Abstract
Introduction
Pneumonia presents a significant risk to person’s health 1 with a mortality rate that accounts for about 30% of all respiratory causes of death. 2 Pneumonia is commonly misdiagnosed because of its similarities to several pulmonary conditions.3–6 Moreover, low sensitivity and specificity of diagnostic criteria, radiological and microbiological culture findings represent additional diagnostic challenge. 7
Efforts have been made to develop systems supporting physicians in the diagnosis of pneumonia.8–13 However, knowledge of pneumonia diagnosis that these systems rely on is rudimentary and often sufficient for a proof-of-concept demonstration rather than use at the point of care. In recent years, codification of medical knowledge with a help of ontologies gained ground due to their ability to represent medical concepts and relationships among them in a structured and formal manner. 14 Thus, an ontology can help with a representation of a complex knowledge, like one about diagnosing pneumonia, by providing a standard vocabulary that helps in integrating heterogeneous biomedical data sources. The only and a very basic pneumonia diagnosis ontology was developed as part of an ontology-driven clinical decision support system (CDSS) 15 to identify patients with pneumonia in the emergency department. However, this ontology did not capture the breath of the knowledge needed to diagnose pneumonia.
Research on medical ontologies grew over the last decade, 16 but many developed ontologies suffer from quality and content problems14,17 that can be mitigated by reusing existing high-quality ontologies. An empirical analysis of ontology reuse is described in Ochs et al. 18
The distinguishing feature of ontology development methodologies is dynamic, collaborative, and distributed character that involved using so-called foundational ontologies to ensure precision and interoperability. Foundational ontology includes upper ontologies which represent the very general concepts common across different domains, and upper domain ontologies which represent general concepts common for one specific domain. A big advantage of these methodologies is the fact that they facilitate interoperability among ontologies through their alignment to the same foundational ontology.
Building a high-quality ontology in a medical domain is a challenge further amplified by the breadth of clinical knowledge to be captured, diversity of knowledge sources, and a large number of other ontologies that cover sub-topics from a domain of interest. For example, concepts related to diagnosing pneumonia are partially covered in such ontologies as Systematized NOmenclature of MEDicine Clinical Terms (SNOMED-CT), Human Phenotype Ontology (HPO), or Disease Human Ontology (DDO). However, many of these concepts are differently represented in those ontologies making their reuse rather challenging. This challenge can be mitigated by an informed choice of the most appropriate representation of the concept to be reused so such representation is aligned with the intended use of the ontology.
This paper describes how we created a comprehensive Pneumonia Diagnosis Ontology (PNADO). Our PNADO development process started with using pneumonia diagnosis clinical practice guidelines (CPGs) as a primary clinical knowledge source (see Phase 2: Building corpus and extraction of terms for detailed description). CPGs are disease-specific textual, evidence-based documents that summarize the medical knowledge required to diagnose, manage, and treat the disease in question. 19 The preliminary PNADO created from multiple CPGs has been subsequently refined through the reuse of the elements of other ontologies. Indeed, we proposed an approach to reusing concepts coming from multiple ontologies and having different representations. Finally, we evaluated PNADO using medical data and by involving clinical domain experts. Our research fills a knowledge gap by not only creating a comprehensive and quality ontology for diagnosing pneumonia but also by showing how to reuse related ontologies in a consistent and formal manner in a process of ontology development.
