Abstract
In this article, we examine work with communal data in the context of clinical genetic testing. Drawing from prior research on digital research infrastructures and from the analysis of our empirical data on genetic testing, we describe how data generated in laboratories distributed all over the world are shared and re-used. Our research findings point to six different human-driven activities related to expanding, disambiguating, sanitizing and assessing the relevance, validity and combinability of data. We contribute to research within Health Informatics with a framework that foregrounds human-driven activities for data interoperability.
Keywords
Introduction
Starting more than 3 decades ago, different scientific communities within molecular biology used database management software and computer networks to create communal data repositories advancing collaborative genetic research.1–3 Before the advent of communal repositories, scientists in the domain relied on their own data that they complemented with data gathered from literature or through direct contact with scientists from other laboratories. Nevertheless, the rapid rate of findings and the establishment of teams all over the world rendered “traditional sources of information—such as books and journals inadequate” 2 and motivated the creation of shared electronic databases. The aim was to “hook our individual computers into the worldwide network that gives us access to daily changes in the database and also makes immediate our communications with each other.” 4 Communal data repositories made possible the rapid dissemination of information and the coordination of work across different sites5,6 and soon became key components of the infrastructure for scientific work in the domain.
Scientists within human genomics pioneered data sharing through the implementation of several mechanisms to institutionalize rapid data release. For instance, the need to coordinate globally distributed teams and to advance progress in the Human Genome Project (1990–2003, so far the largest collaborative project in biology) led to establishing the Bermuda Principles in 1996. These principles stipulate that DNA sequence data should be published (i.e. uploaded to public repositories), and this should preferably happen within 24 h of production. Nowadays, the big journals in the field demand disclosure of software code, algorithms and sequence data upon publication. Human Mutation was the first journal to adopt a full data sharing requirement (in 2010), and the European Journal of Human Genetics has gone further, hiring curators to check that each article’s variants have been transmitted to a public database. 7 In a sense, work within human genomics can be viewed as a quintessential project on collaborative work; 8 the contributions of participants from disparate geographical locations and the use of information and communication technologies made it possible to produce results that otherwise could not be realized.
The role of data as a communal resource is today being widely discussed both in the context of innovation policies and strategies for growth and development and in the context of strategies for research advancement and knowledge generation.9–12 Overall, the discussions at the policy and strategy level are fuelled by the increasing availability of digital data and it is common to refer to data as general purpose resources. In that sense, data are conceptualized as long-term assets that can be pervasive in their benefits.13,14 Data are instrumental for informating and automating work, 15 and also for desegregating work when shared across localities. In the research reported in this article, we investigat what is entailed in making data communal within a scientific domain focusing on the human-driven activities required. We reviewed prior research on digital infrastructures for scientific collaboration and synthesized the insights of this literature stream by placing them within a concise framework covering activities related to both data sharing and data re-use. Furthermore, we conducted empirical work with a view to elucidating the literature-derived framework and refining it for the clinical genetics testing domain studied. Specifically, our research examines “how data generated in distributed laboratories are becoming communal resources?” Answering this question, we point to the different dimensions of the work entailed in making data shareable and re-usable and contribute to research on data interoperability within Health Informatics by providing insights that foreground the human-driven activities required.
The remainder of the article is structured as follows. First, we describe our methodological approach and provide an overview of the empirical case investigated. Then, we present key insights from prior research focusing on the human-driven activities required for making data shareable and re-usable. We continue with the empirical study on communal data for genetic testing related to the BReast CAncer genes (BRCA genes) which are associated with breast and ovarian cancer. Subsequently, we discuss the insights from our analysis, we point to the contribution of our research and we conclude by pointing to limitations of our study and further research directions.
Empirical background and research method
The impetus for our study comes from our involvement in a research and development project within genetics. This is a collaborative project between the Department of Medical Genetics in Oslo University Hospital and the University of Oslo. The aim of the project was to develop a secure information technology (IT) platform that facilitates distributed collaboration and access to a high-performance analysis and storage facility accommodating the increased demand for “personalized medicine.” As one of the research activities in this project, we conducted interviews and observations of how molecular biologists and other specialists conduct their work. Of special interest has been the work of interpretation after the DNA sequencing is conducted, where scientists assess the clinical significance of variants found in the patient’s DNA. During our observations, we were struck with the role of shared databases and we delved into the significant body of research on digital infrastructures for scientific collaboration within the Computer-Supported Collaborative Work (CSCW) and Information Systems (IS) fields. Within this body of literature, our interest was focused on human-driven activities entailed in making data communal within a scientific domain.
We turned to infrastructure studies for scientific collaboration, as this stream of literature adopts a socio-technical perspective with an attention to “the work to make things work.” 16 We selected studies on infrastructures specifically oriented toward distributed scientific collaboration where data sharing is a central aim and requirement. These studies explore different scientific domains ranging from life sciences,17–20 to engineering,21,22 and even papyrology and archeology.23,24 In most studies, researchers explored the processes through which technological arrangements have been shaped and built. Nevertheless, research has also explored activities that are not purely technical pointing to challenges of, for example, establishing shared taxonomies, data models and standards,17,25,26 the need for cleaning data, adding metadata and in general curating the information resource 18 and the need for facilitating the actual collaboration and linking of information.27–29 Going through this literature, we traced and synthesized insights related to activities required for making data communal in a scientific domain. We first mapped the activities associated with producing shareable data (“upstream”), and then the activities associated with re-using shared data (“downstream”). The outcome of this work is a literature-derived framework that maps activities of both sharing and re-using shared data. Drawing from this framework, we analyzed the empirical case with a view to elucidating and refining the insights from prior research for the clinical genetics testing domain studied.
