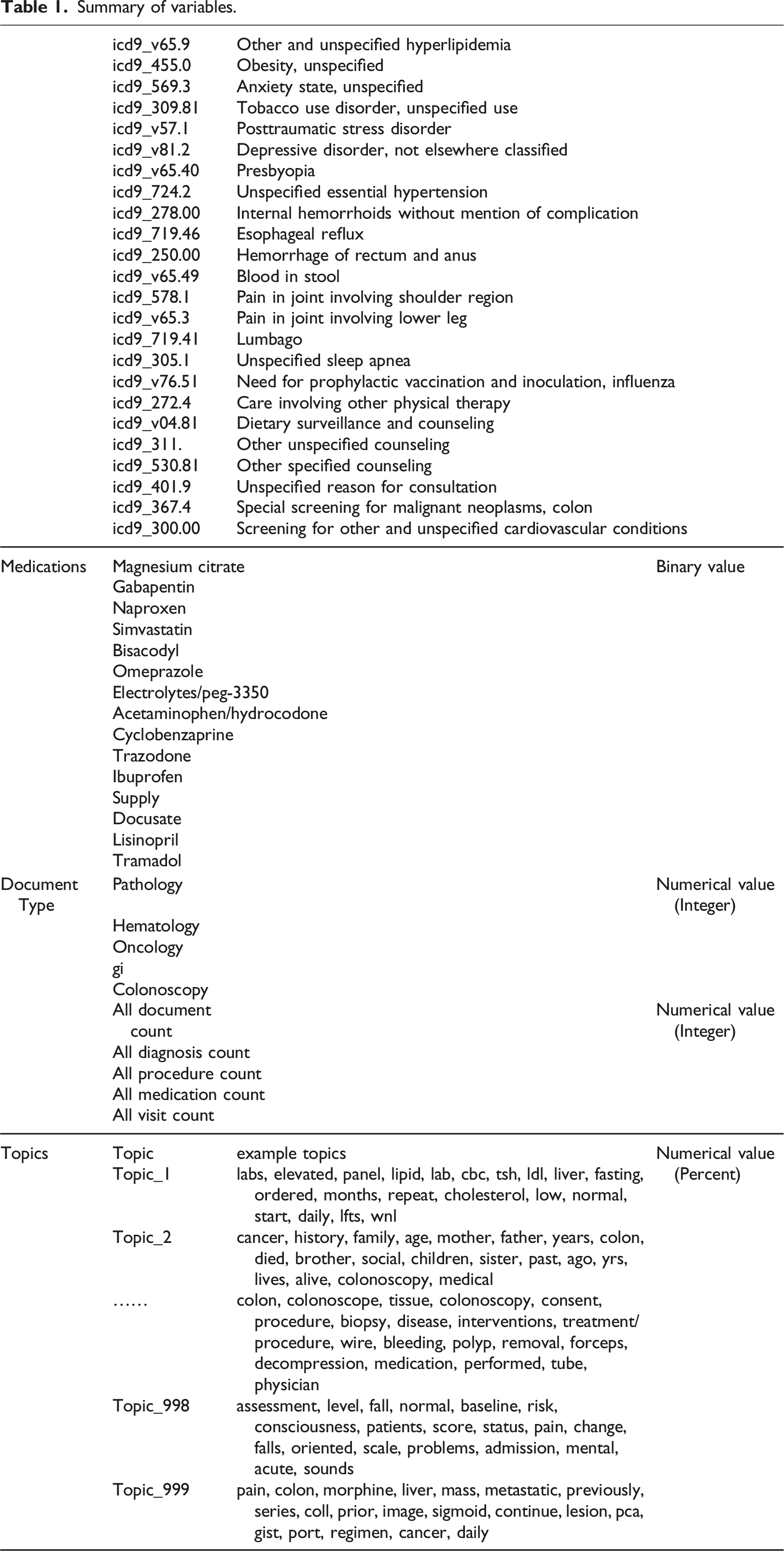
Features from unstructured data
The Latent Dirichlet allocation (LDA) algorithm 22 was used to generate 1000 topics based on clinical notes (n = 33,135) between 1 year before and 1 month after the index dates of the case and control patients. To increase the size of the training set, an additional 2576 documents were included from 70 patients diagnosed with CRC and 10,634 documents from 1188 patients who underwent colonoscopies from the CDW. Documents were converted to lowercase, and words in a stopword list or words occurring 10 times or less were excluded. The stopword list used was a general-purpose list of 524 common English words, to which we added the 25 words most common in our set of clinical documents including “patient,” “date,” and “time.” Table 1 shows examples of some of the topics that were discovered. We did not filter or select the LDA topics based on their content or relation to the outcomes of interest. As such, the LDA served as an unsupervised feature extraction tool. The number of topics (1000) was chosen empirically after reviewing the topic content, after experimenting with smaller numbers of topics (250, 500, and 750). Because the documents were not only focused on colon cancer, many topics including diseases, social history, symptoms, medications, procedures, and tests are covered in the clinical notes, especially in the control group. Only when the number of topics reached 1,000, were we able to find topics specifically focused on colon and colonoscope.