Material and methods
The process of PNADO development is composed of five phases illustrated in Figure 1 and described in greater detail below. During the process, we followed well-established Open Biological and Biomedical Ontology (OBO) Foundry principles
20
and the ARCHitecture for ONTological Elaborating (ARCHONTE)
21
method to create preliminary PNADO (see Phase 3: Building preliminary pneumonia diagnosis ontology). Unlike other methods that do not fully adhere to the Institute of Electrical and Electronics Engineers (IEEE) standards,21,22 ARCHONTE supports a rigorous process of structuring the ontological concepts and facilitating modularization. PNADO development process.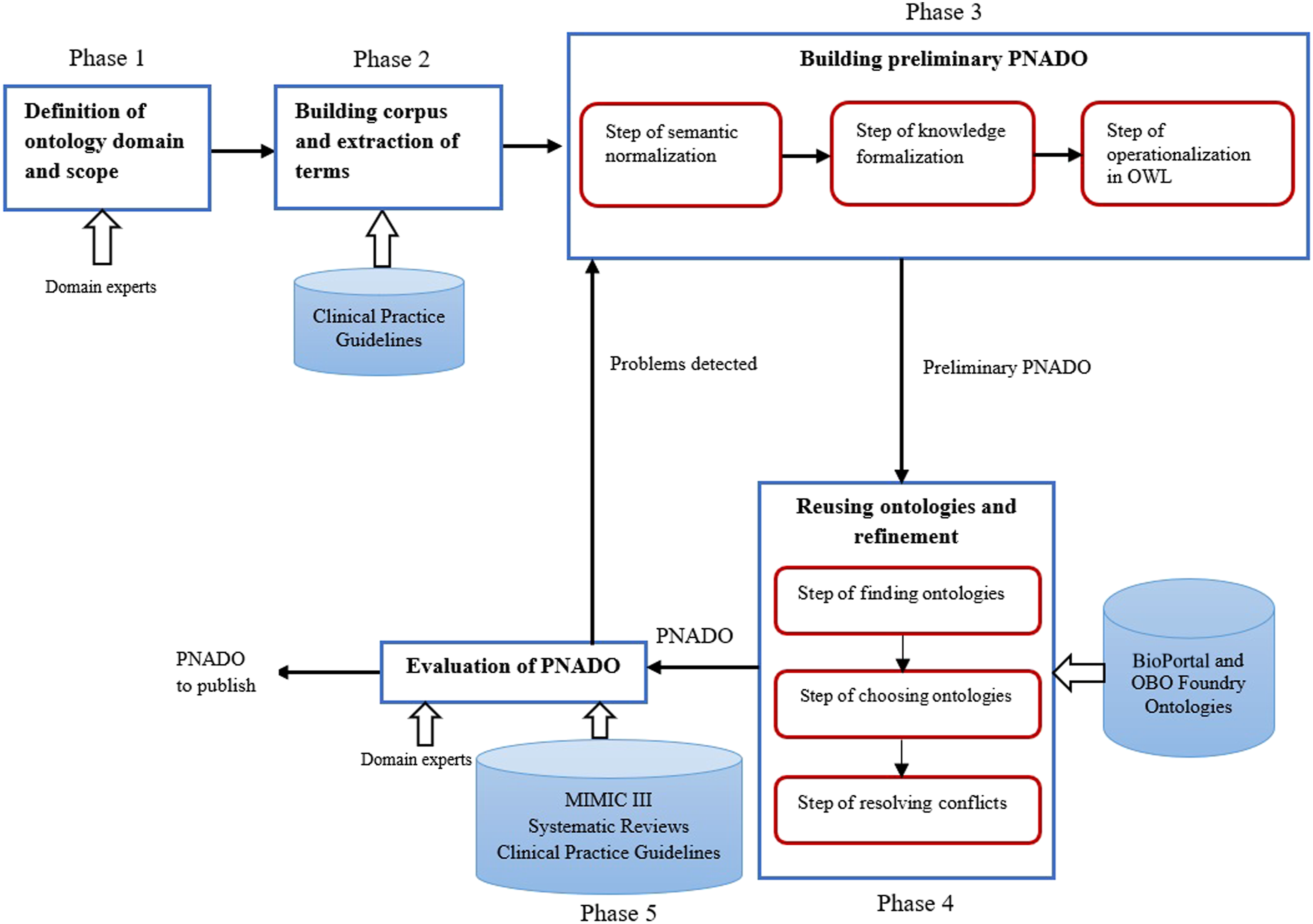
Phase 1: Definition of ontology domain and scope
We defined the ontology domain and its scope during sessions with collaborating emergency medicine physicians from the Gatineau Hospital, Gatineau, Quebec, Canada. Practical requirements for PNADO were established by a set of competency questions (CQs) created by physicians and derived from the CPGs covering main aspects of pneumonia diagnosis. The role of the CQs was to establish the scope of knowledge that physician needs in order to categorize and diagnose pneumonia. Representative examples of the CQs are: 1. What are the symptoms and clinical signs of pneumonia? 2. What are the types of pneumonia? 3. What are the pathogens of pneumonia? 4. What is the clinical history of a patient? 5. What laboratory tests are needed to diagnose pneumonia? 6. What are the results of these tests? 7. What are the results of the physical examination of a patient? 8. What is the result of lung imaging of a patient? 9. What are the potential complications of confirmed pneumonia?
Phase 2: Building corpus and extraction of terms
CPGs represent a reliable source for building the corpus of knowledge.23,24 In our study, we used 13 CPGs published in English and covering different pneumonia diagnosis-related clinical questions. These CPGs came from Cochrane Collaboration, 25 National Institute for Health and Care Excellence (NICE), 26 Infectious Diseases Society of America (IDSA), 27 European Society of Clinical Microbiology and Infectious Diseases (ESCMID), 28 Australian Society for Infectious Diseases (ASID), 29 Canadian Respiratory Guidelines (CTS), 30 Pulmonary and Critical Care Medicine (PulmCCM), 31 American Thoracic Society (ATS), 32 and British Thoracic Society (BTS). 33
We used Text2Onto34,35 tool to build the corpus from the CPGs. The CQs were used to identify terms in the corpus that are relevant to pneumonia diagnosis. These terms were then classified as symptoms, clinical signs, imaging tests and results, laboratory tests and results, pathogens, antecedents, types of pneumonia, differential diagnosis, and complications.
Phase 3: Building preliminary pneumonia diagnosis ontology
In this phase, we built the first version of the ontology called preliminary PNADO (p-PNADO) that covered general concepts. In building p-PNADO, we followed the ARCHONTE 21 method. The text below describes ARCHONTE steps (ontological concepts are in italic).
