Abstract
A common activity carried out by healthcare professionals is to test various hypotheses on longitudinal study data in an effort to develop new and more reliable algorithms that might determine the possibility of developing certain illnesses. The INnovative, Midlife INtervention for Dementia Deterrence project provides input from a number of European dementia experts to identify the most accurate model of inter-related risk factors which can yield a personalized dementia-risk quotient and profile. This model is then validated against the large population-based prospective Maastricht Aging Study dataset. As part of this overall goal, the research presented in this article demonstrates how we can automate the process of mapping modifiable risk factors against large sections of the aging study and thus use information technology to provide more powerful query interfaces.
Introduction
Dementia is a serious loss of cognitive ability beyond what might be expected from normal aging. Worldwide, the number of people with dementia is currently estimated to be 44 million and expected to reach approximately 76 million by 2030 and 135 million by 2050. 1 While dementia is one of the most feared age-related conditions and a chronic and progressive illness with no cure, there is now strong evidence that dementia can potentially be delayed by adopting lifestyle changes in mid-life aimed at improving cardiovascular health, addressing low mood and poor diet and increasing physical and cognitive activity. Given the huge social and economic costs of dementia, even a delay of 1 year would make such interventions cost-effective.2,3 Increasingly, the importance of dementia prevention and the need to take prevention measures based on existing knowledge are being highlighted at an international level.
The INnovative, Midlife INtervention for Dementia Deterrence (In-MINDD) project, 4 funded by the European Union (In-MINDD FP7/2009–2013), is taking place against this backdrop. It has three main objectives. First, it seeks to create a multi-factorial model for dementia risk, taking into account a broad spectrum of factors including cardiovascular risks, mood, physical and cognitive activities. A key element in creating this model is to test its validity against the MAAS dataset, which is a large population-based prospective dataset. Second, it aims to develop a state-of-the-art online profiler for use in primary care to assess the risk that individuals in mid-life (40–60 years) have of developing dementia in later life. The project will also develop personalized strategies to reduce risks to participants’ future cognitive health via a supportive online environment which has access to the best available online strategies and locally sourced options for delaying the onset of dementia. Third, it will evaluate the use of the In-MINDD profiler and online support environment with practitioners and patients. The article relates to the first of these three objectives.
In some clinical research projects, sensors can be used to harvest data 5 with the effect of generating very large datasets. In other approaches, large studies are compiled over significant periods of time 6 using a question-and-answer type system to compile knowledge on individuals with different demographics, lifestyles, and so on. In both cases, the next step is often to build the ontology to provide context and generate new knowledge from the existing dataset. Previous research efforts on constructing medical ontologies have provided frameworks for incorporating data from operational medical systems 7 or tackling the general issue of interoperability across medical applications.8,9 One of the requirements of our project is to develop and test a number of different dementia-risk hypotheses. Currently, testing must be performed manually using spreadsheets or statistical software, but the role of data management researchers on the project is to automate this process. The ontological approach has been shown to be effective in areas such as intensive care 10 and even in broader healthcare like the Lifeline project. 11 In terms of research into dementia, one of the earliest approaches that involved ontology construction was in Malhotra et al. 12 where they sought to describe formally concepts and the relationships between them. Each of these projects demonstrated the impact of a formal approach to classifying terms and relationships and how the ontologies can be exploited for a greater understanding of data in different domains.
Research focus and contribution
This article describes our approach to linking knowledge from existing longitudinal studies to risk factors identified by health specialists for the area of dementia. What may appear as a relatively straightforward task (and currently performed manually) is in reality quite a difficult problem. When specialists devise a series of hypotheses to be tested using one or more longitudinal studies it requires the interaction and manipulation of potentially hundreds of questions from the original study. The problem is in the quick identification of those areas of the study that best test the hypothesis. Given the manual nature of this approach, it is often difficult to ensure that all the relevant questions and answers are used, and thus, the accuracy of the results can be difficult to measure. Our method matches sections of the clinical study to defined ontology risk factors, in a four-step process of Concept Name Match, Concept Property Match, Vocabulary Match, and Structural Match. For the In-MINDD project focusing on dementia, these risk factors are as follows: low cognitive activity, physical inactivity, depression, mid-life obesity, cholesterol, alcohol, hypertension, coronary disease, renal impairment, diabetes, diet, smoking, and functional impairment (which was later dropped). However, simple keyword matching between these risk factors and the keywords used in question-based studies results in a very low level of matching. While ideas such as ontologies for managing healthcare surveys as proposed in Donnelly et al. 13 could greatly assist in matching new concepts to older datasets, the reality is that this type of structured approach to medical studies does not exist. In earlier work, 14 we used WordNet15,16 to generate synonyms at three descriptive levels for risk factors: risk factor name, risk factor properties, and vocabulary associated with a risk factor. These synonyms together with the original terms were matched against all questions in the clinical study. While there were a number of desirable outcomes that emerged from this approach, it resulted in a high number of false positives where inappropriate questions were matched to some risk factors. As a result, a new approach was necessary where the goal was to reduce the false positives as much as possible, while still using the same methodology of matching in four iterations, our previous article 14 can be referenced for what terms are used in each step.
This focus on the creating links between any longitudinal studies focused on dementia and our ontology risk factors has three broad contributions:
We provide a framework in which ontologies can be populated with data from clinical studies.