Empirically, we studied a paradigmatic case 30 of data sharing for scientific collaboration. Specifically, we explored the use of data generated in distributed laboratories as communal resources for BRCA gene testing. BRCA1 and BRCA2 are two genes which influence humans’ susceptibility to breast and ovarian cancer. This is a domain where global communal data repositories have been in use since the 1990s. Furthermore, this is a significant domain within genetics as “a large proportion of the work in genetic services is the management of familial breast and ovarian cancer, and this clinical area exemplifies both the opportunities and challenges to increasing access to gene testing.” 31 We collected empirical material for the BRCA case using a combination of fieldwork, documents’ analysis and hands-on inspection of shared data (Table 1). We observed and interviewed scientists as they were (re)using data from communal data sources. We also inspected one key communal repository in order to learn about the processes of data sharing. This repository is the Breast Cancer Information Core database (BIC) which is a globally shared database for BRCA1 and BRCA2 variants established in 1995. BIC hosts data deposited by individual investigators, research, hospital-based and commercial labs and publishes deposited data after having them examined and edited by several members of its steering committee. After observing the work performed for BRCA variant assessment, reading related documents to familiarize ourselves with the domain and inspecting the BIC database, we interviewed BRCA scientists with a focus on data sharing and re-use.
Data sources for the empirical case.
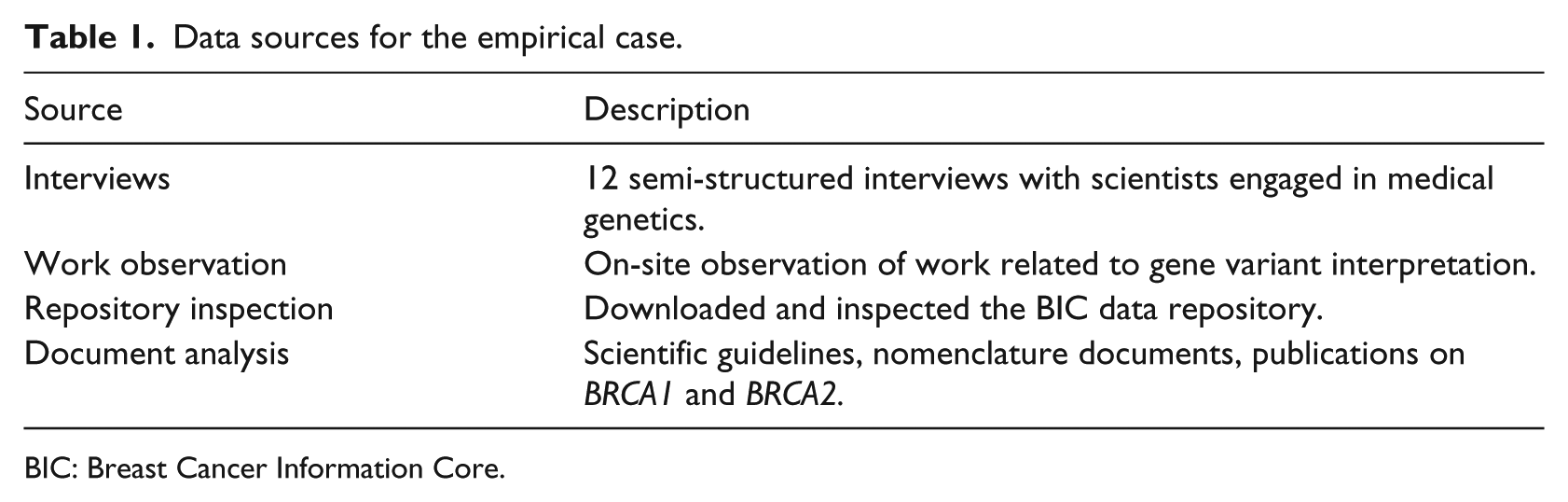
BIC: Breast Cancer Information Core.
We analyzed our empirical data using the framework that was derived from the literature. This allowed us to foreground the case specifics without losing sight of the multiple aspects of work entailed in making data communal. In Figure 1, we provide a schematic overview of the approach followed. Finally, we returned to the field and checked our understanding of BRCA data sharing and re-use by discussing the different data related tasks included in the framework.