Step of semantic normalization
In this step, we used differential semantics principles 21 to describe in natural language the meaning of each concept retrieved from the CPGs, including similarities and differences to its parents and siblings’ concepts. Semantic normalization involved creating a hierarchy of retrieved concepts and relationships organized according to their similarities and differences. For example, concepts of bacterial pneumonia, fungal pneumonia, pneumonia due to parasitic infestation, and viral pneumonia are all sibling concepts because of their similarity (all are infectious diseases) and at the same time, they are different concepts because represent diseases caused by different pathogens.
Step of knowledge formalization
Here we added axioms that define relationships between concepts and their instances and we defined object properties of the concepts. The result of this step was an ontology that covered differentials concepts, relations, instances, and axioms for the pneumonia diagnosis domain. For example, for the concepts of bacteria and viruses, we added an axiom stating that these concepts are disjoint: bacteria disjoint with viruses. For the concepts of antigen test and pathogen, we added an object property
Step of operationalization in Web Ontology Language (OWL)
We used Protégé 36 and the upper medical ontology called Ontology of General Medical Science (OGMS) 37 to operationalize p-PNADO in the OWL language. The choice of OGMS is discussed later. We used MetaMap 38 tool to extract definitions, synonyms, acronyms, and other annotations from the Unified Medical Language System (UMLS). 39 If no definition was found in UMLS, then we left the concept not defined. At this last step of the ARCHONTE method, we obtained the final version of p-PNADO.
Phase 4: Reusing ontologies and refinement
In this phase, we added and detailed concepts in p-PNADO. For this purpose, we reused other ontologies. These ontologies had to be evaluated for the suitability of their content coverage and the depth of knowledge they represent. We identified two types of reuse: hard reuse when one imports the entire ontology, and soft reuse when one imports only the selected concepts and relations from another ontology.
Step of finding ontologies
We searched for these other ontologies in two established open content repositories of biomedical ontologies40–42: OBO Foundry and BioPortal. An ontology was considered as a candidate to reuse if: • it was reused in other ontologies and was described in a peer-reviewed publication; • it included axioms that were related to any of the requirements defined in Phase 1 of ARCHONTE method.
Using these two inclusion criteria, we identified 26 ontologies for potential reuse.
Step of choosing ontologies
For each CQ defined in Phase 1, we identified ontologies that could help with answering this CQ. The selection of the ontology was based on the assessment of its precision and accuracy. Here we measured precision by calculating the number of unnecessary concepts and we measured accuracy by calculating the number of omitted concepts. An ontology with minimal values of these two measures was being selected for reuse.
43
In case when multiple ontologies could be used to answer the same CQ, we selected ontology as follows: - In case of more than one precise ontology, reuse all these ontologies (hard reuse if the intersection of the covered concepts is empty). - In case where there is no precise ontology, then for any chain of ontologies T1,…,Tn such that the set of unnecessary concepts of Ti is included in the set of unnecessary concepts of Ti + 1 and the set of omitted concepts of Ti is included in the set of omitted concepts of Ti + 1, only reuse ontology T1 (hard reuse). - If none of the above, then the ontologies are incomparable with regards to accuracy and precision, consider soft reuse.
Following the process outlined above, we identified the following ontologies for reuse: OGMS because it describes high-level terms used across medical disciplines; Symptom (SYMP) 44 because of its standardized description of the concepts related to symptoms; Human Disease Ontology (DOID) 45 because of its standardized description of human disease concepts; Logical Observation Identifiers Names and Codes (LOINC) 46 because of the terms related to clinical documents; Radiological Lexicon (RadLex) 47 because of its repository of radiology terms; Relation Ontology (RO) 48 because of its relations between entities; Computer-Based Patient Record Ontology (CPRO) 49 because of patient profile description; Human Phenotype Ontology (HPO) 50 because of a controlled vocabulary for the phenotypic features in hereditary and other diseases of humans; Infectious Disease Ontology (IDO) 51 because of covering the infectious diseases, SNOMED-CT 52 because of its comprehensive coverage of clinical knowledge, and NCBITaxon 53 because of its classification and nomenclature of all the living organisms in the public sequence databases.
Step of resolving conflicts
Once the ontologies for reuse have been identified, we proceeded to manually identify the concepts to enrich p-PNADO and derive at a final version of PNADO. Note that these concepts might be in conflict as they are differently represented in selected ontologies. To resolve such a conflict, we defined conflict resolution questions (CRQs). The CRQs pertained to the meaning of a concept and the structure of the ontology concept tree (the hierarchy of classes, intersection and union of classes, equivalent classes, universal classes, universal and existential quantification, has-value restriction, and cardinality restriction).