We introduce a new matching method which uses word stemming and comparison at the phrase level as opposed to single word matches.
We audit every comparison between the risk factors and questions in the clinical study so that we can record at which of the four steps matching is found. This has the benefit that simple queries can detect the precise point at which a false positive occurred. This in turn, provides input for the next iteration of the algorithm to determine selection thresholds for links to the clinical study. The results are shown in the summary statistics of Tables 1 to 3.
Synonym-based match with thresholds Tt1 = 0.3, Tt2 = 0.3, Tt3 = 0.3, and Tt4 = 0.2.
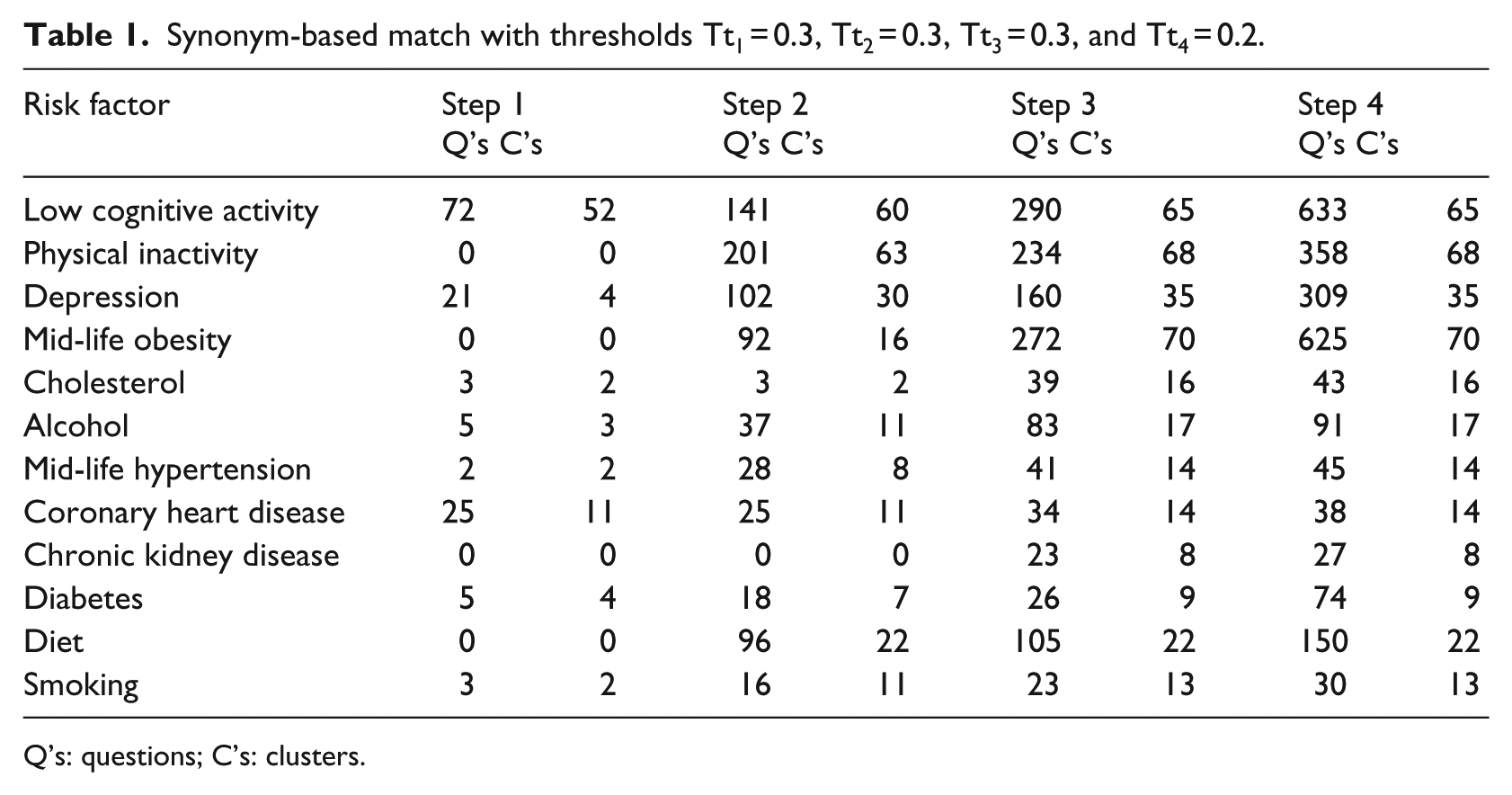
Q’s: questions; C’s: clusters.
Lucene matching results.