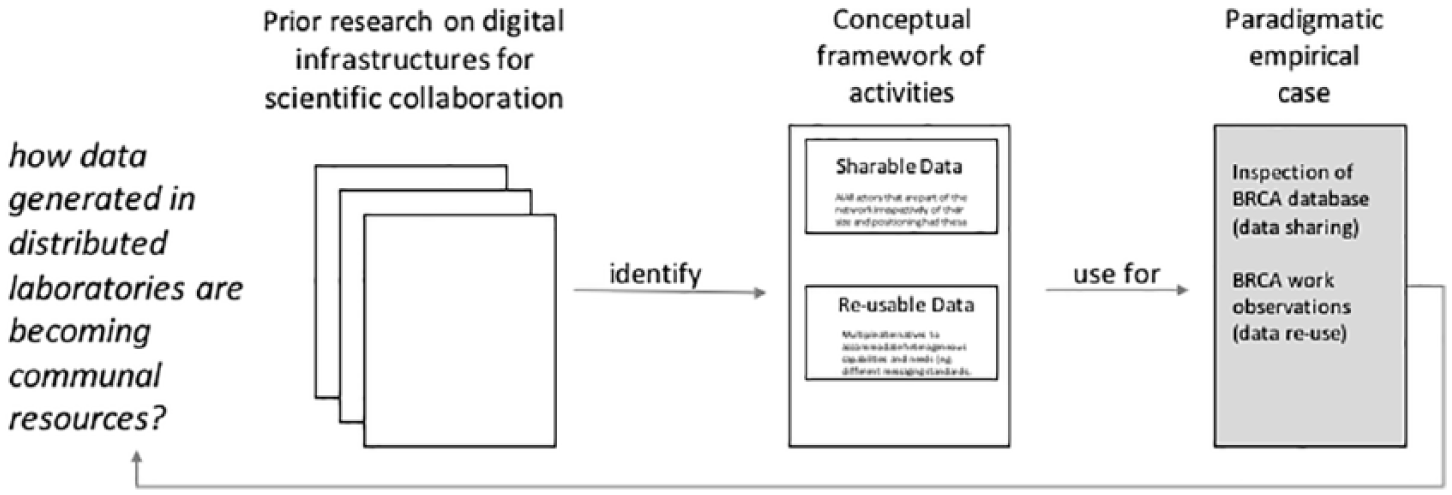
Methodological approach.
From local data to communal resources: a framework
Prior research on digital infrastructures for scientific collaboration has shown that although it is nowadays possible to pool together unprecedented volumes of scientific data generated by measurements, analyses and simulations performed all over the world, exploiting the data pools is far from straightforward. Data can cross local scientific teams’ boundaries physically by being deposited in common depositories, but do not automatically become intelligible and useful for other users. Humans and machines often face distinct barriers when attempting to find and process data that have been produced in different settings. 32 Additional work is required both for successful sharing and for re-using data, including both machine-driven and human-driven activities. Our study is focused on human-driven activities.
There are a number of steps that need to be taken before sharing data beyond local settings. First of all, data captured during ongoing scientific work need to be extended to include information customarily implied without being articulated. This can be done by including a richer representation of the context from which the data were taken. 33 Furthermore, data need to be disambiguated to allow as little uncertainty as possible about their meaning. Disambiguation can be achieved by adopting well-defined nomenclatures or by converting data to different coding formats. For instance, ambiguous geographical references can be converted to geographical coordinates via geoparsing. The minimization of ambiguity is an important prerequisite for data sharing; as Baker and Millerand 20 note, the reluctance of scientists to allow their data to travel can be attributable to “scientifically salient concerns about the lack of maturity of data classification efforts, the risk of misinterpretation of complex data.” Moreover, data need to be sanitized before being deposited to common repositories. Sanitization can include anonymization and treatment of data gaps and data outliers. Ensuring anonymity is a concern salient to scientific areas where data are sensitive and personal in nature (e.g. health-related data at the individual’s level). Anonymizing takes more than simply stripping-off data from all personal identifiers; in order to preserve data linkability, it is common practice to create new identifiers that link different records while secreting the persons’ identity, this process (known as pseudonymization) permits scientists to combine anonymized data. 34 Furthermore, data are often pre-processed before being shared because data anomalies (such as missing values or discontinuities due to addition or deletion of parameters) are commonplace 18 and scientists responsible for data production are the ones best positioned to make sense and treat anomalies (e.g. by fixing or filtering existing data). To ensure meaningful use by remote others, data are logically sanitized before being shared.
On the “downstream” side (data re-use), further work is required. Before using shared data, scientists need to assess their relevance and understandability. 21 Data can only be used if it is possible to make sense of them. In some cases, software and algorithms may be needed to interpret datasets. 35 Furthermore, the data need to be trusted before being re-used. For this assessment, different strategies are employed: inspecting data and exercising professional judgment about their pertinence and the scientific congruence of methods, drawing from the reputation of scientific teams and communicating with the scientists that deposited the data in common repositories to get clarifications and confirm assumptions. Assessing if the information content can be trusted requires significant experience and an eye for detail. For instance, a senior cancer researcher from Australia questioned the veracity of results reported in a series of journal publications on a specific gene in 2015 after realizing that the nucleotide sequences reported were not right. 36 She also identified unexpected similarities between these unconvincing publications, flaws in experimental design and mismatches between some described experiments and the reported results. 37 Following communications with journal editors, some of these gene-specific publications have been retracted. Another particularly challenging aspect of this work relates to the assessment of the combinability of data. Sampling and measurement compatibility have to be ensured to avoid drawing false conclusions out of pooled data. For example, differences between sites due to calibration of medical imaging equipment could lead analysts to false conclusions about population differences 25 or differences in the timing of pH measurements (e.g. immediately during fieldwork or at the lab during a later time) would incorrectly indicate differences in acidity. 17 Assessing data combinability brings forward “complexities of scale due to data heterogeneity, semantic relations and interdisciplinary collaborations.” 38
Overall, prior research indicates that data accessibility is not sufficient for scientific collaboration. In addition, work is required to make the data “shareable” and “re-usable.” We summarize the different data-related tasks entailed in creating shareable and re-usable infrastructural resources out of mere data tokens in Table 2.