Phase 5: Evaluation of pneumonia diagnosis ontology
After creating PNADO, we evaluated its quality. Different approaches to ontology evaluation were discussed in the literature.35,54–56 Also, several authors have proposed automated tools to evaluate ontologies.57,58 However, most of these tools are either prototypes or under development, with only OOPS! 59 and OntoMetrics 60 being sufficiently robust and ready to use. We evaluated PNADO according to the criteria proposed by Vrandecic and Zhu et al.61,62: accuracy (measured in terms of precision and recall), completeness, conciseness, consistency, clarity, computational efficiency, and adaptability. Considering that PNADO is not part of an application and in light that there is no gold standard for diagnosing pneumonia ontology, we followed data-driven evaluation approaches 56 and approaches involving clinical domain experts. 54 For a data-driven evaluation we used OOPS! and Pellet Reasoner for “debugging” PNADO. As domain-specific data sources, we used a clinical dataset Multi-parameter Intelligent Monitoring in Intensive Care III (MIMIC-III), 63 43 systematic reviews and two recently published CPGs64,65 that were not used in Phase 1 of PNADO development.
The evaluation of accuracy and completeness usually relies on having access to a gold standard built by domain experts. While such a gold standard did not exist in our case, we had to proceed differently. To calculate recall, we constructed a new corpus from 2 new CPGs and 43 systematic reviews related to pneumonia diagnosis and also used the MIMIC-III. In the corpus, we created 336 text fragments (each fragment had on average twenty lines of text) and manually extracted 710 concepts related to pneumonia diagnosis. Linking extracted concepts to concepts in PNADO, we found a match for 570 concepts giving a value of recall of 80%. The remaining 140 concepts that could not be matched were added to PNADO.
The MIMIC III contains 38,597 distinct and de-identified adult patient records and 49,785 hospital admissions. We identified 7702 records pertaining to pneumonia diagnosis for 32 types of pneumonia. The number of records pertaining to the ICD09 pneumonia code of 860 was more than half of all cases. To make sure that the evaluation covered all pneumonia types and the number of cases was still manageable, we applied the logarithm function to reduce proportionally the number of records to consider and randomly choose records for each type of pneumonia. During the random selection, we ignored records of patients with multimorbid conditions (as it is difficult to identify the concepts solely related to pneumonia). We ended up with 43 records. For each selected record, we manually annotated terms related to pneumonia diagnosis and verified whether they are covered in PNADO. For those that did not appear in PNADO, we checked if they were synonyms of existing concepts or if they were new concepts.
As a result of this manual evaluation, we found that 36 out of 988 concepts found in the MIMIC-III are not covered in PNADO. The analysis of each concept revealed that 16 of them were synonyms (for example pulmonary embolus and pulmonary embolism) and 20 were subclasses of existing concepts (for example recurrent cough and occasional cough are subclasses of cough). Thus, the recall value calculated earlier from the corpus remained unchanged.
To calculate the precision of PNADO, we used 3 clinical domain experts: a pneumologist from Charles-Le Moyne Hospital, Montreal, Quebec, an emergency medicine physician from the Gatineau Hospital, Gatineau, Quebec, and a family physician from the University of Quebec in Outaouais medical clinic. To explain to the domain experts their task, we prepared a brief document describing the ontology and the CQs. We conducted a number of training sessions using mock ontologies. Once the domain experts were comfortable with the evaluation task, we created accounts in WebProtégé, 66 and shared PNADO. As part of the evaluation, domain experts were asked to evaluate each concept (its relevance, position in the ontology tree, its relations with the other concepts) and to comment directly in WebProtégé. As a result of this expert evaluation, 23 changes were made in PNADO, including adding concepts such as radiological signs of consolidation: air bronchograms, ill-defined, fluffy opacities, air alveologram; moving concepts of fever to general symptom, purulent tracheobronchial secretions to respiratory system and chest symptom, and removing concepts such as transitory tachypnea of the newborn.
Finally, clinical domain experts together with the knowledge engineer (SA) evaluated the clarity of PNADO. Specifically, they evaluated how well PNADO captured pneumonia diagnosis knowledge by assessing how effectively it communicated the meaning of the concepts.
Results
Results of PNADO evaluation.