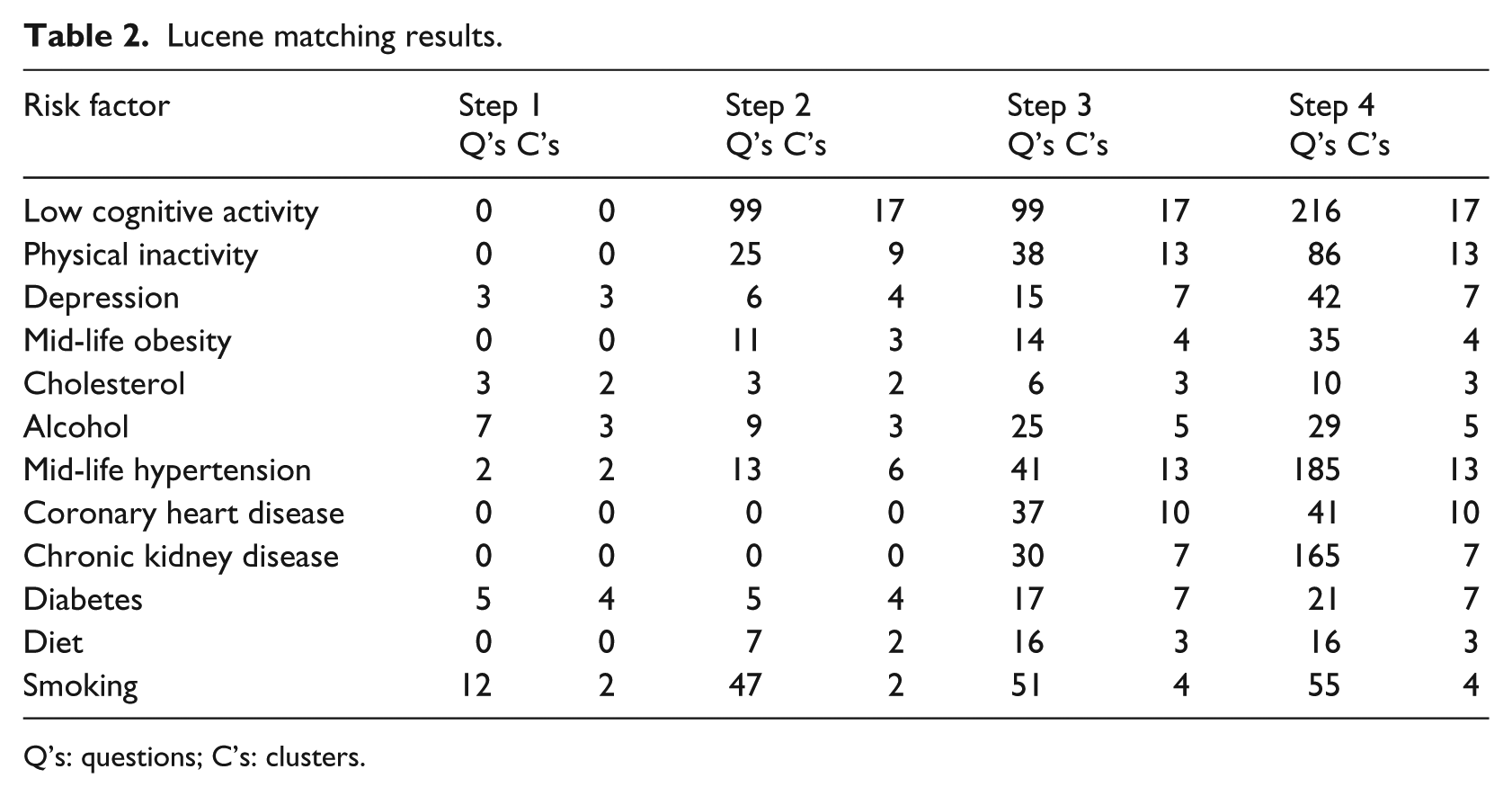
Q’s: questions; C’s: clusters.
Synonym-based match with thresholds Tt1 = 0.3, Tt2 = 0.8, Tt3 = 0.8, and Tt4 = 0.1.
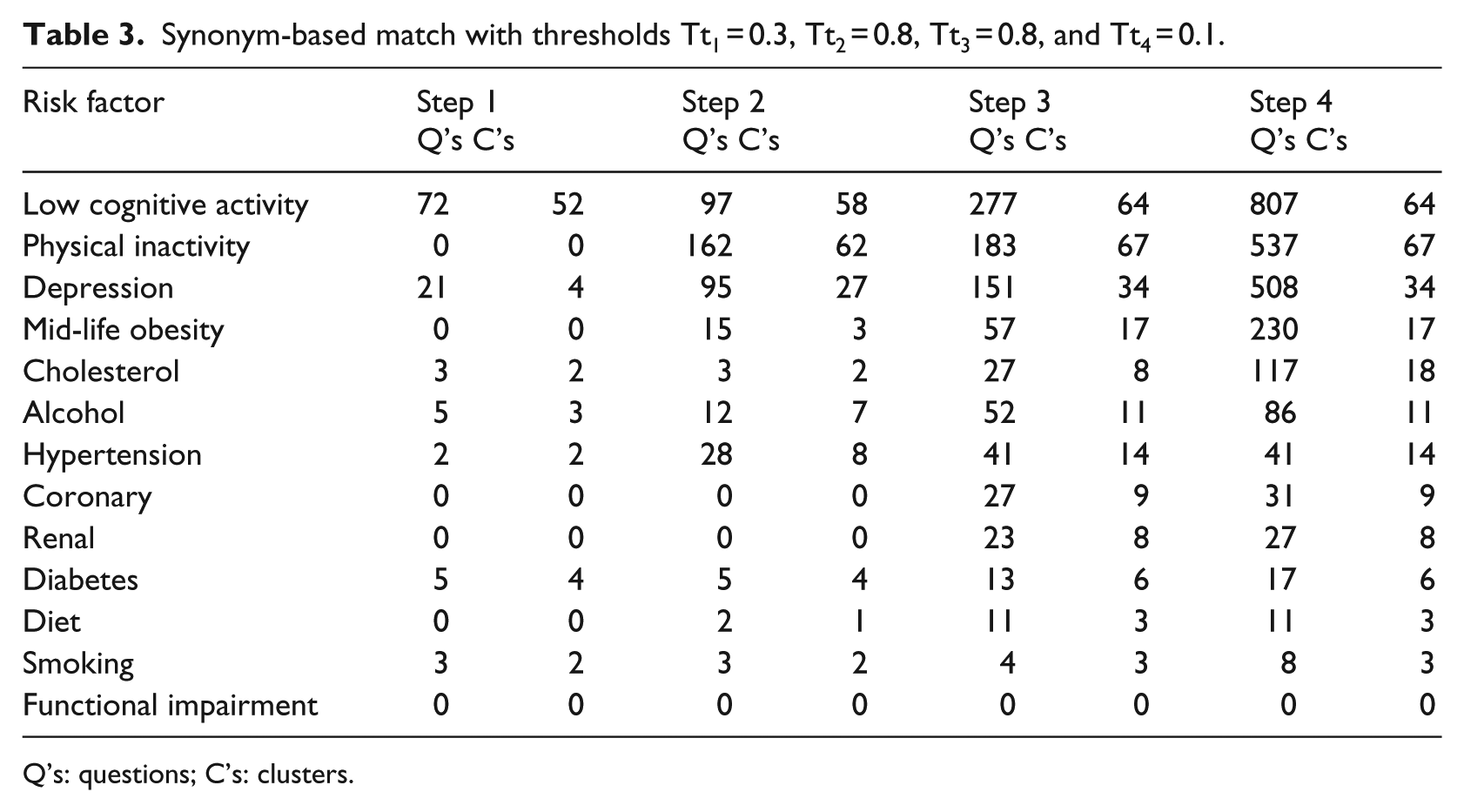
Q’s: questions; C’s: clusters.
Article structure
This article is organized as follows: in Section “Related research,” we discuss related work in this area; in Section “The In-MINDD architecture and ontology,” the In-MINDD architecture and ontology are presented; in Section “Mapping risk factors,” two mapping strategies are described; in Section “Evaluation,” we discuss how the process can provide useful metrics for fine-tuning the process, and we present an evaluation of both approaches and demonstrate why our current method outperforms the previous version; and finally, in Section “Conclusion” we offer some conclusions.
Related research
Automating ontology construction,17,18 population,19,20 and reuse 21 have long been topics of research. There have been many different applications, but none seem to have the overall aim of the creation of a system to quickly understand the scope of a longitudinal study and identify relevant areas of interest to a clinical specialist. The MedLingMap tool 22 is built with a similar goal in mind to ours—finding sections of interest in a corpora—but instead of clinical trial data it is the published literature of a niche research area.
There are research streams18,23 that overlap with the goals of this research, but instead aim to construct and extend ontologies rather than semantically enrich and query them. The authors of the first project 18 take pre-existing ontologies and use natural language processing (NLP) techniques, specifically semantic analysis and subject indexing, to completely overhaul existing ontologies and give them greater semantic depth. They process natural language (NL) descriptions of attributes and turn them into subject term descriptions for concept attributes. Through this semantic analysis and subject indexing, they extract concepts from the NL description and create non-taxonomical links between concepts instead of only having inheritance links. We do not adopt this approach as we are enriching the ontology with external knowledge rather than enriching the links between concepts. Other work 23 links to external sources of information where they mine domain texts, glossaries