The topic model was applied to all clinical notes on each patient recorded from 1 year before the index date to 1 month after. The proportions of each topic’s representation in each patient note were averaged to give an overall proportion for each topic for each patient. These probabilities were used as the values for the topic features in the ML data set.
Machine Learning
Four supervised ML methods were applied and evaluated for their ability to classify patients as cases or controls. These included Logistic Regression (LR), Support Vector Machine (SVM), Random Forest (RF), and Deep Neural Network (DNN) mechanisms. For the first three methods (i.e., LR, SVM, RF), we used a Java ML library called Weka. We also used the default settings in Weka for the hyperparameters: LR: 24 ridge parameter=1.0E-8; SVM: 25 Kernel = Linear, complexity constant = 1.0; RF: 26 Size of each bag, as a percentage of the training set size = 100, Number of trees = 100, number of attributes to randomly investigate = 0, minimum number of instances = 1, minimum variance for split = 0.001, Seed for random number generator = 1, maximum depth of the tree = unlimited. Because of the large number of parameters, we cannot explain each one in detail but included the references for the algorithms.
For DNN, we implemented a Python deep learning library named Theano 27 together with a helper library called Lasagne. 28 The DNN was constructed of 5 hidden layers of sizes 200, 300, 200, 300, 200, and a single output using sigmoid activation, 300 epochs, batch size 100, and Nesterov momentum 29 with a constant learning rate of 0.001 and momentum of 0.9.
To avoid overfitting in the DNN training, the data was partitioned into 70% training, 20% validation, and 10% testing groups, with performance being measured on the testing group only. Training, validation, and testing groups were created using stratified sampling. After each pass over the whole training set, the DNN model was evaluated in the validation set. When performance on the validation dataset starts to degrade, we stopped further training the DNN model and applied it to the 20% testing set to assess final performance. This strategy is called “early stopping.”
LR, SVM, and RF did not require a separate validation dataset. The evaluation was performed with 10-fold cross-validation in all cases, each fold consisting of 3905 training (including validation for DNN) patients (650 cases, 3255 controls) and 434 testing patients (72 cases, 362 controls). All performance measures including the confusion matrix results were computed as the micro average of the 10 evaluations.
Results
Machine learning models and their performance in classifying colorectal cancer cases versus controls. Values are represented as mean.
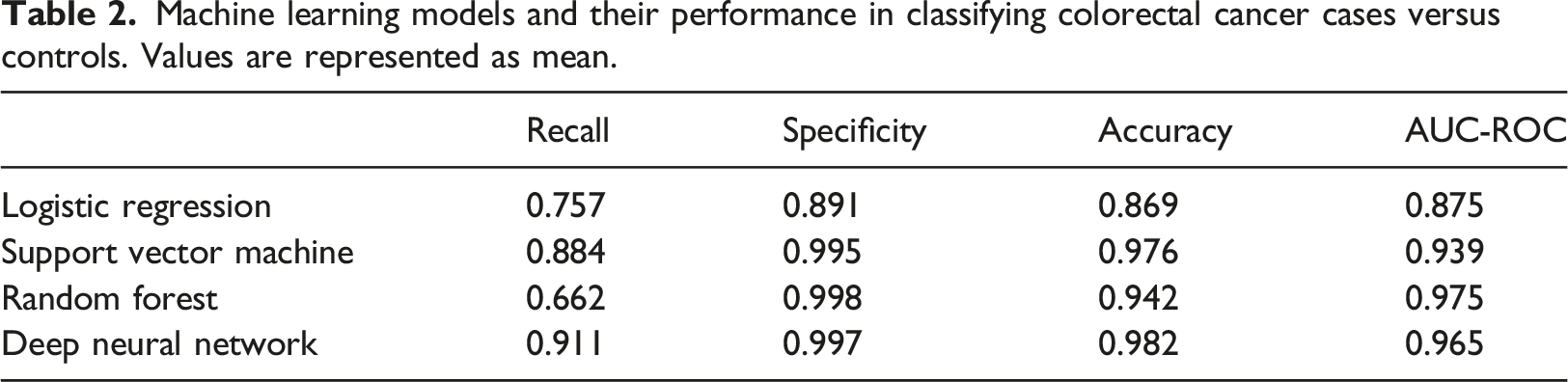
Confusion matrixes of LR, SVM, RF and DNN models.