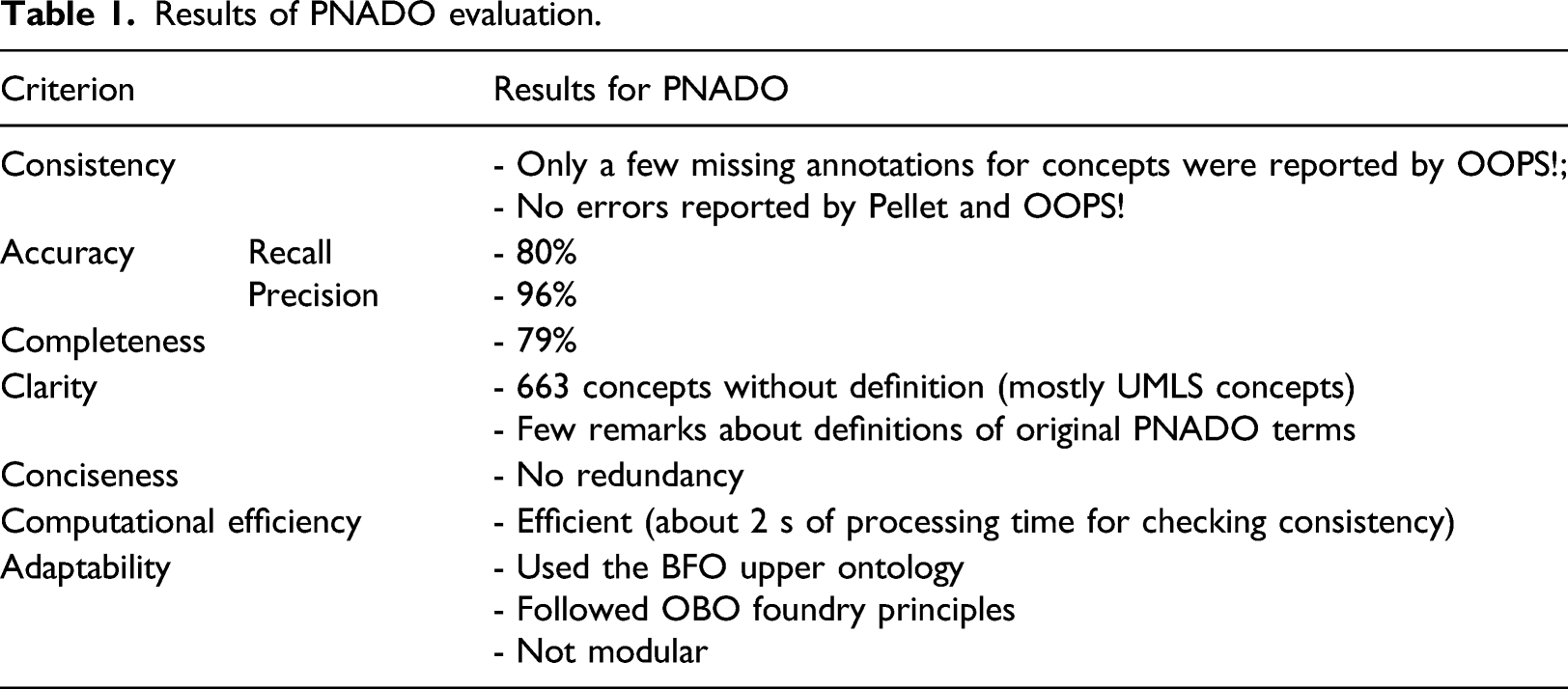

PNADO describes pneumonia diagnosis knowledge using 1640 unique concepts, 1598 classes, 42 object properties, 1591 logical axioms, and 83 annotation properties. The latest version of PNADO is published in the BioPortal repository and can be accessed at https://bioportal.bioontology.org/ontologies/PNADO.
Reused concepts and conflict resolution.
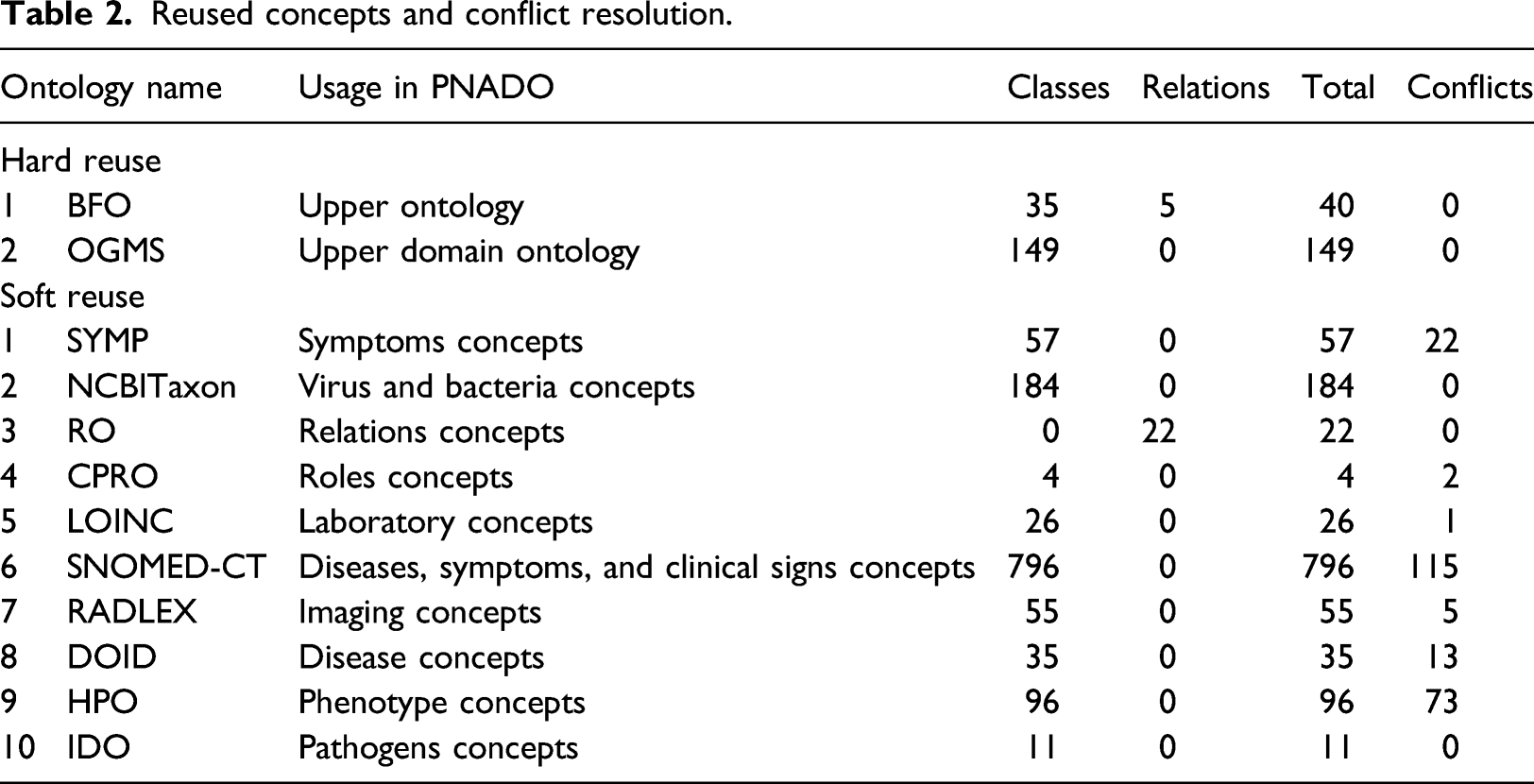
Figure 2 illustrates the top-level structure of PNADO. The main classes of PNADO are represented under the relevant classes of upper