Making data shareable and re-usable.
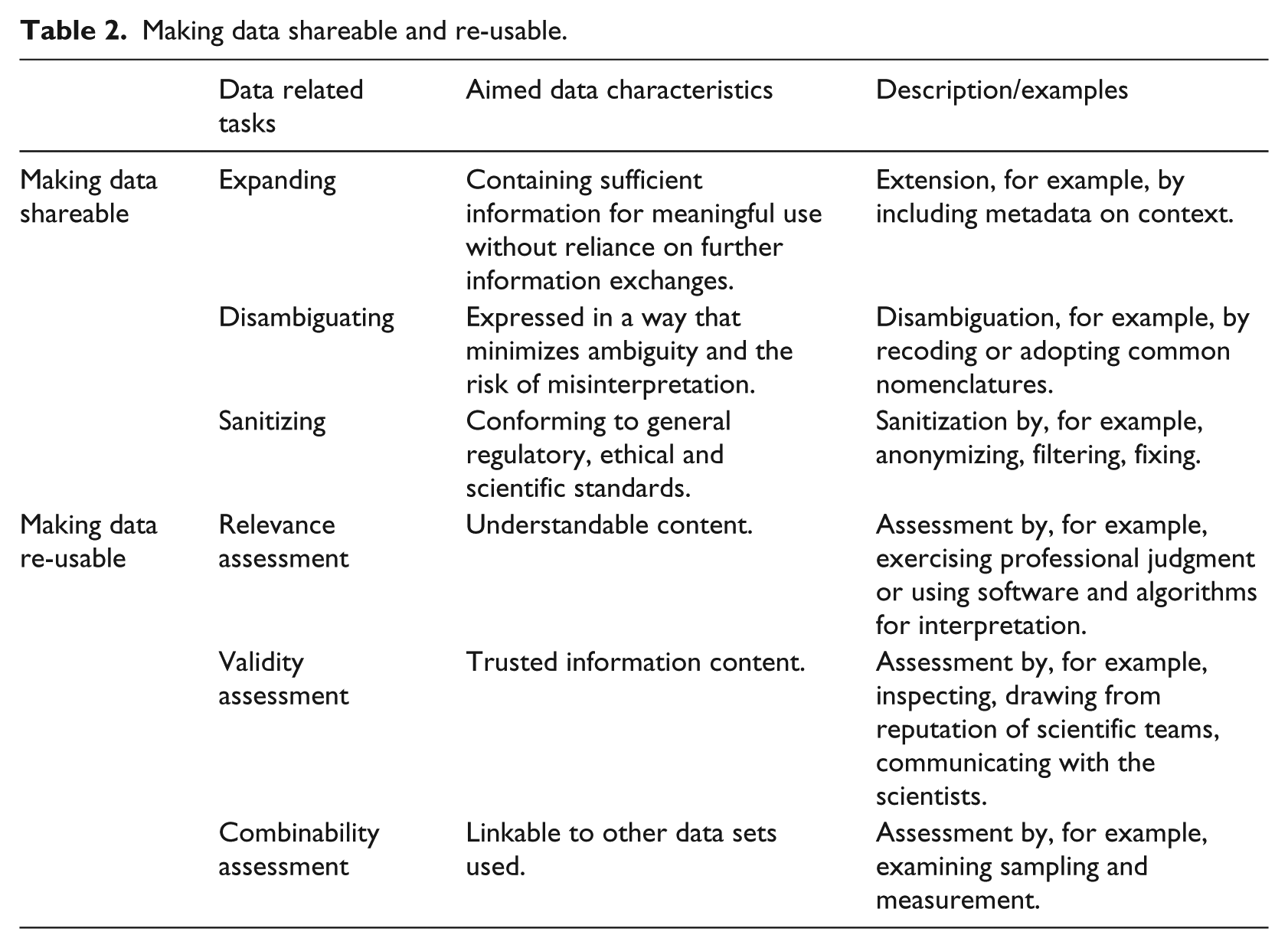
Case vignettes and analysis
In the paragraphs that follow, we present two vignettes from our empirical material. First, we present a vignette of data work in a medical genetics lab where molecular biologists and other specialists assess the clinical significance of genetic variants for the BRCA1 and BRCA2 genes. The work described in the vignette is performed after gene sequencing is completed with the identification of variants (i.e. differentiations from the common sequence). After a variant is identified, it can be assessed as a variant that indicates pathogenicity (clearly or most probably), a variant that does not indicate pathogenicity (clearly or most probably), or a variant of uncertain/unclassified significance (VUS). During the assessment, specialists search for past variant classifications in datasets containing anonymized test results for the same gene. Search can be performed in local datasets (containing results from tests performed in the lab) or in shared ones (where many labs deposit data), including both structured data repositories and published scientific articles and reports. Hence, the first vignette presents data re-use in the BRCA variant assessment context. Then, we present a second vignette which offers a snapshot of the BIC repository (a globally shared database for BRCA1 and BRCA2 variants established in 1995). The second vignette illustrates the work entailed in data sharing by exposing the content of the different fields needed for creating a new entry in the shared database.