and dictionaries to extract feature and glossary groups found through the input of a seed-ontology. This is a semi-automated process as these groups aid an ontology creator in extending the ontology by presenting possible additions instead of automatically creating links to an external source. Again, this project is different to the work we have carried out in that it is mining domain corpora and linking them to an ontology rather than linking them to a clinical trial. They extract feature groups by stripping the stop words extant in the domain text as well as words irrelevant to the domain, and only after this, do they extract features (words) depending on their lexical co-occurrence within similar contexts
Researchers in Ding et al. 21 focus on automated ontology reuse instead of construction and enrichment, also using NLP to complete this task. NL web pages are evaluated to figure which bests fit their criteria through matching elements in the web pages, that is, concept names and concept values, to those that are in the ontology. The process here is the reverse of the work that we have carried out. They use NLP to establish the link between concepts, relationships, and attributes in documents with existing ontology sub-trees, whereas we use NLP to link an ontology and a sub-section of the NL elements in a longitudinal study to determine how the study can best be queried in order to test dementia-risk hypotheses.
There are two other bodies of work19,20 whose aim is to populate ontologies with the use of NLP, but they differ to this research in that they are not linking the ontologies to external sources in order to find the best areas for domain specialists to query. Ontology population consists of instance identification and maintenance (adding new concepts to the ontology that were not previously present). The first 20 uses hidden Markov models (HMMs) to recognize instances of a particular ontological concept. The HMMs are trained on sparsely and semantically annotated corpora, and the algorithm is used at runtime to identify matches. The second 19 combines two different techniques that are NLP—as previously mentioned—and information extraction (IE). These techniques are combined in the fashion that NLP identifies instance candidates, and IE is used to construct a classifier and then classify the instances.