ontology Basic Formal Ontology (BFO) and upper domain ontology OGMS. For example, symptom class (9) is a class from OGMS and it is populated with new PNADO classes and classes that are reused from SYMP, HPO, and SNOMED-CT ontologies. Top-level hierarchy of PNADO.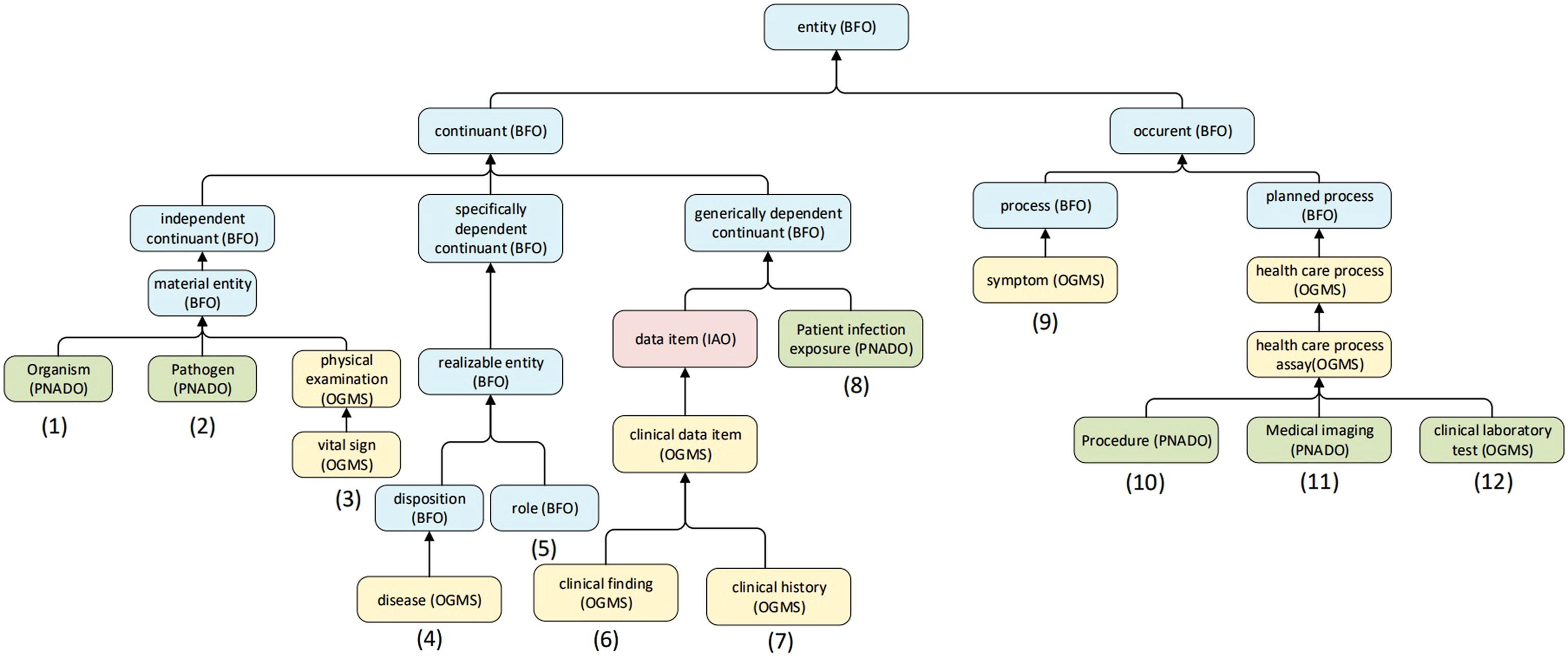
Figure 3 shows a fragment of the infective pneumonia class hierarchy and, in particular, the subclass viral pneumonia that represents the different types of viral pneumonia. This disease is caused by some Viruses (subclass of Organism), also called Pathogen. Viral