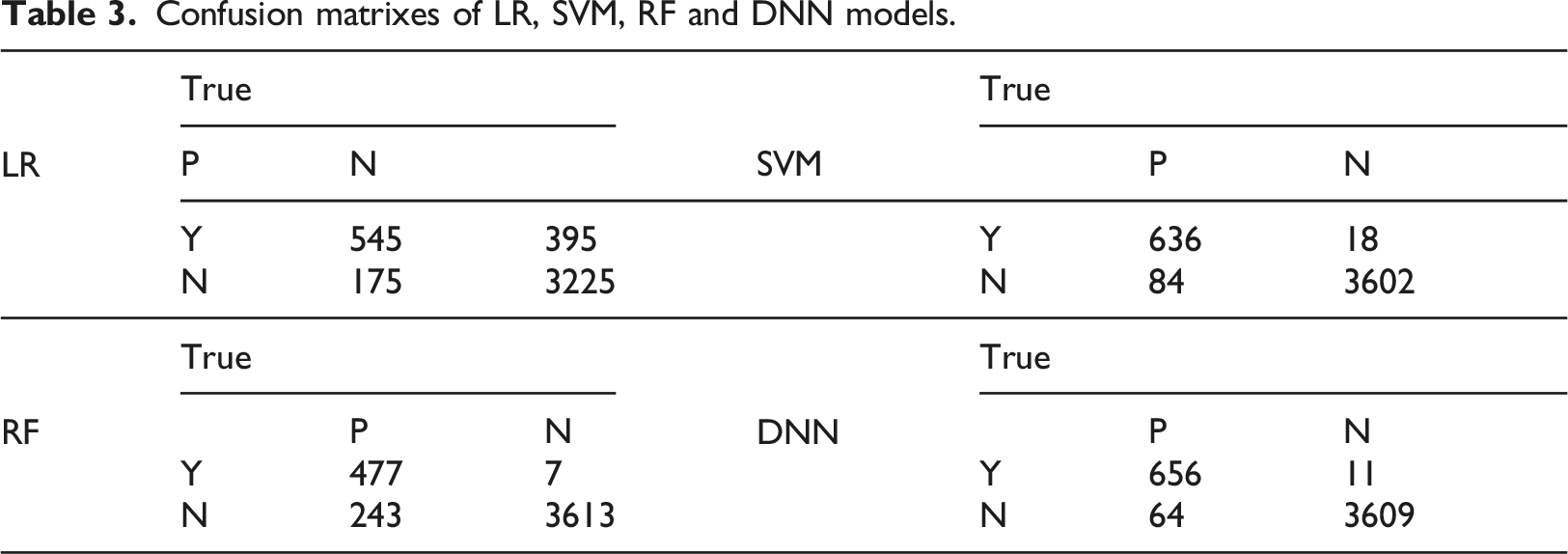
Discussion
Main findings
This study demonstrated that multiple ML models can correctly classify patients ages 35–49 with colorectal cancer using a combination of structured and unstructured medical data. This process can greatly impact future colorectal cancer studies by reducing the effort required to perform a large-scale chart review.
Among the 3 ML methods (SVM, RF, and DNN), DNN can be considered to be the most powerful and complex while the least interpretable. The fact that all three achieved an excellent and comparable performance level (in terms of F measure and AUC) is not very surprising, as ML methods' performance tends to be task and dataset-specific.
Compared to the most pertinent prior study by Xu et al., 19 our work focused on a separate age group (<50 years old). The low prevalence of CRC in the younger patient limited the number of cases we could use for learning. On the other hand, we utilized more features including procedures, medications, and comorbidities as well as a large number of topics as opposed to CRC ICD codes and keywords. As a result, were achieved a higher AUC (97.5% vs 93%). Even though we did not explore the feature importance or contribution in this paper, it is well known that models like DNN benefit from a large number of features.
Beyond CRC, there have been a number of prior studies that have utilized a combination of structured data and/or free text notes for case identification, including our own work30,31 and work by other researchers.32,33 Such efforts are often driven by the need for more accurate or consistent phenotype definitions beyond what diagnostic codes (often assigned for billing purposes) have to offer. This study, consistent with prior studies, showed that free text notes could be mined and combined with structured data to achieve high distinguishing power.