In work most similar to that presented here, researchers 17 use existing ontologies (Australian Medical Terminologies (AMT), Systematized Nomenclature Of Medicine—Clinical Terms (SNOMED CT)) and link them to the Australian Imaging, Biomarker and Lifestyle (AIBL) study of aging. Our goal was to enrich the ontology from an existing dementia study rather than enriching the clinical trial data itself. They identified instances of a class in the ontology, like that of the drug paracetamol via the OpenClinica data standard meta-data (used in the creation of the study to give meaning to the trial data) and a two-phase mapping process. Also suggested in the article was a Linked Clinical Data Cube for more exhaustive querying of the clinical trial data through its links with ontologies. Our approach used NLP to identify inexact matches as opposed to pre-existing meta-data inherent in the trial. Longitudinal studies are not always constructed in a framework with such exhaustive meta-data; therefore, we think the NL approach is very pertinent. Also, we do not link clinical trial concepts with other class instances to see how they interact, but instead to allow clinical specialists to explore if the clinical, and in this case dementia, risk factors that were proposed in the literature are corroborated in the dataset. We also use a data cube in our validations, but we chose to develop in structured query language (SQL) rather than resource description format (RDF); it contains the results of the matching instead of results in the trial for efficient exploration and validation.
The In-MINDD architecture and ontology
In this section, we provide an outline view of the In-MINDD ecosystem and briefly describe the main components. While there are different approaches to constructing ontologies, 24 a common approach is to identify the basic (dementia) concepts, describe the properties of these concepts, and generate a vocabulary of concept-related keywords. This process here is referred to as Ontology Initialization, see Figure 1, and the terms in each stage of initialization map directly to those used in each of our three stages of matching. As the In-MINDD project is focused on dementia, these concepts are the associated (dementia) risk factors and the properties are those characteristics used to describe or measure a particular risk factor. When a clinical study has been identified as a candidate for knowledge extraction or any form of query processing, it is first necessary to import all of the questions presented in the study into the system. In effect, this is a process of generating meta-data. Most studies will have some form of structure where questions are asked in a specific order, or the study is sub-divided into identifiable sub-sections. This process is known as the Model Clinical Study phase as all questions are imported and are then sectioned into clusters as pre-determined by the study.
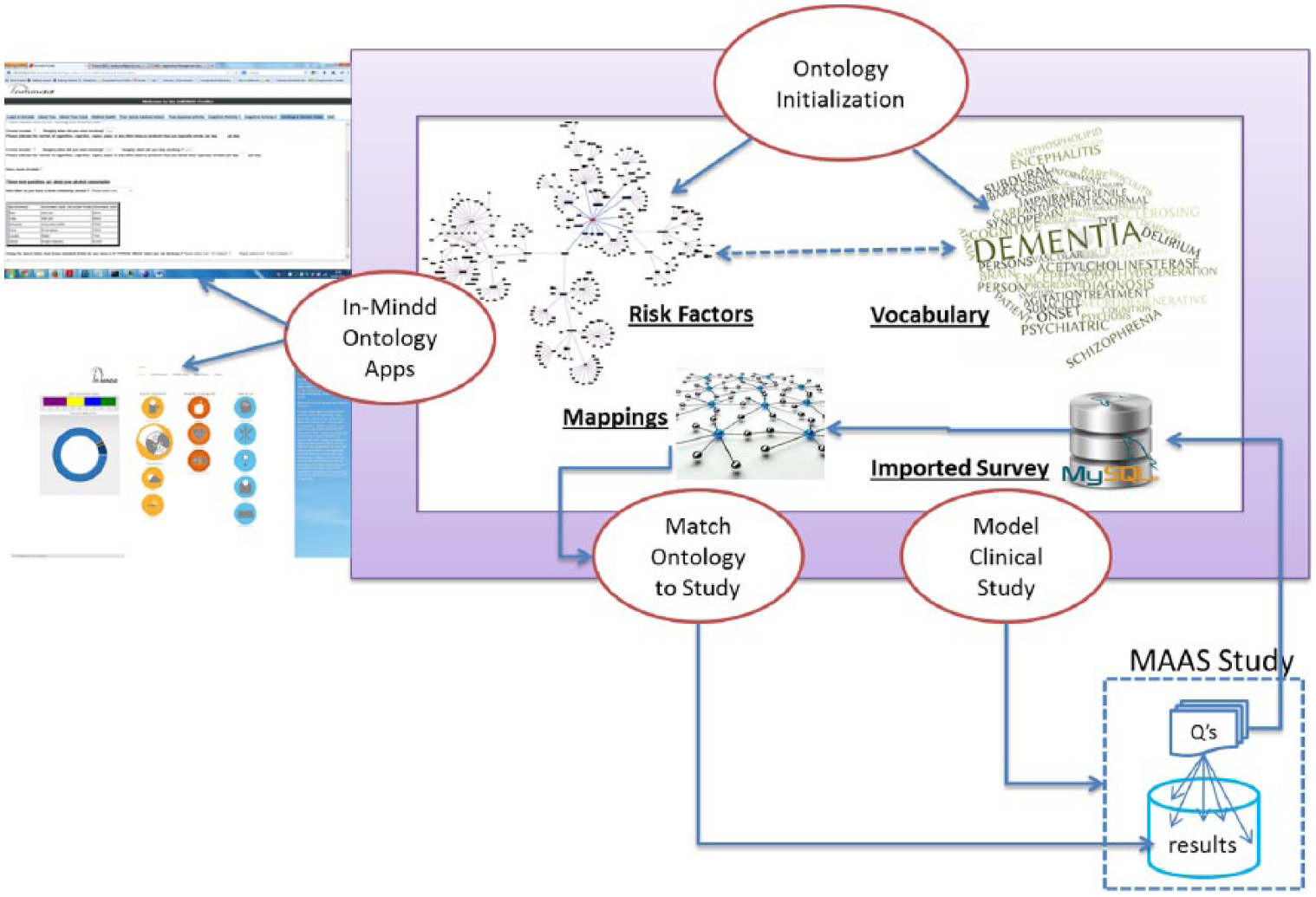
Ontology population and applications.