pneumonia class is mainly populated from SNOMED-CT and we created an axiom to represent associated complications of viral pneumonia. It is represented under the class disease. An example of a viral pneumonia subclass.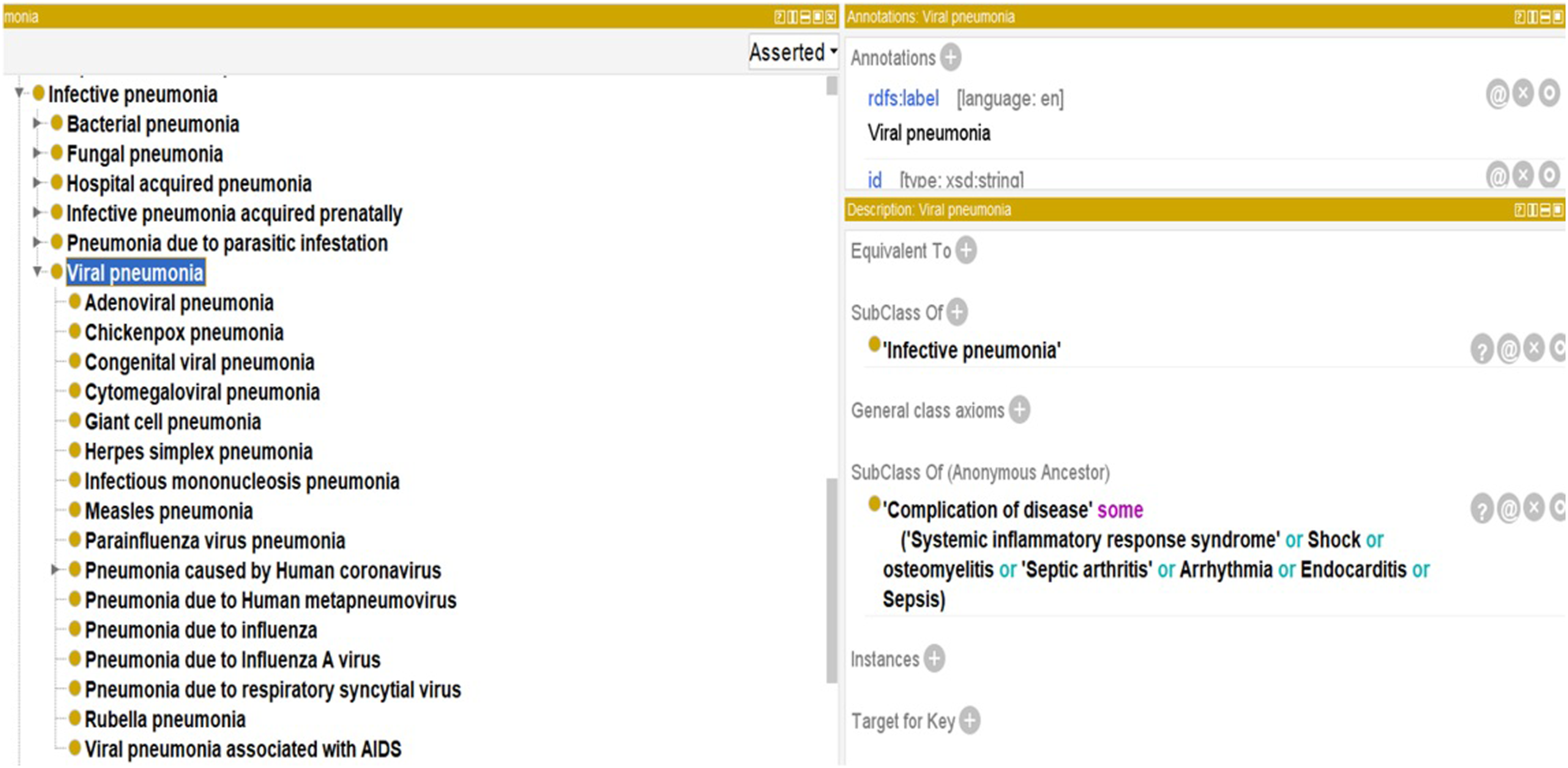
Discussion
Research described in the paper resulted in development of PNADO—an ontology that codifies knowledge of diagnosing pneumonia. PNADO covers different types of pneumonia, its symptoms, clinical signs, pathogens, laboratory tests and imaging, clinical findings, complications, and diagnoses.