Vignette 1: data re-use: genetic variant evaluation in a medical genetics lab
The following vignette is based on our on-site observations and illustrates how the genetic specialist draws on multiple information resources, including data from a number of global, shared repositories during the evaluation of genetic variants.
Alice wants to show us how she works with a VUS. It is a variant that was found in the BRCA1 gene of a patient, in DNA position 5504 of the gene. Usually, there would be a “G” nucleotide in the DNA sequence, but here it has changed to an “A.” This is captured by the G > A notation. “This is a missense variant, they are always uncertain,” Alice says. The reason is that these variants result in an amino acid change in the corresponding protein (the product of the gene). She opens a supporting analysis software, and searches for information about the variant. She clicks the button called “Frequency” and the system queries an external database. The query response is “No frequency data available.” This means that there have not been enough observations of this variant to estimate a frequency in the general population. The frequency of the variant can therefore not be used to exclude it as a common (and therefore benign) variant. Alice decides to change strategy and clicks a button that links to a BRCA1 specific database and a new window opens. Here, she finds information about a previously observed variant in the same position, but this one involves a change to “C,” not “A” as in the current patient. The G > C variant results in a different amino acid with different chemical properties in the corresponding protein product than G > A, and is therefore not likely to be relevant. She leaves this database and clicks on “Human Gene Mutation Database” (HGMD). This presents data from the large HGMD to her. Here, she chooses “missense” and looks for 5504, but does not find anything. She goes back to her software and clicks on a button with the Google logo. A Google search result page opens up, with results of a search string of multiple alternative names for the G > A variant. From the Google search, Alice identifies a number of relevant articles where the specific G > A variant is mentioned. For one of them, she sees a note in a local database that she also consulted in order to see whether the variant had been encountered in house earlier. Here another specialist at the department had noted that the Human Genome Variation Society (HGVS) and BIC notations (two alternative ways to express the position within a gene) were mixed up in the article. She explains this could indicate that the article has to be interpreted with caution. Two of the studies relate to Malayan and Indonesian patients. When she opened the PDF of one of these articles, she searches through the text for the number 1835, which is the amino acid position corresponding to the DNA variant. She finds it in a table, where it is accompanied by a question mark and a comment that it is “possibly important, may influence.” Even though these studies are non-conclusive, Alice says she has still learnt something; that the variant is in the BRCT domain which is an important part of the protein. She goes back to her software and chooses “BIC.” In the new window that opens, she selects “Search Database,” writes “exon 22” and “codon 1835” and pushes “search.” She finds nothing. She then goes back to do the final step and check whether the variant is located near to or in a splice site. This is important for the protein product. The software she uses integrates several prediction tools, and all of them predict “no effect on splice site.” Alice concludes by classifying this as VUS. She explains that she was led to this conclusion because she cannot find frequency data, there are some articles describing a possible relation to disease but with no clear conclusions and there is likely no impact on the splice site. Alice records her classification in the local database, as it may be useful if she or her colleagues later encounter the same variant. However, the entry is not automatically shared beyond the laboratory. To share the data, a special preparation and submission process is required which is not part of Alice’s everyday work tasks. In the second vignette, we turn into examining the accompanying information required for submitting a variant assessment to one globally shared BRCA database. The first vignette reflects the work required for searching and assessing the found information, and shows how Alice approaches the communal data repositories with an interest in using them for genetic variant evaluation in a clinical context.
Vignette 2: data sharing: genetic variants as entries in the BIC repository
In this vignette, we investigate what is required for data to be shared in common data repositories. We do this by examining the content and procedures of one such repository, the BIC. The content of BIC is both accessible via an interactive web interface and as downloadable flat files. We performed a full download of the database instance of September 2014 and investigated its content. Each entry in the database relates to a specific genetic variant identified during genetic testing for a specific patient. For each entry, a number of fields need to be filled-in (Table 3). The fields relate to the variant itself (e.g. its location), the assessment of the variant’s importance and a number of contextual details. Going through the database, it is easy to observe that not all fields’ content is standardized. For instance, the field “detection method” in the BRCA1 dataset downloaded included 98 “distinct” methods. A simple visual inspection reveals that the big number of methods is not only attributable to the heterogeneity of testing arrangements among laboratories, but also to different levels of specificity adopted for method description, non-standardized data entry (e.g. “FCCM” vs “FCCM (Fluorescent Ch)”) and entry slips (e.g. a space makes the records “SSCP” and “SSCP” to be distinct, “PPT” instead of “PTT”). Most searches within the database are performed on the basis of variants. When a particular variant is identified, the additional information about the specific entry (e.g. the “detection method”) may become of interest. Genetic experts can make sense of the non-standard descriptions as they have a good sense of their semantics.
List of fields per BIC entry on the clinical importance of BRCA1 and BRCA2 variants.