The result of this process is that all questions are given unique identifiers (many studies will already contain this information), and clusters are also given unique identifiers. In the case of clusters, many studies will already have these labels (e.g. Family History or Details of Activity/Exercise), although it is not necessary for the system to have meaningful labels. In other words, the system need not understand labels as they are merely used to classify questions into clusters.
The final phase of matching the ontology to the target clinical study is the primary focus of this article and is described in depth in the following section and builds on previous work. 14 In brief, the goal is to link each ontological concept (dementia risk factor) with all relevant questions in the clinical study. In Figure 1, a process for comparing risk factors with questions from the clinical study results in the generation of mappings or links between them. This removes the need for human pre-processing as required querying or data mining operations can now exploit the links to auto-generate query expressions. The In-MINDD apps pertain to an online profiler for dementia risk, a support environment for those with high risk, and the interface (not yet implemented) to the system described here to allow clinical researchers from the project to query dementia datasets in order to identify variables relevant to testing hypotheses.
The In-MINDD ontology contains 12 major concepts which are risk factors related to dementia: alcohol, coronary heart disease, physical inactivity, chronic kidney disease, diabetes, cholesterol, smoking, mid-life obesity, mid-life hypertension, diet, depression, and low cognitive activity. Each concept has a series of properties (measures), a method to calculate a score for each risk factor and an associated vocabulary which helps with matching questions from clinical studies. Due to space limitations, the properties and vocabulary cannot be described here. There have been two cloud-based applications developed using Google App Engine and the ontology: the profiler which populates the properties of each risk factor and computes the scoring, and the support forum which offers a means of reducing the score for selected risk factors.
Mapping risk factors
In this section, we present the two methods that match risk factors to clusters of questions in a clinical study. Method 1 uses Natural Language Tool Kit (NLTK) 26 and Method 2 uses Lucene 25 . Both use the same four steps as outlined in our previous article. 14 These are Concept Name Match, Concept Property Match, Vocabulary Match, and Structural Match. The first three steps focus on matching ontology keywords to terms used in the questions in clinical studies, each at a different level in the ontology, while the fourth uses the structure of the study to match further questions. In essence, each step adopts the same approach but uses different keywords for matching with the questions. For Step 1, we use the risk factor names (see Table 1); for Step 2, we use the properties that describe the risk factor; and for Step 3, we use all keywords that are contained in the vocabulary and are associated with the risk factor.
NLTK method using synonyms
The NLTK approach was described in Roantree et al. 14 and is briefly discussed again here for comparison with our more recent approach (discussed in Section “Lucene method using stemming and phrases”). The method was to find synonyms for all keywords and use this larger keyword set to map to as much of the clinical study as possible. The underlying NL technology used was WordNet 16 which allowed us to set thresholds for increasing or decreasing the matching proximity. For each iteration of keyword matching, we begin with a set of terms that represent the risk factor

Each term RFti is passed to the WordNet system 16 together with a threshold Tti which represents the level of synonym match to be used. WordNet then returns set(s) of word matches, which are combined so that for each term RFti, there is now a set of terms. In some cases, this will be a singleton set where only the term itself is returned

The goal at this point is to reduce multiple sets of synonyms to a single set for each risk factor as argument for the comparison algorithm. For risk factor RFi, we refer to the set of all possible synonyms as RFTi. A union operation is used to create a single set so that for risk factor RFi, all synonyms are present in RFTi

At this point, we have a single set of terms to represent each risk factor RFi.
As each clinical study is imported into the system as a series of clusters (representing sub-sections) of questions, each cluster has an identifier Cm, and within any cluster each question is identified by Qn. Thus, every question in the clinical study has a unique identifier provided by {Cm, Qn} where Cm represents the cluster identifier and Qn the question identifier. Each word in each question can then be addressed by the triple {Cm, Qn, Wo}.
Every term RFij is matched against each word Wo (excluding stop words) in each question {Cm, Qn} contained in the clinical study. If any terms {RFij, CmQnWo} match, then that question Qn is linked to the risk factor RFi in the ontology.
Lucene method using stemming and phrases
Due to the high number of false hits 14 with NLTK, our second method did not use synonyms alone. Instead, we also adopted a stemming approach and used the Lucene 25 library to develop our algorithms. Lucene 25 is an extremely rich and powerful full-text search library distributed by the Apache Software Foundation. At Lucene’s core is the Analyzer, which was used to create an inverted index for the MAAS dataset. The algorithms selected were part of the Snowball analyzer and facilitated the usage of word stems (a search on the word “smoke” will return entries in the document for “smoking,” “smoked,” etc.) and will match whole word phrases like “average units per week.” This led to fewer but more accurate synonyms found as well as different suffixes for each word.