Limitations and future work
We selected the DNN parameters based on past experience and used the default settings in Weka for the hyperparameters for LR, SVM, RF. Since all ML methods reached a fairly high performance level, there was no need for extensive hyperparameter tuning. Without tuning the hyperparameters, we did not need to use a validation dataset for LR, SVM, and RF and went directly from training to testing. Before any ML models are used in a clinical context, additional validation would need to be performed. It is probable that these models will continue to perform well on a VA population meeting the same inclusion/exclusion criteria. Applying these models to other populations would require adaption and additional validation, adding examples from the target population to the training set or using a hierarchical method to augment the original results.
In the real world, CRC under 50 years of age is extremely uncommon (33 per 100,000 in 45–49-year-olds vs. 59.5 per 100,000 in 50–54-year-olds) 34 but is associated with worse outcomes than in older patients. For that reason, we sought to identify high-risk patients for targeted screening. In terms of training a model, having extremely imbalanced classes is a known problem. We chose the 1:5 match to allow more effective learning from positive and negative cases. We also want to note that the AUC values we reported are not dependent on the prevalence of cases in a sample, while accuracy rates are. In future studies, we would like to apply this model to a large set of clinical data and evaluate the positive predictive values using different thresholds.
We utilized a large number of topics (n = 1000) and a smaller number of structured data elements (n = 52). We did not experiment with a model using only the topics or structured data, which can be explored in the future. Features from the topic modeling cover all findings in a clinical note, from personal history to medications and from mental health to functional status. Because we performed unsupervised topic modeling, the topics do not exactly correspond to a specific diagnosis or treatment. As such, we expect the topics to overlap but not completely overlap with the structured data variables.
From the perspective of interpreting a model, having fewer features is desirable. On the other hand, one of the key strengths of DNN is its ability to handle a large number of features and does not require feature engineering. In a related analysis not described in this paper, we explored the features contributing to the different models by examining the feature coefficients in LR, weight in SVM, importance measure 26 for random forest, and impact score 35 for DNN. Different topic features were among the highest contributing features. For example, the top feature in the DNN is a topic on colon and colonoscopy. What was intriguing is that there was almost no overlap in the top 15 features utilized by the 4 different models. Further studies are needed to interpret the models and understand why and how other models use different features.
This study focused on the classification of CRC cases; however, an informative follow-on study could investigate predictive modeling. This may be accomplished by restricting the data period to only include measurements from 3 months to 1 year prior to the diagnosis. With a predictive model, it may be possible to identify features corresponding to risk factors. This could allow validation by features mapping to known risk factors and also identifying possibly unknown risk factors.
Conclusion
Patients with colorectal cancer can be identified with a highly reliable performance from structured and unstructured medical records using ML. These ML models can be used to decrease the manual effort required for chart review to identify cases in clinical research.
Summary table
• The ML and statistical model identified colorectal cancer cases using EHR with the best AUCs over 97% and best accuracy over 98%. • Overall, the DNN model performed the best. • There was a very low correlation among the four models in terms of the top features used.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a Veterans Administration Merit Review Grant (IIR 14-011).
Ethical approval
Due to the retrospective character of the study and because data were collected during routine healthcare procedures and patients were not subject to intervention, the Indiana University Institutional Review Board waived the need for informed consent (#1511734340).
Data availability
The datasets generated during and/or analyzed during the current study are not publicly available to protect the privacy of research participants. Still, aggregated datasets are available from the corresponding author on reasonable request.
Appendix
Inclusion and exclusion criteria for cases, colonoscopy controls, and clinic controls. Note that for this study, the primary and secondary control groups (Clinic Controls and Colonoscopy Controls) are merged into a single control group.
Group
Inclusion
Exclusion
Cases
Colorectal cancer
• CRC diagnosis during 2008–2015
• Ages 35-49 at CRC diagnosis• Inflammatory bowel disease
• Hereditary polyposis or non- polyposis syndrome
• Surgical resection of any part of the colon prior to CRC diagnosis
• High-risk family history of colorectal cancer
Colonoscopy controls
Colonoscopy without colorectal cancer
• Ages 35–49
• Colonoscopy during 2008–2015• Same exclusions as cases
• Screening colonoscopy performed on patients at average risk for CRC
• Colonoscopies that failed to reach cecum, terminal ileum, or appendiceal orifice or with bowel prep quality insufficient for adequate examination
Clinic controls
No colonoscopy and no colorectal cancer
• Ages 35–49
• No history of colonoscopy
• Received at least some of their health care through VA-based primary care clinics• Same exclusions as cases
• Prior colonoscopy