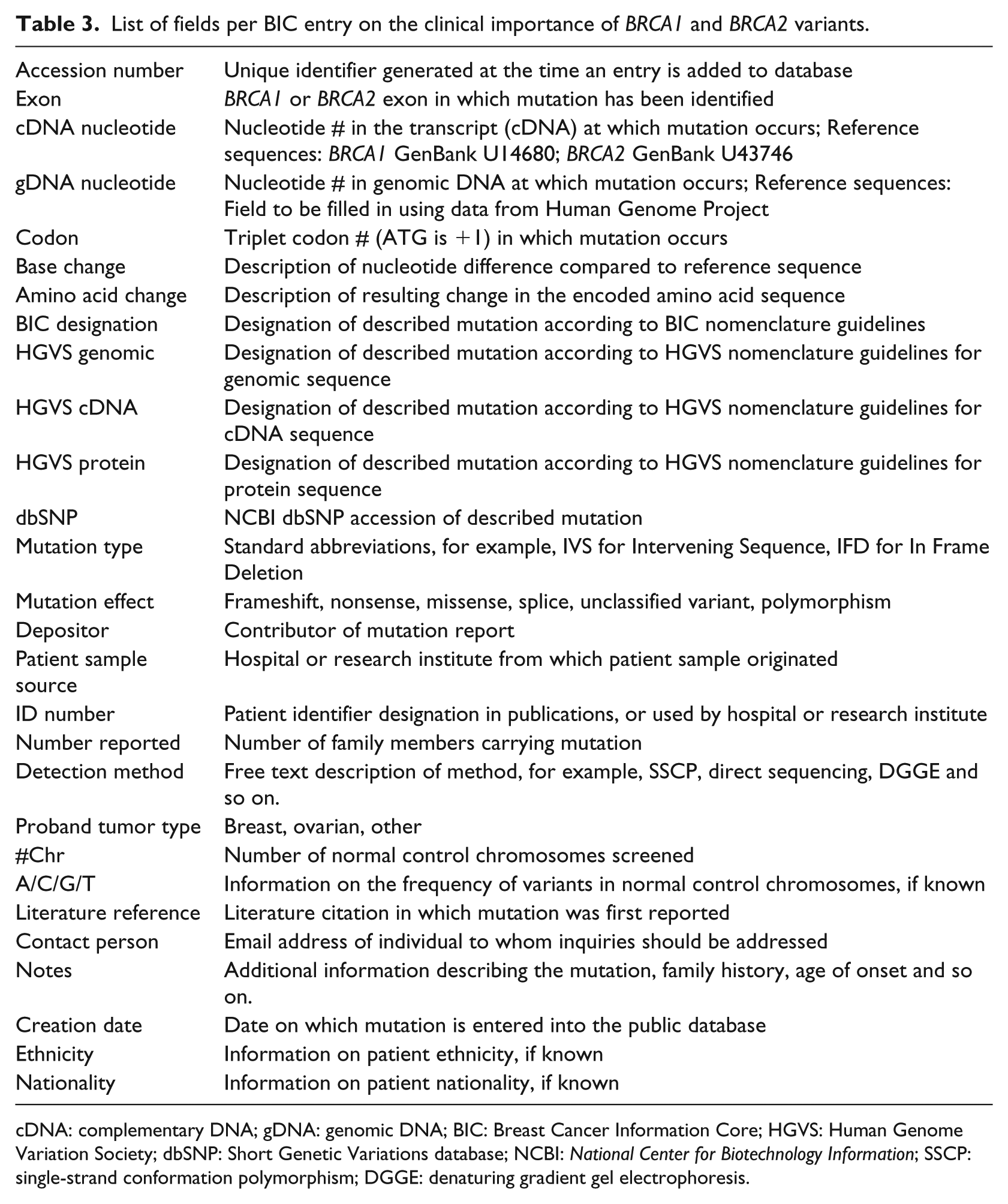
cDNA: complementary DNA; gDNA: genomic DNA; BIC: Breast Cancer Information Core; HGVS: Human Genome Variation Society; dbSNP: Short Genetic Variations database; NCBI: National Center for Biotechnology Information; SSCP: single-strand conformation polymorphism; DGGE: denaturing gradient gel electrophoresis.
As the search within BIC is variant oriented, the most important piece of data that facilitates use is the variant designation itself. Interestingly, there are many different fields for the variant designation. There is one field following the BIC nomenclature and also, fields following the HGVS nomenclature (Table 3). To make sense of this redundancy in the database, we had to trace the roots of the different nomenclatures. Since the identification of variants is of critical importance in the use of the database, BIC introduced a specific nomenclature for the designation of BRCA variants to ensure consistency and unambiguity. Nevertheless, in the years that elapsed since BIC’s introduction (back in 1995), the HGVS developed a universal nomenclature for the description of sequence variants in any gene (not only the BRCA genes). The first version of the HGVS nomenclature was published 5 years after BIC’s nomenclature, in 2000. 39 Since then, the HGVS nomenclature has been continuously updated and amended to include special cases and new insights and soon became the preferred standard for variant descriptions as it is more unambiguous than the BIC one. The two different nomenclatures use different reference points for nucleotide numbering (e.g. 1184G > A in BIC is c.1065G > A in HGVS) and have a number of syntactic differences (e.g. 1014delGT in BIC is c.895_896delGT in HGVS). To facilitate variant look-up within BIC, each variant has to be described both in the “legacy” BIC nomenclature and in HGVS. Built-in redundancies in data entry facilitate variant look-up and validation.