As with the WordNet approach, we performed the same four-step matching process, although this time with a far smaller set of terms per risk factor. We took the terms at each of the three levels of the ontology, stemmed each term and found the appropriate synonyms, which lead to more accurate matching and better input for the structural matching phase. As part of the evaluation, the same audit of all matching operations was performed with one difference: with WordNet, RFij was a single term, while with the Lucene method, RFij could be a set of terms.
Structure-based mapping
Due to the nature of the questions in clinical studies, there remain many unmatched questions after the first three rounds of word matching. Example 1 shows a sample question (a) and statement (b) for participants to provide input. However, there is no context with which to associate either with a particular risk factor. Our approach is to associate this type of question with other questions that are richer in context or have clear keywords and an existing match to a risk factor. The second phase in the matching process uses the inherent structure in clinical studies to attempt to match remaining questions.
Example 1. No-Context Questions
Did you ever feel that it is all a bit too much? Choose option 1/2/3/9 as described in item 1
For most people it is easier to remember interesting facts than uninteresting facts. Answer from 1–9.
This stage begins with the creation of a matrix of clusters by risk factor. Recall that we use clusters to group sets of questions. The matrix is populated with the percentage of questions matched so far, for each cluster against each risk factor. For example, if cluster Ci has a total of 10 questions of which 5 are matched for risk factor RF1 and 8 for RF2, then
RF1Ci = 0.5
RF2Ci = 0.8
The algorithm is simple in approach. If any pairing {Ci, RFj} exceeds a set threshold Ts, then all of the remaining questions in cluster Ci are mapped to RFj. For the purpose of this analysis, the setting was Ts = 0.3.
Evaluation
The In-MINDD project uses the MAAS dataset for hypothesis testing. MAAS is an epidemiological study into biological, medical and psychosocial aspects of normal and pathological cognitive aging, 6 with 2372 questions spread across 79 clusters (or questionnaire sub-sections). Clusters had between 3 and 121 questions, with an average of 30 questions across each cluster. The In-MINDD ontology was generated using Intel Core 2 Duo processor CPU E8400 running at 3 GHz on a 64-bit Ubuntu 12.04 LTS platform. The NLTK 26 and WordNet 16 technologies were incorporated into a Python 2.7 application. We used the MySQL database with a data warehouse schema model to capture the result of every comparison operation and its result (matching status and threshold score).
Evaluation methodology
Recall that each sub-category or cluster has an identifier Cm, and within any cluster each question is identified by Qn. Thus, every question in the clinical study has a unique identifier provided by {Cm, Qn}, where Cm represents the cluster identifier and Qn the question identifier. A word or phrase is identified by the triple {Cm, Qn, Wo}. Every RFij is matched against each Wo (excluding stop words) in each question {Cm, Qn} contained in the clinical study. For all four steps (concept match, property match, vocabulary match, and structure match) in the matching process, a fact table FTwm with details of each comparison {RFij, CmQnWo} is created inside the data warehouse with the structure shown in Definition 1.
Definition 1. Fact table structure.
FTwm = {CID, QID, RFID, Step, RFti, CmQnWo, Tti, Result}
CID is cluster identifier; QID is the question identifier; RFID is the risk factor identifier; Step has a value of 1, 2, 3 or 4 depending on which step the comparison occurs; RFti is the risk factor; CmQnWo is the word or phrase identifier; Tti is the threshold (used for synonym match only); and finally, Result is the Boolean and indicates if the comparison was true or false. As the structure suggests, we adopt a purely relational approach to data mining queries for performance reasons and as we are dealing with a single relational style dataset. However, we also have an extensible markup language (XML)-based approach used 19 for web-based source data.
Evaluation forms are automatically generated for all risk factors. The database of matched questions is queried using SQL with the rand() function to randomly select 16 questions per risk factor, together with details of which (and how many) of the four steps results in the match. This sample of questions matched to risk factors were presented to dementia experts, and they were asked to indicate those matches which were relevant (true positives) and those which were not (false positives). Our fact table can easily be queried to determine at which of the four steps, the question was matched. For those matched correctly, we would like it to be matched as early as possible; for those matched incorrectly, we must determine which step provided the false hit.
Evaluation: degree of matching
For the WordNet approach using synonyms, we set initial threshold low in order that risk factors could be linked to as many questions in the clinical study as possible. Clearly, this has the risk of a high number of questions incorrectly linked to risk factors, but our analytical tools allow us to quickly identify the step at which the hit occurred and even the keyword. The purpose was to empirically determine the optimum thresholds for all four steps in matching links. The goal is to maximize matched questions to risk factors, while minimizing the number of false hits. The results of the initial run are shown in Table 1.
For Lucene, there were a smaller number of matches as the matching criteria were higher to some degree. While stemming was used to increase matches, no synonyms were used and where phrases were found in the ontology, phase-based matching was used. While thresholds were not required at Steps 1–3, the threshold of 0.1 was used for structural matching, meaning that if 10 percent of the questions in a cluster were matched, then the entire cluster was matched.