In our opinion, a primary challenge of creating a new ontology (such as PNADO) and integrating it with the existing ones is the evolutive, distributive, and heterogeneous nature of the knowledge to be represented in the modules of an ontology. The interfaces between the modules are hard to define, the knowledge they represent evolves so even modules with a high level of abstraction—so-called upper domain ontology—require frequent updates. In this section, we reflect on the structured approach illustrated in Figure 1 that we followed while creating PNADO.
We noticed that while OGMS was used as an upper domain ontology, it did not include many concepts (classes and relations) required for the description of a diagnostic process. PNADO covers the concepts related to pneumonia diagnosis thus the concepts related to the diagnostic process should be available in an upper domain ontology (OGMS), but this was not our case.
A different example concerns the concept of a patient. While high-quality upper domain ontology that describes this concept does not exist, there are several domain ontologies where these descriptions can be found (e.g., CPRO, PatientSafetyOntology 67 ) but they model concept of a patient differently.
During PNADO development we realized that most of the reused ontologies do not respect the principles of modular design. 18 We note that it would be useful to design several levels of upper domain ontologies to get a better module’s coupling and cohesion, and at the same time, to reduce the complexity of reuse and interoperability issues.
Another challenging part of ontology development is ontology reuse. Our experience with PNADO development revealed the following: - An ontology to be reused may have the representation of a concept that is not relevant considering the requirements of the target ontology. - An ontology to be reused is selected based on its ability to answer a set of pertinent CQs; however, its reuse may introduce conflicts among the concepts associated with other CQs. For example, reusing DOID ontology because of its coverage of the concepts related to diseases required from us resolving new conflicts among concepts related to symptoms and clinical signs. - Having a concept to be reused represented in multiple ontologies, may require manual fine-tuning as illustrated by representing the concept of medical complication (discussed below). - Most of the reused ontologies did not respect the principles of modular design.
18
This added to the complexity of reuse and interoperability issues.
A medical complication is a phenomenon that may occur during or after an onset of a disease, procedure, and treatment. This concept is represented as a class in many ontologies. In SNOMED-CT, the complication is a subclass of disease and represents disease complications such as complications due to diabetes mellitus, or complication due to Crohn’s disease. Diabetes mellitus and Crohn’s disease are subclasses of disease. The diseases causing complications are also represented as subclasses of the class disease. Such a representation creates a heritage chain disease → complication → disease and does not cover all aspects of complications due to disease, procedure, treatment, or function. In PNADO we took a different approach and represented complication as an object property named complication of with four sub-properties: 1) complication of disease that associates disease to disease; 2) complication of procedure that associates procedure to disease; 3) complication of treatment that associates treatment to disease; and 4) complication of function that associates function to disease. Axioms were progressively added as needed to represent complications. We posit that our representation is cleaner and improves interoperability and reusability.
Limitations
Our work has some limitations. First, we were not able to evaluate PNADO with a help of a gold standard as such does not exist and there is no reference ontology describing similar medical conditions to be used for comparison. We mitigated this limitation by using systematic reviews, new CPGs and MIMIC III data. Nevertheless, this allowed for evaluating the concepts (including relationships) of PNADO and not the logical axioms. Furthermore, some evaluations were conducted by the knowledge engineer who developed PNADO, which may have introduced unintentional bias. PNADO, similarly to most of the ontologies published in the BioPortal repository, lacks modular structure. While our intention was to follow modular ontology design, we were constrained by lack of the tools supporting the creation of modular ontologies.
Conclusions
PNADO was developed starting with the textual knowledge representation captured in multiple CPGs that was subsequently expanded and verified through a rigorous ontology engineering process. PNADO is a comprehensive pneumonia diagnosis ontology that provides a standard vocabulary for biomedical entities and integrates multiple knowledge resources to improve interoperability, reusability, and portability. It is the first pneumonia diagnosis ontology to represent different aspects of pneumonia in a formal logical format. A comprehensive evaluation of PNADO included a number of data-driven approaches and clinical domain expert evaluations. PNADO is now available in the BioPortal repository for consultation and reuse. Its current form can be used by the developers working on systems supporting a diagnosis of pneumonia or it can be reused by ontology engineers developing ontologies describing similar diagnostic situations.
Footnotes
Acknowledgements
We would like to thank the CISSSO (Centre Intégré de la Santé et des Services Sociaux de l’Outaouais) for supporting this research. We are grateful to Dr Sylvain Croteau, Dr Yasmine Lisa Rebaine, and Dr Serge Chartrand for teaching us about pneumonia diagnosis and for their work on evaluating PNADO. We thank Professor Veronique Nabelsi for her advice during PNADO development process.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was partially supported by CISSSO.