The broad range of fields required for creating a single entry in the common database shows the extent of work required for sharing the lab findings. The data need to include sufficient contextual information (e.g. information on sample source, detection method, tumor type). Furthermore, they need to be expressed in a way that minimizes ambiguity and the risk of misinterpretation (hence, the multiple fields for designation, the identification of exon, codon etc.). Finally, while being anonymous, BIC entries also include identifiers to link with different records in other databases in case further investigations need to be performed. This vignette illustrates the effort required for ensuring that the data deposited in the repository can be re-used by other scientists.
Analysis: sharing and re-using BRCA data
The medical genetics community for BRCA testing is an exemplary case of a technology-enabled collective that is built and sustained around rich communal data sets. Every new analysis is based on the outputs of laboratories distributed around the world, and as these past outputs are continuously reworked, evaluated and amended, the knowledge basis keeps evolving, becoming more mature and comprehensive. We use this case to revisit and elaborate on the literature-derived framework.
Expanding
The assessment of clinical importance can only have one out of five possible values: positive (indicating pathogenicity clearly or most probably), negative (indicating no pathogenicity clearly or most probably), or unknown (i.e. VUS). Nevertheless, a significant volume of accompanying context-related data is required for each variant assessment entry. The accompanying information is meant to support genetic experts to make better sense of the variant assessment and is not provided in standardized format, but as free text. This indicates that the additional data are not intended for computer processing, but for being interpreted by humans. Issues of scale (large volumes) and speed (dynamics of change) are not present in the handling of BIC data. Although there are thousands of entries in the database (more than 30,000 in the downloaded instance), the average number of records per variant is less than eight. The small dataset is rich in contextual information to support a critical assessment process. A balance is needed in providing rich enough information for downstream work, while not imposing a too heavy workload for the submission of information.
Disambiguating
Gene variants are complex scientific objects, and it is no surprise that the development of suitable nomenclatures is an ongoing project that has not been concluded yet. To minimize the risk of misinterpretation, BIC includes redundancies in variant descriptions. These redundancies serve the practical purpose of quick data retrieval and at the same time, help the members of the community to quickly identify data contributors that are not rigorous enough and contribute inconsistent information (this relates also to the assessment of the validity of data entries). At the same time, the ambiguity in the description of detection methods can be tolerated and compensated by expert knowledge of the domain. So, the level of rigorousness and standardization varies among the different data fields. Ambiguity in variant description is unacceptable as it may lead to wrong conclusions. What we find in the BRCA case is that disambiguating is not a universal aim for all data shared, but it is specific to each different piece of information. While some level of ambiguity is tolerated for supplementary information, disambiguation is key for the critical data (variant designations).
Sanitizing
The BIC database is variant-centric, not patient centric. The data relate to the variant itself, the assessment and contextual details including contact information. To assess BRCA variants for susceptibility to breast and ovarian cancer, there is no need to cross-examine the information about other genes for the same patient. This holds for studying diseases related to monogenic alterations (such as for BRCA1 and BRCA2). In general, information about single variants is considered anonymous. While being anonymous, BIC entries also include identifiers to link with records in local databases, but these identifiers are of no practical use without the depositor’s collaboration. Overall, the BIC database does not include any data beyond those that are strictly necessary for assessing variants based on our current knowledge. This allows the dataset to conform to general regulatory and ethical standards, but bounds its generative potential and limits its combinability with other datasets.
Relevance assessment
Following Alice in her work, we find that relevance assessment is facilitated by searching in gene-specific databases. If there were no special-purpose archives, she would have to spend significant time for data discovery and assessment. The use of specific repositories allows her also to familiarize with their structure and conventions. Furthermore, the precise description of the variants with the use of specific notations allows her to screen quickly search results for relevance even when using a search engine on the web and to put together a set of informative resources (e.g. about the frequency and location of the variant). In the case studied, standardization plays a significant role for relevance assessment.
Validity assessment
As observed during Alice’s work, when data retrieved indicate sloppiness or inadequate mastery of notations (as in the case of mixing HGVS and BIC notations) scientists can be reluctant to re-use the data. In this way, a distinction between credible and less credible members of the community can be made. Other ways to assess the credibility of the data include the appraisal of the reputation of scientific teams that have deposited the data and also, direct communication. For this reason, the details of the depositing laboratories are included in the databases. Furthermore, in the genetic domain, most repositories are “curated.” Curation includes a range of activities to validate, manage and maintain the information deposited in the repository. In the case of BIC, the data deposited are assessed for relevance by the BIC scientific committee before becoming accessible by all. Alice still has the responsibility for assessing the validity of the data she uses, but her work is facilitated by central curation.
Combinability assessment
For the limited combinatorial usages of BRCA datasets, the variant designation is key. BRCA datasets tend to be linked mostly with other BRCA datasets and not with other types of data (e.g. other health-related data for the same individuals). Furthermore, for BRCA testing, there is no need to use longitudinal data analyses. Another interesting aspect of these data is that they are continuously updated and renewed based on new scientific findings. The more recent assessments normally include also the latest findings in the domain.