Evaluation: quality of matching
Querying this fact table is used as part of the validation process that determines both the accuracy of the links created between risk factors and clinical studies, and in cases where false hits occurred, to quickly drill down and determine the process which resulted in the false hit.
Definition 2. Result Analytics Sub-expression
(QID = <Query ID> | CID = <Cluster ID>)
The expression in Definition 2 is a standard SQL expression with three variables automatically extracted from the validation results, depending on the type of analytics required. Query Type can be one of Step, Risk_Factor_Term, Question_Term or Threshold. For example, if we wish to determine at which step a question was linked to a risk factor. The Risk Factor, Query ID, and Cluster ID variables are extracted from the report for those matches that are marked as “Not Appropriate.” The clause with QID provides more detailed analysis, while the clause with CID provides a more abstract analysis. Example 2 shows a query expression generated by the system.
Example 2. Result Analytics Sub-expression
select Step
WordNet/synonym approach
The system was used to run query expressions for all false positives found in the dementia expert’s validation report in order to conduct a high level analysis. In all, the number of false hits from the initial set of thresholds came to just over 70 percent for the matches shown in Table 1. The analysis of false hits can be summarized as follows: Step 1 had 11 percent, Step 2 had 42.5 percent, Step 3 had 46 percent, and Step 4 had no false hits. As a result of this process, we modified the thresholds for three of the four steps as shown in the captions for Table 3. The threshold for Step 1 remained the same; thresholds for Steps 2 and 3 were significantly higher; while the threshold for Step 4 was lowered. As Table 3 shows, although a lower number of matches were found, the number of false positives (found through random sampling and evaluation of a dementia expert) decreased to 23 percent.
Lucene stemming/phrase approach
The Lucene approach had a superior quality of matching as a lower number of false positives were found and a second run was not required. Less than 1 percent of the matches were found to be false positives in most of the factors, although within some, this was not the case. Some risk factors had a significant (>60%) number of false positives, that is, questions having no direct relation to the risk factor. When these were examined, it was found that some of the words used for the match did relate to concepts that were close to the risk factor but not directly, while a small number were not all related. An example of the latter were words like “intelligence,” “thinking,” and “remembering” that were used when searching for questions related to the risk factor “low cognitive activity.” These words were matched against many questions, but these questions were related to “cognitive ability” and “cognitive testing” but not necessarily the stated dementia risk factor, that is, cognitive inactivity. Similar situations were found in other risk factors, and this was found to be the cause of many false positives throughout the application.
Removing these types of keywords from the vocabulary eliminated most of the false positives in the risk factors where they were present. Similarly, use of words like “activity” when searching for matches against the risk factor “physical inactivity” produced matches against questions concerning the concept of “social activity” or “physical health.” It was harder to eliminate words from the vocabulary that produced such false positives in this case.
When searching for questions related to risk factors such as diet and obesity, we initially identified some questions found as false positives, but as the question did not have to be strictly related to the risk factor, many of these false positives became positives. For example, searches for questions related to obesity resulted in questions about weight, physical activity, lists of conditions that include those associated with obesity, appetite, and food consumption. Whereas weight is strictly related to obesity, the rest are broadly related to obesity, and once both types of questions were identified as positives, the number of false positives dropped sharply.
As the quality of matching was superior for the Lucene-based approach, it is worth examining each of the results for the risk factors in this case. After making modifications to the rules and vocabulary, the matching was as follows: depression, obesity, cholesterol, alcohol, renal, diabetes, and diet had no false hits; smoking and coronary had less than 10 % while low cognitive activity and physical inactivity had almost 50 percent false hits due to the reasons provided above. To reduce the false hits for the poorest performing risk factors, we adopted a system which looks at how matching occurred at each of the four levels. A threshold was set for specific risk factors whereby unless matching was made at three steps or more, the match was removed. This has the effect of removing a significant number of the false hits for the two worst risk factors.
Conclusion
Many of the clinical studies into the various effects of aging commenced 10 or 20 years ago. As a result, they are not well suited to modern approaches of defining risk factors and ontologies as a mechanism for better processing data and extracting knowledge. As these studies provide a wealth of information, a strategy for matching older clinical studies with new representations for knowledge is necessary. In this article, we presented an approach which maps older clinical studies to modern ontologies by building a small set of algorithms on top of existing NL utilities. Using our evaluation framework, we compared an approach used in earlier work 14 with a new approach and different technology, presented in this article. Unlike 14 where a high volume of false positives were identified, our current approach (with results in Tables 2 and 3) creates a high level of matching between the ontology’s risk factors and knowledge in clinical studies. Together with clinical experts, we demonstrated a low level of false matches through a system of automated evaluation forms and easy detection of where erroneous matches occurred.
While our ontology also provides an opportunity for interoperability across clinical studies and for an XML-based integration with online clinical data, we do not present a discussion here. Instead, this forms part of current research building upon the optimization strategies presented in Liu and colleagues.27,28 While only a brief description of the In-MINDD ontology was discussed here, it has evolved from the initial ontology in Roantree et al. 14 and is now stable to the point where it is used as a foundation for the profiler and support forum cloud-based apps. This forms part of a second stream of current research 13 where cloud-based medical data are anonymized.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research leading to these results has received funding from the European Union Seventh Framework Programme (FP7/2012) under grant agreement no. 304979.