Discussion
The case we studied is special as it relates to a scientific domain where the data to be shared are well defined and with relatively little heterogeneity. This facilitates the creation and use of communal data repositories, especially when comparing with situations where scientists engage with less well-defined scientific objects as, for instance, in the case of communities of archeologists facing much greater challenges in data-sharing. 24 The BRCA datasets are deep and not wide. 40 Deep datasets have big numbers of records, but not a lot of diversity. In other words, they are characterized by “trimness” as they include only objects from a specific, common scientific subject pool. 19 Studying such a trimmed dataset allowed us to get a comprehensive view on the different aspects of work entailed in making data shareable and re-usable.
The characteristics of the BRCA domain facilitated the actual establishment of communal data resources. Relevant aspects of the domain are the descriptive particularities of the variants, the limited categories for classifying their impact, the fact that the phenomenon under study (diseases related to hereditary genetic alterations) is invariant over time, the monogenic nature of the BRCA alterations and the fact that there are no privacy and anonymity issues to be addressed. These characteristics can explain how it was possible for BIC (which is the first communal data repository in the BRCA domain) to emerge out of a scientific initiative that established an infrastructure with maximum simplicity adapted to the characteristics of BRCA data. 2 One key characteristic of the data is that they do not raise privacy and anonymity issues. The non-sensitive nature of single-gene data and the focused type of tests allowed the community to evolve without being challenged by issues related to patient consent, confidentiality and incidental findings. As genome-wide testing is becoming more common, data sharing is increasingly relying on complex arrangements for patient consent,41,42 but these are not integral parts of BRCA sharing configurations today.
Although the datasets in the domain are well defined, relatively narrow and non-sensitive, the work involved in making BRCA data shareable and re-usable is still significant. The amount of effort required is one of the reasons explaining why the data in communal repositories are only a small part of what is generated in laboratories. The number of entries in BIC is significant (more than 30,000), but far from all-inclusive. It has been reported that more than 100,000 BRCA tests are performed annually in the United States alone. 43 For the growth of communal data resources, the work of data curators and repository managers is instrumental. These new roles can perform some of the work that is required for primary data depositors and data users. Specifically, in the BRCA case, data curators and repository managers can significantly contribute in disambiguating, sanitizing and validating entries becoming part of the overall communal data infrastructure.
In the empirical case we followed, variant data were retrieved, processed and evaluated by human experts. This creates requirements that are different to those for automated processing. When aiming for automated processing, it is important to ensure that data are consistently expressed in a way that is decipherable by computers. Computer decipherability has some common characteristics with human-oriented disambiguation and data expansion (making data sets understandable and self-contained), but is also quite different. For instance, while elaborate domain models are required for automated data integration and analysis, 25 humans can make good inferences from complete and coherent, but unstructured text. The term “data interoperability” is used in scholarly work pointing to requirements for making data re-usable and itinerant within multiple sites. Nevertheless, frequently there is no distinction between requirements for making data more suitable for remote collaboration versus requirements related to automated processing. The two different aims are not always compatible. Requiring scientists to use rigid vocabularies and coding schemes can impede collaboration by making coding and decoding information more challenging: “rigid data standardisation may even diminish semantic interoperability … information that is readable but not understood may lead to incorrect conclusions.” 44 Even more significantly, rigid coding could direct scientists to be more specific and definitive than they are able to be or limit their capacity to express emerging understandings that are not yet concrete.
Conclusion
The significance of machines in data-rich research environments will be continuously increasing, while the role of human experts in critical domains such as healthcare is also expected to remain strong. One of the grand challenges of data-intensive science is to assist both humans and computational agents, in the identification and analysis of data. 32 Our research contributes to the literature on health informatics interoperability by foregrounding the human-driven activities complementing prior health informatics research that has mostly focused on the machine-driven aspects.45,46 We complement this technically oriented research stream by synthesizing insights about human-driven activities required for data interoperability drawing both from prior related research and from an empirical case of established sharing and re-use arrangements in clinical genetics testing. Our research shows how data generated in distributed laboratories become communal resources that contribute to everyday work by pointing to six different activities for making data shareable and re-usable. These activities are shaped by the characteristics of data that need to be shared and re-used.
In our study, we were limited by exploring one specific domain (BRCA testing). We expect that further interesting and complementary insights can be found in related fields. Of special interest are areas where considerations of privacy and confidentiality prevail. In such areas, the activities related to sanitizing and assessing data combinability are expected to be increasingly complex. Unlike BRCA-specific data, sharing and re-using comprehensive individual genome–phenome datasets entails significant privacy issues47,48 and there is a growing interest for empirical research in this area. Another significant area for further research is the exploration of work allocation between genetic experts that generate and use data and data managers and curators. Studies in this direction can provide insights for expediting and sustaining communal data sharing through arrangements that include both the scientific laboratories and data custodians (curators, data managers, or other data-related roles). As the findings of our study demonstrate, there is significant effort required related to expanding, disambiguating, sanitizing and assessing the relevance, validity and combinability of data. The distribution of this effort among multiple actors is instrumental for the growth and sustainability of communal data resources.
Footnotes
Acknowledgements
We gratefully acknowledge the constructive input by Dr Morten Cristoph Eike on earlier drafts of the article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Norwegian Research Council (Norges Forskningsråd), projects no. 210622 and no. 237766.
